Abstract
Radiotherapy is an important treatment option for non-small cell lung carcinoma patients. Despite the appropriate use of radiotherapy, radioresistance is a biological behavior of cancer cells that limits the efficacy of this treatment. Deregulation of microRNAs contributes to the molecular mechanism underlying resistance to radiotherapy in cancer cells. Although the functional roles of microRNAs have been well described in lung cancer, their functional roles in radioresistance are largely unclear. In this study, we established a non-small cell lung carcinoma Calu-1 radioresistant cell line by continuous exposure to therapeutic doses of ionizing radiation as a model to investigate radioresistance-associated microRNAs. Our data show that 50 microRNAs were differentially expressed in Calu-1 radioresistant cells (16 upregulated and 34 downregulated); furthermore, well-known and novel microRNAs associated with resistance to radiotherapy were identified. Gene ontology and enrichment analysis indicated that modulated microRNAs might regulate signal transduction, cell survival, and apoptosis. Accordingly, Calu-1 radioresistant cells were refractory to radiation by increasing cell survival and reducing the apoptotic response. Among deregulated microRNAs, miR-29c was significantly suppressed. Reestablishment of miR-29c expression in Calu-1 radioresistant cells overcomes the radioresistance through the activation of apoptosis and downregulation of Bcl-2 and Mcl-1 target genes. Analysis of The Cancer Genome Atlas revealed that miR-29c is also suppressed in tumor samples of non-small cell lung carcinoma patients. Notably, we found that low miR-29c levels correlated with shorter relapse-free survival of non-small cell lung carcinoma patients treated with radiotherapy. Together, these results indicate a new role of miR-29c in radioresistance, highlighting their potential as a novel biomarker for outcomes of radiotherapy in lung cancer.
Introduction
Lung cancer remains the leading cause of cancer-related death in men and women worldwide, surpassing colorectal, breast, and prostate cancer combined.
1
Histologically, lung cancer is divided into non-small cell lung carcinomas (NSCLC) and small-cell lung carcinomas (SCLC). NSCLC accounts for more than 80% of lung cancer diagnoses, and nearly 70% of the patients are diagnosed with advanced disease with a 5-year overall survival of less than 15%. Moreover, roughly 20% of all NSCLC diagnoses belong to the locally advanced setting. These patients are candidates for multimodal treatment, including chemotherapy, radiotherapy, and surgery, with curative intent. Despite the improvements in diagnosis and therapy in recent years, the prognosis and outcome of patients with NSCLC are still unsatisfactory.
2
Radiation therapy is an effective treatment option for lung cancer patients.3,4 Approximately 20% of lung cancer patients undergo thoracic radiotherapy with curative intent in the locally advanced setting. Unfortunately, even with multimodal treatment, a subpopulation of cells within the primary tumor avoids the cytotoxic effect of radiotherapy, permitting the clonal selection of cancer cells with a more malignant phenotype, and thus limiting the effect of treatment. A fraction of this clonal selection is represented by radioresistant tumor cells, which contribute to disease progression, tumor recurrence, and cancer mortality.1,4 Massive cell death is a mechanism by which radiation therapeutically functions to reduce tumor volume. In cells, ionizing radiation (IR) induces DNA damage directly or indirectly, resulting in cell cycle checkpoint activation, DNA damage response, breakdown cell proliferation pathways, and, finally, apoptosis activation. However, tumor cells with a radioresistant phenotype avoid these biological responses. Cancer cells undergo inherent molecular changes as a consequence of carcinogenesis that favors radioresistance; furthermore, cancer cells may also develop radioresistance by undergoing molecular and genetic alterations following radiation treatment.5,6 The molecular mechanisms associated with intrinsic or acquired radioresistance in lung cancer are poorly understood. Accumulating evidence indicates that aberrant expression of microRNAs (miRNAs) as a result of carcinogenesis confers inherent radioresistance to cancer cells. Moreover, miRNA expression has been shown to be altered in cancer cells during treatment, which contributes to the development of radioresistance.7,8 MiRNAs are a class of naturally occurring small non-coding RNA molecules that have emerged as key regulators of genetic expression. This regulatory control is exerted through sequence-specific complementary binding to the 3′ untranslated region (3′ UTR) of target messenger RNA (mRNA) resulting in translational repression or degradation of the target.
