Abstract
Objectives
To assess the association between polymorphism in the interleukin (IL)-10 promoter region of 1082 G/A and the risk of cervical cancer and/or cervical intraepithelial neoplasia (CIN), using meta-analysis.
Methods
The electronic literature databases PubMed®, Embase®, Web of Science, CBMdisc and CNKI were searched for relevant studies. The strength of association between IL-10 gene −1082 G/A polymorphism and cervical cancer and/or CIN was measured using pooled odds ratios with 95% confidence intervals in four genetic models: allelic model (A allele versus G allele); additive model (A/A versus G/G); recessive model (A/A versus G/A+G/G); dominant model (A/A+G/A versus G/G).
Results
Eight studies involving 1983 cases and 1618 controls were identified and included in the meta-analysis. No significant associations were found between IL-10 gene −1082 G/A polymorphism and cervical cancer and/or CIN in any of the genetic models.
Conclusions
IL-10 gene −1082 G/A polymorphism does not appear to be associated with the risk of cervical cancer and/or CIN.
Introduction
Cervical cancer is the second most common malignant tumour among women and is thought to arise as the result of a combination of host and environmental factors. 1 Persistent infection with high-risk human papilloma virus (HPV) plays a key role in the development and progression of cervical intraepithelial neoplasia (CIN), which can develop into squamous cell carcinoma of the cervix. 2 It is likely that host genetic differences influencing the response to viral infection may determine the risk of developing cervical lesions and progression to invasive carcinoma, suggesting that polymorphisms present in genes related to viral infection, immune response or DNA repair systems or in tumour suppressor genes may influence the risk of developing cervical cancer. 3
Interleukin (IL)-10, a multifunctional cytokine, plays a key role in the inflammatory response associated with the progression of HPV infection and neoplasia or cancer formation. 4 Studies have reported that polymorphism in the IL-10 promoter region of –1082 G/A is associated with IL-10 gene transcription and secretion. 5 A number of epidemiological studies have focused on the association between IL-10 gene –1082 G/A polymorphism and the risk of cervical cancer and/or CIN. However, the findings have been inconsistent. The present meta-analysis was designed to give a more precise estimation of the strength of this association.
Materials and methods
Identification of studies
This meta-analysis followed the preferred reporting items for systematic reviews and meta-analyses (PRISMA) criteria. 6 Eligible studies published up to 2 December 2013 were identified by searching the electronic literature databases PubMed®, Embase®, Web of Science, CBMdisc and CNKI. The following keywords and search strategies were used: (“interleukin-10” OR “IL-10”) AND (“polymorphism” OR “mutation” OR “variant” OR “genotype”) AND (“cervical cancer” OR “cervical intraepithelial neoplasia” OR “CIN”). In addition, reference lists of reviews or original articles analysing the association between IL-10 gene −1082 G/A polymorphism and cervical cancer and/or CIN were also searched, to identify additional relevant articles. No language restrictions were set in this meta-analysis.
Inclusion criteria
To be included, studies had to meet all of the following criteria: (i) human studies; (ii) studies on the relationships between IL-10 gene –1082 G/A polymorphism and cervical cancer and/or CIN; (iii) unrelated case–control studies; (iv) genotype frequency details provided; (v) not republished data.
Data extraction
Data were extracted from eligible studies by two authors (S. Z. and Y.W.Y.) independently in duplicate, using a standard data collection form. The results were reviewed by a third author (Y-L.K.). The following information was collected from each study: last name of the first author; publication year; ethnicities and countries of participants; source of controls; number of cases and controls; genotype and allele data; the result of Hardy–Weinberg equilibrium (HWE) testing in the control group (P < 0.05 was considered to be a significant deviation from the HWE).
