Abstract
Lung cancer causes huge mortality worldwide, and targeting new pathway may provide an alternative in modulating signaling in cancer. Nuclear factor erythroid 2–related factor 2 is the major regulator of endogenous and exogenous stress by activating numerous antioxidant genes critical in cancer, Alzheimer’s, Parkinson’s, and inflammatory bowel diseases. Ganoderic acid is a triterpene from basiodiomycetes fungus Ganoderma lucidum with numerous therapeutic effects. In this study, ganoderic acid and its 50 isomers and natural activators were docked by receptor-based molecular docking using Maestro 9.6 (Schrödinger Inc.) in the Kelch-like ECH-associated protein 1-nuclear factor erythroid 2–related factor 2 signaling pathway. The receptor-based molecular docking reveals the best binding interaction of nuclear factor erythroid 2–related factor 2 and ganoderic acid A with GScore (−9.69) (kcal/mol), Lipophilic EvdW (−1.83), Electro (−0.72), Glide emodel (−73.369), H bond (−1.1), molecular mechanics/generalized Born surface area (−75.541) with Leu 718, Asp 800, Cys 797 residues involved in hydrogen bonding. The calculated docking energy highlights the lipophilic, hydrogen bonding, pi–pi stacking interactions, and non-covalent bonding. Analysis showed the involvement of cysteine and serine residues which were crucial in the activation and translocation from cytoplasm to the nucleus in the nuclear factor erythroid 2–related factor 2 signaling process. The molecular docking tool QikProp analyzed the absorption, distribution, metabolism, excretion, and toxicity but needs some modifications in their structure to satisfy Lipinski’s rule. Ganoderic acid A is a best docked isoform which inhibits the cell proliferation, viability, migration, and reactive oxygen species and messenger RNA expression of nuclear factor erythroid 2–related factor 2 in H460 cells.
Keywords
Introduction
Lung cancer causes huge mortality worldwide, mainly among smokers and few among non-smokers. Non-small-cell lung carcinoma (NSCLC) is common cancer among smokers, and non-smokers focus on its specific subtypes, squamous cell carcinoma or adenocarcinoma (AC). The reactive oxygen and nitrogen species causes oxidative stress and prove to be harmful, whereas controlled level of oxidant regulates signaling pathways. 1 The defense system is counterbalanced by oxidant system using different mechanisms regulating physiology of the body. The Kelch-like erythroid-derived Cap-n-Collar Homology (ECH)-associated protein 1 (Keap1)-Nrf2 pathway is the main regulator of cytoprotective responses caused by reactive oxygen species (ROS). Nuclear factor erythroid 2–related factor 2 (Nrf2) is the transcription factor which under normal condition remains in the cytoplasm with the association of Keap1 and Cullin 3 proteins which degrade Nrf2 ubiquitination. 2 Keap1 is a cysteine-rich protein substrate adapter which promotes Cullin 3 to ubiquitinate Nrf2. The process of degradation ubiquitinated the Nrf2 and recycled by translocation of the proteasomes. Oxidative stress conditions disrupt the cysteine residues in Keap1, which results in disruption of Keap1-Cul3 ubiquitination system. 3 Non-ubiquitination of Nrf2 results in the accumulation in the cytoplasm which further translocates it into the nucleus. In the nucleus, it forms a complex with small transcription factor (Maf) protein that binds to the promoter region of antioxidant response element (ARE) of antioxidative genes which initiates cascade for activation of transcription. 4 Activation of Nrf2 pathway leads to NAD(P)H-quinone oxidoreductase 1 (Nqo1), glutamate–cysteine ligase, sulfiredoxin 1 (SRXN1), and thioredoxin reductase 1, glutathione S-transferase (GST), and uridine 5′-diphospho-glucuronosyltransferase (UDP)-glucuronosyltransferase (UGT). ROS are mutagenic and may thereby promote cancer. 5 Activation of Nrf2 signaling protects against cancer, aging, diabetes, cardiovascular disease, neurodegenerative diseases, inflammation, pulmonary fibrosis, and acute pulmonary injury. 6 Different natural products such as quercetin, 7 curcumin, 8 ursolic acid, 9 and celastrol 10 have potential to modulate Nrf2 signaling cascade. 11 One of the neglected natural products is Ganoderma lucidum, used as medicine since ancient time, hence known as herb of spirtual potency.12,13 Previously, our research group carried out the anticancer property of G. lucidum and active compounds in different signaling pathways in cancer. The methods use in this study was taken from our previous studies.14–16 Among different bio-constituents in G. lucidum, ganoderic acid was explored for revealing mechanism in cancer signaling. 17 In this study, molecular docking was performed for the first time on Nrf2 with 50 isoforms of ganoderic acid and their natural inhibitors to reveal the mechanistic binding of Keap1-Nrf2 pathway. After molecular docking, best isoforms of ganoderic acid on the basis docking score was checked for the biological activity in the cell line.
