Abstract
Zinc-fingers and homeoboxes 1 (ZHX1) is a nuclear transcription repressor and known to be involved in cell differentiation and tumorigenesis. However, the pathophysiological roles of ZHX1 have not been characterized in glioblastoma. We examined ZHX1 expression in glioblastoma patients’ tissues and analyzed overall survival of the patients based on expression level of ZHX1. We also examined the effects of ZHX1 on proliferation and motility of glioblastoma cells. In silico analysis and immunohistochemical studies showed that the messenger RNA and protein expressions of ZHX1 were higher in the tissues of glioblastoma patients than in normal brain tissues, and that its overexpression was associated with reduced survival. In vitro, the downregulation of ZHX1 decreased the proliferation, migration, and invasion of glioblastoma cells, whereas its upregulation had the opposite effects. In addition, we showed ZHX1 could contribute to glioblastoma progression via the regulations of TWIST1 and SNAI2. Taken together, this study demonstrates that ZHX1 plays crucial roles in the progression of glioblastoma, and its findings suggest that ZHX1 be viewed as a potential prognostic maker and therapeutic target of glioblastoma.
Introduction
Glioblastoma is a common malignant brain tumor derived from astrocytes. 1 Despite therapeutic advances, brain tumors remain the most lethal form of brain disease. The incidence of glioblastoma is 3.19 per 100,000 in the United States and its 5-year survival rate is ~10%;2,3 even for radiotherapy plus temozolomide, median overall survival was only 14.6 months. 4 These survivals demonstrate that the identification of tumor targets related to the pathogenesis of glioblastoma is important in terms of the development of novel effective treatments.
Glioblastoma is a grade 4 astrocytoma. According to the World Health Organization (WHO) grading system, astrocytomas can be classified into four grades. Grade 1 astrocytomas grow slowly and can be treated curatively by surgical excision, whereas grade 2 astrocytomas tend to recur. 5 Grade 3 and 4 astrocytomas are considered severe and require gross total resection, chemotherapy, and radiation therapy. 5 However, grade 3 and 4 astrocytomas exhibit recurrence, invasiveness, and regenerative proliferation after surgical excision, which means that to understand the mechanism of glioblastoma progression and establish curative treatments, proliferation, migration, invasion, and chemosensitivity studies are required.
The zinc-fingers and homeoboxes 1 (ZHX1) belongs to the ZHX family, which is a group of nuclear transcription repressors. 6 ZHX1 was first discovered in a murine endothelial-adipose stromal cell line, and then found to be highly expressed in mouse brain, but barely expressed in liver or kidneys. 7 ZHX1 possesses several motifs, that is, a zinc-fingers region, a homeobox DNA binding region, a nuclear localization signal region, and an acidic region. 8 ZHX1 can form homodimers with itself or heterodimers with ZHX2 or ZHX3.9–11 ZHX1 regulates transcription by interacting with NF-YA, BS69, and DNMT3.12–14 Recently, studies have described the contrary roles for ZHX1 during tumorigenesis. Some studies have suggested tumor-suppressive roles of ZHX1. The proliferations of hepatic and gastric cancer cells were decreased by ZHX1.15,16 Moreover, ZHX1 inhibited the migration of gastric cancer cells.15,16 However, ZHX1 suppressed B-cell differentiation and was positively associated with lymphoid malignancy. 17 Thus, studies indicate ZHX1 plays different roles in different cancers.
In this study, we examined the relationship between the expression of ZHX1 and overall survival in glioblastoma patients, and investigated the roles played by ZHX1 in the proliferation, migration, and invasion of glioblastoma cells.
Materials and methods
Data analysis
Glioblastoma patient gene expression profiling data (GSE7696) was obtained from the National Center for Biotechnical Information (NCBI) Gene Expression Omnibus (GEO) database. The data included the results from normal (n = 4) and glioblastoma (n = 80) tissues. To analyze overall survival, the patients were classified into two groups according to ZHX1 messenger (mRNA) expression levels in glioblastoma tissues. Kaplan–Meier analysis and the log-rank test were performed using the R survival package.
