Abstract
RNA-binding motif 5 is a putative tumor suppressor gene that modulates cell cycle arrest and apoptosis. We recently demonstrated that RNA-binding motif 5 inhibits cell growth through the p53 pathway. This study evaluated the clinical significance of RNA-binding motif 5 expression in gastric cancer and the effects of altered RNA-binding motif 5 expression on cancer biology in gastric cancer cells. RNA-binding motif 5 protein expression was evaluated by immunohistochemistry using the surgical specimens of 106 patients with gastric cancer. We analyzed the relationships of RNA-binding motif 5 expression with clinicopathological parameters and patient prognosis. We further explored the effects of RNA-binding motif 5 downregulation with short hairpin RNA on cell growth and p53 signaling in MKN45 gastric cancer cells. Immunohistochemistry revealed that RNA-binding motif 5 expression was decreased in 29 of 106 (27.4%) gastric cancer specimens. Decreased RNA-binding motif 5 expression was correlated with histological differentiation, depth of tumor infiltration, nodal metastasis, tumor–node–metastasis stage, and prognosis. RNA-binding motif 5 silencing enhanced gastric cancer cell proliferation and decreased p53 transcriptional activity in reporter gene assays. Conversely, restoration of RNA-binding motif 5 expression suppressed cell growth and recovered p53 transactivation in RNA-binding motif 5–silenced cells. Furthermore, RNA-binding motif 5 silencing reduced the messenger RNA and protein expression of the p53 target gene
Introduction
Gastric cancer is very common globally, especially in Japan. 1 Although the mortality of gastric cancer has declined, mainly due to early detection, the clinical outcome of patients with advanced gastric cancer remains poor.2–4 Surgical excision remains the mainstay treatment for gastric cancer,5–7 although endoscopic treatment for early gastric cancer is gaining ground. 8 The outcome of patients after radical gastrectomy is influenced by many tumor- and patient-based factors. 6 Tumor progression is considered a multifactorial process that involves the dysregulation of various oncogenes and tumor suppressor genes.4,9,10 The accumulation of these multiple abnormalities confers growth advantages to gastric cancer cells.11–13 Therefore, understanding the molecular mechanisms associated with the progression of gastric cancer is critical for improving clinical outcomes.
RNA-binding motif 5 (RBM5) is a putative tumor suppressor gene involved in differentiation, apoptosis, cell proliferation, and carcinogenesis.14–16 However, the molecular mechanisms underlying RBM5-mediated tumor suppression remain unclear. RBM5 can modulate apoptosis by regulating the alternative splicing of multiple target genes.17–20 We previously demonstrated that RBM5 is required for normal p53 transcriptional activity and biological function by augmenting p53 protein expression in the absence and presence of DNA damage. 21 We also suggested that RBM5 exerts its tumor-suppressing functions by affecting p53 signaling pathways. 21 Although RBM5 protein expression is frequently reduced in various cancers,22–26 its protein level in gastric cancer remains uninvestigated. Despite the potential importance of RBM5 in carcinogenesis, 14 its pathogenic role in gastric cancer remains unclear.
In this study, we hypothesized that RBM5 may be involved in gastric cancer progression. We examined RBM5 expression in tumor tissue specimens obtained from patients with resected gastric cancer and evaluated the relationship of RBM5 protein expression with clinicopathological parameters. Furthermore, we also investigated the effect of RBM5 silencing on gastric cancer cell proliferation and p53 transcriptional activity to define the function of RBM5 that might be linked to gastric cancer progression. Our observations suggest that RBM5 behaves as a tumor suppressor gene in gastric cancer and that decreased RBM5 expression may be associated with the malignant potential of gastric cancer cells.
Materials and methods
Patients
Primary gastric cancer tissues and corresponding normal adjacent tissues were collected from 106 patients who underwent curative gastrectomy for gastric cancer at Hokkaido University Hospital between 1990 and 2000. This period was selected to provide a sufficient period of follow-up to determine patient prognosis after radical resection. All patients were postoperatively classified according to the Union for International Cancer Control (UICC) stage classification. 27 Relevant clinicopathological parameters were acquired from medical records. The patients included 76 males and 30 females (age range, 35–90 years; mean, 64.4 years). Based on the UICC tumor–node–metastasis (TNM) staging system, 60 (56.6 %), 13 (12.2%), 15 (14.2%), and 18 (17.0%) patients had stage I, II, III, and IV lesions, respectively. All lesions were pathologically diagnosed as adenocarcinomas. Histologically, we categorized well-differentiated and moderately differentiated adenocarcinomas as differentiated tumors, and poorly differentiated adenocarcinoma, signet-ring cell carcinoma, and mucinous adenocarcinoma as undifferentiated tumors. Relevant clinicopathological information was acquired from medical records and the pathological diagnosis. The clinicopathological characteristics of patients and tumors are summarized in Table 1.