Methodology
Preparation of ligands and protein molecule
The three-dimensional structure of protein crystallized structure was downloaded from Protein Data Bank (PDB) site with Nrf2 (PDB: 4L7D) 18 from which water molecules were removed in concern to the domain topology that usually interacts with hydrophobic regions. Subsequently, polar hydrogen, wherever required, were incorporated to fill the inappropriate valency of the protein atoms, ensuing in increased polarizability of bonds. The increased polarizability enhanced the probability of ligand–protein interactions. Later, processed protein structure was checked for the stereochemical quality by residue-by-residue geometry as well as overall structural geometry. The preparation of different isoforms of ganoderic acid and other natural activator structures was carried out by software ChemBioDraw Office 17 (licensed at Cambridge’s soft). Literature survey highlighted that some of the natural activators such as quercetin, 7 curcumin, 8 ursolic acid, 9 and celastrol 10 target the Nrf2 signaling. These compounds were subjected in the LigPrep for the ligand preparation by Maestro 9.3. LigPrep was corrected by addition and optimization of hydrogen bonds for correcting the valency, creation of disulfide bonds, and fixing of missing residues. These modifications prepares the ligands prior to the docking. Furthermore, prepared structure was minimized and optimized with liquid simulations (OPLS-2005) force field to acquire an energetically stable geometry.19–22
Receptor grid formation
Grid pre-calculates grid maps of interaction energies by scoring electrostatic, dispersive energies of surrounding target with a macromolecule before the docking. 17 The grid maps are used to determine the total interaction energy for a ligand with a macromolecule. Grid mapping computes the crucial coordinates data and assigns the coordinates of the Nrf2 prior to the receptor based docking. Furthermore, grid mapping affords an appropriate surface topology for the ligand atoms for interaction with the Nrf2 domain. Grid mapping is a pre-requisite to direct various isoforms of ganoderic acids to bind for their region of the firm affinity of the Nrf2 domain. Grid results in favorable interaction and best post during binding which represents conformation, position, orientation in the receptor-ligand complex. Other parameters such as sites, constraints, rotatable groups, and excluded volume, which are the default settings of the Maestro 9.6, are used. 23
Glide receptor based molecular docking
Molecular docking procedures were carried out after preparing the ligand, protein, and the grid on the active site of the protein. Glide docking tools predict the systematic and computational simulation method for predicting best binding orientation to the protein target. Glide molecular docking outputs GScore (empirical scoring function) which is a combination of various parameters relevant to binding energy. The GScore is calculated by calculating ligand–protein interaction energies, root mean square deviation (RMSD), hydrogen bonds, hydrophobic interactions, internal energy, pi–pi stacking interactions, and desolvation. 17 Glide module of the XP visualizer analyzes the specific ligand–protein interactions. Ligands were docked with the X-ray crystal structure of Nrf2 (PDB: 4L7D) using Glide. The best possible fit compounds’ results after docking were analyzed for thermodynamic optimal energy value, the potential of bonding and conformations, types of interactions, residues involved in the interaction, and the distance between different residues.24,25
Prime/molecular mechanics/generalized Born surface area simulation (free energy calculation)
The proper solvation facilitates in the process of molecular recognition. The solvation effect estimates the binding affinities by molecular mechanics/generalized Born surface area (MM-GBSA) scoring, where MM means molecular mechanics, GB means Generalized Born, and SA means solvent-accessible surface area. In MM-GBSA simulation, electrostatics of ligand–receptor complex and solvation were analyzed which has the edge over prior methods of Coulomb-based terms. The Prime/MM-GBSA approach calculates free energy and calculates the binding affinity of the receptor and the ligands. 26 The simulation was based on the structure obtained by receptor-based ligand molecular docking. The ligand poses were minimized by Prime, whereas energy of the complex was obtained by OPLS-2005 force field with VSGB2.0 solvent model in GB/surface area continuum solvent model 27
where ΔE is the minimized energy of complex, protein, and ligand.
where ΔG is the solvation free energy of the complex protein and ligand.
where ΔGSA is surface area energy of the complex protein and ligand.
