Abstract
Evidence is accumulating in estimating the potential role of human papillomavirus infection in prostate carcinogenesis. However, the results remain inconclusive. We measured the serostatus of antibodies to one of the high-risk human papillomaviruses, human papillomavirus 16, with a newly developed peptide microarray. Serum samples were collected from 75 untreated prostate cancer patients, along with 80 control subjects. We identified 12 peptides with significant differences in prostate cancer samples from all 241 peptides derived from human papillomavirus 16. Our results showed human papillomavirus 16 infection in 64.0% of prostate cancer serum samples, which is significantly different compared with the controls (p < 0.01) because only 17.5% of the control serum was considered seropositive. The area under the receiver operator characteristic curve was 0.793 (95% confidence interval 0.721–0.864), indicating that the new microarray technique may have diagnostic value. The results showed an association between serological evidence for human papillomavirus 16 infection and risk of prostate cancer. The different serostatus of antibodies in the two subgroups indicated that human papillomavirus 16 infection might occur and play a potential role of progression in a minority of prostate cancer.
Introduction
Prostate cancer is among the most common urological malignancies in men. The exact mechanisms underlying cancer development in the prostate gland have not been well characterized. Over the past few decades, studies have found positive associations of prostate cancer with sexual activity and sexually transmitted infections.1–6 Furthermore, substantial studies on epidemiological research suggested that several viruses might have the oncogenic potential in prostate cancer progression, such as human papillomaviruses (HPVs), polyomaviruses (SV40), and members of the herpes virus family.7–12 HPV, an important infectious agent, contains two genes E6 and E7 that encode oncogenic proteins that can interact with and inhibit the tumor suppressor proteins Rb and p53, respectively. 13 Previous studies showed that the E6 and E7 oncoproteins of HPV types 16 or 18 can immortalize prostate epithelial cells.14,15 It has been hypothesized that the prostate gland can be infected because of anatomic proximity of direct exposure to HPV particles through urethra, as penile cancer and urethral HPV lesions have also been described.16,17
HPV types 16 and 18 are present in the normal and cancer tissues of the human prostate. 18 A series of laboratory studies reported that HPV was detectable in prostate cancer cells, as shown by Southern blot and polymerase chain reaction (PCR) analysis,19–24 although other investigations have not reproduced this finding.25,26 These inconsistent results may be attributed to different sensitivities of the various assays for detection of HPV DNA. 27 Serological studies, however, offer an alternative method for the detection of HPV in both cases and controls and can provide an integrated, readily standardized measurement of long-term HPV exposure to biological samples. However, the divergent findings reported from serological studies also indicate that the link between prostate cancer and HPV infections is still open to debate.2,28–30
The past few decades have witnessed a fascinating growth in the microarray technology, which has become a crucial tool for large-scale and high-throughput biology, resulting in a new era of the collection and analysis of information. We previously developed a peptide microarray with a novel solid supporting material, polymer-coated initiator integrated poly(dimethylsiloxane) (iPDMS) or SJ membranes 31 (iPDMS was later introduced by Suzhou SJ Biomaterials Co. Ltd.), which could specifically interact with autoantibodies (AAbs) in serum. 32 This newly developed technique could achieve an absolute zero background for large-scale and high-throughput seroscreening in a short period of time with a small amount of serum samples. HPV-16 has a unique association with prostate cancer both in DNA detection 23 and serological screening. 2 To verify the hypothesis that HPV-16 may act as an antigen in prostate cancer patients, we conducted an enzyme linked immunosorbent assay (ELISA)–based microarray with HPV-16 peptides synthesized and spotted onto SJ membranes as probes to immobilize HPV-16 antibodies. This study is designed to investigate the different antibody serostatus against HPV-16 of prostate cancer cases in comparison with controls, taking into account its correlation, if any, with various risk factors, such as mean age and Gleason scores.
Materials and methods
Patients and serum samples
A total number of 155 blood samples were analyzed in this study. Seventy-five prostate cancer serum samples were collected from the First Affiliated Hospital of Soochow University (Suzhou, China). All patients had been pathologically diagnosed for the first time and were not receiving any antitumor treatments. Eighty volunteers served as controls and were obtained from the same hospital. All controls underwent diagnostic testing through ultrasound-guided prostate biopsy for reasons such as high prostate-specific antigen (PSA) levels or suspicious digital rectal examination (DRE). These patients were eligible as controls because all their biopsy specimens showed no evidence of cancer. Detailed subject information is given in Table 1. All samples were kept frozen at −20 °C prior to analysis. The study was approved by the ethics committee of the First Affiliated Hospital of Soochow University. Informed consent was obtained from all participants in the study. All the experiments were performed in accordance with the approved guidelines.
