Abstract
Breast carcinogenesis is a multistep process, involving both genetic and epigenetic modification process of genes, involved in diverse pathways ranging from DNA repair to metabolic processes. This study was undertaken to assess the role of promoter methylation of GSTP1 gene, a member of glutathione-S-transferase family of enzymes, in relation to its expression, polymorphism, and clinicopathological parameters. Tissue samples were taken from breast cancer patients and paired with their normal adjacent tissues. A total of 51 subjects were studied, in which the frequency of promoter methylation in cancerous tissue was 37.25% as against 11% in the normal tissues (
Keywords
Introduction
Belonging to a heterogeneous group of diseases, breast cancer is characterized by the disordered cellular programming, wherein cells loose the normal growth control regulation. It ranks as the leading cancer in females worldwide, including India, where 41.3% of all cancer types in females account for breast and cervical cancers. 1 Similar trend is observed in Kashmir also, with 1797 cases reported between 2000 and 2013, accounting for 16.83% of all the cancers among women. 2 Over the past decade or two, significant progress has been made in determining the risk factors for the breast cancer. However, the established risk factors including the mutations in BRCA1 and BRCA2 genes account only for 27% of the breast cancer risk. 3 More recently, focus has been shifted to the second piece of the puzzle, the epigenetic causes. 4 These epigenetic modifications, usually not inherited, change the way in which the gene functions and are mediated by the number of interconnected molecular mechanisms like histone modifications, polycomb/bithorax protein complexes, small non-coding or anti-sense RNAs, imprinting of genome, and methylation of the DNA bases. 5 DNA methylation, the most common epigenetic modification commonly occurring in the CpG dinucleotide clustered into CpG islands near the promoter region of the most genes, is characterized by the enzyme-driven addition of the methyl group to the 5-carbon of the cytosine base 6 in the otherwise hypomethylated promoters in the normal cells; DNA methyl transferases (e.g. DNMT1, DNMT3a, and DNMT3b) are the enzymes that catalyze the covalent addition. 7 This hypermethylation event is associated with silencing the transcription of the many cancer critical genes, either by interfering with the transcription activation or by remodeling the chromatin structure.8,9
Glutathione-S-transferase (GST) family of enzymes, of which GSTP1 is a member, is the phase-II xenobiotic metabolizing enzymes known to play a significant role in the detoxification of various endogenous as well as exogenous xenobiotics, including carcinogens, by their conjugation and subsequent reduction to reduced glutathione. 10 Located on chromosome 11q3, GSTP1 has been reported to play a role in the regulation of cell signaling by interacting with various signaling proteins like c-Jun NH2 terminal kinase, AJK-1, and TRAF-2, thereby regulating their downstream effects. 11 Epigenetic silencing of the GSTP1 gene mediated by promoter hypermethylation and subsequent loss of protein expression 12 has been reported in many human cancers like human neoplasia, 13 prostate cancer, 14 and breast cancer. 15 Hypermethylation of the CpG dinucleotide in the GSTP1 promoter has further been associated with aggressive phenotype 16 and poor prognosis. 17
Reduced gene activity has also been seen in the mutant alleles of the GSTP1 genotype. The single-nucleotide polymorphism in codon 105 of the exon 5 results in the A to G transition at nucleotide position 313, with the subsequent substitution of isoleucine with valine in the enzyme product. 18 Individuals with the mutant GSTP1 genotype have been reported to show reduced reaction kinetics, substrate specificity, catalytic efficiencies, and conjugation abilities for its substrate.19,20
Recent evidences have shown that the hypermethylation profile of the cancer critical genes depends on the tumor type and is specific to the ethnic populations.21–23 To our knowledge, few studies of GSTP1 promoter methylation profile in relation to the breast cancer have been carried out on Indian population,24,25 and no such study has been carried out on the Kashmiri population which is ethnically different from the rest of India. The focus of this study was to examine DNA methylation status of the GSTP1 gene promoter and its effect on the expression of protein in patients with breast cancer. Furthermore, the gene promoter methylation profile and its protein expression have been statistically correlated with the GSTP1 genotype, to check the hypothesis whether the two events occur simultaneously. The hypermethylation, expression, and the genotype of the GSTP1 gene have also been studied in association with the various clinicopathological characteristics of the patient population.