Absorption, distribution, metabolism, and excretion properties
QikProp is an important tool that calculates properties of the significant descriptors and pharmaceutically relevant molecules by comparing their values with those of 95% of already known pharmaceutical drugs. Absorption, distribution, metabolism, excretion, and toxicity (ADME-T) properties of the docked ligand molecules were subjected to QikProp tool. It analyzes and predicts different properties of drugs, about the use and aftermath of drug intake, that is, ADME. It gives the information about QPlogPo/w, QPlogBB, overall central nervous system (CNS) activity, Caco-2, Madin–Darby Canine Kidney (MDCK) cell permeability, logKhsa for human serum albumin binding, and the percentage of human oral absorption. 28
Materials and methods
Ganoderic acid A (≥98%) was purchased from Sigma-Aldrich and dissolved in dimethyl sulfoxide (DMSO) at a concentration of 50 mM and stored at −20°C. Lung cancer cell line (H460) used in the study was procured from National Centre for Cell Sciences, Pune, India. The cell line was carefully maintained in RPMI media supplemented with 10% heat-inactivated fetal bovine serum (FBS), 1% penicillin (units/mL), streptomycin (100 mg/mL). Cells were maintained at 37°C in a humidified atmosphere with 5% CO2.
Total RNA isolation, complementary DNA synthesis, and reverse transcription-polymerase chain reaction
About one million cells were seeded and supplemented with culture media having 10% FBS and 1% penicillin/streptomycin, incubated at 37°C. H460 cells were treated with 80 µM of ganoderic acid for 48 h, and then, total RNA was isolated and extracted using Trizol (Life Technologies, Gaithersburg, MD, USA). RNA was quantified, and later, complementary DNA (cDNA) preparation was carried out. Approximately 1 µg of total RNA was reverse transcribed into cDNA using PrimeScript 1st strand cDNA Synthesis Kit (Takara Bio Inc., Shiga, Japan). Real-time quantitative polymerase chain reaction (PCR) was performed with Assays-on-demand (Applied Biosystems, USA) with primers Nrf2 forward primer 5′-TCTCCTCGCTGGAAAAAGAA-3′, reverse primer 5′-AATGTGCTGGCTGTGCTTTA-3′, GAPDH forward 5′-ACGGATTTGGTCGTATTGGGCG-3′, reverse primer 5′-CTCCTGGAAGATGGTGATGG-3.′ Each data point was repeated in triplicates. Quantification values were obtained using threshold PCR cycle number (Ct), where the increase in signal results in an increase in PCR product. The normalization of relative messenger RNA (mRNA) level in each sample was done with β-actin. The relative expression target gene levels equaled ΔCt = Cttarget gene − Ctβ − actin.
3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide cell proliferation assay
Effect of different treatments of ganoderic acid A on the growth of lung cancer cells (H460) cells were assessed using the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay in triplicates. Approximately, 10,000 cells were seeded in the 96-well plate in RPMI media containing 10% FBS and incubated 37°C overnight, followed by serum starvation for 24 h. Cells were treated with different concentrations of ganoderic acid A (5, 10, 20, 50, 80 µM/mL) in serum-free media which was incubated for 48 h. After, 48 h, the medium was replaced with 100 µL of MTT (0.5 mg/mL in phosphate-buffered saline (PBS)) in 10% FBS containing media, and incubated at 37°C for 4 h in the dark. Then, the supernatant was removed and the reduced MTT, formazan complex, was solubilized in 200 µL/well DMSO. Absorbance was measured at 570 nm using microplate reader.
Evaluation of cytotoxic/apoptotic effect
The rate of cell death in response to ganoderic acid A with treatment was assessed for cytotoxic effect by trypan blue exclusion test. Around 2 × 105 H460 cells were seeded in six-well culture plates. The adhered cells were treated with ganoderic acid A (5, 10, 20, 50, 80 µM/mL) for 48 h. After treatment, both floating cells in the medium and adhered cells on the plate were collected and concentrated by centrifugation. The cell viability was estimated after staining with 0.4% trypan blue for 15 min. Both live (unstained) and dead (stained) cells were counted in three replicates using Automated Cell Counter (Invitrogen, India). Percent data of dead cells were calculated and used as an indicator of the degree of cell death.
