Abstract
The zinc-finger transcription factor PRDM1 (PR domain containing 1) plays key roles in the development of malignant lymphoma, leukaemia and some non-haematopoietic cancers, including breast cancer, colorectal cancer and glioma. However, little is known regarding the function of PRDM1 in the progression of lung cancer. Here, we found that PRDM1 is expressed in normal human lung epithelium but is downregulated in lung cancer cells. Decreased expression of PRDM1 correlates with poor prognosis in lung cancer. Depletion of PRDM1 in lung cancer cells promotes cellular invasion and anoikis resistance in vitro and lung metastasis in vivo. PRDM1 is silenced by an ectopically expressed lymphocyte-specific transcription factor Aiolos. The transcription of these two genes is negatively correlated in 206 lung epithelial cell lines. Our results indicate that PRDM1 functions as a tumour suppressor in lung cancer.
Introduction
Lung cancer is the most prevalent cancer worldwide in terms of incidence and also has the highest cancer mortality rate, thus creating a large global public health burden. 1 Common treatments include a combination of surgery and postoperative chemotherapy and/or radiotherapy. However, fewer than 20% of patients diagnosed with lung cancer survive 5 years after diagnosis. 2 Tumour metastasis is the main cause of death in patients with lung cancer. Therefore, detailed research on molecular mechanisms of tumour metastasis is urgently needed to improve the accuracy of diagnostic methods for lung cancer and to effectively treat it.
The zinc-finger transcriptional repressor PRDM1 (PR domain containing 1), also known as Blimp1, is a representative member of the PRDM family. 3 PRDM1 was originally identified as a silencer of β-interferon gene expression 4 and has subsequently been identified as a master regulator of plasma cell terminal differentiation.5,6 PRDM1 governs cell fate decisions in the developing embryo and adult tissues. 7 Although the exact mechanism by which PRDM1 represses transcription is not fully understood, PRDM1 possesses DNA-binding activity and recruits histone deacetylases and histone methyl transferases, as well as the corepressor Groucho. 8
PRDM1 plays important roles in haematopoietic tumourigenesis. 9 PRDM1 deficiency is frequently reported in different malignant lymphomas and leukaemia.10–14 Recently, dysregulated PRDM1 expression has also been detected in non-haematopoietic cancer cells. Via induction of the PRDM1 gene, RelB promotes a more invasive phenotype in oestrogen receptor alpha (ERα)-negative breast cancers. 15 Knockdown of PRDM1 by using small-interfering RNA (siRNA) results in both apoptosis and growth arrest in colorectal tumour cells. 16 However, a tumour-suppressive role of PRDM1 has been reported in glioma; PRDM1 negatively regulates Wnt/β-catenin signalling and decreases glioma malignancy. 17 Whether or how PRDM1 affects lung cancer progression is unknown.
In this study, we show that PRDM1 is downregulated in lung cancer cells, owing to ectopic expression of Aiolos. Decreased PRDM1 expression correlates with poor prognosis in lung cancer. Depletion of PRDM1 in the lung cancer cell line A549 promotes lung cancer cell invasion through Matrigel, anoikis resistance and grape-like colony formation in three-dimensional (3D) culture. In addition, PRDM1 depletion promotes lung metastasis in vivo. Our study uncovers a tumour suppressor function of PRDM1 in lung cancer progression.
Materials and methods
Cell lines
Human umbilical vein endothelial cells (HUVEC) and human bronchial epithelial cells (HBEC) primary cells were purchased from Lonza. Phoenix-293, A549, Beas-2B, H209, H69 and H1155 cells were obtained from the American Type Culture Collection (ATCC). Phoenix-293 and Beas-2B cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Hyclone) containing 10% foetal bovine serum (FBS; Hyclone). A549, H209, H69 and H1155 cells were cultured in RPMI-1640 medium (Hyclone) containing 10% FBS. HUVEC cells were cultured in endothelial cell growth medium-2 (EGM-2; Gibco). HBEC cells were cultured in bronchial cell growth media (BEGM) with supplemental factors (Lonza).
