Abstract
BAP1 is an emerging tumor suppressor whose inactivating mutations have been found to play critical roles in tumor development. This study was conducted to elucidate the potential value of BAP1 mutation in guiding prognostic prediction and clinical stratification. We conducted a comprehensive analysis of relevant studies from multiple databases, to determine the impact of BAP1 mutation on the overall survival and disease-free survival of patients in various cancers. A total of 2457 patients from 21 studies were included in the final analysis. Although the pooled results demonstrated that BAP1 mutation was a negative indicator of overall survival (hazard ratio = 1.73; 95% confidence interval = 1.23–2.42) and disease-free survival (hazard ratio = 2.25; 95% confidence interval = 1.47–3.45), this prognostic value was only applicable to uveal melanoma and clear cell renal cell carcinoma, but not to malignant pleural mesothelioma or cholangiocarcinoma. Consistently, BAP1 mutation was correlated with critical clinicopathological features only in uveal melanoma and clear cell renal cell carcinoma. In uveal melanoma, BAP1 mutation and SF3B1/EIF1AX mutations were negatively correlated, and BAP1-mutant tumors indicated significant worse prognosis than SF3B1/EIF1AX-mutant tumors (p = 0.028). While in clear cell renal cell carcinoma, BAP1 mutation was mutually exclusive with PBRM1 mutations, and BAP1-mutant clear cell renal cell carcinomas also showed significantly worse prognosis than PBRM1-mutant clear cell renal cell carcinomas (p = 0.001). Our study revealed a unique tissue-specific significance of BAP1 mutation in prognostic prediction among different types of cancer. Clinically, combining detection of BAP1 mutation and other driver mutations may further allow for a more precise molecular taxonomy to stratify patients into distinct subgroups in uveal melanoma and clear cell renal cell carcinoma.
Introduction
The BAP1 gene (BRCA1-associated protein 1) is located on chromosome 3p21.1 and encodes a 90-kDa, nuclear localized deubiquitinase (DUB), which contains an N-terminal ubiquitin carboxyl hydrolase (UCH) domain and a C-terminal nuclear localization signal (NLS) domain. The BAP1 protein was first identified to interact with BRCA1 and mediate its growth-suppressive properties. 1 Later studies unraveled the multifaceted functions of BAP1 by interacting with various proteins.2–10 Relying on its DUB activity, BAP1 was found to play pivotal roles in epigenetic modification, 4 transcription regulation,2,10,11 and DNA damage response.6,7,9,12–14
In recent years, BAP1 emerged as a critical tumor suppressor as its inactivating mutations were recurrently identified in several cancers. 15 At first, germline mutation of BAP1 was associated with an inherited cancer syndrome characterized by uveal melanoma (UM), malignant pleural mesothelioma (MPM), clear cell renal cell carcinoma (CCRCC), and cholangiocarcinoma (CCA).16–19 Later on, somatic mutations of BAP1 were also identified in corresponding sporadic cases.20–23 Furthermore, loss of BAP1 indeed promoted cancer development in genetic engineered mouse models of these cancers.24–27 Clinically, the tissue-specific manner of BAP1 mutation showed potential applicability in routine diagnosis of these cancers.28–31 Meanwhile, accumulating studies reported that BAP1 mutations were also significantly correlated with tumor metastasis and poor prognosis, indicating a potential metastasis suppressor role of BAP1.22,32 To date, there has been no consensus on the prognostic role of BAP1 mutation in cancers. Although a recent meta-analysis reported that BAP1 deficiency was associated with poor prognosis in multiple cancer types except for a protective role in MPM, 33 there are several limitations in that study. First, the author used BAP1 deficiency (including both BAP1 mutation and BAP1 immunohistochemistry loss) as the parameter, which generated a certain heterogeneity as BAP1 mutation was not the only factor impacting BAP1 expression. Second, the studies enrolled were relatively small, especially in MPM (two studies), which is not sufficient to draw convincing conclusions.
For these considerations, in this study, we intend to exclusively analyze the prognostic value of BAP1 gene mutation in multiple cancer types. Furthermore, based on the antagonistic effect of BAP1 mutation with other key gene mutations, we will further establish a mutation-defined classification which may shed light on the prognostic stratification for BAP1-mutated cancer types.
Method
Study design and literature search
This study was performed strictly according to Reporting Recommendations for Tumor Marker Prognostic Studies (REMARK). 34 We searched the PubMed, Web of Science, and Embase comprehensively for relevant studies until July 1, 2016. The search terminologies were as follows: (“BAP1” OR “BRCA1 associated protein 1” OR “UCHL2” OR “hucep-6” OR “HUCEP-13”) AND (“survival” OR “mortality” OR “fatality” OR “death” OR “hazard ratio” OR “prognosis” OR “prognostic” OR “recurrence” OR “progression”). These terminologies were used in all possible combinations without language restriction. The reference lists of included studies were also scanned manually for identification of additional potentially relevant publications.