Wound healing assay
H460 cells were seeded in a six-well plate and allowed to grow up to 75%–80% confluence level. The monolayer was scratched to form a wound, using a pipette tip, of 1.5 mm diameter. Subsequently, the cells were then washed twice with PBS to remove the detached cells. After that the cells were treated with different concentrations of the ganoderic acid A and incubated for 48 h. After this, the cells were monitored at different time intervals to observe their invasive behavior. Cell invasion into the wounded area was determined under Olympus inverted microscope (Olympus, Tokyo, Japan).
Nitro blue tetrazolium reduction assay
In this assay, superoxide ion production was measured by colorimetric nitro blue tetrazolium (NBT) assay which converts water-soluble NBT to water insoluble, NBT diformazan. Around 8 × 103 cells/well were seeded into the 96-well plates in RPMI media containing 10% FBS and 1% penicillin/streptomycin and then incubated at 37°C overnight followed by serum starvation for 24 h. After 24 h, the media was changed with fresh complete medium (200 µL) and exposed to different ganoderic acid concentrations (5, 10, 20, 40, and 80 µM). After 48 h, the medium was removed and incubated with 100 µL 0.1% NBT, incubated for 4 h. The reduced NBT was solubilized with 100 µL 2 M KOH and 100 µL DMSO for 30 min, and absorbance was taken at 570 nm.
Intracellular ROS assay
Around 10,000 cells were counted using trypan blue assay and seeded in 96 wells of the microtiter plate. Treatment strategies of ganoderic acid with hydrogen peroxide are represented in Figure 1. H460 cells were incubated for 24 h followed by treatment with compounds alone in different strategies. After the treatment, 10 mM of 2′,7′-dichlorodihydrofluorescein diacetate (H2DCFDA) dye was used in the cultured cells and kept in the dark for 30 min, followed by absorbance at the emission wavelength of 530 nm.

Schematic representation of strategies employed for the determination of ROS production by ganoderic acid.
Statistical analysis
The results were expressed as the mean ± standard deviation of experiments performed in triplicates. Data obtained were subjected to one-way analysis of variance (ANOVA), and significant differences of the mean were determined statistically using Tukey’s test using SigmaPlot 11.
Results and discussion
Stress exacerbates cancer, cardiovascular disease, chronic inflammation, and neurodegenerative diseases. 29 To counteract the detrimental effects of stress, cells evolved the adaptive, dynamic, endogenous antioxidants by Nrf2 signaling mechanism. The activation of the cytoprotective enzyme by Nrf2 signaling provides the basis of chemopreventive and chemoprotective activities. 30 During stress conditions, phosphorylation of the cysteine residue in Keap1 senses the cellular redox action. The oxidant modification in the cysteine residues results in transportation from cytoplasm to the nucleus. In addition to the decisive role of cysteine residues, serine or threonine is also considered important in the activation of Nrf2 signaling. 30 Among different protective agents targeting Nrf2, ganoderic acid along with its 50 isomers were docked using receptor-based molecular docking by Maestro 9.6 (Schrödinger Inc.). The molecular docking identifies the maximum energy, protein–ligand interactions, orientations, and conformations best suited for effective drug design. 31 Molecular docking predicts the best suitable fitted orientation to form a stable complex and finds the strength of association of binding affinity by different docking parameters. 17
This study was carried out by ganoderic acid and its 50 isoforms which were docked with X-ray crystal structures of Nrf2 retrieved (PDB: 4L7D) from PDB. Triterpene of G. lucidum, ganoderic acid, has lanosterol scaffold, and variation in its isoforms varies in the functional group or in the side chain. 32 The binding interaction between receptors complexed with various isoforms of ganoderic acid highlighted the lipophilic, salt bridges, electrostatic, and hydrogen bonding interactions. In molecular docking of Nrf2, ganoderic acid A exhibits best docking parameters as GScore (−9.69), Lipophilic EvdW (−1.83), Glide emodel (73.369), and H Bond (−2.68), and hydrogen bonding forms basis of the stability of the complex. Different residues involved in hydrogen bonding were Leu 718, Asp 800, and Cys 797 with 2.45, 2.23, and 3.61 Å hydrogen bond length, respectively. Also, the stability of complex depends on activation of other covalent and non-covalent bindings, as shown in Figure 2. The binding affinity was calculated by MM-GBSA, which assesses the orientation of ligands to the receptors using scoring and thereby predicting binding interaction (Table 1).