9
Several miRNAs involved in increasing sensitivity or resistance to radiotherapy by inhibiting key functional proteins have been identified and characterized.7,8 Thus, miRNA-targeting therapy has been proposed as a novel approach to radioresistant tumor management. Many examples of miRNAs involved in radioresistance have been reported in breast cancer,10,11 laryngeal carcinoma,
12
prostate cancer,
13
nasopharyngeal cell carcinoma,14,15 and liver cancer,
16
among other cancers. In lung cancer cells, miRNAs have been described as potential regulators of radiation response.17–22 Thus, many efforts to discover and identify novel miRNAs associated with radioresistance have been undertaken. Although these findings suggest that deregulation of miRNAs expression plays a pivotal role in radioresistance mechanisms in cancer cells, more extensive investigation is required to elucidate the functions of miRNAs in the radioresistant phenotype of lung cancer. In this study, we uncovered a novel set of miRNAs in radioresistant lung cancer cells not previously reported in resistance to radiotherapy. In particular, we showed for the first time that downregulation of tumor suppressor miR-29c promotes radioresistance in lung cancer cells
Materials and methods
Cell lines
Human NSCLC Calu-1 cell line was obtained from American Type Culture Collection (ATCC; ATCC® HTB-54™). Calu-1 and Calu-1 radioresistant (Calu-1RR) cell lines were routinely cultured in Dulbecco’s Modified Eagle Medium-F12 (DMEM-F12; Invitrogen) supplemented with 10% fetal bovine serum and 100 U/mL penicillin and 100 µg/mL streptomycin at 37°C in a 5% CO2 atmosphere.
Development of radioresistant lung cancer cells
Subclones displaying significant resistance to radiotherapy (hereafter named Calu-1RR) were established from the surviving populations of their irradiated parental lung cancer cell lines Calu-1 as described below. Calu-1 (1 × 106) cells were exposed to 15 sequential fractions of 2 Gy/week of IR using a linear accelerator (Clinac 600, Varian) for a total dose of 30 Gy. The novel radioresistant Calu-1RR cells were established from the surviving populations of their parental irradiated Calu-1 cells. After the final irradiation, a group of cell clones isolated from the Calu-1RR cells were cultured individually, and the Calu-1RR cells were established. Non-irradiated controls were handled identically to the irradiated cells without radiation exposure. All experiments were performed between 4 and 10 passages after the establishment of individual clones.
Clonogenic survival assays
The
RNA isolation and miRNAs polymerase chain reaction arrays analysis
Total RNA was isolated using TRIzol reagent (Invitrogen) according to the manufacturer’s protocol. RNA integrity and quantification was assessed by agarose gel electrophoresis, spectrophotometry (Nanodrop 2000, Thermo Scientific), and bioanalyzer (2100 bioanalyzer system, Agilent Technologies). Expression analysis of 1,170 miRNAs in Calu-1 and Calu-1RR cells was assessed by quantitative real-time reverse transcription polymerase chain reaction (qRT-PCR) using the miRNA PCR arrays (Qiagen). RT was carried out using 100 ng of total RNA and the miScript II RT Kit. The detection of mature miRNAs was evaluated by using the miScript PCR arrays Kit according the manufacturer’s protocol. PCR was carried out in a GeneAmp System 9700 (Applied Biosystems).