Inclusion and exclusion criteria
Four investigators (X.-Y.W., Z.W., X.-D.R., and J.-B.H) independently evaluated the included studies according to the following criteria.
Inclusion criteria were as follows: (1) studied BAP1 mutation in any type of human cancers; (2) mutations of BAP1 were detected on tumor tissues, rather than in blood cells; (3) comparison group was the population with wild-type of BAP1 gene; and (4) reported data sufficient to calculate the association of BAP1 mutation with patient survival and other clinicopathological features.
Exclusion criteria were as follows: (1) reviews, case reports, conference records, comments, and guidelines; (2) animal studies, familial studies, and cell line studies; (3) not written in English; (4) detected germline mutations but not somatic mutations of BAP1; (5) investigated the protein expression of BAP1; (6) not able to provide sufficient data to calculate the impact of BAP1 mutation on patient survival; and (7) duplicated information of previous publications. Two reviewers determined study eligibility independently. Disagreements were solved by group meetings.
Quality assessment and data extraction
Two investigators (X.-Y.W. and Z.W.) independently made the assessment of literature qualities according to the Newcastle–Ottawa Scale (NOS; http://www.ohri.ca/programs/clinical_epidemiology/oxford.htm). Only the publications with the score more than six were included in the final analysis. Disagreements were solved by consensus.
Key data were extracted from the included articles based on the guidelines proposed by the Meta-analysis of Observational Studies in Epidemiology (MOOSE) 35 and Preferred Reporting Items for Systematic reviews and Meta-Analyses (PRISMA) statement. 36 For each paper, the following information was obtained: (1) first author name, (2) cancer type, (3) year of publication, (4) study region, (5) patient number, (6) male-to-female ratio, (7) mean age, (8) comparison group, (9) gene mutation rate, (10) specimen type, (11) mutation detection method, and (12) the hazard ratio (HR) and 95% confidence interval (CI) of BAP1 gene mutation for overall survival (OS) and disease-free survival (DFS). If the individual patient-level survival data were available, HR was calculated by a Cox regression model. If the survival information was only available in Kaplan–Meier curves, HR was obtained by the method of Parmar et al. 37
Statistical analysis
We used Stata version 12.0 (STATA Corporation, College Station, TX, USA) to perform the meta-analysis of the enrolled studies. HRs with their 95% CIs obtained from each article were used to calculate pooled HRs. The χ2 test was applied to test for statistical heterogeneity, and the I2 test was used to assess the extent of variability attributable to statistical heterogeneity across trials. p > 0.05 for the χ2 test and I2 < 50% were interpreted as signifying low-level heterogeneity. Either a fixed-effects model (if I2 < 50%) or a random-effects model (if I2 > 50%) was applied to estimate the summary HRs. To investigate the source of heterogeneity, we performed predefined subgroup analysis based on cancer type. Furthermore, subgroup analysis on the relationship between BAP1 mutation and clinicopathological features was conducted in different cancer types. Begg’s test and Egger’s test were both performed to evaluate potential publication bias. In the mutation-defined classification, OS of different mutational subgroups was compared with the Kaplan–Meier method, and the significance was determined by the log rank test. This analysis was performed with the Statistical Package for the Social Sciences (SPSS) version 15.0 software for Windows (SPSS, Chicago, IL, USA). All p values were two-sided.
Results
Prevalence and distribution of BAP1 mutation among cancers
After the identification, screening, and eligibility assessment procedures (Figure 1), a total of 21 non-duplicated articles covering 2457 patients were included in the final study (Table 1). All these articles were published during 2010–2016. According to NOS assessment, all 21 articles were of high quality (score ≥6). Geographically, the 21 included studies were mainly from the United States (12/21), Asia (4/21), and Europe (5/21). The sequencing methods varied among different papers, including Sanger sequencing (SS), Access Array sequencing (AAS), multiplex ligation-dependent probe amplification (MLPA), Integrated Mutation Profiling of Actionable Cancer Targets (IMPACT), next-generation sequencing (NGS), whole genome sequencing (WGS), and whole exome sequencing (WES). Interestingly, all of these articles were eventually confined to four cancer types (seven about UM, six about CCRCC, four about MPM, and four about CCA). This tissue-specific manner of BAP1 mutation was also confirmed by COSMIC and cBioPortal databases as we re-organized in Figure S1. BAP1 mutation rates ranged from 6.4% to 57.7% in all the studies enrolled here, with highest in MPM (22.6%–57.7%) and UM (20.6%–50.0%) and relative lower in CCRCC (6.4%–14.5%) and CCA (8.4%–28.6%) (Table 1).