Correlations between RBM5 expression and clinicopathological variables.

RBM5: RNA-binding motif 5.
Immunohistochemistry
Histopathological studies were performed as previously reported. 28 Briefly, tissues were fixed in 10% buffered formalin and embedded in paraffin. Three-micrometer-thick sections were stained with hematoxylin–eosin or analyzed for RBM5 expression via immunohistochemistry. RBM5 immunostaining was performed using standard procedures with a specific mouse monoclonal antibody. 21 RBM5 staining was evaluated using a semiquantitative score, which was calculated as the sum of the percentage of positively stained tumor cells and the staining intensity. The percentage of positive staining was scored as 0 (0%–9%, negative), 1 (10%–25%, sporadic), 2 (26%–50%, focal), or 3 (51%–100%, diffuse), and the intensity was scored as 0 (no staining), 1 (weak staining), 2 (moderate staining), or 3 (strong staining) compared with noncancerous gastric mucosa. The total immunostaining score was calculated as the positivity score multiplied by the staining intensity score. Five areas were randomly selected, and the average score in each case represented the final positive staining points. Thus, we separated the patients into high (total score ⩾3) and low (total score <3) expression groups. Two pathologists who were blinded to clinical outcome separately evaluated all samples. They evaluated the RBM5 score of the immunostaining slides under an optical microscope at ×400magnification. The average value of two independent scores was presented in this study.
Cell culture
MKN45 human gastric cancer cells were obtained from the Japanese Collection of Research Bioresources Cell Bank (Osaka, Japan) and maintained in RPMI 1640 medium supplemented with 10% (v/v) heat-inactivated fetal bovine serum, 100 U/mL penicillin, and 100 µg/mL streptomycin, and cells were incubated in a humidified atmosphere of 5% CO2 and 95% air at 37°C. Cells in the logarithmic growth phase were used for further experiments.
RBM5 plasmids and stable cell lines
RBM5 short hairpin RNA (shRNA) plasmids, which are double-stranded oligonucleotides encoding small interfering RNA (siRNA) that specifically target the
Subcellular fractionation and western blot analysis
Cytoplasmic and nuclear fractions from MKN45 cells were prepared using an EzSubcell Fraction Kit according to the manufacturer’s instructions (Atto, Tokyo, Japan). Total cell lysates were prepared in lysis buffer (10 mM Tris–HCl, pH 7.5, 150 mM NaCl, 5 mM ethylenediaminetetraacetic acid (EDTA), 0.5% Nonidet P-40) supplemented with protease inhibitors. Western blotting was conducted using standard protocols as previously described. 21 RBM5 was detected using a mouse monoclonal antibody raised against a partial recombinant protein of the human carboxy-terminus. 21 Other specific antibodies were used to detect p53 (DO-1; Santa Cruz Biotechnology, Santa Cruz, CA, USA), p21 (Ab-1; EMD Millipore, MA, USA), p27 (DCS-72.F6; Thermo Fisher Scientific, Fremont, CA, USA), α-tubulin (DM 1A; Sigma-Aldrich, St. Louis, MO, USA), and β-actin (AC-15, Sigma-Aldrich).
Immunofluorescence
MKN45 cells were plated on six-well culture plates with coverslips. Cells were fixed with 4% paraformaldehyde in phosphate-buffered saline (PBS) for 10 min and then permeabilized with 0.5% Triton X-100 in PBS for 5 min. Cells were blocked with 2% bovine serum albumin (BSA) in PBS and then incubated with anti-RBM5 mouse monoclonal antibody 21 and anti-β-actin goat polyclonal antibody (I-19; Santa Cruz Biotechnology) for 1 h in PBS with 2% BSA, followed by incubation for 1 h with Rhodamine Red X-conjugated anti-mouse antibody (Jackson ImmunoResearch, West Grove, PA, USA) and Cy2-conjugated anti-goat antibody (Jackson ImmunoResearch) in PBS with 1% BSA. To visualize nuclei, 1.0 µg/mL 4′,6-diamidino-2-phenylindole dihydrochloride was added during the last wash. The images were observed under a fluorescence microscope (Biozero BZ-8000; Keyence, Osaka, Japan).