Meta-analysis
The strength of associations between IL-10 gene −1082 G/A polymorphism and cervical cancer and/or CIN were measured using odds ratios (ORs) with 95% confidence intervals (CIs). Pooled ORs were combined for the allelic model (A allele versus G allele), the additive model (A/A versus G/G), the recessive model (A/A versus G/A+G/G) and the dominant model (A/A+G/A versus G/G). Between-study heterogeneity was assessed using Cochran’s Q statistic and the I2 statistic, with a Pheterogeneity < 0.1 or I2 > 50% indicating the existence of heterogeneity.7,8 If between-study heterogeneity existed, the random-effects model was used to calculate the pooled ORs; otherwise the fixed-effects model was used.9,10
Subgroup analysis was performed, based on ethnicity. In addition, sensitivity analysis was conducted to confirm the reliability of combined analyses (based on studies showing HWE only). Publication bias was assessed using Begg’s funnel plot and Egger’s regression test (P < 0.05 was considered to be representative of statistically significant publication bias). 11 All statistical analyses were performed using STATA® software version 11.0 (StataCorp LP, College Station, TX, USA).
Results
Characteristics of eligible studies
The study selection process is shown in Figure 1. In total, eight articles containing a total of 1983 cases and 1618 controls were identified that met the inclusion criteria.12–19 Three studies were performed in Europeans, four in Asians, and one in Africans. Subgroup analysis based on ethnicity was performed for Asians and non-Asians. The genotype distributions among the controls of all studies followed the HWE except for the study of Yu et al.
17
The studies and their main characteristics are given in Table 1.
Flow diagram of the study selection process in a meta-analysis, undertaken to assess the association between polymorphism in the interleukin-10 promoter region of 1082 G/A and the risk of cervical cancer and/or cervical intraepithelial neoplasia. Main characteristics of studies included in a meta-analysis of the association between interleukin-10 gene –1082 G/A polymorphism and cervical cancer and/or cervical intraepithelial neoplasia. HWE, Hardy–Weinberg equilibrium.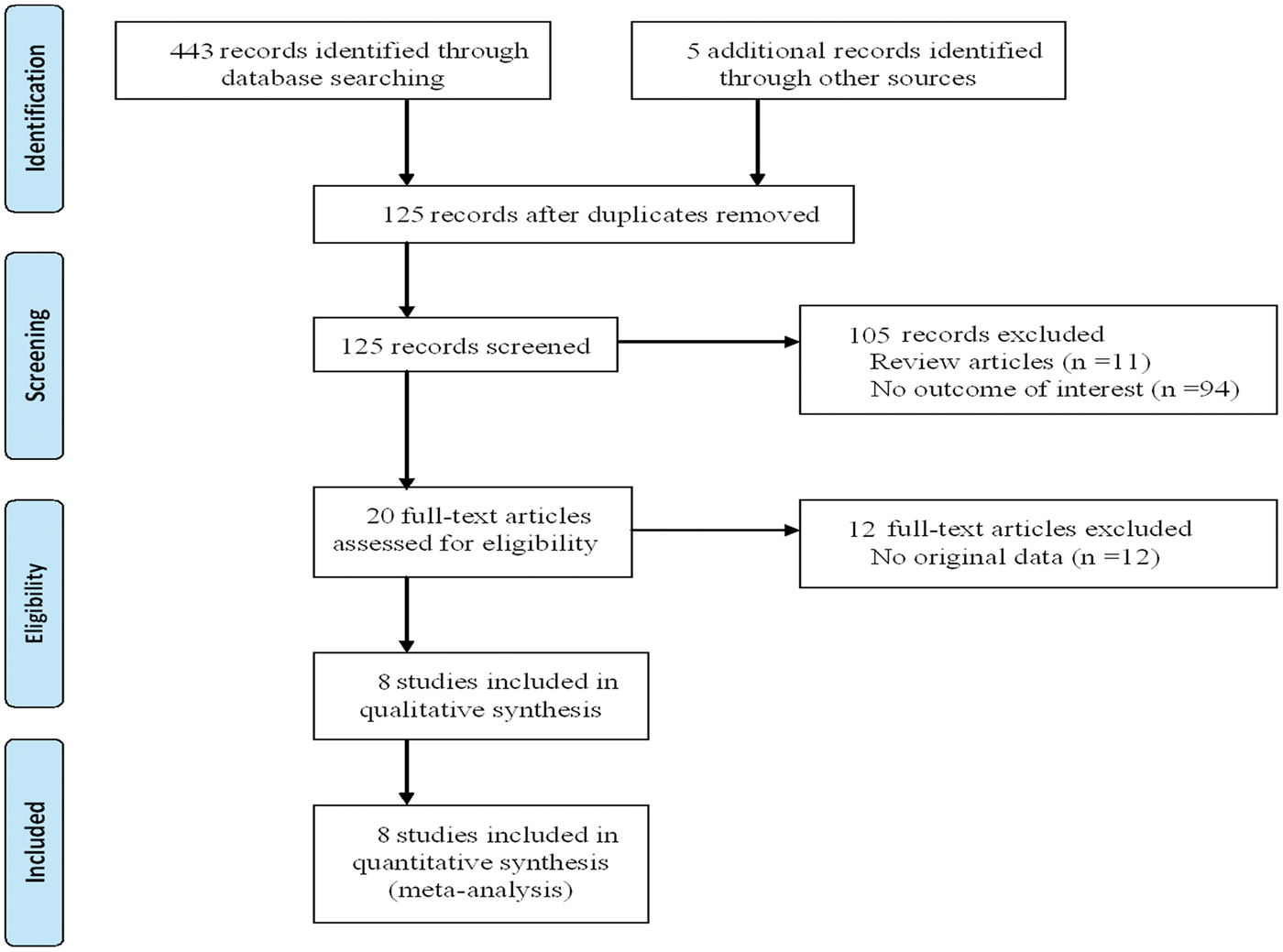

Quantitative synthesis