Data analysis
The level of miRNAs was measured by qRT-PCR using the comparative Ct (2−ΔΔCt) method. SNORD61, SNORD68, SNORD72, SNORD95, SNORD96A, and RNU6-2 were used for data normalization. Ct raw data were obtained using an automatic baseline and a threshold of 0.2. A fold change value ≥ 2 was used to define the differentially expressed miRNAs. An adjusted
Functional and pathway annotation analysis
The experimentally validated miRNAs target genes were obtained from MiRTarBase (http://mirtarbase.mbc.nctu.edu.tw/) and published studies. The predictive miRNAs target genes were obtained using four algorithms: TargetScan (http://www.targetscan.org/); mirWalk (http://www.umm.uni-heidelberg.de/apps/zmf/mirwalk/); miRDB (http://mirdb.org/miRDB/); miRANDA (http://www.microrna.org/microrna/getGeneForm.do) and DIANA (http://diana.cslab.ece.ntua.gr/). Only miRNA-target interactions identified by at least three prediction algorithms were considered for further analysis. Enrichment analyses for KEGG (Kyoto Encyclopedia of Genes and Genomes), panther pathways, and gene ontology (GO) terms were performed with the GeneCodis (http://genecodis.cnb.csic.es/) and DAVID (http://david.abcc.ncifcrf.gov) software programs.
Apoptosis assays
Calu-1, Calu-1RR, and Calu-1RR cells transfected with mir-29c mimic were treated with or without 8 Gy of IR, and IR-induced cell death was then detected by Annexin V assays using the Annexin-V-FLUOS Staining Kit (Roche). Briefly, 1 × 106 cells were seeded in T-25 flasks and treated with IR. The cells were harvested and stained with FITC-conjugated Annexin V and propidium iodide (PI) 24 h after radiation treatment. Following incubation at 37°C for 15 min, cells were analyzed by flow cytometry. Data acquisition and analysis of 10,000 events for each sample were performed using a BD Accuri C6 flow cytometer (BD Biosciences). Annexin V single positive cells were identified as early apoptotic cells, whereas both Annexin V and PI positive cells were identified as end-stage apoptotic cells and PI positive cells as necrotic cells.
Wound healing assays
Calu-1 and Calu-1RR cells (1 × 106) were seeded in triplicated wells in six-well plates and cultured until a confluent monolayer was formed. Cells were serum-starved for 24 h before being scratched. Cell monolayers were scratched with a 200-µL pipette tip to imitate a wound. After washing and removing the floating cells, the distance between cell margins was measured at 24 h and graphed. Images were acquired with an Axiovert 40 CFL fluorescence microscope (Carl Zeiss AG) using a 20× objective.
Cell proliferation assays
Calu-1, Calu-1RR, and Calu-1RR transfected with mir-29c mimic (1 × 104) were seeded in 96-well culture plates. Cells were treated with or without 8 Gy of IR. After 24 h, cells were incubated in MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) solution (0.5 mg/mL) for 4 h. The medium was removed, and formazan dye crystals were solubilized with 50 µL dimethyl sulfoxide (DMSO). Absorbance was measured using a spectrophotometer at a wavelength of 540 nm.
qRT-PCR
The expression of miR-29c was assessed by qRT-PCR with the TaqMan miRNA Assay Kit (Applied Biosystems). Approximately 100 ng of total RNA from culture cells was reverse transcribed using the looped RT primer specific for miR-29c according to the manufacturer’s protocol. MiR-29c detection was performed using TaqMan Universal PCR Master Mix, No AmpErase UNG (Applied Biosystems). PCR was carried out in a GeneAmp System 9700 (Applied Biosystems). The expression of miRNA was determined using the comparative Ct (2−ΔΔCt) method. RNU44 was used as a control for normalization of data.
Transfection of miR-29c
Calu-1RR cells were transfected with miR-29c mimic (Life Technologies #MC10518) and scrambled at a concentration of 30 nM using siPORT amine transfection agent (Ambion #AM4503). The miR-29c mimic was diluted in 25 µL of Opti-Mem (Invitrogen) to concentrations of 30–100 nM and individually added to wells containing cultured cells in 450 µL of DMEM-F12. The expression of miR-29c was evaluated 48 h post-transfection by qRT-PCR, as previously described. Transfected cells were treated with or without 8 Gy of IR 48 h post-transfection. The subsequent analysis, which included clonogenic, proliferation, and apoptosis assays, was performed as previously described.
