Abstract
We read with great interest the paper entitled “Interferon gamma +874 T/A polymorphism increases the risk of cervical cancer: evidence from a meta-analysis” published online by Sun et al. Their results suggest that interferon gamma (IFNG) gene +874 T/A polymorphism might contribute to women’s susceptibility to cervical cancer. They also found that IFNG +874 T/A polymorphism is associated with increased cervical cancer risk in Asian female population. The result is encouraging. Nevertheless, several key issues are worth noticing. We re-evaluate the association between IFNG +874 T/A polymorphism and cervical cancer risk by performing an updated meta-analysis based on 2777 cases and 2542 controls of 11 studies. We found that IFNG +874 T/A polymorphism was not significantly associated with cervical cancer risk in overall population. We also observed that the polymorphism was associated with enhanced cervical cancer risk in Asian population and was relevant to increased squamous cell cervical cancer risk.
Dear Editor
With great interest, we recently read the article entitled “Interferon gamma +874 T/A polymorphism increases the risk of cervical cancer: evidence from a meta-analysis” published online by Sun et al. 1 In the paper, the authors conducted a meta-analysis to assess the association between interferon gamma (IFNG) gene +874 T/A polymorphism (rs2430561) and cervical cancer risk on the basis of a total of nine studies published before January 10, 2014, including 1116 cases and 1290 controls. In overall population, they observed that IFNG +874 T/A polymorphism was associated with elevated cervical cancer risk in the dominant genetic model (TA + AA vs TT) and in the TA versus TT model, with odds ratio (OR) (95% confidence interval (CI)) being equal to 1.399 (1.097–1.784) and 1.471 (1.137–1.903), respectively. In their subsequent stratified analyses by ethnicity, Sun et al. found significant association of IFNG +874 T/A variant with enhanced cervical cancer risk in Asian population (OR (95% CI) = 1.455 (1.062–1.933) in the dominant model and 1.494 (1.069–2.087) for TA vs TT model). The findings provided by Sun et al. are encouraging.
Nevertheless, after careful examination of the data supplied by Sun et al. (shown in Table 2 in their original text), 1 we found that several key issues should be noticed. First, two eligible case-control studies concerning IFNG +874 T/A polymorphism and cervical cancer risk,2,3 published before 2014, were not included in Sun et al.’s analysis. 1 Second, three studies included in Sun et al.’s paper examined the association between IFNG +874 T/A polymorphism and cervical intraepithelial neoplasia (CIN) risk. It is widely recognized that CIN is a precancerous lesion of cervical cancer; however, not all patients with CIN will eventually develop into cervical cancer, and thus, it is appropriate to explore the relationship of IFNG +874 T/A polymorphism with cervical cancer and/or CIN risk, respectively. Third, two eligible studies about IFNG +874 T/A polymorphism and cervical cancer risk have been published after 2014.4,5 Therefore, the conclusions induced from Sun et al.’s data are not entirely reliable. It is necessitated to summarize the association between IFNG +874 T/A polymorphism and cervical cancer/CIN risk more comprehensively and objectively.
A comprehensive search was carried out in the databases of MEDLINE/PubMed, Science Direct, Embase, China National Knowledge Infrastructure (CNKI), and Wanfang Medical Online with a combination of the following terms: “cervical cancer,” “cervical neoplasm” or “cervical carcinoma” and “Interferon-gamma” or “IFN-γ” or “IFNG” or “rs2430561” and “polymorphism” or “variant.” The references cited in the publications and review articles were also manually searched. Last search was updated on September 30, 2016.
Data inclusion criteria were as follows: (1) the papers reporting cervical cancer risk and IFNG +874 T/A polymorphism, (2) case-control studies or cohort studies, and (3) sufficient data to estimate the OR and 95% CI. For overlapping studies, the results including more information were included. Accordingly, papers lacking essential information and review articles were excluded.