Materials and methods
The following methodology was used to test the above-stated hypothesis.
Sample collection
Surgically removed and histopathologically confirmed tumor and normal tissues were collected from 51 breast cancer patients, who consented for the study during the period of 2014–2016, from Department of General Surgery, Sheri Kashmir Institute of Medical Sciences (SKIMS), Kashmir. The patients had no previous history of any type of cancer and had not received any chemotherapy. All the patients were females. The tissue samples were appropriately labeled and stored at −80 °C till further processing. Complete clinical data of the patients were collected. Staging of the breast cancer into four stages (I, II, III, and IV) was done applying tumor–node–metastasis (TNM) system of the American Joint Committee on Cancer/the Union Internationale Contre le Cancer (AJCC/UICC)as reference, 26 whereas the Scarff–Bloom–Richardson classification 27 was used to grade the tumors histopathologically as GI, GII, and GIII. The study was approved by the Ethical Committee of the SKIMS.
DNA isolation
Genomic DNA was isolated from the tissues (both tumor and normal) using phenol chloroform extraction. The genomic DNA was dissolved in the DNA storage buffer (Tris-EDTA), aliquoted, and stored at 4°C and −20°C. The purity of the isolated DNA was checked by measuring its absorbance at 260 and 280 nm, with the ratio of 1.8 taken to be pure. Also, the integrity of the isolated DNA was checked on 0.8% agarose gel and visualized by ethidium bromide staining.
Protein extraction
Total protein was extracted from all tissue samples using whole cell lysis buffer (50 mM KCl, 1% NP-40, 25 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES; pH 7.8)), supplemented with protease inhibitor cocktail (Sigma-Aldrich, USA). The extracted protein was quantified by Bradford assay, aliquoted, and stored at −80°C.
Bisulfite modification of DNA and methyl-specific polymerase chain reaction
Isolated genomic DNA (up to 2 µg) was subjected to bisulfite modification using EZ DNA-Methylation Kit (Zymo Research Corp, USA), according to the manufacturer’s protocol, which converted the unmethylated cytosines into uracil, whereas the methylated cytosines remained unchanged. The efficiency of the bisulfite modification was checked on 2.5% agarose gel and visualized by the ethidium bromide staining. Bisulfite-converted DNA was subjected to methyl-specific polymerase chain reaction (MS-PCR), using primers targeted for the promoter region of the GSTP1 gene. Primers were designed for the methylated and the unmethylated alleles of the promoter region based on the available literature. 28 The following primer pairs were used:

The DNA was amplified using thermal cycler (Mastercycler; Eppendorf, Germany) in 25 µL volume, containing 3 µL of bisulphate-converted DNA, 3.0 µL of 10 × PCR buffer, 0.5 µL of 10 pm of each primer, 0.5 µL of 10 mM deoxynucleotide triphosphate (dNTPs; Thermo Fisher Scientific, USA), and 1 unit of Taq polymerase (Sigma-Aldrich, USA). The amplification after the initial denaturation at 94°C for 5 min was carried out under the following set of conditions: denaturation at 94°C for 1 min, annealing at 66°C for methylated allele and at 63°C for unmethylated allele for 50 s, and extension at 72 °C for 1 min. The reaction was set for 40 cycles, followed by the final extension of 40 s at 72°C. Methylated control DNA (Sigma, USA) was used as the positive control, whereas DNA from whole blood was used as a control for the unmethylated reaction. For negative PCR control, water was used in both reactions.