Characteristics of the serum samples included in the study.

PSA: prostate-specific antigen.
Preparation of peptide microarray
The microarray was prepared in a 100,000 grade clean room. HPV-16 protein candidates were selected and translated into a peptide library. Experimental peptides (20 mers), synthesized by GL Biochem (Shanghai, China), were first dissolved in 30% acetonitrile solution (v/v, in Milli-Q water) to 1 mg/mL and then diluted to 200 µg/mL with printing buffer (0.3 M PB, 0.2% glycerin, 0.01% Triton, and 1.5% mannitol) for further printing. Using the contact printer Smart 48 (Capitalbio, Beijing, China), synthetic peptides were spotted onto iPDMS to form a 9 × 9 × 6 array. Each sub-array included a positive control with human IgG (DGCS-Bio, Beijing, China) at the concentration of 50 µg/mL and a negative control with PB.
Serological screening
HPV-16 is a non-enveloped double-stranded DNA virus and encodes seven functional proteins named E1, E2, E4, E6, E7, L1, and L2. Based on the function of these proteins and a novel method for predicting linear B-cell epitopes, 33 we assumed that epitopes derived from seven HPV-16 antigen proteins were immunodominant and could interact with antibodies. The amino acid sequence information for all seven proteins were obtained from the National Center for Biotechnology Information (NCBI) website. Antigen sequences were then cut into 20-mer peptides containing overlaps of 10 amino acids, after having been printed onto iPDMS, resulting in a total of 241 HPV-16 peptide microarrays. Serum samples were first diluted 1:100 in serum-dilution buffer, and 200 µL was included in each microarray for incubation for 2 h on the shaker (150 r/min, 4 °C). The microarray was then rinsed three times with Tris buffered saline with Tween 20 (TBST) (20 mM Tris–HCl, pH 6.8, 137 mM NaCl, 0.1% Tween 20) and incubated with 200 µL horseradish peroxidase (HRP)–labeled goat anti-human IgG (ZSGB-Bio, Beijing, China) at a 1:25,000 dilution with peroxidase conjugate stabilizer/diluent (Thermo Fisher, Waltham, MA, USA) for another 1 h at the same temperature, followed by the same washing steps as described above. Chemiluminescence substrate (15 µL) was added to the microarray, and chemiluminescence signals were captured by a charge coupled device (CCD) camera using a LAS4000 imaging system (GE, USA). Signals were finally saved as images in TIFF format and then processed using GenePix Pro 6.0 software to calculate the median chemiluminescence intensity of each peptide spot and the local background (i.e. the area surrounding a spot) at a 635 nm wavelength. The digitized image from each microarray was then imported into R for further analysis.
Statistical analysis
Statistical analyses were performed by R version 3.1.0. Chemiluminescence intensity was converted to signal-to-noise ratio (SNR) for further analysis: SNR = (signal intensity − background)/background. A “pheatmap” package was used for the cluster analysis and heatmap performing. Significant analysis of microarray (SAM) was performed using “samr” package to identify differential expression between the two groups. The receiver operator characteristic curve (ROCC) and chi-square test were used when applicable, and a p <0.05 was considered statistically significant.
Results
Comparison of antibody profiles of serum samples
With the use of microarray containing 241 overlapped peptides, 155 serum samples (75 prostate cancer subjects and 80 control samples) were included in the serological search. The workflow of the seroscreening is shown in Figure 1. Antibody binding to individual peptides was identified and converted to the SNR, resulting in a serum/peptide original data. The unique feature of the iPDMS membrane could provide a near-“zero” background for serological assays, making the acquisition of data and subsequent analysis simple and easy. Reactions with SNR ≥2 were identified as seropositive. Specific antibody–epitope interactions performed as high signal intensity region and methods should be taken in identifying these peptides regarding the specific area for further diagnostic use. Here, we used “three-mode analysis,” which was introduced in our previous study, 32 to facilitate the process in identifying these specific peptides. Coverage of all 241 peptides in prostate cancer samples and the controls was shown with the three-mode analysis (Figure 2). Only 15 of 241 peptides, indicated by arrows in Figure 2(a), had a coverage greater than 20% in prostate cancer samples. The three-mode analysis gave a direct visualization of the result of all the peptides responding to the sera samples. However, not all the high responsive peptides showed significant difference between the two groups. SAM, once used in DNA microarray analysis, has its shortcomings in performing direct analysis of such a large-scale original data. However, SAM was successfully used here in extracting peptides that showed a statistically significant difference among serum groups of the peptide microarrays. With the data of SAM, 12 peptides were identified as statistically significantly higher responding peptides in prostate cancer samples (Figure 3(a); p < 0.001). Meanwhile, the 12 peptides and their coverage in two groups are shown in Figure 3(b). SNR distributions of these 12 peptides responding to prostate cancer and controls also showed a significant difference between the two sample groups (Figure 3(c)). The heatmap of responses of 155 serum samples (from 75 prostate cancer patients and 80 control samples) to the identified 12 peptides is displayed in Figure 4. All SNR values have been subtracted by 2 to get the SNR′ (SNR′ = SNR − 2), and reactions of SNR′ <0 were converted to 0. The responses were obtained and clustered by SNR′. Responses of SNR′ ≥0 in the heatmap were regarded as seropositive responses. The heatmap showed that we could easily distinguish prostate cancer samples from healthy samples using these 12 peptides.