Flow diagram of the study selection process.
Characteristics of the studies included in the meta-analysis.

CCA: cholangiocarcinoma; CCRCC: clear cell renal cell carcinoma; MPM: malignant pleural mesothelioma; UM: uveal melanoma; FFPE: formalin-fixed paraffin-embedded; SS: Sanger sequencing; NGS: next-generation sequencing; MLPA: multiplex ligation-dependent probe amplification; IMPACT: Integrated Mutation Profiling of Actionable Cancer Targets; WES: whole exome sequencing; WGS: whole genome sequencing; OS: overall survival; DFS: disease-free survival; FAAS: fluidigm access array sequencing.
Impact of BAP1 mutation on patient outcome in different cancers
Altogether, there were 15 and 11 studies with available data about the impact of BAP1 mutation on OS and DFS, respectively. On the whole, the forest plot demonstrated that BAP1 mutation was indeed correlated with worse OS (HR = 1.73; 95% CI = 1.23–2.42, Figure 2(a)) and DFS (HR = 2.25; 95% CI = 1.47–3.45, Figure 2(b)) in this all-cancer cohort. However, the results showed significant heterogeneity both in the OS group (I2 = 65.7%; p < 0.001) and the DFS group (I2 = 54.9%; p = 0.009).
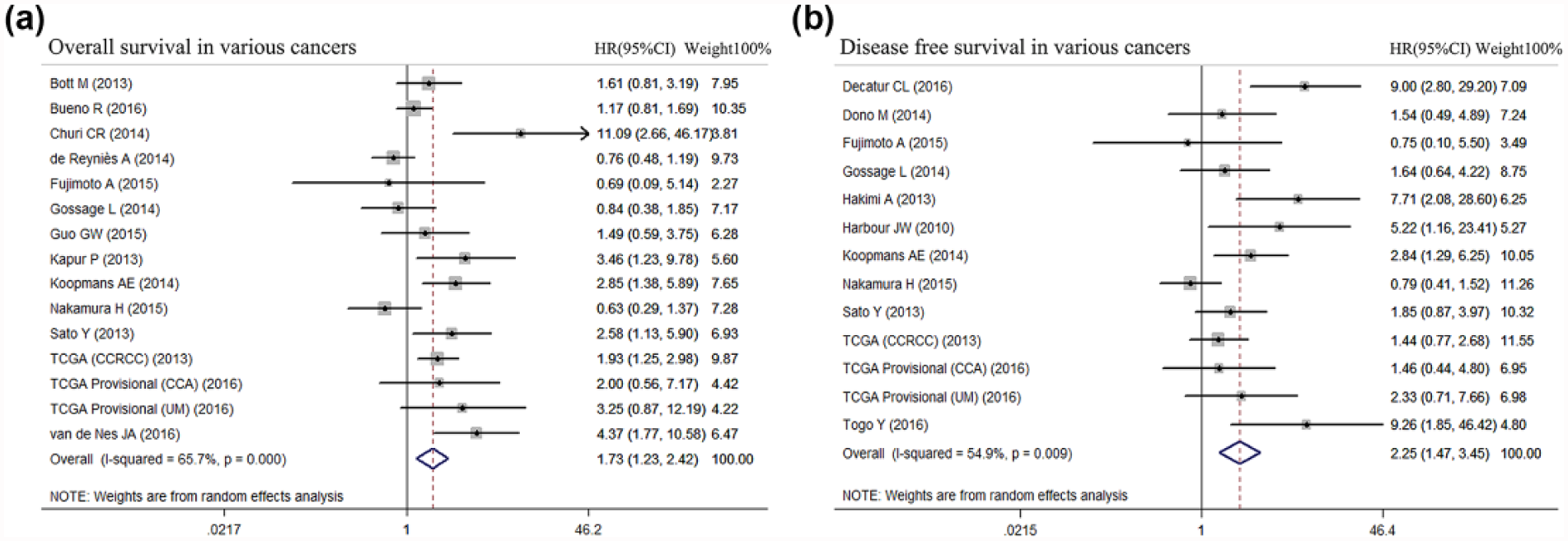
Prognostic value of BAP1 mutation status in patients with cancer. (a) Forest plots of the pooled HRs for the association between BAP1 mutation status and OS in various cancers and (b) forest plots of the pooled HRs for the association between BAP1 mutation status and DFS in various cancers. Squares and horizontal bars indicate the point estimates (HRs) with 95% CIs for each individual study. Diamonds indicate the summary estimates for the hazard ratio. The width of the diamond corresponds to the 95% CI.