Cell proliferation assay
Cells were seeded in four 96-well plates at a density of 1.5 × 103 cells/well. At the indicated time points, cell growth was determined using a CellTiter-Glo Luminescent Cell Viability Assay Kit according to the manufacturer’s instructions (Promega, Madison, WI, USA). Luminescence was measured using a GloMax-Multi+ Detection System (Promega). All experiments were performed in triplicate.
Reporter gene assay
The Dual-Luciferase Reporter Assay System (Promega) was used as described previously.
21
Briefly, cells were co-transfected with
Quantitative real-time polymerase chain reaction
Quantitative real-time polymerase chain reaction (qPCR) was performed as described previously.
21
Briefly, total RNA was isolated from clones stably transfected with shRBM5 plasmids or control vectors using RNAiso reagent (TaKaRa, Otsu, Japan). qPCR was performed with TaqMan probes for
Statistical analysis
The chi-square test was used to analyze the relationships between RBM5 protein expression and various clinicopathological parameters. Disease-specific survival rates were calculated using the Kaplan–Meier method and assessed using the log-rank test. The Cox proportional hazard regression model was used for multivariate analyses to evaluate the suitability of clinicopathological variables and RBM5 expression as independent predictors of patient survival. Multivariate logistic regression analysis was performed to assess the interrelation between RBM5 expression and the variables for overall survival. Two-tailed
Results
Subcellular localization of RBM5 in gastric cancer cells
To assess the biological role of RBM5 in gastric cancer, we first examined its protein expression in gastric cancer cells via immunofluorescence analysis and western blotting with a specific mouse monoclonal antibody we originally prepared. 21 Immunofluorescence analysis revealed that RBM5 was localized in the nucleus (Figure 1(a)) as previously reported in other cancer cells.26,29 The nuclear localization of RBM5 was also confirmed by fractionating the cytosolic and nuclear compartments via western blotting (Figure 1(b)).

RNA-binding motif 5 (RBM5) protein expression in gastric cancer cells. (a) Analysis of RBM5 expression via immunofluorescence analysis in gastric cancer cells. β-actin and the nucleus (6-diamidino-2-phenylindole dihydrochloride (DAPI)) were also stained and (b) subcellular localization of RBM5 in gastric cancer cells. MKN45 cells were fractionated into cytosolic (C) and nuclear fractions (N). Each fraction and whole cell lysate (W) were subjected to western blotting for RBM5 expression. HSP90 (cytosolic) and hnRNP C1/C2 (nuclear) expression was analyzed to authenticate the purity of the individual fractions.
The association between RBM5 expression and clinicopathological characteristics based on immunohistochemical staining
To obtain insight into the clinicopathological value of RBM5 expression in gastric cancer, we performed immunohistochemical analysis of RBM5 protein expression in gastric cancer specimens. Immunohistochemical staining revealed that RBM5 expression was predominantly detected in the nucleus as predicted in the immunofluorescence study (Figure 2(a)). RBM5 expression as assessed by immunohistochemistry in 106 primary gastric cancer specimens is shown in Table 1.