Cloning
Human genomic DNA was extracted from HUVEC cells using the rapid isolation of mammalian DNA method. Primers bearing restriction sites were used to amplify the 5′ flanking region of the PRDM1 gene from the extracted human genomic DNA. The polymerase chain reaction (PCR)-amplified fragment was excised with XhoI and KpnI (NEB) and was ligated into the equivalent site of the pGL3-Basic Vector (Promega) to form the PRDM1 promoter-luciferase reporter construct. The full-length Aiolos vector was constructed in our laboratory. For vector-based RNAi (short hairpin RNA (shRNA)), oligos encoding shRNA specific for PRDM1 were ligated into pSUPER.retro.puro, and the fragment containing the H1 promoter and hairpin sequences was subcloned into the lentiviral vector pCCL.PPT.hPGK.GFP.Wpre. The PRDM1 shRNA target sequences were as follows: 5′-GAT CTG ACC CGA ATC AAT G-3′ (PRDM1 shRNA1) and 5′-GCA ACT GGA TGC GCT ATG T-3′ (PRDM1 shRNA2). 16 For a shRNA control construct, we used shRNA against firefly luciferase (shRNA luciferase).
The specific primer pairs were as follows: PRDM1, forward: 5′-GGA GAA TGT GGA CTG GGT AGA GAT G-3′, reverse: 5′-TCC ACA GAG TCA TAT CCG CAT CCT-3′; p66Shc, forward: 5′-AAT GAG TCT CTG TCA TCG CTG GAG-3′, reverse: 5′-GGC GAT GAT CTG TTT GCA GTC TGC GG-3′ and GAPDH, forward: 5′-GTC AAC GGA TTT GGT CGT ATT-3′, reverse: 5′-AGT CTT CTG GGT GGC AGT GAT-3′.
Transfection and viral infection
Transfection with polyethylenimine (PEI) was performed as previously described. 18 Lentivirus was produced in Phoenix-293 cells with the packaging plasmids pMD2.BSBG, pMDLg/pRRE and pRSV-REV. After 24 h of transfection, lentivirus supernatant was supplemented with 5 µg/mL of polybrene filtered through a 0.45-µm syringe filter and used to infect target cells.
Luciferase assays
For luciferase assays, cells were seeded into a 24-well plate (105 cells each well) before transfection. Approximately 800 ng of DNA was complexed with 1 µL of Lipofectamine 2000 (Invitrogen) and then added to cells. Renilla reniformis luciferase (pRL-TK) was used as the transfection control. Cells were harvested and lysed approximately 24 h after transfection. The luciferase activity was measured using a Dual-Luciferase Reporter Assay System (Promega) in a Promega GloMax 20/20 Luminometer.
Cell death assays
After lentivirus infection, cells were dissociated with trypsin, resuspended and counted. Single cells (105 per well) were seeded into a 24-well ultra-low attachment surface plate (Corning) or CellBIND surface plate (Corning) with 0.5 mL of appropriate culture medium and were incubated at 37°C for 24 h. Cells were harvested and washed with phosphate-buffered saline (PBS). Cell death was assessed as DNA oligonucleosome fragmentation using Cell Death Detection ELISAPLUS (Roche). In brief, cells were lysed with lysis buffer, and the supernatant and immunoreagent were transferred into the streptavidin-coated microtiter plate and incubated at room temperature for 2 h. The solution was decanted, and each well was rinsed three times with incubation buffer. Absorbance was measured at 405 nm after the addition of 2,2′-Azinobis (3-ethylbenzothiazoline-6-sulfonic acid)-diammonium salt (ABTS) solution.
Transwell invasion assays
Invasion assays were performed as previously described with minor modifications. 19 In brief, 104 cells were seeded in 2% FBS/DMEM (Sigma) in the upper chamber of 24-well Transwell plates (pore size: 8 µm; BD Falcon), which had been pre-coated with a layer of Matrigel (BD Biosciences). The lower chamber was filled with 20% FBS/DMEM. After incubation for 20 h at 37°C, cells in the upper chamber were removed with a cotton swab. Cells that had traversed the membrane were fixed in 4% paraformaldehyde/PBS and quantified with 0.1% crystal violet staining at 100× magnification in three separate fields using light microscopy.
3D Matrigel culture
These assays were performed as previously described. 20 Single cells (2 × 103 per well) were cultured in suspension in 24-well ultra-low attachment surface plates for 8–14 days in 500 µL of RPMI-1640 medium supplemented with 2% FBS and 5% Matrigel. The Matrigel-containing medium was replenished every 3 days. Colonies were counted at 200× magnification in 10 separate fields using light microscopy.