Western blot assays
Calu-1, Calu-1RR, and Calu-1RR cells transfected with miR-29c mimic were harvested by tripsinization and lysed in a buffer containing 25 mM Tris-HCl pH 7.5, 150 mM NaCl, 1% NP40, and protease and phosphatase inhibitors (Roche). Proteins were separated on 12% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) gel and transferred to nitrocellulose membranes (Bio-Rad). Membranes were probed with the specific primary antibodies (MCL1 (1:1000 Santa Cruz); BCL2 (1:1000 Cell Signaling); GAPDH (glyceraldehyde 3-phosphate dehydrogenase; 1:2500 Cell Signaling)), followed by horseradish peroxidase (HRP)-conjugated secondary antibodies (1:2000, Zymed). The bands were visualized using an enhanced chemiluminescence (ECL) western blotting detection system (GE Healthcare).
The Cancer Genome Atlas data analysis
Level 3 miRNA-Seq data from 221 lung tumor samples were downloaded from The Cancer Genome Atlas (TCGA) lung cancer data set portal (https://tcga-data.nci.nih.gov/). Patients were divided into those showing lung adenocarcinoma (LUAD) and those showing lung squamous cell carcinoma (LUSC). MiR-29c expression levels (reads per million (Rpm)) were obtained from 91 matched tumors and normal tissues (46 LUAD and 45 LUSC paired samples). Then, 79 LUAD and 51 LUSC samples from patients who received radiotherapy as primary and/or additional treatment and with available RFS data were divided into two groups according to whether they presented high (> median) or low (≤ median) miR-29c expression levels. RFS curves were analyzed by the Kaplan–Meier method, and a log-rank test was used to assess significance. A Cox regression model was applied.
Statistical analyses
All results derived from three independent experiments were represented as mean ± standard deviation (SD). For comparison between groups, Student’s
Results
Development of radioresistant lung cancer cells
The inherent radioresistance of Calu-1 cells was assessed by clonogenic survival assays by administering a range of single radiation doses (2, 4, 8, and 10 Gy). The IR caused a decrease in surviving clones in a dose-dependent manner relative to non-irradiated controls (Figure 1(a)). When the survival fraction (SF) was calculated, we found a significant survival difference between Calu-1 at 8 Gy and non-irradiated cells. These results indicate that 8 Gy of IR significantly reduced the SF of Calu-1 cells (Figure 1(b)). An IR dose of 8 Gy was used for all following experiments. Based on fractionated treatment protocols used in clinic practice, and to ensure that these

Establishment of acquired radioresistant lung cancer cell model. Calu-1 cells were exposed to 2, 4, 8, and 10 Gy of X-ray irradiation; cells were seeded and cultured. (a and b) Cell survival after irradiation was examined with a clonogenic survival assay in parental Calu-1 cells. The median lethal dose was 8 Gy. (c) Diagram of fractionated treatment protocol for establishment of Calu-1RR cells. Calu-1 parental cells were exposed to 30 Gy of fractionated irradiation (2 Gy) to obtain a radioresistant subclone Calu-1RR. (d) Representative images of the morphological changes in Calu-1RR cells compared with Calu-1 cells. (e) Representative image of clonogenic assays of Calu-1 and Calu-1RR cells exposed to 8 Gy of X-ray irradiation to validate radioresistance of Calu-1RR cells. (f) Survival fraction of irradiated Calu-1 and Calu-1RR cells (IR+) was normalized by the survival fraction of non-irradiated cells (IR−). Error bar, SD from three independent experiments.