Immunohistochemistry
A glioblastoma tissue array was obtained from US Biomax (Rockville, MD, USA). It contained 3 normal brain tissues, 2 cases of astrocytoma (WHO grade 2), and 33 cases of glioblastoma. On receipt, the array slide was deparaffinized and rehydrated to remove paraffin. Antigen retrieval was performed using sodium citrate buffer, and endogenous peroxidase was inactivated using 3% hydrogen peroxide for 15 min. The microarray was then treated with 1% bovine serum albumin (BSA) in tris-buffered saline (TBS) for 30 min, with anti-ZHX1 antibody (1:75, Protein Tech #13903-1-AP, Chicago, IL, USA) overnight at 4°C, horseradish peroxidase-conjugated secondary antibody for 40 min at room temperature, and then stained with 3,3′-diaminobenzidine (DAB; 1:50) and counterstained with hematoxylin for 3 min at room temperature. Histopathologic diagnoses of brain tumor were performed at the Department of Pathology, Pusan National University Hospital. The percentage of ZHX1-positive cells in given tissues was graded on a scale of 1–4 (1, <25%; 2, 25%–49%; 3, 50%–74%; and 4, 75%–100%). The intensity of ZHX1 was decided relative to that observed in adjacent brain tissues (0, 1, 2, 3 for basal, weak, moderate, and strong, respectively). The overall staining score was determined by a composite score (product of the above two scores, ranging from 0 to 12). Twofold increase in the staining score defined the overexpression of ZHX1.
Cell culture
U373MG and GBM05 glioblastoma cells were cultured in Roswell Park Memorial Institute (RPMI) medium 1640 or Dulbecco’s Modified Eagle Medium (DMEM), respectively, containing 10% fetal bovine serum (FBS), 25 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES), and 100 µg/ml of penicillin/streptomycin. Cells were incubated in 5% CO2 incubator at 37°C.
siRNA transfection and generation of a stable cell line
Scrambled (SCR) siRNA and ZHX1 siRNA were purchased from Dharmacon (Lafayette, CO, USA) and Bioneer Corporation (Daejeon, Korea). The siRNA sequences used were as follows: ZHX1 siRNA, 5′-CUGACUUUUGAUGGUAGUU(dTdT)-3′, 5′-GAAAGUAAUGCAGGUAGUU(dTdT)-3′, and SCR siRNA, 5′-CAGUUCAUCAUAACUCAGU(dTdT)-3′ and 5′-GAUCCGCAAAAGAGCGAAA(dTdT)-3′. For ZHX1 knockdown, glioblastoma cell lines were seeded at 1500 cells/well in a 96-well plate or 70,000 cells/well in a 6-well plate. Cells were transfected with 100 nM of SCR or ZHX1 siRNA within 1 day of seeding. DharmaFECT 1 (Thermo Scientific, Kafayette, CO, USA) was used for the transfection as directed by the manufacturer. The ZHX1 gene was obtained from Invitrogen and was amplified using ZHX1 primers including NotI and XbaI restriction site. The amplified ZHX1 gene and the 3XFlag CMV-10 vector (Sigma, E4401) were digested by NotI and XbaI restriction enzyme, and then ZHX1 was inserted into the vector. To create a stable cell line, U373MG cells were seeded at 100,000 cells/well in a six-well plate and transfected with empty vector as control or the vector containing the ZHX1 using ViaFectTM (Promega, Madison, WI, USA), according to the manufacturer’s instructions. G418 (30 µg/ml; Roche, 04727894001) was used to choose empty control vector-expressing (Mock) and ZHX1-overexpressing (ZHX1-over) clones.