Western blotting
Total protein extracted from the tissues after quantification and normalization was resolved on 12% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE), transferred to nitrocellulose membrane of pore size 0.2 µm, blocked with 5% bovine serum albumin (BSA) in 0.2% tris-buffered saline with Tween 20 (TBST), and then probed with polyclonal antibody against the GSTP1 gene (Cell Signaling Technology, USA), followed by the fluorescent-labeled anti-mouse secondary antibody (Cell Signaling Technology). For the loading control, membrane was also probed with the polyclonal antibody against α-tubulin (Cell Signaling Technology), followed by the fluorescent-labeled secondary antibody against it. The blots were visualized by imaging systems from LI-COR Biosciences.
GSTP1 polymorphism detection
Codon 105 of exon 5 of the GSTP1 gene was amplified using the following primer pair: forward primer: 5′-CAGTGACTGTGTGTTGATCA-3′ and reverse primer: 5′-TGCTCACATAGTTGGTGTAGATGAGGGATA-3′. The DNA was subjected to PCR amplification in 25 µL volume after the initial denaturation at 95°C for 5 min under the following reaction conditions: denaturation at 95°C for 30 s, annealing at 53°C for 30 s, and amplification at 72°C for 30 s. The reaction was set for 35 cycles, followed by the final extension at 72°C for 7 min.
The single-nucleotide polymorphism A → G transition occurring at 313th nucleotide, which results in the substitution of isoleucine with valine at codon 105, was detected by restriction digestion of the PCR product using 1U
Statistical analysis
Results were statistically analyzed by the specific statistical tests using SPSS software. Analysis of the association, if any, between GSTP1 polymorphism, methylation status, gene polymorphism, and expression as well as any correlation between clinicopathological parameters (lymph node, estrogen receptor (ER), progesterone receptor (PR), human epidermal growth factor receptor 2 (HER-2), tumor size, and age at diagnosis) of the patients with GSTP1 polymorphism, methylation status, and expression were examined by chi-square test. Fisher’s exact test and Pearson’s chi-square test were employed to analyze GSTP1 protein expression. Value of
Observations and results
GSTP1 promoter methylation
Of the 51 histopathologically confirmed cases, GSTP1 promoter was methylated in 19 samples (37.25%) and unmethylated in 32 tissues (62.7%). In the adjacent normal tissues of the same patients, promoter was methylated in 6 tissues (11%; Figure 1). Samples that showed methylation of the promoter in the normal tissue were also methylated in the tumor tissue. High frequency of the promoter methylation of GSTP1 gene was significantly associated (

Graphical distribution of methylated and unmethylated alleles in cases and control tissues.

Representative picture of MS-PCR assay.
GSTP1 expression using western blot
Of the 51 tumor tissues, 18 cases showed downregulation of the protein expression, whereas the expression was comparable for tumor and normal tissues in the remaining 33 cases. Of the 18 cases that showed protein expression loss, 12 (66.7%) were hypermethylated for the GSTP1 promoter and 6 (33.3%) were unmethylated for the GSTP1 promoter. Of the 33 cases that showed no loss of expression when compared to the normal adjacent tissue, 7 (21.2%) were methylated and 26 (78.2%) cases were unmethylated in the GSTP1 promoter region. The correlation between GSTP1 promoter methylation and the protein expression of the gene is shown in Table 1. Figure 3 shows the western blot analysis of the gene expression.
Correlation between GSTP1 promoter methylation and protein expression.