Adhesion assays
Single cells were washed with PBS and suspended in RPMI-1640 medium at a final concentration of 2.7 × 105 mL−1. Next, 1.5 mL of the cell suspension was added into a 35 mm dish containing a dried glass coverslip coated with 10 ng/mL of fibronectin (R&D Systems) and incubated at room temperature for 15 min. Cells were washed gently and fixed in 4% paraformaldehyde/PBS. Attached cells were counted at 40× magnification in four separate fields using light microscopy.
Real-time quantitative PCR
Total RNA was isolated using Trizol reagent (Invitrogen). The complementary DNA (cDNA) was synthesized from 1000 ng total RNA with Oligo dT Primer and Thermo Scientific RevertAid First Strand cDNA Synthesis Kit. Quantitative PCR (qPCR) was performed with specific primers using the Power SYBR Green PCR Master Mix (Invitrogen). Gene expression was normalized to GAPDH. All reverse transcriptase (RT) reactions, including no-template controls, were run in a 7900 Real-Time PCR System (Applied Biosystems). Each sample was tested in triplicate.
Sequences of primer pairs for real-time qPCR were as follows: CDH1, forward: 5′-ACA CTG CCA ACT GGC TGG AGA TTA-3′, reverse: 5′-TGA TTA GGG CTG TGT ACG TGC TGT-3′; CDH2, forward: 5′-CAT CAA GCC TGT GGG AAT CC-3′, reverse: 5′-AAT GAA GTC CCC AAT GTC TCC AG-3′; FN1, forward: 5′-CCA GTC CTA CAA CCA GTA TTC TC-3′, reverse: 5′-CTT CTC TGT CAG CCT GTA CAT C-3′; VIM, forward: 5′-TTC CAA ACT TTT CCT CCC TGA ACC-3′, reverse: 5′-TCA AGG TCA TCG TGA TGC TGA G-3′; GAPDH, forward: 5′-GGT GGT CTC CTC TGA CTT CAA CA-3′, reverse: 5′-GTT GCT GTA GCC AAA TTC GTT GT-3′.
Western blotting analysis
For immunoblotting analysis, total cells were mixed with Laemmli sample buffer containing 0.05% 2-mercaptoethanol and heated at 95°C for 10 min. Equivalent amounts of protein were electrophoresed in a 12% gels and transferred to polyvinylidene difluoride (PVDF) membranes (Millipore). The membranes were blocked with 5% bovine serum albumin in Tris-buffered saline (pH 7.4) containing 0.1% Tween 20 and were incubated with primary and secondary antibodies according to the manufacturer’s instructions. The blot was probed with an anti-fibronectin (1:1000; Abcam), anti-caspase-3, anti-caspase-9 (1:1000; Cell Signaling Technology) or anti-β-actin (1:1000; Millipore), followed by anti-mouse or anti-rabbit IgG conjugated with peroxidase (1:5000; Bio-Rad).
Tail vein injection experiment
Twelve 6-week-old female BALB/c nude mice were obtained from the animal centre of the Cancer Institute of the Chinese Academy of Medical Science. A suspension of A549 cells (2 × 106) with control shRNA or PRDM1 shRNA was injected into the tail vein of each mouse. Surviving mice were killed 2 months after the injection. Lungs were fixed with formalin and observed using immunohistochemistry. All procedures of animal experiments were approved by Animal Care and Use Committee of Tianjin Medical University and conformed to the legal mandates and national guidelines for the care and maintenance of laboratory animals.
Statistical analysis
Statistical analyses were conducted using GraphPad Prism 5 and SPSS 17.0 software. The significance of the differences between the groups was determined using analysis of variance and t-tests. For all tests, p values < 0.05 were considered to be statistically significant.
Results
Reduced PRDM1 expression correlates with poor prognosis in lung cancer
We studied expression of PRDM1 in various human lung epithelial cells. As shown in Figure 1(a), PRDM1 messenger RNA (mRNA) was detected in the immortalized human lung epithelial cells Beas-2B and HBEC, human umbilical vessel endothelial cells (HUVECs), and the lung adenocarcinoma cell lines A549 and H1155. The small-cell lung cancer (SCLC) cell lines H209 and H69 did not express PRDM1.