Calu-1RR cells exhibit a differential miRNome expression
To identify miRNAs modulated in lung cancer cells involved in radioresistance, we compared the miRNAs expression profiles between Calu-1 and Calu-1RR cells. The expression of 1170 miRNAs was analyzed using high-throughput qRT-PCR arrays, and differentially expressed miRNAs (p ≤ 0.001; twofold change) were identified by comparative Ct analysis (2−ΔΔCt). Our results indicate that 50 miRNAs were modulated (16 upregulated and 34 downregulated) in Calu-1RR (Figure 2(a); Table 1). Chromosomal localization of miRNAs modulated in Calu-1RR cells is indicated in Table 1. No relation between miRNAs deregulation and their chromosomal localization was observed, suggesting that changes in miRNAs expression could not be the direct result of genomic instability. Remarkably, miR-450a, miR-96, and miR-29c have been associated with the response to radiotherapy in certain types of cancer, including lung cancer (Table 1), whereas miR-197, miR-29c, and miR-21* have been established as prognostic and risk factors in lung cancer (Table 1).

Calu-1RR cells trigger differential expression profile of microRNAs, higher cell proliferation and lower apoptosis levels. (a) MiRNome was analyzed by high-throughput quantitative real-time PCR (qRT-PCR) arrays in Calu-1RR and Calu-1. Comparative Ct analysis (2−ΔΔCt) was used to identify miRNAs differentially expressed in Calu-1RR cells (p ≤ 0.001; fold change: 2). MiRNAs differentially expressed (16 upregulated and 34 downregulated) in Calu-1RR are shown. (b) Cell proliferation was evaluated by MTT assays in Calu-1 and Calu-1RR cells at 24 h after cell exposure to 8 Gy of X-ray radiation (IR+). Results were measured and normalized by using values obtained from non-irradiated cells (IR−). (c) Apoptosis was measured using Annexin-V-FITC assays. Calu-1 and Calu-1RR cells were exposed to 8 Gy of X-ray radiation (IR+) and 24 h after Annexin-V-FITC assays were conducted by flow cytometry. Graphical representation of Annexin-V-FITC assays of irradiated cells (IR+) compared with non-irradiated cells (IR−). (d) Representative images of flow cytometry results. Data are presented graphically as the mean ± SD of three independent experiments.
MiRNAs modulated in radioresistant lung cancer cells Calu-1RR.

Identification of novel miRNAs implicated in radioresistance