To uncover the tissue-specific significance of BAP1 mutation, we further analyzed the prognostic impact of BAP1 mutation in UM, MPM, CCRCC, and CCA, respectively. Interestingly, in UM and CCRCC, BAP1 mutation was significantly correlated with both OS (HR = 1.86, 95% CI = 1.34–2.58 in CCRCC, Figure 3(a); HR = 3.36, 95% CI = 2.00–5.64 in UM, Figure 3(c)) and DFS (HR = 2.48, 95% CI = 1.31–4.68 in CCRCC, Figure 3(b); HR = 3.20, 95% CI = 1.97–5.20 in UM, Figure 3(d)). However, in MPM and CCA, BAP1 mutation has no impact on OS or DFS (Figure 3(e)–(g)). All these results suggested that BAP1 mutation predicts cancer patient outcome in a tissue-specific pattern.
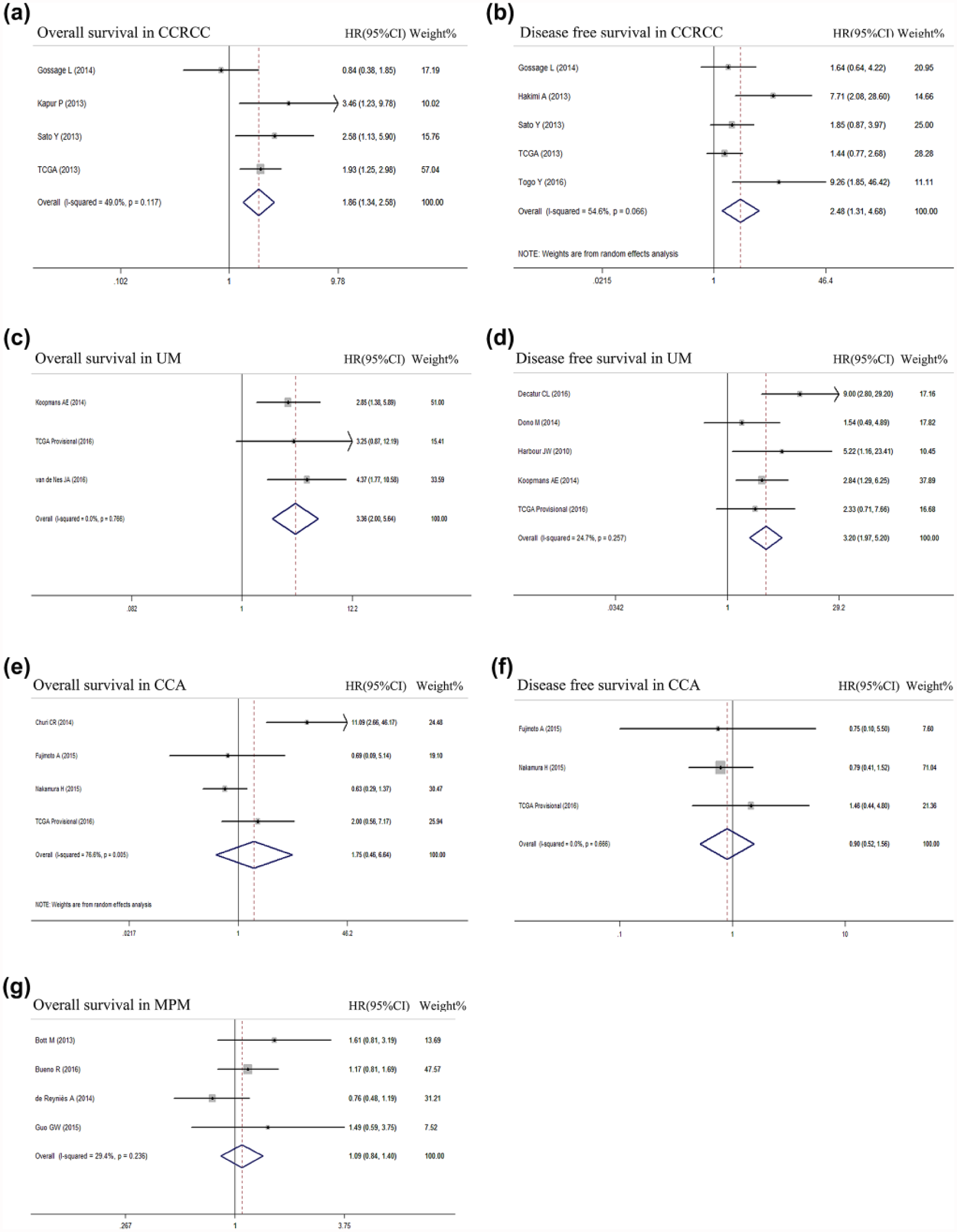

Prognostic value of BAP1 mutation status in CCRCC, UM, CCA, and MPM. (a) Forest plots of the HRs and 95% CIs for OS in CCRCC patients. (b) Forest plots of the HRs and 95% CIs for DFS in CCRCC patients. (c) Forest plots of the HRs and 95% CIs for OS in UM patients. (d) Forest plots of the HRs and 95% CIs for DFS in UM patients. (e) Forest plots of the HRs and 95% CIs for OS in CCA patients. (f) Forest plots of the HRs and 95% CIs for DFS in CCA patients. (g) Forest plots of the HRs and 95% CIs for OS in MPM patients.