PRDM1 is downregulated in lung cancer. (a) PRDM1 expression in lung cancer cell lines was analysed using RT-PCR. (b) Expression of PRDM1 was assessed in human lung epithelial cell lines, including airway epithelial cells, such as immortalized human bronchial epithelial cells and immortalized human small-airway epithelial cells and non-immortalized human small-airway epithelial cells. NSCLC subtypes included adenocarcinoma, squamous cell and large cell. (c) Kaplan–Meier survival analysis of the relationship between survival time and PRDM1 signature in lung cancer was performed using an online tool (http://kmplot.com/analysis/).
To assess whether PRDM1 is involved in the regulation of tumourigenesis in human lung cancer, we analysed the Gene Expression Omnibus database GSE32036 for PRDM1 expression in human normal lung epithelial cells (n = 59), non-small-cell lung cancer (NSCLC) cell lines (n = 118) and SCLC cell lines (n = 29). 21 Cells derived from lung cancers, compared with normal lung epithelium, showed reduced expression of PRDM1 (p < 0.001; Figure 1(b)). Notably, PRDM1 expression was significantly lower in SCLC than in NSCLC (p < 0.001; Figure 1(b)), consistent with the above results. Notably, SCLC is associated with an extremely poor prognosis (median survival < 2 years) because of its strong tendency to disseminate early and present with established metastatic foci. Therefore, we assessed the prognostic significance of PRDM1 downregulation. Kaplan–Meier survival analysis of PRDM1 was performed with an online tool (http://kmplot.com/analysis/), and the results revealed that lower PRDM1 expression (p = 0.00047) was associated with worse overall survival of patients with lung cancer (n = 2437; Figure 1(c)). These data indicated that PRDM1 is expressed in normal epithelium but downregulated in lung cancer cells. Downregulation of PRDM1 predicts poor prognosis in lung cancer.
Knockdown of PRDM1 enhances cell invasion
To determine the effect of PRDM1 expression on lung cancer cell behaviour, we performed an in vitro invasion assay on Matrigel. We transduced lentiviral vectors carrying either PRDM1 shRNAs or luciferase shRNA into A549 cells. The effect of shRNA knockdown on PRDM1 expression is shown in Figure 2(a).

Downregulation of PRDM1 enhances cellular invasion potential. (a) Semi-quantitative RT-PCR was performed to examine PRDM1 expression level in shRNA-transfected A549 cells. A threefold serial dilution of the cDNA reaction was used as the template for PCR. (b) Transwell invasion assays assessed the effect of PRDM1 knockdown on invasion of A549 cells. (c) Adhesion assays assessed the effect of PRDM1 knockdown on A549 cell adhesion. (d) Morphologies of 3D-cultured cells (n = 3; *p < 0.05).
PRDM1 knockdown A549 cells or control cells were seeded on the upper surface of a Transwell filter coated with a reconstituted basement membrane (Matrigel). Invasive cells invaded through Matrigel, passed through the pores and migrated onto the lower surface of the filter. More PRDM1 shRNA-expressing A549 cells invaded through Matrigel, as compared with control cells (p < 0.05; Figure 2(b)), thus indicating that PRDM1 depletion promotes cancer cell invasion.
Adhesion to the extracellular matrix is an important prerequisite for cancer cell invasion. 22 Thus, we tested the effect of PRDM1 downregulation on cell adhesion to fibronectin, an important component in lung stroma. PRDM1 knockdown A549 cells or control cells were plated on fibronectin-coated plates. After 15 min, attached cells were counted. As compared with control cells, more PRDM1 knockdown cells adhered to fibronectin-coated plates (p < 0.05; Figure 2(c)), thus indicating that reduced PRDM1 expression promotes cellular adhesion to fibronectin.
The use of 3D culture models to study aspects of normal and malignant phenotypes in culture has been widely recognized. 23 PRDM1 knockdown cells or control cells were seeded in 5% Matrigel. As demonstrated in a previous study, the 3D morphologies of cells cultured for 8–10 days can be classified into two distinct morphological groups: round and grape-like. Representative examples of each group and colony numbers of grape-like cells are shown in Figure 2(d). PRDM1 shRNA-expressing A549 cells exhibited more grape-like colonies than round colonies, and the number of grape-like colonies in PRDM1 shRNA-expressing A59 cells was approximately twice that as in control cells (p < 0.05; Figure 2(d)). Formation of a grape-like colony with poor cell–cell contacts and a disrupted boundary is associated with the ability to break away from the primary tumour.24,25 Therefore, depletion of PRDM1 in lung cancer cells increases cancer cell invasive capacity.