Real-time polymerase chain reaction
Total RNAs were extracted using the RNeasy Mini kit (Qiagen, Valencia, CA, USA). cDNA was synthesized from purified RNA using oligo-dT, dNTP, RNasin, and MMLV reverse transcriptase (Promega). The cDNAs obtained were used as template for real-time polymerase chain reaction (PCR). The prepared cDNA, specific primers, and FastStart Essential DNA Green Master (Roche, Nonnenwald, Germany) were mixed together to determine relative ZHX1 mRNA expression. Real-time PCRs were performed on a LightCyclerTM 96 Real-Time PCR system (Roche). Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as the internal control. Three separate experiments were performed. The primer sequences for real-time PCR were as follows: GAPDH, 5′-TGG TGA CCA GGC GCC CAA TAC G-3′ and 5′-GCA GCC TCC CGC TTC GCT CT-3′; ZHX1, 5′-TCC CTT ACC CAA CAA TGT CA-3′ and 5′-TTG TTT CCT TCT TGC CTC CT-3′; SNAI1, 5′-GAG GCG GTG GCA GAC TAG-3′ and 5′-GAC ACA TCG GTC AGA CCA G -3′; SNAI2, 5′-TAG GAA GAG ATC TGC CAG AC-3′ and 5′-CCC CAA GGC ACA TAC TGT TA-3′; TWIST1, 5′-CGG GAG TCC GCA GTC TTA-3′ and 5′-TGG ATC TTG CTC AGC TTG TC-3′; TWIST2, 5′-CTT ATG TTT GGG GGG AGG TT-3′ and 5′-TAG CCA AGC AAT CAC GGA GA-3′.
Western blot
Cells were first lysed with radioimmunoprecipitation assay (RIPA) buffer (Thermo Scientific) containing β-mercaptoethanol and protease inhibitor cocktail. Total protein concentrations were determined using the Bio-Rad protein assay kit (Bio-Rad, Hercules, CA, USA). Protein samples (20 µg or 40 µg/well) were subjected to sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and separated proteins were then transferred to nitrocellulose membranes (Amersham Biosciences) overnight at 4 °C. Membranes were blocked with 5% nonfat dry milk in phosphate-buffered saline + 0.1% Tween-20 (PBS/T) for 1 h, and treated with ZHX1 or actin antibodies diluted in PBS/T containing 5% BSA overnight at 4 °C. Horseradish peroxidase–conjugated anti-mouse or anti-rabbit was used as the secondary antibody. A chemiluminescent agent (EZ-Western Detection Kit, Seoul, Korea) and the LAS-3000 imaging system (FUJIFILM) were used to detect ZHX1 and actin bands. The ImageJ program was used to quantify ZHX1 protein expressions. β-Actin was used as the internal control, and three separate experiments were performed. The antibodies used were anti-ZHX1 (1:1000; Protein Tech #13903-1-AP) and anti-β-actin (1:5000; Santa Cruz Biotechnology #sc-47778).
Cell proliferation assay
Glioblastoma cell proliferation was determined using the Ez-Cytox proliferation assay kit (ITSBIO, Seoul). Briefly, 3–5 days after SCR or ZHX1 siRNA transfection and after seeding empty control vector-expressing (Mock) cells or ZHX1-over cells, 10 µl of Ez-Cytox was added to each well and incubated for 1–2 h in a CO2 incubator. Optical density (OD) values were obtained at 450 nm using an enzyme-linked immunosorbent assay (ELISA) reader (TECAN, Mannedorf, Switzerland). SCR or Mock was used as the control to determine relative cell proliferations. Three separate experiments were performed.
Soft agar assay
Cells were prepared within 2 days after SCR or ZHX1 siRNA transfection. The base agar layer was composed of 0.7% soft agarose in culture medium containing 10% FBS, and the top agar layer, which contained glioblastoma cells (5 × 103), was composed of 0.35% soft agarose in culture medium containing 10% FBS. Colonies exceeding 40 µm in diameter were counted after 13 days of incubation. Experiments were performed in triplicate.
Boyden chamber assay
To examine FBS-induced cell migration, 50 µl of culture medium containing 10% FBS was added to bottom chambers, which were covered with collagen-coated membranes. Cells were diluted in culture medium containing 0.05% FBS and 5 × 104 cells (50 µl) were added to upper chambers. After 5–8 h, cells that had migrated through the membrane were fixed and stained using Diff-Quik solution (Sysmex, Kobe, Japan). Total numbers of migrated cells were counted using Adobe Photoshop CS6 software. Experiments were performed in triplicate.