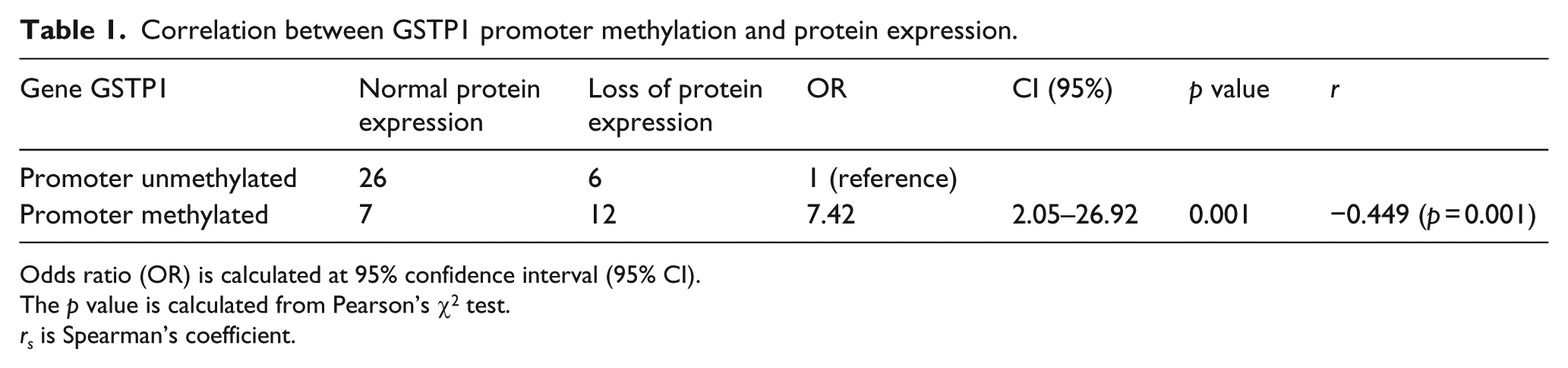
Odds ratio (OR) is calculated at 95% confidence interval (95% CI).
The

Western blot analysis showing the expression of GSTP1 protein using GSTP1 antibody and corresponding loading control (α-tubulin).
Clinicopathological findings
The study included 51 histologically confirmed breast tumor tissues and their adjacent corresponding normal tissues. The mean age of the patients was 50.137 ± 11.366 years, which for all statistical purposes was divided into two groups, with 58.82% (30 cases) in the age group of 50 years or above. The mean body mass index (BMI) of the patient population was 20.54 ± 1.21. The cases were also grouped on the basis of their dwelling (urban and rural) with 70.58% (36 cases) of the patients being rural. Anatomically, the tumor was divided into four stages (I, II, III, and IV) having the distribution of 17.64% (9 cases), 45.09% (25 cases), 27.45% (14 cases), and 5.8% (3 cases), respectively; but for all statistical purposes, stages III and IV were combined. The tumors were also assessed for the presence of ER, PR, and HER-2/neu receptor, and their distribution is shown in Table 2.
Univariate logistic regression for promoter methylation status–unmethylated (reference) versus methylated.

BMI: body mass index.
Bold values indicate statistical significance (
Odds ratio (OR) is calculated at 95% confidence interval (95% CI).
Specific-level
Overall variable
Correlation between methylation of the GSTP1 promoter and clinicopathological features
As summarized in Table 2, no association was observed between GSTP1 promoter hypermethylation and the clinical factors such as age, menopausal status, dwelling, BMI, and HER-2/neu status. However, significant association was observed between the higher GSTP1 methylation frequency and positive nodal status when compared to the negative lymph node involvement, significant association was observed between tumor stages II, III, and IV when compared to tumor stage I, and significant association was observed between tumor grades II and III when compared to tumor grade I. Furthermore, the univariate analysis revealed the positive association of the higher GSTP1 promoter methylation with the PR negative tumor (OR = 3.33, 95% confidence interval (CI) = 0.10–11.12,
Correlation between GSTP1 expression and clinicopathological parameters
Table 3 summarizes association between the clinical parameters and gene expression. Taking normal protein expression as the reference, no significant association was found between the loss of the protein expression and the most of the clinicopathological parameters like age, menopausal status, BMI, dwelling, and Her2Neu status. The parameters that showed positive correlation with the GSTP1 promoter methylation like nodal status, tumor stage, tumor grade, and progesterone negative receptor tumors were also not significantly associated with the loss of the protein expression, except for the ER negative tumors (OR = 9.06, 95% CI = 2.23–36.71,
Univariate logistic regression for GSTP1 expression—loss of protein expression versus expression (reference).