Downregulation of PRDM1 expression promotes anoikis resistance
Normal epithelial cells have to attach to the extracellular matrix to survive and they will undergo apoptosis, also known as anoikis, upon detachment. Anoikis resistance has been suggested to be a critical step in the survival of cancer cells during circulation, thereby facilitating cancer metastasis. To assess the effect of PRDM1 depletion on anoikis, we cultured PRDM1 knockdown A549 cells or control cells in ultra-low attachment multi-well plates to force them into suspension and tested for apoptosis using a Cell Death Detection ELISA assay. Reduced apoptosis was detected in PRDM1 knockdown A549 cells in comparison with control cells at 24 h of suspension (p < 0.05; Figure 3(a)). No statistically significant difference in the apoptosis rate was observed between PRDM1 knockdown cells and control cells in adherent culture (p > 0.05; Figure 3(a)). Therefore, PRDM1 depletion promotes anoikis resistance.
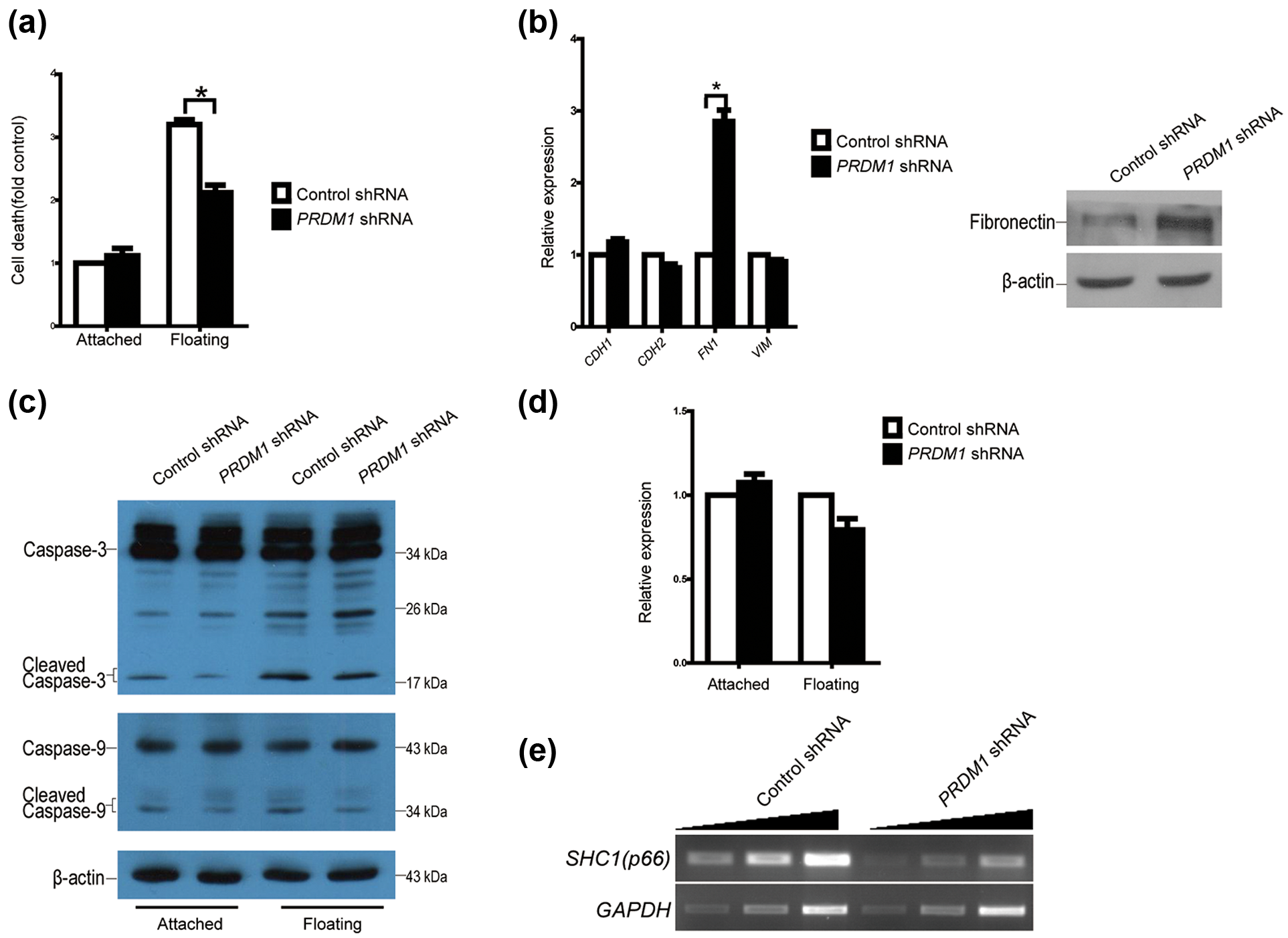
Downregulation of PRDM1 expression increases anoikis resistance. (a) PRDM1 shRNA was transduced into A549 cells using a lentivirus system, and the extent of cell death in cultures (attached and floating cells) was assessed at 24 h (n = 3; *p < 0.05). (b) Expression levels of epithelial–mesenchymal transition–related genes were measured by quantitative RT-PCR and western blotting (n = 3; *p < 0.05). (c) Caspase activity was monitored by western blotting. (d) Expression level of Bcl-2 was measured by quantitative RT-PCR. (e) Downregulation of PRDM1 expression decreased p66Shc mRNA levels.