Thirty-six novel radioresistance-associated miRNAs were identified in Calu-1RR, including miR-523, let-7d*, miR-937, miR-450b-3p, miR-154*, miR-548o, miR-617, miR-18b*, and miR-1208 (Table 1). Remarkably, certain passenger strands of miRNAs (miR*) of the better-characterized oncomiRs were modulated in Calu-1RR, including miR-154*, miR-21*, miR-196b*, miR-23a*, miR-18b*, miR-20a*, miR-130a* let-7d*, and miR-27a*. Notably, their corresponding leading strands play important roles in cancer and radiotherapy resistance, but the role of the passenger strand is unknown in lung carcinogenesis and tumoral radioresistance.
GO processes and biological pathways modulated by radioresistance-associated miRNAs
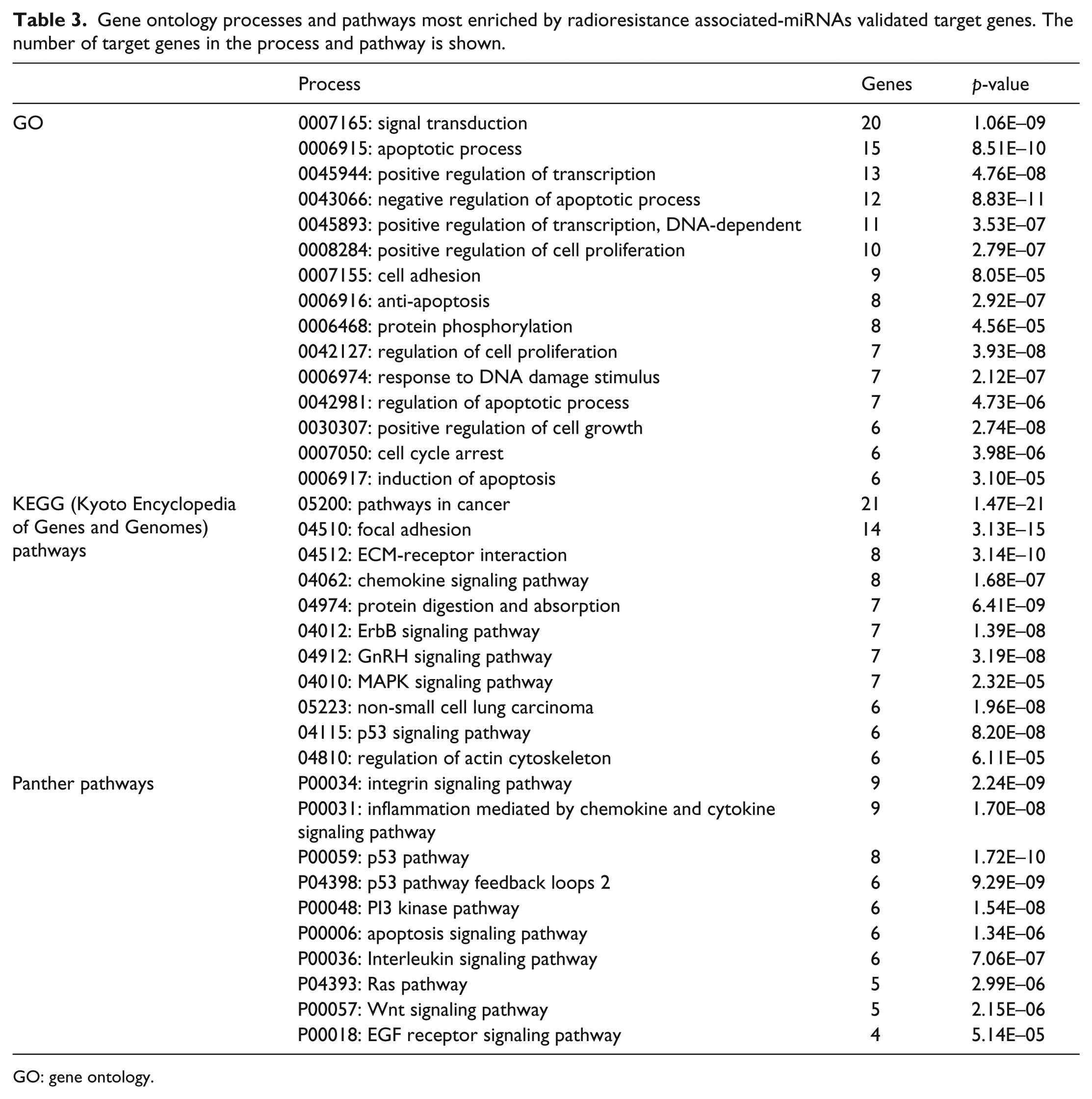
To understand the function of miRNAs, the putative target genes of the 50 differentially expressed miRNAs were predicted by integrating at least three algorithms. Results predicted 850 and 2879 putative target genes for the upregulated and downregulated miRNAs, respectively. GO terms of the target genes for the upregulated miRNAs indicated that they are involved in transcriptional processes, signal transduction, cell differentiation and apoptotic process, whereas for the downregulated miRNAs, the target genes have functions in signal transduction, cell adhesion, apoptotic process, and protein phosphorylation (Supplementary Table S1). The target genes of the miRNAs modulated in Calu-1RR cells were also uploaded into the KEGG and Panther databases for pathway enrichment analysis. Results showed that five pathways were statistically enriched, including the mitogen-activated protein kinase (MAPK) pathway, focal adhesion, the Wnt pathway, cytokine-cytokine receptor interaction, and regulation of actin cytoskeleton (Supplementary Table S1). However, validated target genes for certain miRNAs modulated in Calu-1RR cells have been previously reported (Table 2). Interestingly, these genes include oncogenes and tumor suppressors with functions in apoptosis, DNA repair, signal transduction, transcription, and epigenetic master regulation (Table 2). Results obtained for the GO and biological pathways most enriched with respect to validated transcriptional target genes showed that these pathways are implicated in biological processes such as signal transduction, apoptosis, and cell and focal adhesion (Table 3). Our results suggest that radioresistance-associated miRNAs could regulate survival, migration, and apoptosis pathways, which have been reported as the most important processes involved in the radioresistance of cancer cells.