(a) RNA-binding motif 5 (RBM5) protein expression in gastric cancer surgical specimens as assessed by immunohistochemistry. Representative examples for the low (left) and high (right) RBM5 expression groups. Hematoxylin and eosin staining (upper) and immunohistochemical staining (lower) and (b) overall cumulative survival rates of patients according to RBM5 expression in 100 patients with gastric cancer after radical resection (Kaplan–Meier method). RBM5 expression was classified as low (
Twenty-nine tumors (27.4%) exhibited reduced RBM5 expression compared with the corresponding adjacent normal tissues, whereas the remaining 77 specimens (72.3%) displayed strong RBM5 expression (Table 1). We analyzed the relationship between RBM5 expression in gastric cancer tissues and clinicopathological features. Decreased RBM5 expression was significantly associated with histological differentiation (
RBM5 expression and clinical outcome
We used Kaplan–Meier analysis to determine whether RBM5 is a significant prognostic factor for disease-specific survival in patients (
RBM5 silencing enhances gastric cancer cell proliferation
To examine the impact of reduced expression of endogenous RBM5 in gastric cancer cells, we generated MKN45 cells stably expressing plasmids for shRNA specifically targeting the
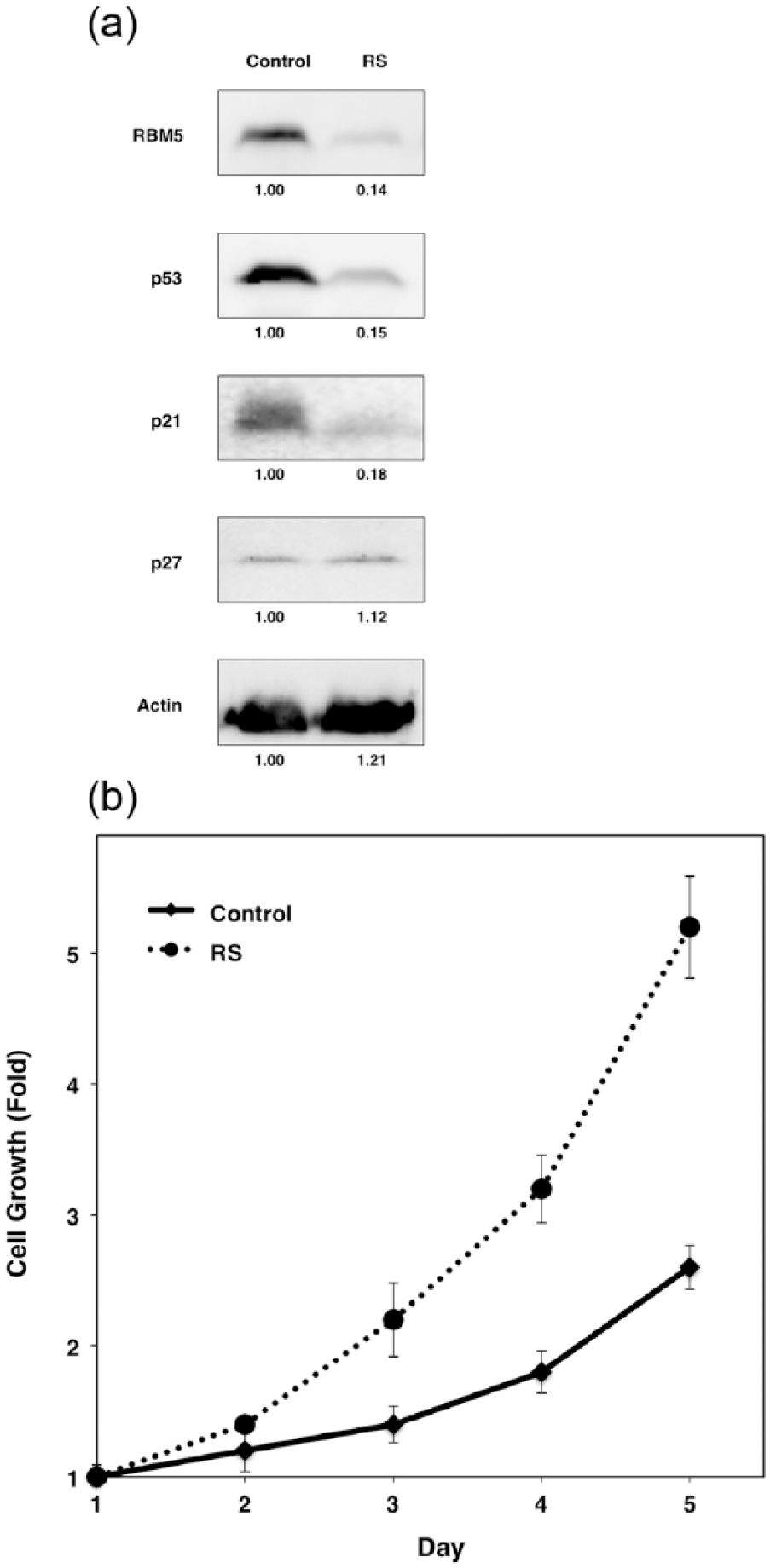
(a) Endogenous p53 and p21 protein levels are influenced by the RBM5 expression level. Western blotting uncovered the expression levels of RBM5, p53, p21, p27, and β-actin in RBM5-silenced (RS) and control (Control) cells. The expression levels were normalized to β-actin levels, and relative intensities are presented at the bottom of each panel and (b) RBM5 silencing stimulates cell growth. Cell proliferation assay reveals the enhanced proliferation of RBM5-silenced cells (RS) compared with control cells (Control). Data are shown as the mean ± standard deviation of three independent experiments.