Combined analyses for relationships between IL-10 gene −1082 G/A polymorphism and cervical cancer and/or CIN are shown in Figure 2 and Table 2. Significant heterogeneity existed in the allelic model (Pheterogeneity < 0.001, I2 = 76.6%) and dominant model (Pheterogeneity < 0.001, I2 = 76.6%); therefore, the random-effects model was used to synthesize these data. No heterogeneity was found in the additive model (Pheterogeneity = 0.416, I2 = 1.1%) and recessive model (Pheterogeneity = 0.734, I2 = 0%), so the fixed-effects model was used to analyse these data. Overall, there were no significant associations between IL-10 gene −1082 G/A polymorphism and cervical cancer and/or CIN in any of the genetic models (Table 2). When analysed by ethnicity, there were again no significant associations between IL-10 gene −1082 G/A polymorphism and cervical cancer and/or CIN in either Asians or non-Asians (Table 2).
Forest plots of the association between interleukin-10 gene –1082 G/A polymorphism and cervical cancer and/or cervical intraepithelial neoplasia in eight studies, using different genetic models: A, allelic model (A allele versus G allele); B, additive model (A/A versus G/G); C, recessive model (A/A versus G/A+G/G); D, dominant model (A/A+G/A versus G/G). OR, odds ratio; CI, confidence intervals; p, Pheterogeneity value. Studies: Stanczuk 2001,
18
Zoodsma 2005,
19
Farzaneh 2006,
12
Matsumoto 2010,
15
Wang 2011,14 Yu 2011,
17
Barbisan 2012,13 Roh 2002
16
Meta-analysis of data from eight studies of the association between interleukin-10 gene –1082 G/A polymorphism and cervical cancer and/or cervical intraepithelial neoplasia, using different genetic models: allelic model, A allele versus G allele; additive model, A/A versus G/G; recessive model, A/A versus G/A+G/G; dominant model, A/A+G/A versus G/G Significant heterogeneity demonstrated using random-effects model. Based on Hardy–Weinberg equilibrium (studies without Hardy–Weinberg equilibrium were excluded). OR, odds ratio; CI, confidence intervals Differences between the groups were not statistically significant (P ≥ 0.05).

Sensitivity analysis
Sensitivity analysis was performed to evaluate the stability of the overall results. When the study of Yu et al., 17 which was the only study that did not meet the HWE, was excluded, the corresponding pooled ORs were not materially altered in any of the genetic models (Table 2), suggesting that the results of the meta-analysis were stable and that the study without HWE was not influencing the overall results.
Publication bias
There was no evidence of publication bias using Begg’s funnel plots (Figure 3). In addition, there was no statistical evidence of publication bias among studies using Egger’s regression test (P ≥ 0.05 for all four genetic models).