Interaction displayed in receptor-based molecular docking profile of Nrf2 (PDB: 4L7D) with ganoderic acid A and quercetin. Protein–ligand interactions involved in ganoderic acid A binding were Leu 718, Asp 800, and Cys 797, whereas Gln 791, Met 793, Asp 800, and Cys 797 were involved in quercetin.
Binding affinities, scores, and interaction of Nrf2 with different isoforms of ganoderic acid along with natural activators.

The structure of ganoderic acid A has a tetracyclic ring with one double bond at a different position, and the branch ends with a carboxyl group while others with some modification. In molecular docking of ganoderic acid and its isoforms, –COOH group has a crucial role in different activities such as inhibitory effects, whereas –C=O or –OH group also has a role in signaling through different receptors. The lanosterol moieties in the ganoderic acid A have involvement of cysteine residue which leads to the activation and translocation of Nrf2 to the cytoplasm to the nucleus.
Among different natural activators, quercetin exhibits best-docked docking parameters with GScore (−7.8), Lipophilic EvdW (−4.69), Glide emodel (62.453), and Gln 791, Met 793, Asp 800, and Cys 797 residues participate in hydrogen bonding (Figure 1). The binding interaction of Nrf2 with quercetin exhibits participation of both phenyl and chromone moieties during docking process which makes it vital to design an effective drug. Different parameters of other isoforms and natural activators were presented in Table 1. The active participation of Cys and Ser residues during molecular docking assists in translocation and activation of different antioxidant genes in Nrf2 signaling pathway.
ADME properties
Binding affinity prediction is useful for analyzing the pharmacokinetic and pharmacodynamic properties. ADME-T predicts physicochemical, effective permeability in jejunum, blood–brain barrier permeation, pKa (ionization constants a multi-protic model), intrinsic clearance solubility, toxicity risks, optimal dose in human, fraction absorbed in human, likely sites of metabolic attack, logD, logP, and MDCK apparent permeability. QikProp is an important tool that calculates properties of the valid descriptors and pharmaceutically relevant molecules by comparing their values with those of 95% of already known pharmaceutical drugs. Ganoderic acid and its 50 isoforms were checked for ADME-T properties. Most interesting aspects of these compounds are their admirable QPlogPo/w, QPlogHERG K+ channels, QPlogBB, QPlogKP, and QPlogKhsa values that satisfy Lipinski’s rule of five (Table 2). Some of the isomers in different receptors did not satisfy all aspects and needed some modifications in the basic lanosterol structure (Table 2). Predicted IC50 value for blockage of HERG K+ channels (acceptable range: above −5.0); QPP Caco—predicted apparent Caco-2 cell permeability in nm/s. Caco-2 cell is a model for the gut–blood barrier (nm/s) (<25: poor, >500: great); QPlogBB—predicted brain/blood partition coefficient; QPP MDCK—predicted apparent MDCK cell permeability in nm/s. MDCK cells are considered to be a good mimic for the blood–brain barrier (nm/s) (<25: poor, >500: great); QPlogKP—predicted skin permeability; percentage of human oral absorption (<25% is poor and >80% is high).
Evaluation of drug-like properties of the different isoforms of ganoderic acid and another natural activator of Nrf2 by QikProp.

Effects of ganoderic acid A on cancer cells
The cytotoxicity of ganoderic acid A in cancer cells was performed by MTT assay. Lung cancer cells H460 were treated with different concentrations (5, 10, 20, 50, 80 µM) of ganoderic acid A at different time intervals (24, 48, and 72 h). Ganoderic acid A marked 50% of inhibition at 50 µM and higher concentration and best marked in 48 h. Ganoderic acid A exhibited a remarkable reduction in cell proliferation in cancer cells in a concentration-dependent manner (Figure 3(a)).