RBM5 restoration abrogates the augmentation of cell proliferation by RBM5 silencing. RBM5-silenced (RS) and control cells (Control) that were transfected with an additional RBM5 expression plasmid or an empty plasmid were cultured with G418 for 2 weeks to pool transfected cells. (a) Western blotting was performed to evaluate RBM5 and β-actin expression. Protein expression was normalized to β-actin expression, and relative intensities are presented at the bottom of each panel and (b) the cells were seeded, and cell growth was examined after 5 days. The bars represent the relative numbers of viable cells along with standard deviations calculated from three independent experiments.
RBM5 silencing decreases p53 transcriptional activity
We previously demonstrated that RBM5 affected the p53 pathway by regulating p53 protein levels and transcriptional activity in various cancers, including breast cancer, cervical cancer, and osteosarcoma.
21
We also revealed that RBM5 augmented the growth-suppressive functions of p53.
21
To investigate the molecular role of RBM5 in the suppression of gastric cancer cell growth, we first evaluated the effect of RBM5 silencing on p53 protein expression and transcriptional activity in p53 wild-type MKN45 cells. A corresponding decrease in p53 expression was observed by western blotting upon RBM5 silencing (Figure 3(a)). Next, p53 reporter gene assays were performed to examine whether p53 transactivation is influenced by RBM5 in gastric cancer cells. RBM5 silencing reduced p53 transcriptional activity (Figure 5(a), lanes 1 and 3). We further examined whether RBM5 silencing affects p53 transcriptional activity using a native promoter of
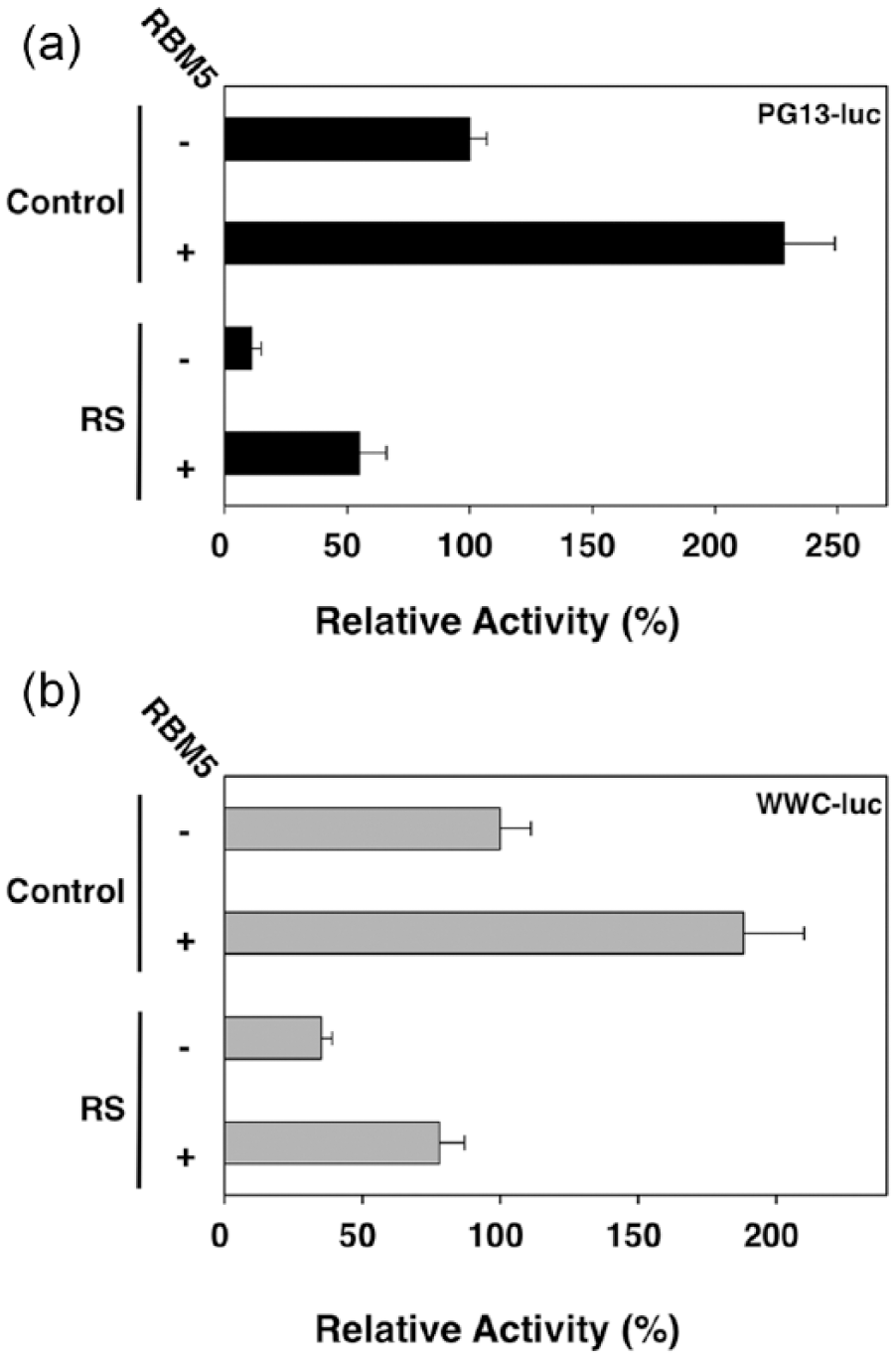
RBM5 expression in gastric cancer cells affects p53 transcriptional activity in reporter gene assays. RBM5-silenced (RS) and control (Control) cells were co-transfected with firefly luciferase reporter plasmids (a: PG13-Luc and b: WWC-Luc), Renilla luciferase plasmids, and RBM5 expression or empty control plasmids to ensure equal amounts of transfected plasmids. Firefly luciferase activity, adjusted for transfection efficiency using Renilla luciferase activity, was determined 24 h after transfection. The adjusted luciferase activity of control cells transfected with reporter plasmids and without RBM5 plasmid was set as 100%. Shown are the means and standard deviations for three independent experiments.
RBM5 silencing decreases p21 expression
We next evaluated the effect of RBM5 ablation on the expression of the endogenous p53 target gene

Discussion