Matrigel invasion assay
To examine FBS-induced cell invasion, we used 24-well BioCoatTM Matrigel chamber inserts (BD Bioscience, San Jose, CA, USA). Upper surfaces of inserts were covered with 30 µL of 0.5 mg/mL of growth factor–reduced Matrigel (BD Bioscience), and lower surfaces of inserts were covered with 20 µL of 0.5 mg/mL fibronectin (Sigma-Aldrich). Coated inserts were first rehydrated with culture medium for 4 h in a CO2 incubator. Within 1–2 days after SCR or ZHX1 siRNA transfection and immediately after preparing Mock or ZHX1-over cells, 5 × 104 cells in 500 µL of media containing 0.1% FBS were added to inserts. To observe chemotactic cell migration, 700 µL of culture medium containing 10% FBS was added to each well. Mitomycin C (100 or 300 ng/mL; Sigma-Aldrich) was used to inhibit glioblastoma cell proliferation. After incubation for 70 or 90 h, cells on upper surfaces of inserts were removed by scraping. Cells that had penetrated inserts were fixed and stained using Diff-Quik solution (Sysmex). Total numbers of invasive cells were counted using Adobe Photoshop CS6 software. Experiments were performed in triplicate.
Statistical analysis
Results are presented as means ± standard error of the means (SEMs). Student’s t-test was used to determine the significances of differences, and p values < 0.05 were considered statistically significant.
Results
Expression of ZHX1 in glioblastoma tissues
To analyze the expression of ZHX1 in glioblastoma tissues, we first obtained a tissue microarray data from the GEO database and analyzed ZHX1 expressions. The analysis showed that ZHX1 mRNA levels were greater in glioblastoma tissues than in normal brain tissues (Figure 1(a), n = 80, Student’s t-test p = 0.008). To examine ZHX1 protein expression, immunohistochemistry was performed using a paraffin-embedded glioblastoma tissue array. Normal brain, normal adjacent, and glioblastoma tissues were stained with ZHX1 antibody. ZHX1 was found to be overexpressed in glioblastoma tissues as compared with normal tissues or adjacent non-cancerous tissues (Figure 1(b); n = 19/33). Next, we examined whether ZHX1 overexpression was correlated with patient outcome using data from the GEO database. Kaplan–Meier analysis showed that patients with ZHX1-overexpressing glioblastoma had significantly poorer survival (Figure 1(c), log-rank test, p = 0.005).

ZHX1 was overexpressed in glioblastoma and its expression was related to patient prognosis. (a) RNA microarray data sets (GSE7696) were obtained from the NCBI Gene Expression Omnibus. ZHX1 mRNA expression in glioblastoma tissues (n = 80, p = 0.008) was compared with that in normal brain tissues (n = 4). (b) Immunohistochemistry showed ZHX1 protein was overexpressed in glioblastoma tissues as compared with normal brain tissues. The scale bar represents 20 µm. (c) Patients with glioblastoma were divided into two groups based on ZHX1 expression level (low or high). A Kaplan–Meier curve was plotted and the log-rank test was performed using the R survival software package (p = 0.005).