Correlation of BAP1 mutation with clinicopathological features in different cancers
To explore the relationship between BAP1 mutation and clinicopathological parameters, we performed subgroup analysis according to different cancer types. As Table 2 suggested, BAP1 mutation was significantly associated with multiple critical clinicopathological factors in CCRCC and UM, but not in MPM and CCA. In CCRCC, BAP1 mutation status was significantly associated with larger tumor size (odds ratio (OR) = 2.92, 95% CI = 1.43–5.98), higher Fuhrman grade (OR = 2.32, 95% CI = 1.46–3.67), sarcomatoid differentiation (OR = 4.12, 95% CI = 1.74–9.74), metastasis (OR = 2.04, 95% CI = 1.23–3.38), and higher TNM stage (OR = 3.32, 95% CI = 1.99–5.55). In UM, BAP1 mutation was significantly associated with older age (OR = 1.70, 95% CI = 1.04–2.78), ciliary body involvement (OR = 2.50, 95% CI = 1.41–4.44), histological epithelial phenotype (OR = 3.79, 95% CI = 1.98–7.27), chromosome 3p loss (OR = 45.59, 95% CI = 21.16–98.22), chromosome 8q gain (OR = 9.40, 95% CI = 3.90–22.64), metastasis (OR = 4.65, 95% CI = 2.70–8.01), and higher TNM stage (OR = 2.67, 95% CI = 1.22–5.88). In MPM and CCA, however, BAP1 mutation did not correlate with these critical clinicopathologiocal factors, but with some etiological factors, including hepatitis infection in CCA (OR = 2.83, 95% CI = 1.10–7.29) and smoking in MPM (OR = 2.45, 95% CI = 1.39–4.30), indicating that BAP1 mutation was probably associated with certain etiological subtypes in these two cancer types.
Association of BAP1 mutation and clinicopathological features in different cancer types.

CCA: cholangiocarcinoma; CCRCC: clear cell renal cell carcinoma; MPM: malignant pleural mesothelioma; UM: uveal melanoma; FEM: fixed-effects model; REM: random-effects model; OR: odds ratio; CI: confidence interval.
Parameters with statistic significance were emphasized in bold.
All these results revealed that BAP1 mutation was significantly correlated with indicators of poor prognosis in UM and CCRCC. This partially explained why BAP1 mutation has prognostic values only in UM and CCRCC, but not in MPM and CCA.
Prognostic stratification based on the unique antagonistic role of BAP1 mutation with other mutations
In order to have an in-depth insight into the genetic profile underlying BAP1-associated cancer types, we further analyzed the relationship between BAP1 mutation and other most commonly mutated genes. The most frequently mutated genes in these four cancer types were determined according to COSMIC database (Figure S2). In UM, GNAQ, GNA11, BAP1, SF3B1, and EIF1AX were the most commonly mutated genes, and BAP1 mutation was negatively correlated with SF3B1 mutation (OR = 0.11, 95% CI = 0.04–0.33) and EIF1AX mutation (OR = 0.11, 95% CI = 0.04–0.30). In CCRCC, VHL, PBRM1, SETD2, and BAP1 were the most commonly mutated genes, and BAP1 mutation was negatively correlated with PBRM1 mutation (OR = 0.28, 95% CI = 0.17–0.48). In MPM and CCA, no significant association was detected between BAP1 mutation and other commonly mutated genes, indicating a more complicated genetic heterogeneity within these two malignancies (Table 2).