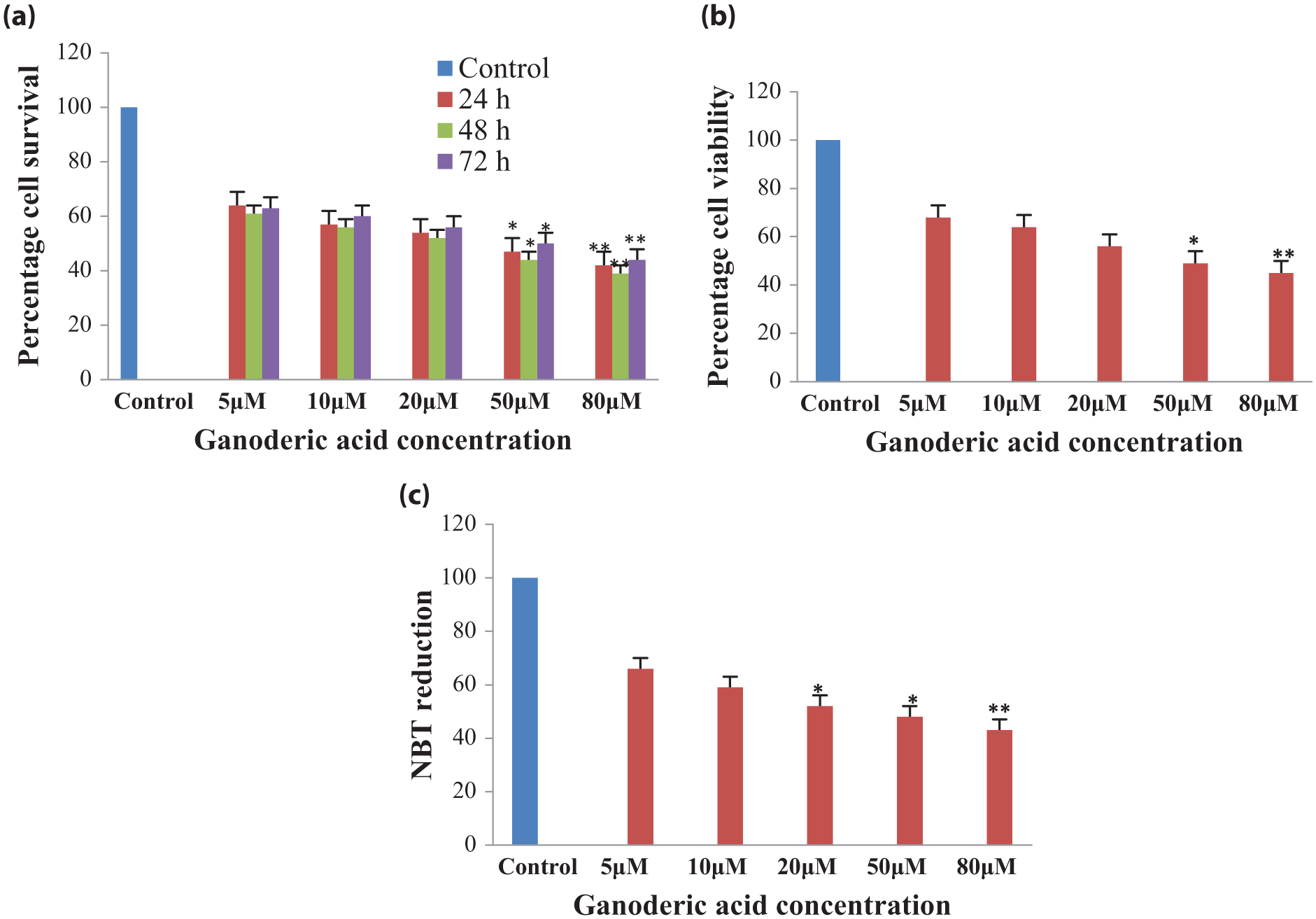
Effect of ganoderic acid A on H460 cells. (a) Ganoderic acid A reduces the cell growth in a dose-dependent manner determined by the MTT assay, (b) reduces the cell number of H460 cells in a dose-dependent manner determined by the trypan blue exclusion test of the cell, (c) significantly reduces the cell number of H460 cells in a dose-dependent manner determined by the NBT reduction assay.