Begg’s funnel plots with pseudo 95% confidence limits for interleukin-10 gene −1082 G/A polymorphism and cervical cancer and/or cervical intraepithelial neoplasia, using different genetic models: A, allelic model (A allele versus G allele); B, additive model (A/A versus G/G); C, recessive model (A/A versus G/A+G/G); D, dominant model (A/A+G/A versus G/G). logor, log odds ratio; s.e., standard error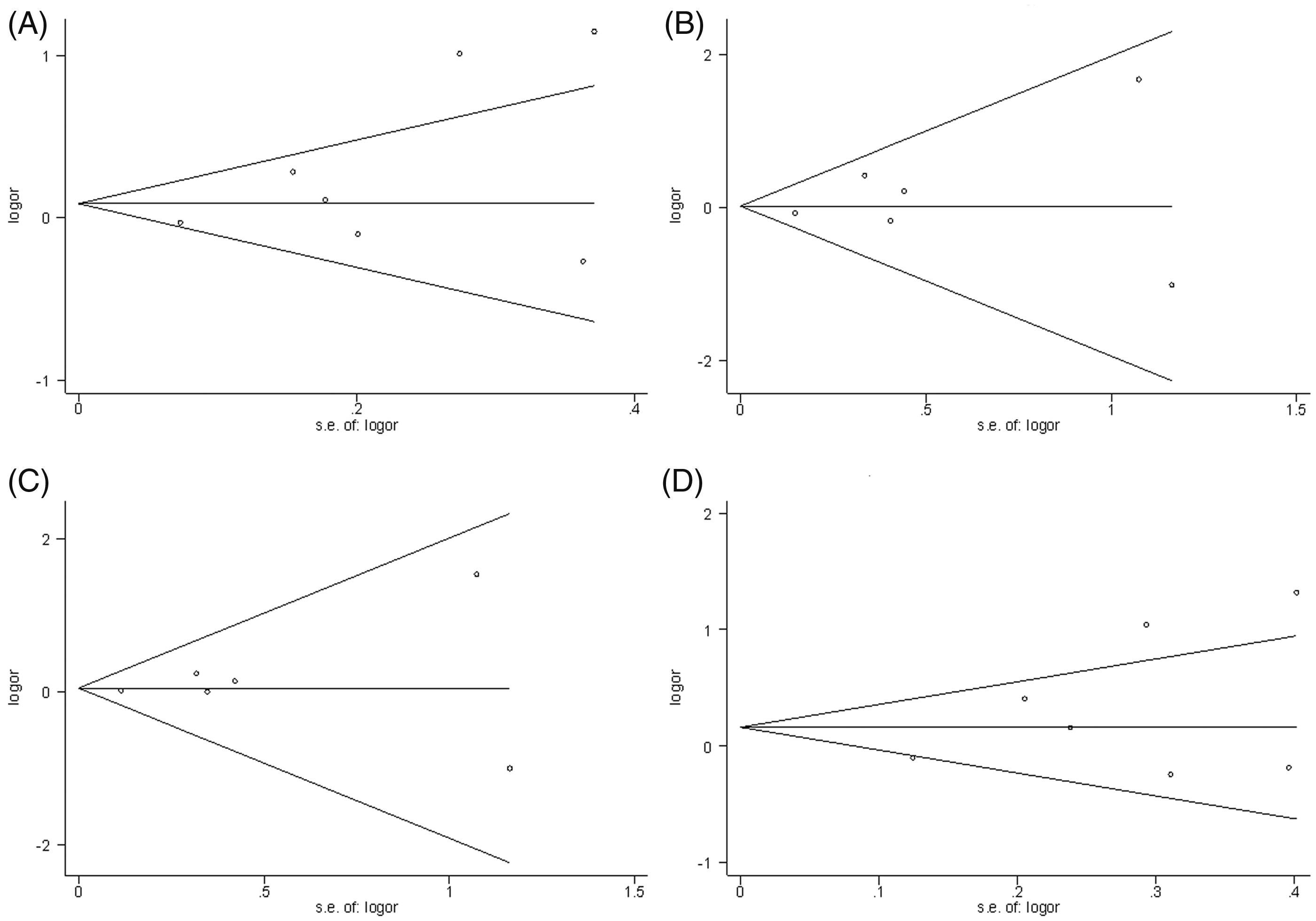
Discussion
The associations between IL-10 gene −1082 G/A polymorphism and cervical cancer and/or CIN have been intensively studied, but the results have been inconsistent. For example, in 2001, Stanczuk et al. 18 found that, compared with the IL-10 A allele, the IL-10 G allele was associated with a significantly higher risk for developing cervical cancer in Zimbabwean populations. In 2010, Matsumoto et al. 15 also resported that the IL-10 G allele was associated with a higher cervical cancer risk in Japanese populations compared with the IL-10 A allele. However, Roh et al., 16 Zoodsma et al., 19 Farzaneh et al., 12 Wang et al.14 and Yu et al., 17 and Barbisan et al.13 found that the IL-10 gene −1082 G/A polymorphism was not associated with cervical cancer or CIN in Korean, Dutch, British, Chinese and Argentinean populations, respectively. The present meta-analysis of data from eight eligible studies, including 1983 cases and 1618 controls, was therefore performed to derive a more precise estimate of the strength of the association.
In the present meta-analysis, IL-10 gene −1082 G/A polymorphism and its relationship with the risk of cervical cancer and/or CIN was examined using four genetic models. Overall, no significant association was found in any of the genetic models. These findings are consistent with most of the studies in our meta-analysis.12–14,16,17,19 Subgroup analysis by ethnicity showed that there were no significant associations between IL-10 gene −1082 G/A polymorphism and cervical cancer and/or CIN in Asians or non-Asians. Because the inclusion of studies without HWE might influence the results of the meta-analysis, 20 a sensitivity analysis restricted to the studies with HWE was performed. Similar results to those of the full meta-analysis were found. Thus, the results of subgroup and sensitivity analyses further strengthened the finding that IL-10 gene polymorphism is not associated with cervical cancer and/or CIN.