Downregulation of PRDM1 expression induces fibronectin expression and inhibits caspase activity
Anoikis resistance is a hallmark of epithelial–mesenchymal transition (EMT). We next examined the expression of key proteins that are involved in EMT, such as E-cadherin, N-cadherin, vimentin and the proteins that are involved in anoikis pathways, such as Bcl-2 and caspase family proteins. The knockdown of PRDM1 induced a dramatic increase in the expression of mesenchymal marker fibronectin in mRNA and protein level, but did not alter the expression of vimentin, N-cadherin and E-cadherin, suggesting that PRDM1 depletion is not able to induce EMT programme (Figure 3(b)). The knockdown of PRDM1 had no effect on Bcl-2 expression (Figure 3(d)), but reduced cleaved caspase-3 and -9 under both non-adherent and adherent culture conditions (Figure 3(c)), indicating that caspase signal pathway is involved in PRDM1-mediated anoikis.
We also examined the expression of p66Shc, which is required for anoikis and functions as a powerful metastasis suppressor in mice. 26 As expected, the knockdown of PRDM1 expression induced a marked decrease in the expression of p66Shc mRNA (Figure 3(e)).
Downregulation of PRDM1 expression promotes lung metastasis in vivo
To examine the effect of PRDM1 knockdown on metastatic capacity in vivo, A549 cells expressing PRDM1 shRNA were injected into the tail veins of 6-week old female BALB/c nude mice. At day 50, at which point the last animal in the PRDM1 knockdown group died, 33% of control animals were still alive. The animals died from lung metastases (Figure 4(b)). Survival was significantly shortened in PRDM1 knockdown A549 cells compared with controls (p < 0.05; Figure 4(a)). Therefore, consistent with the in vitro results, knockdown of PRDM1 increased the metastatic capacity of lung cancer cells in vivo.

Knockdown of PRDM1 shortens survival in mice. (a) Survival curve for nude mice injected with PRDM1 knockdown A549 cells via tail vein injection (*p < 0.05). (b) Gross and histological appearance of lung tumour from mice injected with PRDM1 knockdown A549 cells.
Aiolos represses PRDM1 gene expression
Aiolos (also known as IKZF3), a zinc-finger transcription factor, is normally expressed in lymphocytes. Our previous study has shown that Aiolos is frequently expressed in lung cancer cells, and its expression confers lung cancer cell anoikis resistance. Analysis of the Gene Expression Omnibus database GSE50812 revealed that transient expression of Aiolos in A549 cells represses PRDM1 transcription. 26 This finding was confirmed using semi-quantitative reverse transcription-polymerase chain reaction (RT-PCR; Figure 5(a)), thus suggesting that PRDM1 downregulation might be the consequence of ectopic Aiolos expression. To test whether PRDM1 repression is caused by endogenously expressed Aiolos, we examined expression of Aiolos and PRDM1 in various cell lines by using RT-PCR. We observed a strict negative correlation between the expression of Aiolos and PRDM1 in primary culture cells (HUVEC and HBEC), immortalized HBEC (Beas-2B) and human lung cancer cell lines (A549, H69 and H209; Figure 5(b)). Quantification of transcription levels in 206 human lung epithelial cell lines also revealed a negative correlation between these two genes (r2 = −0.202; p < 0.05; Figure 5(c)).