The above results also shed light on potential mutation-defined taxonomy in CCRCC and UM, which may further stratify patients into different prognostic subtypes. In CCRCC, BAP1 mutations and PBRM1 mutations may define two distinct molecular subtypes. Of the 568 CCRCC patients in the pooled cohort (TCGA cohort 38 and Sato cohort 39 ) included in our analysis, 52 had BAP1-mutant tumors (including six tumors with mutations in both BAP1 and PBRM1) and 167 tumors with mutations exclusively in PBRM1 (Figure 4(a)). Using the Kaplan–Meier model and log rank test, we found that BAP1-mutant CCRCCs showed significantly worse OS than PBRM1-mutant CCRCCs (p = 0.001, Figure 4(b)). In UM, BAP1 mutation was mutually exclusive with SF3B1 and EIF1AX mutations (Table 2 and Figure 5(a)). We used a cohort (n = 80) of UM patients from TCGA (the survival data and gene mutation status were downloaded from the cBioPortal (http://www.cbioportal.org/) 40 ) for further prognostic analysis. Consistently, the Kaplan–Meier model and log rank test demonstrated that BAP1-mutant UMs have significantly worse OS than those with SF3B1/EIF1AX mutations (p = 0.028, Figure 5(b)).
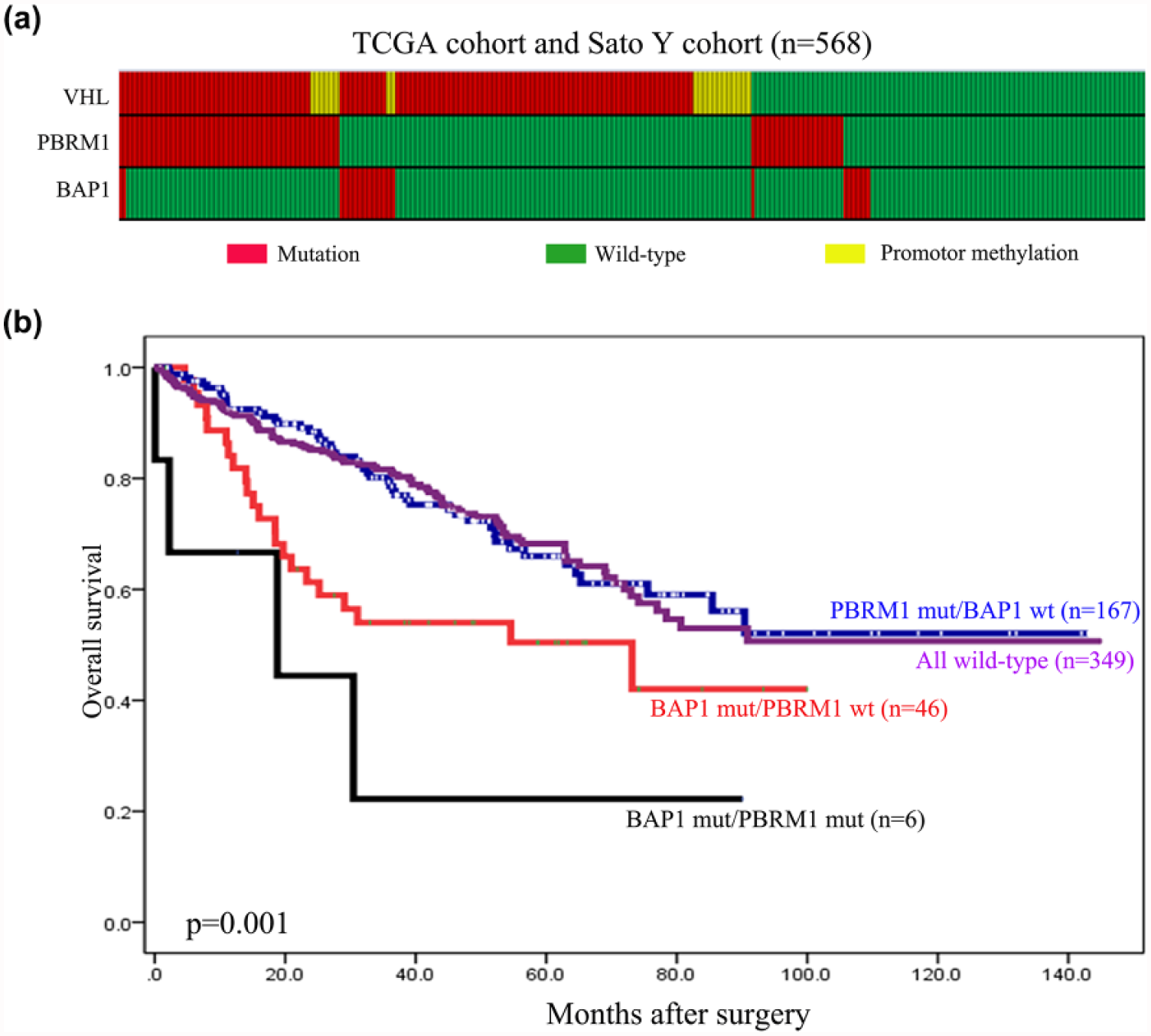
Prognostic stratification based on the mutation-defined molecular taxonomy in CCRCC. (a) The distributing pattern of VHL, PBRM1, and BAP1 mutations in the pooled cohort of CCRCC from the TCGA and Sato studies. (b) Distinct prognosis between BAP1-mutant CCRCCs and PBRM1-mutant CCRCCs by Kaplan–Meier model and log rank test.