We reassess the association between IFNG +874 T/A polymorphism and cervical cancer/CIN risk by performing an updated meta-analysis on the basis of a total of 14 studies containing 3068 cases and 2755 controls.2–13 The Cochrane Q statistics test was used to examine the heterogeneity among studies. A fixed-effect model or a random-effect model was adopted to estimate the combined effects according to the result of a heterogeneity test. 14 A fixed-effect model is chosen while the effects are assumed to be homogeneous; otherwise, a random-effect model is used. We first examine the relationship between IFNG +874 T/A polymorphism and cervical cancer/CIN risk, that is, pooling the cervical cancer and CIN cases together. Then, subgroup meta-analyses were conducted to investigate the association of IFNG +874 T/A polymorphism with cervical cancer risk or CIN risk, respectively. Further stratified analyses were also carried out by ethnicity, Hardy–Weinberg equilibrium (HWE) of the control group, and histological types of cervical cancer. Cumulative meta-analysis was conducted to investigate the tendency of results by accumulating single study year by year, which could be used to determine whether or not new relevant studies are needed. Sensitivity analysis was implemented via sequentially removing each study per time from the meta-analysis, which was employed to evaluate the stability of the results of current meta-analysis. Deviation from HWE in control subjects may bias the estimates of genetic effects in genetic association studies and meta-analysis 15 ; thus, the Pearson chi-square (χ2) test was used to examine whether the genotype distribution in each control group is deviated from HWE. The funnel plot was drawn to assess publication bias visually. In addition, Begg’s test and Egger’s test were used to examine the publication bias.16,17 All the statistical analyses were performed on the STATA/SE 10.1 software package (Stata Corporation, College Station, TX). Statistical significance was determined as a two-sided p value less than 0.05 for any test or model.
The general characteristics of the included studies in this meta-analysis were shown in Table 1. The summary ORs for the association of IFNG +874 T/A polymorphism with cervical cancer/CIN risk were listed in Table 2. Overall, we found a significant association between IFNG +874 T/A polymorphism and cervical cancer/CIN risk in the dominant genetic model, with OR (95% CI) being equal to 1.262 (1.007–1.581; as shown in Figure 1 and Table 2). However, no significant associations of IFNG +874 T/A polymorphism with cervical cancer risk or CIN risk were observed, when stratified meta-analysis by disease category (dividing into cervical cancer and CIN) was performed, with OR (95% CI) being 1.240 (0.952–1.615) for cervical cancer and being 1.382 (0.837–2.282) for CIN in the dominant model (Table 2). Considering that the majority of the included studies explored the association between IFNG +874 T/A polymorphism and cervical cancer risk, we concentrated our analysis on this association based on 11 studies with 2777 cases and 2542 controls (Table 1). In the subsequent subgroup analysis by ethnicity of cervical cancer cases, we observed that IFNG +874 T/A polymorphism was significantly associated with elevated cervical cancer risk in Asian population but not in Caucasian and mixed population in the dominant model, with OR (95% CI) being equal to 1.390 (1.032–1.873), 1.221 (0.567–2.630), and 1.333 (0.879–2.021), respectively (Table 2). The finding in Asian population is similar to the result of Sun et al. 1 In the stratified analysis by histological types of cervical cancer, we found that IFNG +874 AA+TA genotypes were relevant to enhanced squamous cervical cancer risk, with OR (95% CI) being 1.566 (1.178–2.082) (Table 2). No significant associations between IFNG +874 T/A polymorphism and cervical cancer risk were found in stratified analysis based on the studies whose control groups were in HWE (Table 2).
Characteristics of the included studies of the meta-analysis of INFG +874 T/A polymorphism and cervical cancer risk.