PRDM1 transcription is negatively regulated by Aiolos. (a) Semi-quantitative RT-PCR determined PRDM1 expression in control or in Aiolos-expressing A549 cells. A threefold serial dilution of the cDNA reaction was used as the template for PCR. (b) The expression of PRDM1 and Aiolos in lung cancer cell lines was analysed by using RT-PCR. (c) Quantitative analysis showed the correlation of PRDM1 with Aiolos/IKZF3. (d) Luciferase assays were performed with constructs containing 5′-flanking regions of PRDM1 in A549 or H209 cells. (e) Luciferase assays were performed with constructs containing 5′-flanking regions of PRDM1 in Aiolos-expressing A549 cells (n = 3).
To further study the effect of ectopic Aiolos expression on PRDM1, a series of luciferase reporters containing the PRDM1 promoter (−149 to +207, −929 to +207, −1988 to +207 and −3154 to +207) were constructed. The genomic fragment −149 to +207, containing the minimal promoter region, was necessary for PRDM1 transcription. 27 These luciferase reporters in series were transfected into A549 cells that do not normally express Aiolos and H209 cells that express endogenous Aiolos. The luciferase expression was much higher in A549 cells than in H209 cells (Figure 5(d)). Consistently, cotransfection of these constructs with vector containing Aiolos expression cassette into A549 cells significantly reduced luciferase expression (Figure 5(e)), indicating inhibitory function of Aiolos to PRDM1 promoter. Altogether, these results indicated that expression of PRDM1 is strongly repressed by Aiolos in multiple lung cancer cells in vitro.
Discussion
Our findings uncovered a connection between PRDM1 and lung cancer. The results presented here demonstrate that PRDM1 exerts tumour-suppressive functions in lung carcinogenesis. Depletion of PRDM1 in lung cancer cells inhibits anoikis and promotes cellular invasion and lung metastasis in vivo. This study also shows that downregulation of PRDM1 may be the consequence of aberrant activation of Aiolos.
PRDM1 is widely expressed in multiple cell types including B lymphocytes, T lymphocytes, mammary luminal progenitor cells 28 and skin epithelial cells. 29 Our results clearly showed that PRDM1 is also expressed in normal human airway epithelial cells. The function of PRDM1 in normal bronchial epithelial cells is not clear. However, downregulation of PRDM1 in lung cancer cells correlates with poor prognosis in patients with lung cancer.
We found that PRDM1 depletion in lung cancer cells increased the number of grape-like colonies in 3D culture, a result consistent with a previous report that PRDM1 depletion in mammary epithelial cells causes epithelial cell polarity defects. 28 Loss of polarity results in grape-like colony formation in 3D culture. Therefore, PRDM1 might execute a similar function in mammary epithelial cells and lung epithelial cells, although increased PRDM1 expression is observed in a more invasive phenotype of ERα-negative breast cancers.
Aiolos, a lymphocyte lineage-restricted transcription factor, is ectopically expressed in lung cancer cells and confers anoikis resistance to cancer cells. Aiolos directly represses expression of p66Shc. 26 PRDM1 downregulation, another consequence of ectopic Aiolos expression, also promotes anoikis resistance. Because re-expression of p66Shc completely restores anoikis of Aiolos-expressing cells, and because knockdown of PRDM1 expression induces a marked decrease in lymphocyte p66Shc expression, PRDM1 is important for p66Shc silencing in Aiolos-induced anchorage independent growth (Figure 6). As Aiolos and PRDM1 are both closely associated with lymphocyte development, they may cause changes in the inflammatory microenvironment.

Schematic representation of the role of PRDM1 in regulation of Aiolos-associated cancer metastasis. Aiolos has a negative role in PRDM1 and p66Shc transcription and loss of PRDM1 downregulates p66Shc. Thus, lack of PRDM1 promotes cancer metastasis through promotion of cell invasion and at the same time confers cancer cell anoikis resistance through decreasing p66Shc transcription.
Although further work is required, these results suggest that PRDM1 may be a promising therapeutic target for lung cancer and a prognostic marker for survival in patients with early-stage lung cancer.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
This work was supported by the National Natural Science Foundation of China (30801377, 91519331, 31371295 and 81572271), Ministry of Science and Technology of China (2014CB910104) and the Tianjin Municipal Science and Technology Commission (15JCZDJC34800).