Prognostic stratification based on the mutation-defined molecular taxonomy in UM. (a) The distributing pattern of GNAQ/11, SF3B1, EIF1AX, and BAP1 mutations in the cohort of UM from the TCGA study (unpublished) and (b) distinct prognosis between BAP1-mutant UMs and SF3B1/EIF1AX-mutant UMs by Kaplan–Meier model and log rank test.
Publication bias
Begg’s funnel plot and Egger’s linear regression test were used to assess publication bias. Begg’s funnel plot for both OS (p = 0.198, Figure S3A) and DFS (p = 0.246, Figure S3B) revealed no obvious bias. Further confirmation using Egger’s regression test also failed to find evidence of a publication bias for OS (p = 0.104, Figure S3C) and DFS (p = 0.057, Figure S3D).
Discussion
Although the role of BAP1 mutation in tumor development is emerging in recent years, its prognostic significance was not well elucidated. According to the data from COSMIC and cBioPortal, BAP1 was exclusively mutated in UM, MPM, CCRCC, and CCA (Figure S1). Here, we found that BAP1 mutation was indeed correlated with poorer OS and DFS in cancer patients. However, cancer type specific analysis revealed that BAP1 mutation was only associated with worse prognosis and clinicopathological features in UM and CCRCC, but not in MPM and CCA. Moreover, by assessing the mutual exclusivity of BAP1 mutation with other commonly mutated genes, we found BAP1-mutant tumors displayed significantly poorer prognosis than other mutational subtypes in UM and CCRCC.
Frequent mutations of BAP1 were first identified in familial and sporadic UM cases.18,20 Harbour et al. first found that BAP1 mutation was significantly correlated with metastasis and class 2 gene expression profiles suggestive of more aggressive clinical subtype. 20 Here, we found that BAP1 mutation was indeed associated with poor prognosis (both OS and DFS) in UM. Moreover, BAP1 mutation was significantly associated with multiple critical clinicopathological factors in UM, such as older age, ciliary body involvement, histological epithelial phenotype, distant metastasis, and higher clinical stage. In addition, BAP1 mutation was also positively correlated with chromosome 3p loss and 8q gain, indicating an important role of chromosome aberration mechanism in the development of BAP1-mutant UMs. Most of these clinicopathological and molecular features were reported to be important prognostic parameters in UM, indicating that BAP1 mutation status defined a unique subtype of UM with aggressive clinical manifestation and poor prognosis.
Germline or somatic mutations of BAP1 were also frequently found in CCRCCs, and BAP1 mutation was reported to define a unique class of CCRCCs.17,22 We found that BAP1 mutation was also associated with poor prognosis (both OS and PFS) in CCRCC. Moreover, BAP1 mutation was also significantly associated with critical clinicopathological factors, such as larger tumor size, higher Fuhrman grade, sarcomatoid differentiation, distant metastasis, and higher clinical stage.
Similar to UM and CCRCC, germline or somatic mutations of BAP1 were also identified in MPM and CCA.16,21,23 However, BAP1 mutation has no impact on OS or PFS and did not correlate with critical factors in these two cancers. Instead, BAP1 mutation was correlated with some etiological factors, including smoking in MPM and viral hepatitis (HBV or HCV) infection in CCA. Asbestos exposure is the most important risk factor for MPM, 41 and germline mutation of BAP1 indeed accelerates development of asbestos-induced malignant mesothelioma in animal models. We found that BAP1 mutation was not significantly correlated with asbestos exposure, but with smoking history, and this point has already been presented in the previous meta-analysis by Luchini et al. 33 Viral hepatitis infection is a critical risk factor for intrahepatic cholangiocarcinoma (ICC) especially in East Asian countries, 42 and our results suggested that BAP1 mutation was positively correlated with HBV/HCV infection in CCA. Two of the three enrolled studies were from Asian countries and one was from the United States (TCGA provisional), indicating the universality of this correlation among different regions. Moreover, both Begg’s funnel plot (p = 0.296) and Egger’s regression test (p = 0.428) revealed no obvious bias for this result. To our knowledge, this is the first report of the potential association between BAP1 mutation and viral hepatitis infection in CCA. Considering the relatively small patient cohort, larger prospective studies are needed for the relationship between BAP1 mutation and etiological factors in MPM and CCA.