Role of ZHX1 in glioblastoma proliferation
To find the functional roles of ZHX1 in glioblastoma cells, we performed loss-of-function and gain-of-function assays using specific siRNA and cDNA, respectively. Initially, basal ZHX1 mRNA expression levels in five glioblastoma cell lines (U373MG, GBM05, U87MG, T98, and A176) were similar as determined by real-time PCR (data not shown). Next, ZHX1 was knocked down by siRNA transfection in U373MG and GBM05 cells and upregulated by ZHX1 transfection in U373MG (ZHX1-over) cells. Knockdown and overexpression efficiencies were determined by real-time PCR and Western blot. ZHX1 siRNA decreased the mRNA level of ZHX1 by 95.3% and 85.9%, in U373MG and GBM05, respectively, versus SCR siRNA (Figure 2(a)). The protein levels of ZHX1 in U373MG and GBM05 were also reduced by ZHX1 siRNA versus SCR siRNA by 72.2% and 54.9%, respectively (Figure 2(b) and (c)). In ZHX1-over cells, the mRNA and protein levels of ZHX1 were higher than in Mock cells by 327.6% and 320.7%, respectively (Figure 2(a) and (c)). The effect of ZHX1 on glioblastoma cell proliferation was investigated using ZHX1 knocked-down and ZHX1-over cells. Cell proliferation was measured 3–5 days after SCR or ZHX1 siRNA transfection and plating Mock or ZHX1-over cells. The proliferations of U373MG and GBM05 were significantly reduced by ZHX1 siRNA versus SCR siRNA by 30% and 19%, respectively (Figure 2(d)). In contrast, the proliferation of ZHX1-over cells was significantly increased versus Mock cells by 122.9% (Figure 2(d)). To further confirm the effect on the proliferation, we examined effects of knockdown or overexpression of ZHX1 on the cell cycle. ZHX1 knockdown induced an increase in U373MG and GBM05 cells at G0/G1 phase and a decrease in G2/M phase. ZHX1 overexpression in U373MG showed the opposite results (Supplementary Figure 1). Moreover, the change in the level of CCNB1 (cyclin B), an essential molecule regulating G2/M phase progression of the cell cycle, by modulation of ZHX1 supported the result obtained from the cell cycle experiment (Supplementary Figure 1(c)). Next, to examine the effects of ZHX1 on the anchorage-independent growth of glioblastoma cells, soft agar assay for colony formation was performed. Downregulation of ZHX1 significantly reduced the colony numbers of U373MG and GBM05 cells versus SCR siRNA by 71.1% and 47.5%, respectively (Figure 2(e) and (f)).

ZHX1 increased the proliferation of glioblastoma cells. (a) Efficiencies of ZHX1 knockdown and overexpression were determined by real-time PCR. For knockdown, glioblastoma cells were treated with 100 nM SCR or ZHX1 siRNA. SCR siRNA-transfected cells were used as controls to normalize ZHX1 mRNA levels. For ZHX1 overexpression, we used ZHX1-overexpressing U373MG (ZHX1-over) cells and empty control vector-expressing (Mock) cells as controls to calculate relative ZXH1 mRNA levels. GAPDH was used as an internal control. (b) The efficiencies of ZHX1 protein knockdown and overexpression were determined by Western blot. (c) Relative ZHX1 protein levels were quantified using β-actin as the internal control. Values from SCR siRNA-transfected cells and Mock cells were used as controls to calculate relative ZHX1 protein levels. (d) Cell proliferations were measured 3–5 days after SCR or ZHX1 siRNA transfection and 3–5 days after seeding Mock or ZHX1-over cells. SCR siRNA-transfected and Mock controls were used to calculate relative cell proliferations. (e) Effects of ZHX1 knockdown on colony formation in soft agar were examined. U373MG and GBM05 cells were resuspended in 3.5% soft agar 2 days after SCR or ZHX1 siRNA transfection. (f) Colonies were counted and the results obtained are presented as bar graphs, which represent the means ± SEMs of three independent experiments.
Roles of ZHX1 during the migration and invasion of glioblastoma cells
To investigate the effect of ZHX1 on cell motility, we used a Boyden chamber assay. U373MG and GBM05 cells were transfected with SCR or ZHX1 siRNA and the cells were tested 2 days after transfection. ZHX1 siRNA treatment of U373MG and GBM05 inhibited cell migration as compared with SCR siRNA (Figure 3(a)). Numbers of migrated cells were significantly lower for ZHX1 siRNA than for SCR siRNA (54.3% vs 36.2%, respectively) (Figure 3(b)). In contrast, ZHX1 overexpression promoted U373MG migration (Figure 3(c)). The quantification showed that the migration of ZHX1-over cells was significantly greater than that of Mock cells by 151.2% (Figure 3(d)). A cell invasion assay using Matrigel-coated inserts showed knockdown of ZHX1 suppressed the invasivenesses of U373MG and GBM05 versus SCR by 69.2% and 67%, respectively (Figure 4(a) and (b)). In contrast, the overexpression of ZHX1 in U373MG increased the number of invaded cells versus Mock controls by 159.9% (Figure 4(c) and (d)).