Significant between-study heterogeneity existed in the allelic and dominant models. Heterogeneity is a potential problem that may affect the interpretation of the results. It has a wide range of potential sources, such as case definition and sampling, sample sizes, methods of genotyping, and differences in ethnicity. The subgroup and sensitivity analyses performed were unable to identify the sources of heterogeneity. Further analyses were not performed due to the lack of original data. Therefore, all the potential reasons for heterogeneity should still be taken into account.
Publication bias did not exist in the overall comparisons, indicating that the results of the present meta-analysis were statistically robust. However, certain limitations of this work should be acknowledged. First, the sample size in most of the included studies was small, which could increase the probability of false positives or false negatives. Secondly, significant between-study heterogeneity existed in the allelic and dominant models, indicating that the findings should be interpreted with care. Thirdly, cervical cancer and CIN are complex diseases that can be affected by a number of factors such as HPV infection, HPV subtype, infectious state, the balance of pro- and anti-inflammatory cytokines, and other conditions that influence cytokine levels. However, many of the eligible studies included in this meta-analysis did not present sufficient original data related to these factors to allow further subgroup analyses. In addition, meta-analyses have some inherent limitations in that their retrospective nature, meaning that they are subject to the methodological deficiencies of the included studies.
Despite these limitations, the present meta-analysis had some strengths. First, the process of meta-analysis has the ability to detect small effects in human genetic association studies. 21 In the present study, data were extracted from as many different studies as possible, which increased the statistical power of this analysis. Secondly, the corresponding pooled ORs were not materially altered in any of the genetic models when subgroup and sensitivity analyses were performed, and no publication bias was found, indicating that the results of this meta-analysis were stable and reliable.
In conclusion, findings of the present study suggest that there is no association between IL-10 gene −1082 G/A polymorphism and cervical cancer and/or CIN. However, this result should be interpreted with caution because of the limitations of this meta-analysis. Further studies with larger sample sizes, especially multicentric case–control studies, will be needed to confirm these findings.
Footnotes
Declaration of conflicting interest
The authors declare that there are no conflicts of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