Brief illustration of seroscreening process of HPV 16 peptides. 1–4: Preparation of peptide microarray. Seven proteins were selected from HPV 16 and translated into 20/10aa overlapped peptides (1), chemically synthesized (2), and then spotted onto the iPDMS (3) to form a microarray (4). 5 and 6: Two steps of incubation with serum samples (5) and enzyme-labeled second antibody (6). 7 and 8: Antibody binding to target peptide is then acquired (7) and processed to chemiluminescence immunoassay picture by CCD camera for further analysis (8).

The line/area chart was conducted by the three-mode analysis of (a) prostate cancer and (b) control samples. The area chart (gray area) indicates the total coverage of the three modes. An epitope containing peptides (ECPs) is defined as a cutoff value of coverage greater than 20%, as shown by the dashed lines in the figure. Peaks above the dashed line indicated the location of the ECPs. Only 15 of 241 peptides were identified as ECPs using the three-mode analysis.

A total of 12 peptides were identified as highly responsive peptides in prostate cancer group. (a) SAM analysis was used to find the peptides that have significant difference between prostate cancer group and controls. Twelve peptides with significant different SNRs, as shown in red circles, were selected as highly responsive peptides in prostate cancers rather than the controls. Red circles: 12 newly identified peptides. Black circles: the rest of 241 peptides. (b) The 12 newly identified peptides and their responding coverage against prostate cancer and the control samples. (c) SNR distribution of these 12 peptides responding to the prostate cancer and healthy samples. PC: prostate cancer. HC: control samples. Red circles: the 12 newly identified peptides. Black circles: the rest of 241 peptides.

Heatmap of responses of 155 serum samples (75 prostate cancer patients and 80 control samples) to the identified 12 peptides. Reactions of SNR ≥2 were identified as seropositive reactions. All the SNR values have been subtracted by 2 to get the SNR′ (SNR′ = SNR − 2), and reactions of SNR′ <0 were converted to 0. The responses were obtained and clustered by SNR′. Responses of SNR′ ≥0 in the heatmap were regarded as seropositive responses. We could easily identify the differences between the two groups in the serological screening of HPV peptides from the heatmap.
Diagnostic value of the HPV-16 epitope combination
The 12 peptides were finally identified as diagnostic candidates. To assess the predictive value of these 12 peptides, we first defined the universal binary cutoff value of SNR = 2 as a standard value and each peptide with an SNR ≥2 might indicate a positive result. Because the mean SNR of the blank dots (negative control dots printed with buffer) was 0.5 ± 0.3, the value of (mean SNR + 3 standard deviation (SD)), approximately 1.4, was defined seropositive; we chose 2 for redundancy. The previous study with a traditional strategy of multiplexing 34 suggests that any one of the 12 peptides being positive indicates a positive seroreaction. However, the antibodies in serum are so intricate that it could cause mimotopes or molecular mimics during the seroscreening. Furthermore, immune diversity also leads to contradictions between sensitivity and specificity, 35 making diagnosis unclear. None of the 12 peptides could provide an increased sensitivity with a satisfactory specificity (Table 2). We believe that this contradiction between sensitivity and specificity has hindered the realization of the long-predicted diagnostic value of epitopes. A new strategy, binarization/digitization, which has been described before, 32 successfully solved these problems and enabled the 12 peptides to achieve the diagnostic function. Using this strategy, the 12 combinations performed a sensitivity of 64.0% in prostate cancer with a 82.5% specificity when compared with the controls, with the area under the ROCC of 0.793 (95% confidence interval 0.721–0.864). Our data was in accordance with the PSA levels since we also witnessed a sensitivity of 64.0% in prostate cancer by PSA, with the area under the ROCC being 0.748 (95% confidence interval 0.668–0.828; Figure 5). Significant differences were observed between prostate cancer patients and controls (p < 0.001). Furthermore, we also witnessed a significant proportion of HPV-infected cases with high Gleason score >7 (p = 0.014).