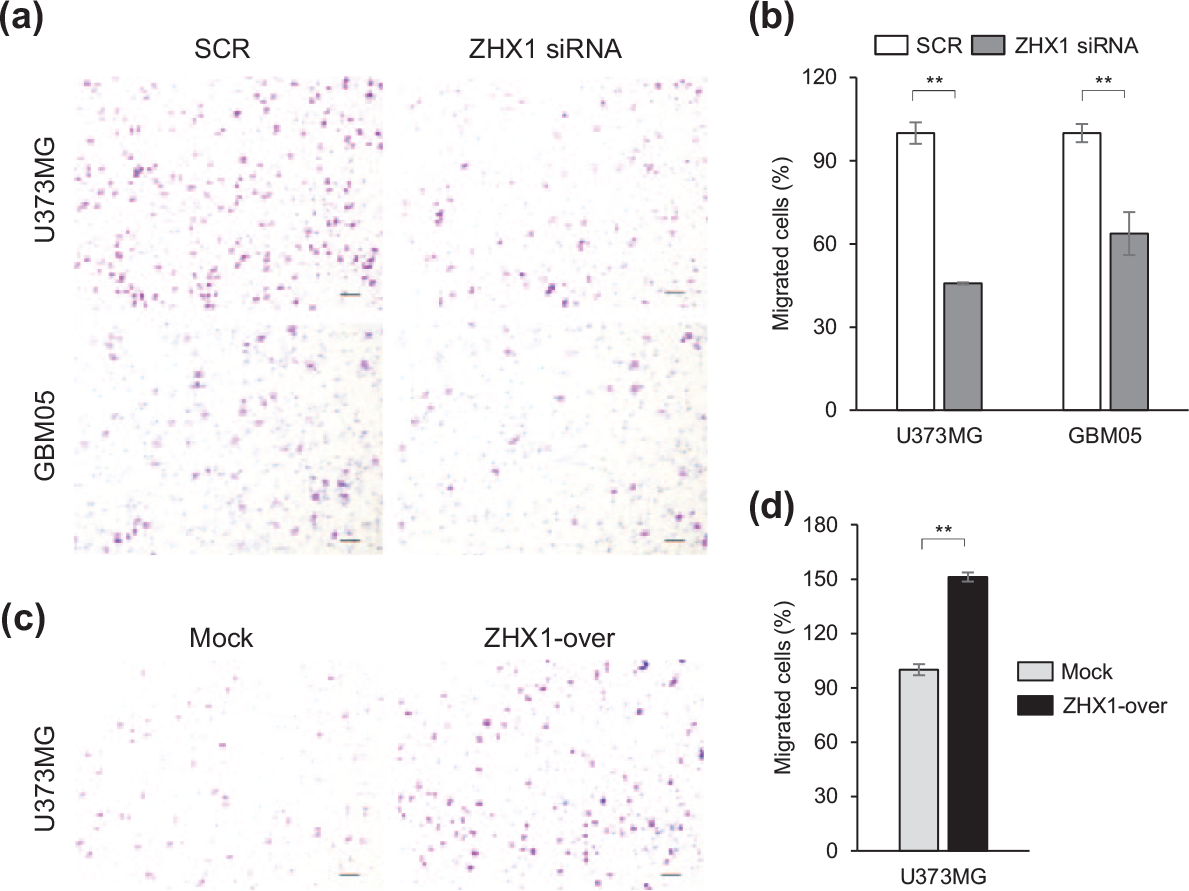
ZHX1 promoted the migration of glioblastoma cells. The migration of glioblastoma cells was examined using a Boyden chamber assay. Migration assays were performed within 2 days of SCR or ZHX1 siRNA transfection and within 2 days after plating Mock or ZHX1-over cells. (a) ZHX1 siRNA treatment suppressed the FBS-induced migrations of U373MG and GBM05 cells versus the SCR siRNA controls. (b) Number of migrated glioblastoma cells were counted and SCR siRNA-treated controls were used as controls to calculate the relative migration rates of ZHX1-knockdown cells. Results are shown as a bar graph and are the means ± SEMs of three independent experiments. (c) ZHX1 overexpression increased the FBS-induced migration of U373MG cells versus the Mock control. (d) Migrated cells were counted and results are presented as bar graphs.