BMI: body mass index.
Bold values indicate statistical significance (
Odds ratio (OR) is calculated at 95% confidence interval (95% CI).
Specific-level
Overall variable
GSTP1 gene polymorphism at codon 105 and its association with promoter methylation
The digestion of 176 bp product (Figure 4) resulted in generation of 95 and 81 bp products for homozygous mutant allele. Heterozygous mutant allele showed three bands of 176, 95, and 81 bp where as PCR product remained undigested for homozygous wild allele (Figure 5). The three allelic forms of the gene, mutant homozygous (G/G), mutant heterozygous (A/G), and the wild type (A/A), had the frequency of 9.8%, 29.4%, and 60.7%, respectively. There was no significant association observed between promoter methylation status of the GSTP1 promoter and genotype of the gene (wild vs mutant), as summarized in Table 4. However, there was statistically significant association (Table 5) between the loss of the protein expression (OR = 0.18, 95% CI = 0.05–0.78,
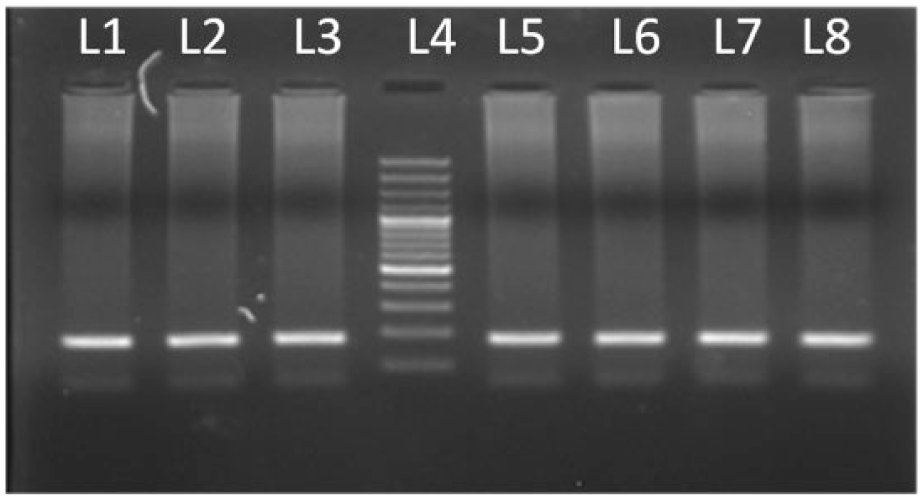
PCR product run on 2.5% agarose. L5 represents 100-bp DNA ladder.
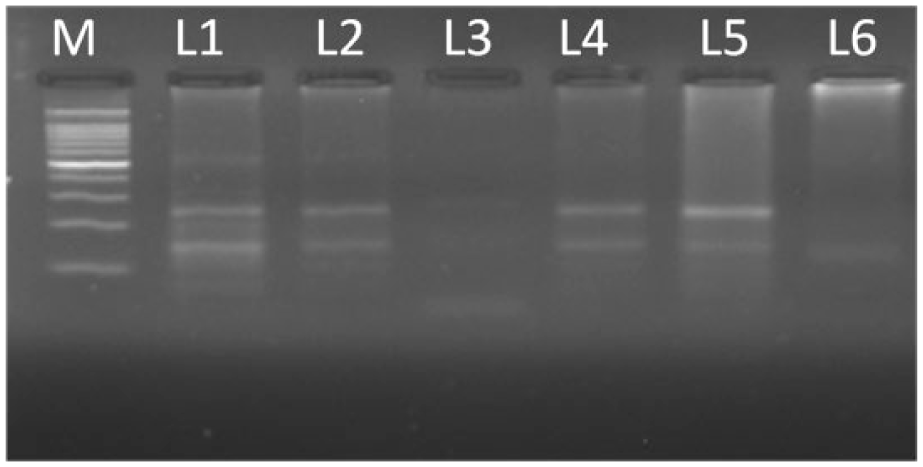
Restriction digestion products resolved on 3% agarose. Lane M represents 100-bp DNA ladder, lanes 1–5 represent heterozygous condition (Ile/Val), and lane 6 represent homozygous mutant condition (Val/Val).