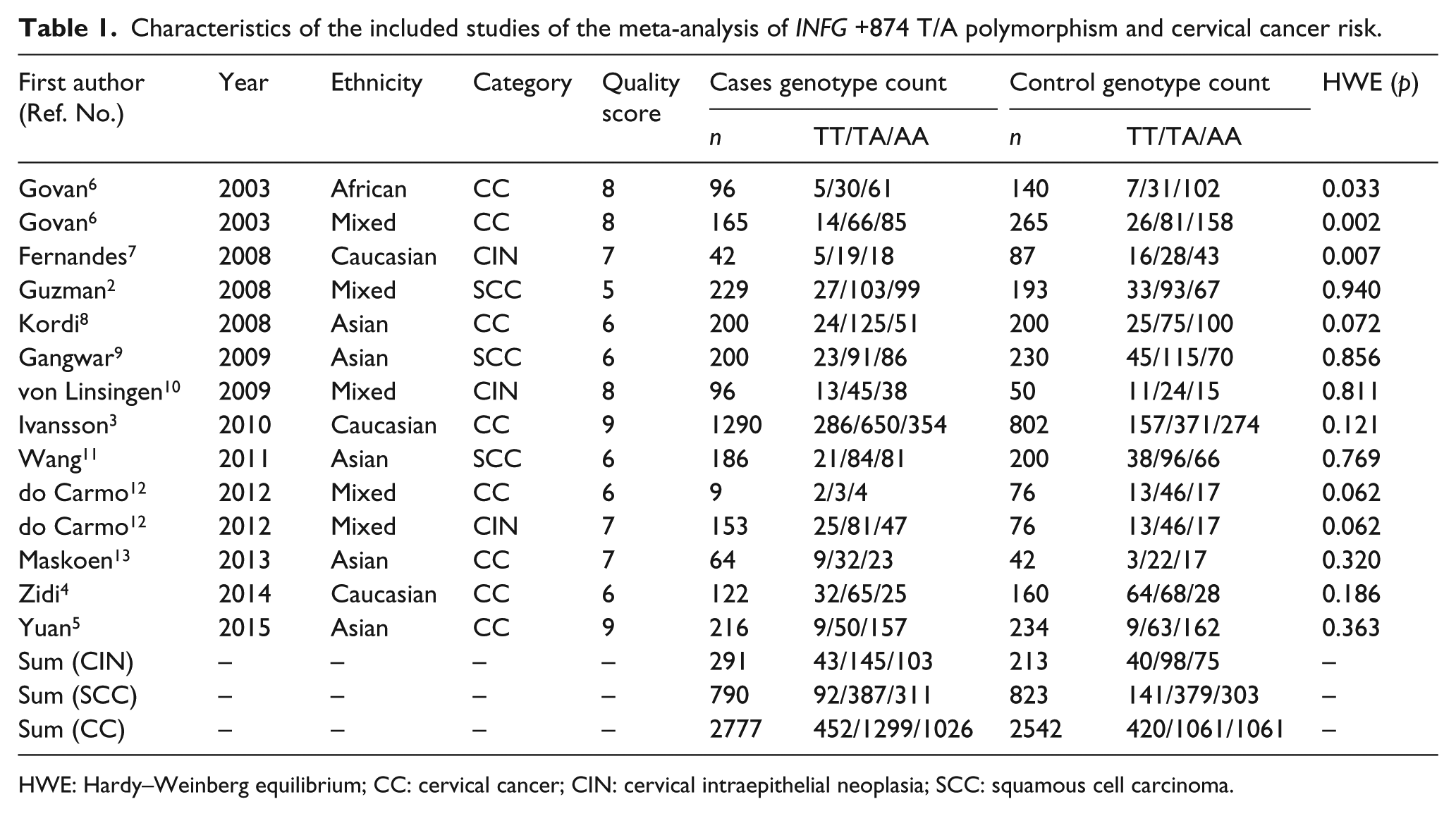
HWE: Hardy–Weinberg equilibrium; CC: cervical cancer; CIN: cervical intraepithelial neoplasia; SCC: squamous cell carcinoma.
Summary ORs for the association between INFG +874 T/A variant and cervical cancer or cervical intraepithelial neoplasia risk.