Validated target genes of miRNAs modulated in Calu-1RR cells.

Bold term significance in order to showed this genes in the table as a targets of miR-29c for to its experimental evaluation in posteriorly assays.
Gene ontology processes and pathways most enriched by radioresistance associated-miRNAs validated target genes. The number of target genes in the process and pathway is shown.
GO: gene ontology.
Calu-1RR cells increase cell proliferation and reduced apoptosis in response to radiation
To further investigate the biological behavior of Calu-1RR cells, cell proliferation, migration, and apoptosis were assessed. Calu-1 and Calu-1RR cells were treated with 8 Gy of IR, and cell proliferation and apoptosis were evaluated 24 h after irradiation by MTT and Annexin V assays, respectively. Results revealed that Calu-1RR cells presented significantly higher rates of cell proliferation than Calu-1 cells (Figure 2(b)). Importantly, Calu-1RR cells were more resistant to radiation-induced apoptosis than Calu-1 cells. Calu-1RR cells showed a clear reduction in the apoptosis percentage compared with Calu-1 cells (23% vs 57%) when treated with IR (Figure 2(c)). As shown in Figure 2(d), the proportion of Annexin V/PI positive cells was lower in Calu-1RR (24%) than in Calu-1 cells (55%) after treatment. These results indicate that Calu-1RR cells were refractory to radiation by increasing cellular proliferation and reducing the apoptotic response. In a wound-healing assay, no significant differences were observed in migration ability between Calu-1RR and Calu-1 cells (Supplementary Figure S1).
MiR-29c overcomes radioresistance in Calu-1RR cells
Among the miRNAs deregulated in the expression profiles analysis, miR-29c was suppressed in Calu-1RR cells (−5.4-fold change). MiR-29c is a tumor suppressor that has been reported as a prognostic factor in lung cancer. To address whether miR-29c might play a key role in radioresistance, miR-29c was overexpressed by transfection of mimic-miR-29c (30 nM) in Calu-1RR cells (Calu-1RR/miR-29c), and its effect on radioresistance was assessed by clonogenic survival and apoptosis assays. The reestablishment of miR-29c expression in Calu-1RR cells was confirmed by qRT-PCR (Supplementary Figure S2). Calu-1RR/miR-29c cells were exposed to IR, and surprisingly, miR-29c caused a significant reduction in the clonogenic survival rate on Calu-1RR compared with that observed for Calu-1RR scrambled transfected cells (Figure 3(a) and (b)). Moreover, miR-29c increased apoptosis in Calu-1RR after exposure to IR compared with the level observed for Calu-1RR control and scramble transfected cells (Figure 3(c) and (d)). The percentage of apoptotic cells in Calu-1RR expressing miR-29c increased significantly (20%) after treatment with 8 Gy of IR (Figure 3(c)). Together, these results indicate that restitution of miR-29c in radioresistant lung cancer cells confers sensitivity to radiotherapy, in part by reducing cellular survival and increasing the apoptotic response.