As BAP1 mutation showed tissue-specific prognostic significance and correlated with critical clinicopathological features especially in UM and CCRCC, we further analyzed the relationship of BAP1 mutation with other most commonly mutated genes in these cancers. In UM, hotspot mutations of G protein subunits GNAQ and GNA11 (GNAQ/11 mutations) account for nearly 90% of UM cases. Their mutations were predicted to be early and initiating events in UM tumorigenesis but are not sufficient for malignant transformation. 43 Our results revealed that BAP1 mutations and SF3B1/EIF1AX mutations tend to be mutually exclusive, indicating that they may represent alternative pathways in tumor progression. Moreover, compared with SF3B1/EIF1AX-mutant UMs, BAP1-mutant UMs have significant worse prognosis. Similarly, in CCRCC, VHL inactivation is an early event and accounts for 70%–90% of CCRCC, but VHL loss did not correlate with patient outcome. 38 Instead, BAP1 mutations were mutually exclusive with PBRM1 mutations, and BAP1 mutations predicted poorer prognosis than PBRM1 mutations, indicating that these two gene mutations define two different subtypes of CCRCC initially driven by VHL inactivation. In MPM and CCA, no significant correlated gene mutations were found, indicating a more complicated genetic profile feature in these two malignancies.
Accumulating studies in recent years have uncovered a novel link between chromatin remodeling and tumor suppression, as recurrent mutations associated with chromatin remodeling have been identified in various cancers. Notably, genes encoding components of the SWItch/Sucrose non-fermentable (SWI/SNF) complexes (including ARID1A, PBRM1, ARID2, SMARCB1, and SMARCA4 genes) were the most prevalent. 44 ARID1A is the most frequently mutated SWI/SNF subunit in cancers, and recent studies have revealed that ARID1A was associated with poor prognosis in various cancer types.23,45,46 Mechanistically, SWI/SNF complexes remodel nucleosome structure and are capable of mobilizing nucleosomes both by sliding and by catalyzing the ejection and insertion of histone octamers in an ATP-dependent manner. 47 Functionally, BAP1 was also found to be tightly associated with chromatin remodeling, while the underlying mechanism seems to be quite different. BAP1 interacts with ASXL1 to form a polycomb group repressive deubiquitinase (PR-DUB) complex, which can specifically deubiquitinate 119 lysine of histone 2A (H2AK119) and antagonize the ubiquitinating effect of polycomb repressive complex 1 (PRC1) complex.4,48 In addition, BAP1 also exhibited multifaceted functions by directly deubiquitinating and stabilizing other proteins involved in DNA damage repair1,6,7,9 and transcription regulation.2,3,49 All these evidences suggested that the oncogenic mechanism of BAP1 mutation is different from that of SWI/SNF subunit mutations. Consistently, recurrent BAP1 mutation occurred only in certain types of cancer, while SWI/SNF subunits genes were extensively mutated in nearly all human cancer types.
Our analysis also has several limitations. First, as a novel tumor suppressor gene whose somatic mutations were only identified in recent years, articles’ availability was somewhat lacking, and further validation was needed especially in CCA and MPM. Second, the method of BAP1 mutation detection is diversified in these included studies, varying from SS, AAS, MLPA, IMPACT, NGS, WGS, and WES. This diversification in mutation detection methods may generate certain heterogeneity of the results. Third, this study only included BAP1 mutations but not BAP1 expression. Whether BAP1 expression detection is superior to BAP1 mutation in prognostic prediction also needs further investigations.
Conclusion
In conclusion, our study revealed the tissue-specific prognostic role of BAP1 mutation, which has practical implications for the clinical prognostic prediction in BAP1-associated cancers. Moreover, based on the integrated gene mutation profiles, we suggested that combining detection of BAP1 mutation and other driver mutations may further shed light on clinical stratification in BAP1-associated cancers.
Footnotes
Acknowledgements
X.-Y.W. and Z.W. contributed equally to this work. L.-X.Q., W.-W.Z., and D.Y. conceived and designed the study; X.-Y.W., Z.W., X.-D.R., and J.-B.H. performed the research; X.-Y.W., Z.W., X.-D.R., and J.-B.H. extracted the data from literature; X.-Y.W. and Z.W. performed data analysis and data interpretation; X.-Y.W. drafted the manuscript; W.-W.Z. and D.Y. helped edit the manuscript. All authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Natural Science Foundation of China (NSFC) General Program, No. 81472677; the NSFC General Program, No. 81522033; the NSFC Program of International Cooperation and Exchanges, No. 81120108016; and the Shanghai “Phosphor” Science Foundation, China, No. 14QA1400600.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