Cell viability was checked by trypan blue exclusion test to determine the number of the treatments of ganoderic acid A. Treatment of ganoderic acid A in H460 cells with different concentrations (5, 10, 20, 50, 80 µM) for the duration of 48 h displayed a reduction in cell viability to exactly 32%, 36%, 44%, 51%, and 55%, respectively (Figure 3(b)). Thus, it can be concluded that ganoderic acid reduced cell viability in a concentration-dependent manner. Furthermore, NBT assay (colorimetric method) was used to analyze the antioxidant potential of ganoderic acid A, which converts tetrazolium chloride to diformazan by superoxide production. Ganoderic acid A causes a reduction in ROS to 50% in 50 µM and higher concentration in NBT reduction assay (Figure 3(c)). Furthermore, treatment of ganoderic acid inhibits the migration of H460 cells in a dose-dependent manner after 48 h. Photomicrographs of H460 cells are represented in Figure 4(a), in which cell migration was analyzed after 48 h as compared to control. Treatment with ganoderic acid (10, 20, and 50 µM) reduced the migration and increased the wound in dose-dependent manner (Figure 4). Thus, this study concludes that ganoderic acid A has potential to inhibit proliferation, viability, and ROS in the H460 cells.

Ganoderic acid A reduces the cell migration in H460 cells determined by wound healing assay. (a) H460 cells treated with different concentrations (10, 20, and 50 µM) of ganoderic acid A are presented with photomicrographs. Cells were seeded in six-well plate and grown overnight, and next day, scratch was made in confluent monolayer, followed by treatment with ganoderic acid for 48 h. (b) The rate of wound closure was analyzed using ImageJ and exhibiting the increase in width of wound after ganoderic acid treatment.
Antioxidant potential of ganoderic acid
The ROS production potential of ganoderic acid A was evaluated by DCFDA strategies represented in Figure 1. Different treatments in H460 cells with different concentrations of ganoderic acid A make them to behave differently. Cell treatment with H2O2 increases free radicals, whereas ganoderic acid scavenges free radicals and enhances antioxidant potential in dose-dependent manner (20, 50, and 80 µM). Potential of ganoderic acid enhances more efficiently when H2O2 treatment was given for 24 h as compared to treatment of H2O2 given for 30 min after 24 h of cells treatment. Thus, it was inferred that ganoderic acid shows antioxidant potential when H2O2 was given in time-dependent manner (Figure 5).

Graphical representation of antioxidant potential of ganoderic acid in different strategies with hydrogen peroxide (H2O2) in the dose- and time-dependent manner.
Effect of ganoderic acid A on Nrf2 mRNA expression in H460 cells
The effect of ganoderic acid A on Nrf2 mRNA expression was determined using quantitative polymerase chain reaction (qPCR). The results show that ganoderic acid A has the potential to reduce the Nrf2 rnRNA expression in H460 cells. The natural inhibitor, quercetin, treatment decreases the Nrf2 mRNA expression by 29%, whereas ganoderic acid A decreases by 22.5% at 80 µM concentration in H460 cells (Figure 6).

Expression analysis of ganoderic acid A and quercetin in qPCR at 80 µM. Natural inhibitor of Nrf2, quercetin, inhibits the mRNA expression (29%), whereas ganoderic acid A inhibits expression (22%) at 80 µM in H460 cells as compared to control.
Conclusion
Nature fulfilled the need of humanity by rich flora and fauna with an inexhaustible reservoir of the natural products. Ganoderma lucidum and its bioconstituents effectively target the expression of numerous signaling proteins in cancer. Various isoforms of ganoderic acid comprise of lanosterol moieties, and variation in its isoforms varies in the side chains or the functional groups. Nrf2 signaling protects from free radicals by activating various antioxidants genes and unbalances in free radicals scavenging forms the cause of the various diseases. This study shows receptor-based molecular docking of Keap1-Nrf2 endowed with an in-depth understanding of docking interaction which assists in designing effective drugs. The active participation of cysteine and serine residues in activation and translocation of the Keap1-Nrf2 pathway in docking with ganoderic acid A highlighted its effectiveness during the process. Ganoderic acid A effectively inhibits cell proliferation, viability, ROS, and mRNA expression of Nrf2 expression in H460 cells and thus provides the platforms to target Nrf2 signaling, but more exploration is still needed in this field. Our present group is engaged in determining the anticancer property of G. lucidum and its bioconstituents.
Footnotes
Acknowledgements
The authors thank Central University of Punjab, Bathinda, for providing the necessary facilities to carry out this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.