Coverage ratio of the 12 identified peptides in the two groups.
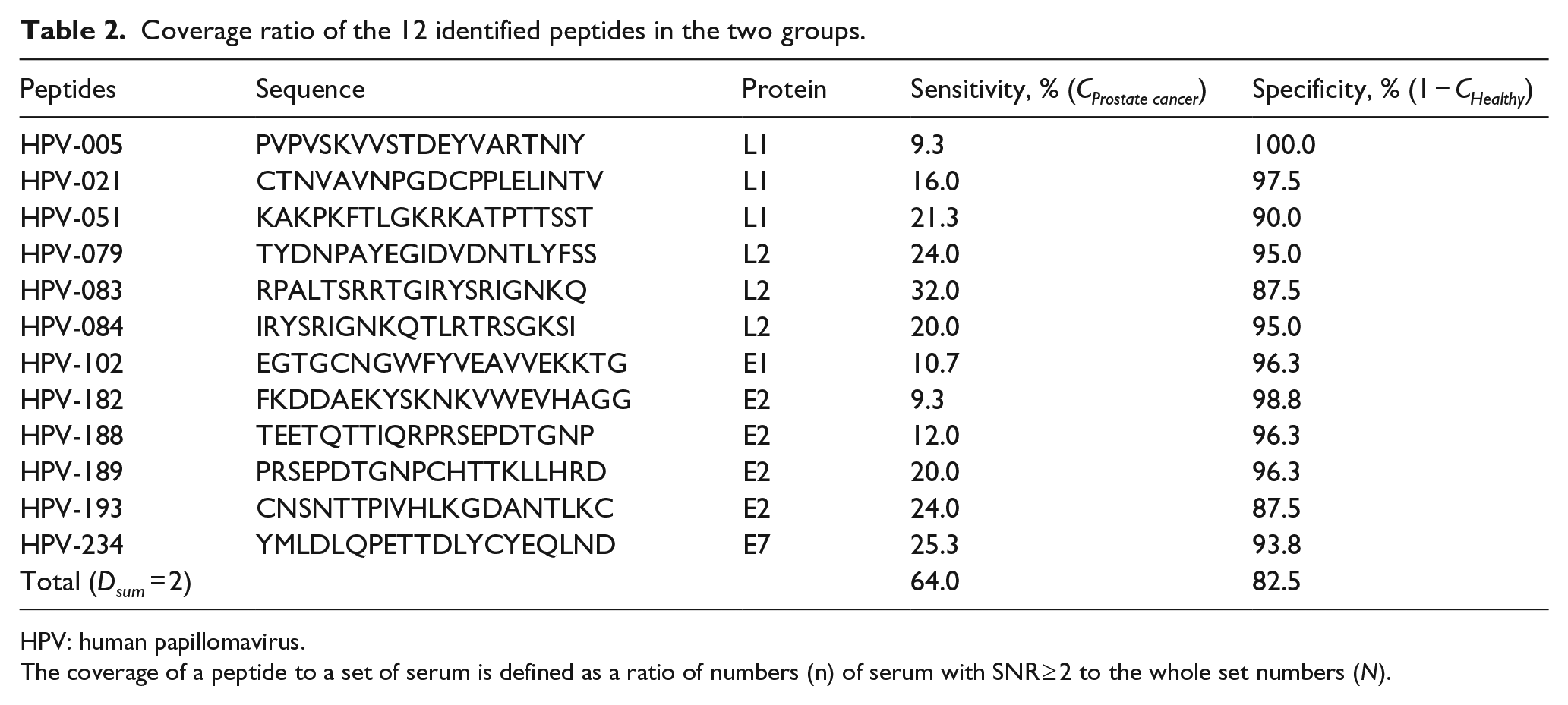
HPV: human papillomavirus.
The coverage of a peptide to a set of serum is defined as a ratio of numbers (n) of serum with SNR ≥ 2 to the whole set numbers (N).

ROC analysis between the (a) identified 12 peptides and (b) PSA. The combination of the newly identified 12 peptides was more efficient at discriminating prostate cancer patients from the controls than the use of PSA, with the area under the curve to 0.793 and 0.748, respectively.