Correlation between GSTP1 promoter methylation genotype.

Odds ratio is calculated at 95%confidence interval (95% CI).
The
Correlation between GSTP1 expression and genotype.

Odds ratio (OR) is calculated at 95% confidence interval (95% CI).
The
GSTP1 polymorphism and clinicopathological parameters of breast cancer patients.

BMI: body mass index.
Bold values indicate statistical significance (
Odds ratio (OR) is calculated at 95% confidence interval (95% CI).
Specific-level
Overall variable
Discussion
GSTP1, a member of the GST family of enzymes, is the phase-II xenobiotic metabolizing enzyme that is involved in the detoxification of the various exotoxins and endotoxins, including many carcinogens and the chemotherapeutic agents, thus conferring a protective role to the cell. 29 It also plays a significant role in the regulation of cell signaling by interacting with various important signaling proteins, including c-Jun NH2 terminal kinase, AJK-1, and TRAF-2, thereby regulating their downstream effects. 30 GSTP1 gene is found to be downregulated mainly by two ways. First is the substitution mutation in codon 105 of exon 5, which causes substitution of the isoleucine with valine at nucleotide position 313 (A to G transition) 31 with the mutant genotype (containing valine allele) effecting the enzyme activity as well as specificity. 19 This observation has been confirmed by the number of published studies concerning the role of this single-nucleotide polymorphism in relation to the individuals’ susceptibility to the various diseases 32 including many cancers, mainly breast cancer,24,25,33–35 bladder cancer, testicular cancer, and prostate cancer.36–38
The epigenetic silencing of this gene in the form of CpG hypermethylation is the second mechanism involved in the cancer progression. GSTP1 is also reported to be methylated in a number of cancers.12,16,39–41 The transcription silencing of this gene has been reported to be mediated through the action of methyl-CpG–binding domain 2 protein, which binds to the hypermethylated promoter. 42 The CpG hypermethylation in the promoter region of GSTP1 gene results in the downregulation of the detoxifying enzyme 12 and has been associated with the aggressive phenotype. 16 Methylation-based inactivation of the GSTP1 gene results in the decreased GSTP1 enzyme formation, thereby decreasing its buffering capacity against the short-term changes in the steroid levels in the body; since it is a caretaker gene, its inactivation further leads to the additional somatic genome alteration, both of which promote tumor growth. 43 Furthermore, the promoter hypermethylation event has been reported to function as the sensitive tumor marker, and based on its property of reversible nature, they have been reported to be excellent targets for the treatment therapies.44,45
The main aim of this study is to find out the promoter methylation profile of GSTP1 gene in breast cancer with respect to the ethnically different population of Kashmir and correlation of this hypermethylation event with the expression of the gene at protein level and also with the various clinicopathological parameters of the patient population. Highly sensitive, more common, and high-throughput technique of methyl-specific PCR 46 was used due to its high efficiency in detecting the methylation changes in heterogeneous cancer cell population. 47 Of the 51 histologically confirmed breast cancer tumors and their corresponding adjacent normal tissues, GSTP1 was found to be methylated in 37.25% of the tumors as compared to 11% of the normal tissues, taken from the same patient. This methylation frequency of the GSTP1 in the population under study is in corroboration with the frequency between 0% and –39% as reported in the most of the studies on different populations.25,48–50 However, the data are not in agreement with the study by Syeed et al., 51 which shows gene to be hypermethylated in 58% of the prostate cancer cases in Kashmiri population. The expression of the gene showed the inverse relation with the hypermethylation frequency, with the high frequency of hypermethylation showing significant association with the loss in the expression of the enzyme, confirming the role of the methylation in gene silencing. The association we found is in agreement with the results by Lin et al. 12 and Parrella et al. 41