ZHX1 accelerated the invasion of glioblastoma cells. Matrigel invasion assays were performed 2 days after transfecting U373MG and GBM05 cells with SCR or ZHX1 siRNA and 2 days after plating Mock or ZHX1-over cells. (a) ZHX1 siRNA inhibited the FBS-induced invasion of U373MG and GBM05 cells versus SCR siRNA. (b) Numbers of invaded cells were counted and SCR siRNA-treated cells were used as controls to calculate relative invasion rates. Results are the means ± SEMs of three independent experiments. (c) Upregulation of ZHX1 promoted the invasion of U373MG cells versus Mock controls. (d) Numbers of invaded cells were counted and Mock cells were used as controls to calculate the relative invasion rate of U373MG cells.
Regulation of epithelial mesenchymal transition–related genes by ZHX1
Previous studies have shown that epithelial mesenchymal transition (EMT)-related genes, such as, SNAI1, SNAI2, and TWIST1, are involved in the proliferation, migration, and invasion of glioblastoma cells.18–20 Because ZHX1 was found to promote the migration and invasion of glioblastoma cells, we investigated whether ZHX1 could regulate EMT-related genes. Using real-time PCR, we examined changes in the mRNA levels of EMT-related genes induced by ZHX1 knockdown or overexpression. ZHX1 knockdown significantly reduced the expression levels of SNAI1, SNAI2, TWIST1, and TWIST2 in U373MG and GBM05 cells (Figure 5), whereas ZHX1 overexpression significantly increased the mRNA levels of SNAI2 and TWIST1, but not those of SNAI1 and TWIST2, in U373MG cells (Figure 5).

ZHX1 regulated EMT-related genes in glioblastoma cells.
Discussion
The pathophysiological roles of ZHX1 during tumorigenesis are poorly characterized. This study shows for the first time that ZHX1 is overexpressed in glioblastoma and that its expression is correlated to patient survival. It also shows ZHX1 increased the proliferation, migration, and invasion of glioblastoma cells, and that the expression levels of TWIST1 and SNAI2 were regulated by ZHX1.
ZHX1 expression is dependent on cancer type. ZHX1 expression was lower in cancerous liver tissues than in paired adjacent cirrhotic liver tissues and lower in gastric cancer tissues than in paired normal tissues.15,16 However, in this study, ZHX1 mRNA expression was higher in glioblastoma tissues than in normal tissues (Figure 1(a)), and ZHX1 protein was more highly expressed in glioblastoma tissues (Figure 1(b)). A previous report showed an inverse correlation between ZHX1 and ZHX2 expression levels in B cells. 17 Therefore, we examined the expression levels of ZHX1 and ZHX2 after the modulation of ZHX1. However, we did not observe such an inverse correlation (Supplementary Figure 2).
The roles of ZHX family members in cell proliferation were first discovered during studies on ZHX2, which was found to inhibit the proliferation of hepatocellular carcinoma cells by stopping the cell cycle at G1. 21 In Hodgkin’s lymphoma, ZHX2 knockdown reduced the expressions of differentiation- and apoptosis-promoting genes, suggesting ZHX2 acted as tumor suppressor. 22 Furthermore, ZHX1 overexpression in hepatocellular carcinoma cells (SMMC-7721) suppressed proliferation and its knockdown in gastric cancer cells (SGC-7901-SH) increased proliferation.15,16 However, other studies have reported that ZHX1 may be an oncogene. In one study, interleukin-2 upregulated ZHX1 expression, which then increased the proliferation of CTLL-2 cells (a cytotoxic T-cell line), 23 and in Hodgkin’s lymphoma, ZHX1 suppressed B-cell differentiation and was positively associated with lymphoid malignancy. 17 In this study, upregulation of ZHX1 increased the proliferation of glioblastoma cells and its downregulation had the opposite effect (Figure 2(d)). These results were consistent with the changes in the cell cycle and the level of CCNB1 by ZHX1 modulation (Supplementary Figure 1). In addition, the anchorage-independent growth of glioblastoma cells was decreased after ZHX1 siRNA treatment (Figure 2(e) and (f)). Taken together, these observations suggest ZHX1 promotes the proliferation of glioblastoma cells.