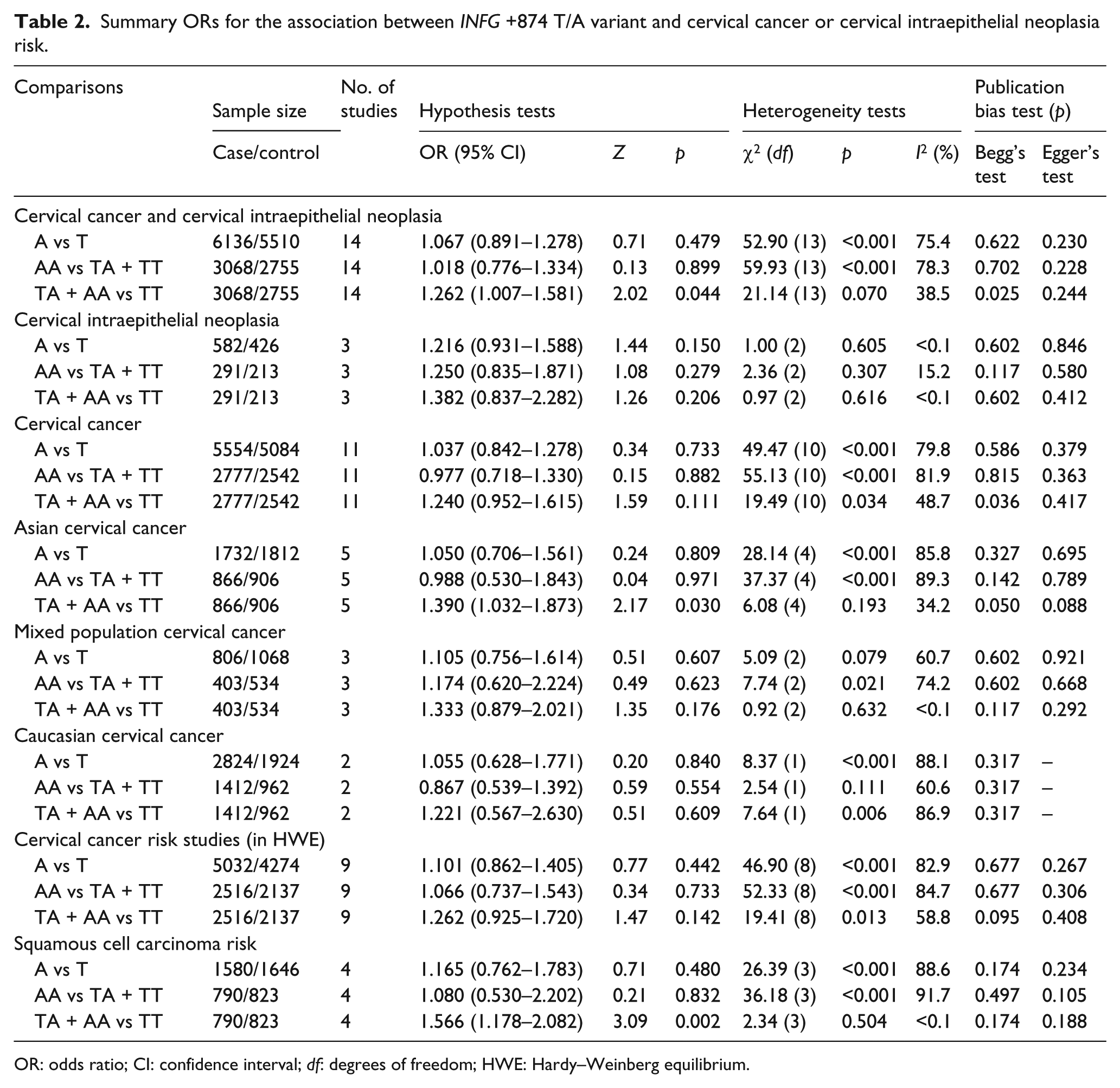
OR: odds ratio; CI: confidence interval; df: degrees of freedom; HWE: Hardy–Weinberg equilibrium.

Forest plots for the association between IFNG +874 T/A polymorphism and cervical cancer or cervical intraepithelial neoplasia risk.