MiR-29c overcomes radioresistance of Calu-1RR cells. (a) Calu-1RR cells transfected with mimic-miR-29c (30 nM) were more sensitive to the cytotoxic effect of IR (8 Gy of X-ray) than were scrambled transfected control cells evaluated by clonogenic assays. The cell survival factor of irradiated (IR+) Calu-1RR transfected cells was measured and normalized by using values obtained from non-irradiated (IR−) Calu-1RR transfected cells. (b) Representative images of clonogenic assay results. (c) Apoptosis of Calu-1RR transfected with mimic-miR-29c (30 nM) was measured using Annexin-V-FITC assays. Calu-1RR transfected cells were exposed to 8 Gy of X-ray (IR+) and 24 h after Annexin-V-FITC assays were conducted by flow cytometry. Graphical representation of Annexin-V-FITC assays of irradiated cells (IR+) compared with non-irradiated cells (IR−). (d) Representatives images of flow cytometry results. (e) Western blot analysis of Mcl-1 and Bcl-2 protein levels in Calu-1RR cells transfected with mimic-miR-29c. GAPDH was used as a control. Images are representative of three independent experiments. (f) Densitometric analysis of bands from western blot results of (e). Data were normalized using GAPDH expression. (g) Illustration of miR-29c overcoming radioresistance through apoptosis activation and suppression of anti-apoptotic genes
MiR-29c downregulates anti-apoptotic genes BCL2 and MCL1
To investigate the mechanism underlying the effect of miR-29c on radioresistance, we evaluated the expression of the apoptosis-related
MiR-29c expression correlates with radiotherapy outcomes in NSCLC patients
To further investigate whether downregulation of miR-29c correlated with radiation resistance in NSCLC patients, we retrieved patient clinical information, including radiotherapy treatment data, and corresponding miRNA expression (miRNAs-Seq) results from the publicly available TCGA database (http://genome-cancer.ucsc.edu/). We used the expression data for miR-29c from a total of 125 primary LUAD and 96 primary LUSC specimens. We first divided 46 LUAD and 45 LUSC samples into paired normal and tumor tissues, and miR-29c levels were compared. The expression of miR-29c was significantly lower in tumor tissues than in paired normal tissues in LUAD (median = 3082.6 Rpm;

Mir-29c is suppressed in NSCLC tumor tissues and correlates with relapse-free survival in NSCLC patients treated with radiotherapy. Mir-29c expression is suppressed in NSCLC tissue tumors. The miR-29c levels from the cancer genome atlas miRNA-seq data of tumor tissues and matched paired normal tissues in (a) lung adenocarcinoma (LUAD) and (b) lung squamous cell carcinoma (LUSC) were compared. Kaplan–Meier analysis according to miR-29c expression in NSCLC patients who received radiotherapy. Relapse-free survival (RFS) in patients with (c) LUAD and (d) LUSC who received radiotherapy as primary and/or additional palliative treatment and with high median or low median expression levels of miR-29c. Curves were compared using a log-rank test. Comparison between groups was performed using Student’s
Discussion
Radiotherapy is an important treatment option for NSCLC patients. Resistance of lung tumor cells to radiotherapy determines the outcome of patients with NSCLC; however, the exact molecular mechanism that underlies NSCLC tumor radioresistance remains unsolved. MiRNAs are small molecules involved in multiple malignant cell behaviors, including radioresistance. Evidence has revealed that miRNA expression profiles are significantly modified in cancer cells undergoing IR.7,8 Moreover, many studies have shown radioresistance by altering miRNA levels in various malignances, such as laryngeal
12
prostate,
13
nasopharyngeal,
14
and breast10,11 cancer
In conclusion, our study emphasizes the systematic identification of novel miRNAs that provide a more comprehensive panorama with regard to how they are involved in radioresistance in lung cancer. We also uncovered potential predictive markers for radiotherapy response or therapeutics targets for overcoming radioresistance. We demonstrated the effect of miR-29c on radioresistance in lung cancer cells
Footnotes
Acknowledgements
The authors are grateful to Biologist Samuel Aréchaga for conducting statistical analyses and to undergraduate students Perla López, Mauricio Orozco, Liliana Nava, and Erick Valdés for their collaboration in producing experimental data. The authors also thank M.S. Christian Adame (Centro Medico Nacional “Siglo XXI,” México) for optimizing the radiotherapy dose for all experimental analyses. EAO is very grateful to Dr. Greco Hernández (Instituto Nacional de Cancerología, México) for his valuable critical comments about this project.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by Consejo Nacional de Ciencia y Tecnologia (CONACYT; 115591, 115552, and 181534), Instituto de Ciencia y Tecnologia del Distrito federal (ICYT-DF; PIUTE10-147), and Programa para el Desarrollo Profesional Docente (PRODEP-SEP; 47310447).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