The role of ZHX1 in the proliferation of glioblastoma cells we found in this study is contradictory to the previous results.16,24 Ma et al. 16 reported that ZHX1 inhibits gastric cancer cell growth by inducing cell-cycle arrest and apoptosis. However, there are many previous reports indicating that the roles of a specific gene during the cancer progression can be different depending on the cancer type. For example, Notch signaling is well known as an oncogenic signaling in some cancers including gastric cancer.25,26 However, in glioblastoma, Sun et al. 27 have reported that Notch signaling showed tumor-suppressive effects. Moreover, Bcl-2 is upregulated in many cancers, and then plays an essential role in cell proliferation and cancer formation.28,29 However, overexpression of Bcl-2 in human lymphoma was associated with a decrease in proliferation. 30 Even in the same cancer cells, a molecule can have dual roles of tumor progression and suppression in time or environment-dependent manner.31,32 In addition, Wang et al. 24 reported that MiR-199a-3p is upregulated in gastric cancer tissues and that it promotes gastric cancer progression by targeting ZHX1. However, Shen et al. 33 showed that the miR-199a-3p was downregulated in glioma tissues compared to normal brain tissues, and suppressed proliferation of glioma cells. These results suggest that a specific gene can regulate cancer progression in the cell context–dependent manner.
Cell migration and invasion are considered causes of treatment failure and tumor recurrence in glioblastoma. In this study, ZHX1 accelerated the motility and invasiveness of glioblastoma cells (Figure 3(c) and (d) and Figure 4(c) and (d)), and ZHX1 knockdown suppressed glioblastoma cell migration and invasion, whereas its overexpression had the opposite effects (Figure 3(a) and (b) and Figure 4(a) and (b)). Our results are consistent with a report that ZHX1 levels were reduced by Pea3 knockdown and that this was associated with reduced invasiveness and motility. 34 Thus, our study suggests that ZHX1 overexpression may be correlated with treatment resistance and tumor recurrence.
Furthermore, this study suggests a relationship exists between EMT and ZHX1, as our ZHX1 knockdown and overexpression studies showed that ZHX1 regulated TWIST1 and SNAI2 (both EMT-related genes; Figure 5). EMT is a process that induces a mesenchymal cell phenotype and promotes migratory and invasive characteristics. 35 In glioblastoma, SNAI1 knockdown inhibited the growth, migration, and invasion of U87MG and GBM05 cells, and SNAI2 overexpression increased the proliferation, migration, and invasion of U251 cells.18,19 In addition, TWIST1 overexpression enhanced the invasivenesses of SNB19 and T98G cells. 20 In addition, Koh et al. 36 recently suggested TWIST2 might be functionally associated with rhabdoid glioblastoma. Thus, our findings suggest that ZHX1 can contribute to glioblastoma progression via the regulations of TWIST1 and SNAI2. This result is consistent with results showing the regulation of cyclin B1 by ZHX1 (Supplementary Figure 1) and previous studies showing that SNAI2 or TWIST1 overexpression increased cyclin B1 expression and then induced EMT and cell growth.37,38
In conclusion, ZHX1 was found to be overexpressed in glioblastoma and to promote the proliferation, migration, and invasiveness of glioblastoma cells, possibly via the regulations of TWIST1 and SNAI2. Importantly, ZHX1 overexpression in glioblastoma was observed to be related to a reduction in overall survival time. Our results suggest ZHX1 be considered a potential prognostic marker in glioblastoma and a novel target for the development of therapeutics.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the Medical Research Center (MRC) Program (#NRF-2015R1A5A2009656) and Basic Science Research Foundation (#NRF-2016R1A2B4014593 and #NRF-2015R1A1A3A04001322) of the National Research Foundation (NRF) funded by the Korean government (MEST).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