The cumulative meta-analysis accumulated the studies in the sequence of year of publications. As illustrated in Figure 2, significant association between IFNG +874 T/A polymorphism and cervical cancer risk was observed for the first time in 2009, when Gangwar et al.’s study was published and a meta-analysis was performed then. We also examined that the summary ORs became statistically insignificant when Ivansson et al.’s study was published in 2010; however, there seemed to be a tendency of significance of association between IFNG +874 T/A polymorphism and cervical cancer risk in the dominant model (Figure 2). Sensitivity analysis was performed to evaluate the stability of the result of the present meta-analysis. As shown in Figure 3, all studies except one did not alter the result of being no significant association between IFNG +874 T/A polymorphism and cervical cancer risk. The exception was Ivansson et al.’s study. The result displayed that the association between IFNG +874 T/A polymorphism and cervical cancer risk became statistically significant when Ivansson et al.’s study was excluded from the current meta-analysis (Figure 3). This seemed to explain why Sun et al.’s study provided a positive association 1 ; as mentioned above, Ivansson et al.’s study was erroneously not included in Sun et al.’s meta-analysis. Meanwhile, the results of sensitivity analysis also revealed that Ivansson et al.’s study seemed to be a main source of heterogeneity of our current meta-analysis, because the statistical significance of heterogeneity (χ2 = 19.49, df = 10, p = 0.034, I2 = 48.7%; Table 2 and Figure 1) altered to be insignificant (χ2 = 8.66, df = 9, p = 0.470, I2 < 0.1%) and the OR (95% CI) was 1.432 (1.155–1.776) in the fixed-effect model for IFNG +874 AA + TA versus TT when Ivansson et al.’s study was excluded from our present meta-analysis.

Cumulative meta-analysis for the association between IFNG +874 T/A polymorphism and cervical cancer risk.

Sensitivity analysis for the association between IFNG +874 T/A polymorphism and cervical cancer risk.
The shape of the funnel plot seemed to be approximately symmetrical among total population (Figure 4). Both Begg’s and Egger’s tests suggested that there was no significant publication bias in current meta-analysis with the exception of the ORs for TA + AA versus TT in overall and cervical cancer comparison group (p = 0.025 and 0.036 in Begg’s test); however, the OR for TA + AA versus TT in cervical cancer risk comparison was not statistically significant, as shown in Table 2. These results indicated that publication biases might have few effects on our current meta-analysis.
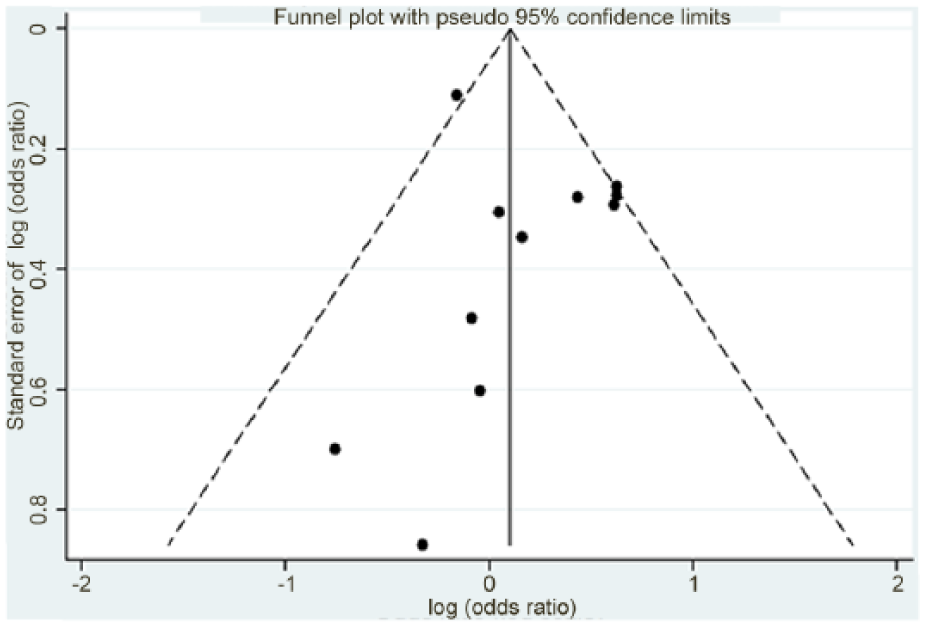
Funnel plot of the association between IFNG +874 T/A polymorphism and cervical cancer risk.
In conclusion, the results supplied by Sun et al. 1 should be taken into account with caution. Further well-designed studies with large sample size are needed to reach a definitive conclusion about the relationship between IFNG +874 T/A polymorphism and cervical cancer risk. We hope our remark will help for more accurate elaboration and substantiation of the findings presented by Sun et al. 1
Footnotes
Acknowledgements
H.Y. and M.Y. contributed equally to this article.
Declaration of Conflicting Interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was financially supported by the scientific research project of Hubei Provincial Department of Education (No. Q20152008).