Discussion
In recent years, together with the development of high-throughput detection platforms such as phage display and protein microarray, the tumor-associated AAbs have been identified and served as minimally invasive diagnostic tools in cancer. In particular, the protein microarray format allows rapid, easy, and economic method to detect all relevant diagnostic parameters in a single experiment. However, every coin has two sides, and both genetic and immunogenic variations in humans could cause the false-negative problems for such antigen-based immunoassays. Motivated by the intrinsic limitations of protein biomarkers, epitopes, as the antibody recognition region of the antigen, were proposed and realized for detection. Our previous study found that the complexity of antibodies in the serum could cause interference due to the mimotopes or molecular mimics, suggesting that one epitope being positive indicates that a positive seroreaction was not reliable and might generate false-positive or false-negative results. We believed an enhanced reliability if one serum could specifically interact with two peptides within one protein during the same screening, that is “Counts.” Obviously, the results seemed more reliable as far as we obtained enhanced Counts. However, in most cases, the linear epitopes within one protein are limited due to the variation in the immunogenicity and the differences in size of the related protein, making the value of Counts limited. Similarly, if one serum could response to different peptides from multiple proteins of one virus, the obtained results could be more reliable. This is the reason we selected 241 peptides from seven proteins of HPV-16 to form an epitope-based microarray in the seroscreening. Using the combination of the three-mode analysis and SAM analysis, we rapidly isolated 12 highly responsive peptides. The newly applied method, binarization/digitization, which delivers a combination of epitopes for the serodiagnosis, successfully helped us to judge the various differences in antibodies between prostate cancer patients and the control subjects.
Previous studies showed the oncogenic potential of high-risk HPVs and their role as a vital etiologic factor in the progression of prostate cancer,14,15 especially the immortalization of prostate epithelial cells with E6 and E7 proteins, which provided a molecular clue for the positive linkage between HPV infection and prostate cancer. This study demonstrated that the antibody serostatus in two groups was significantly different, with many peptides strongly recognized in prostate cancer (64.0%) but only weakly reacted in healthy individuals. Our data were different from a previous study carried out in India that showed the presence of HPV-16 in 32% of prostate cancers. 23 This divergent results might be attributed to sample contamination or PCR condition variance, 27 and to some extent, failing to obtain the suitable samples does not definitely exclude a role for HPV infection in prostate cancer. Unlike the detection of HPV DNA, which could only determine the current infection status, the identification of HPV-related immune response in the serum, as this study showed, indicates both past and present infection and overcomes many limitations. However, serological searching also has its shortcomings. The HPV prevalence in sera did not well correspond to that in tissues of some subgroups because the antibody might be elicited by HPV infection from anatomical sites other than prostate, which could cause the same positive result. The complex link between seropositivity and historical infections and the current disease status could disturb the human immune response in the serum. Thus, detection of HPV DNA and seroscreening both have their merits and demerits. Recently, a meta-analysis showed that the overall HPV prevalence in prostate cancer cases was highest in Africa, 24 followed by Asia, which means the risk estimates varied by study region. This is also similar to our studies, since China and India are two of the biggest countries with the largest populations in Asia.
The ROCC, used to assess the predictive value of the selected peptides, suggested that the 12 combinations were most efficient in discriminating prostate cancer from the healthy controls. The combined model increased the area under the ROCC to 0.793 (95% confidence interval 0.721–0.864), which was in accordance with PSA, indicating that the newly developed microarray technique may have diagnostic value. Despite the limited sensitivity of this study, we also witnessed a higher specificity of 82.5% than the 78.7% of PSA. The high specificity to the PSA might play a subsidiary role in decreasing the false-positive rate in the precancerous diagnosis. Since PSA is organ-specific but not cancer-specific, for the reasons that serum levels may be elevated in the presence of benign prostatic hypertrophy (BPH), prostatitis, and other non-malignant conditions, producing false-positive results and contributing to the unnecessary biopsies. Furthermore, a highly significant correlation was found between HPV-16 infection and the high Gleason scores, implying that HPV-16 might occur and influence the potential role of progression during prostate carcinogenesis.
Together, our study demonstrated that infection with oncogenic HPV-16 might be involved in the etiology of a minority of prostate cancers and we did find the higher antibody serostatus. An important strength of this study is that the epitope-based microarray is a novel method and used in investigating the specific HPV-16-related antibodies in Asian countries such as China. Although the clinical relevance of HPV infection and prostate carcinogenesis is still a matter of debate, large-scale etiological research is needed in finding the causal involvement of HPV infection in prostate cancer.
Footnotes
Acknowledgements
X.Z. and Z.Z. contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