As far as the clinicopathological features are concerned, no significant association was found between the promoter hypermethylation and parameters like age, menopausal status, dwelling, Her2Neu status, and BMI. However, the lymph node metastasis showed the positive association with the high frequency of the promoter hypermethylation, suggesting the role of the CpG hypermethylation of the GSTP1 gene in the metastasis of the primary tumor. Similar results have also been found in the Asian and Caucasian breast cancer patients.16,17 Furthermore, high frequency of the methylated GSTP1 promoters showed the positive association with the tumor grade and tumor stage, suggestive of the late occurrence of the methylation event in the breast cancer progression. These findings are in agreement with the previous published studies.16,17,24
Since both promoter hypermethylation and the substitution mutation of the GSTP1 gene have been implicated in the loss of the protein expression, the association of the GSTP1 genotype was studied in correlation with the promoter hypermethylation. However, in the present cohort study, no significant association of the GSTP1 promoter hypermethylation was found with respect to the mutant GSTP1 genotype, which is in agreement with the previous report on prostate cancer and breast cancer,24,52 suggesting that the promoter hypermethylation in the mutant genotype does not increase the risk of the cancer. However, the expression of the gene was significantly associated with the genotype, with mutant alleles showing decreased expression in relation to the wild allele further implicating that the mutant genotype is associated with the increased cancer risk and the aggressive phenotype. 17 The univariate analysis of GSTP1 genotype with the clinical parameters in the present cohort study showed no significant association of any clinical parameter with the genotype. This finding is in variance with the study by Pongtheerat et al., 53 which showed the positive association of the GSTP1 genotype with the PR status in Thai breast cancer patients
When the hypermethylation profile was correlated with the standard immunohistochemical parameters (Her2, estrogen, ER and progesterone, and PR receptor), the association was statistically significant for the negative ER and PR status. The results are in agreement with the study by A. Saxena, 24 for the north Indian population and for Caucasian women, when compared to the Korean women, 49 presenting the ethnic differences in the ER and PR status and the hypermethylation profile. The expression profile of the gene also showed the significant association with the negative ER status. Since estrogen for long has been implicated in the breast cancer progression 54 and their role in causing the damage to the genome has also been well studied, 55 silencing of the GSTP1 gene further enhances the role of estrogen as the endogenous tumor initiators.
Conclusion
To conclude the study, we found that epigenetic silencing of the GSTP1 gene by CpG promoter hypermethylation is associated with the significant loss of the protein expression, lymph node metastasis, tumor stage, and tumor grade. It is also positively associated with the progesterone and ER status. However, the mutant GSTP1 genotype is not associated with the promoter hypermethylation, and promoter hypermethylation and the substitution mutation of the GSTP1 gene are two independent events, but both leading to the decrease in the protein expression. Based on these observations, we suggest that the early detection of the aberrant methylation pattern may offer an alternative approach to the screening and detection of the tumor which further may be extended to the drug therapy design based on the reversible nature of this epigenetic change and ER/PR status. However, these findings need to be confirmed in the large cohorts for the better understanding of the role of methylation changes in the cancer progression and establishing the therapeutic strategies.
Footnotes
Acknowledgements
We are thankful to DST, India for providing the Inspire fellowship to Aaliya Bhat. We also acknowledge CSIR and UGC, India for providing JRF to Bushra Nissar and Nuzhat Shaheen Khan, respectively.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was financially supported by the Department of Science and Technology (DST), New Delhi, India.