The clinical outcome of gastric cancer is determined by various tumor characteristics, including local tumor growth, distant metastasis, and differentiation grade, which are regulated by a number of related genes.4,9,11 Therefore, identifying gastric cancer–specific determinants involved in tumor progression is important for the treatments of gastric cancer. 32 Here, we examined RBM5 expression in primary gastric cancer tissue and evaluated its prognostic value to postresection survival. The clinical significance and biological functions of RBM5 in gastric cancer have not been explored, although RBM5 expression is frequently reduced in various cancer tissues, including lung, prostate, and pancreatic cancers.22–26 We demonstrated that among 106 paired gastric cancer tissues, 29 tumors (27.4%) displayed downregulated RBM5 expression. Conversely, RBM5 expression was similar between tumors and normal adjacent tissues for more than 90% of T1 gastric cancer tumors. Decreased RBM5 expression was more conspicuous in advanced stages (III and IV) than in early stages (I and II). We also found that decreased RBM5 expression was significantly associated with tumor depth, TNM stage, and lymph node metastasis. In agreement with our observations, reduced RBM5 protein expression was also correlated with lymph node metastasis and TNM stage in pancreatic and lung cancers.22,25 Our results suggest that RBM5 downregulation is involved in tumor progression rather than carcinogenesis in gastric cancer. In addition, we identified lower RBM5 immunoreactivity in undifferentiated gastric cancer tissues than in differentiated tissues, suggesting that decreased RBM5 expression plays a role in the dedifferentiation of gastric tumors. Contrarily, no significant differences were identified between RBM5 expression and cancer cell differentiation in pancreatic cancer. 22 The role of RBM5 in cancer cell differentiation might depend on the tissue of origin.
We found a significant association between low RBM5 expression and poorer clinical outcomes in gastric cancer after radical surgery. However, Cox hazard ratio regression analyses indicated that the RBM5 expression level was not an independent risk factor for disease-specific survival. Although the UICC TNM stage is the most widely used and important prognostic factor for gastric cancer,33,34 the depth of tumor infiltration and lymph node metastasis were not independent prognostic factors for overall survival. Consequently, RBM5 was not a useful prognostic factor in our study. However, the number of genes involved in the clinical outcome of gastric cancer is unknown, and mixed complex effects might have caused the function of RBM5 to be ambiguous in this study. Furthermore, the limited samples of resectable gastric cancer lesions and the retrospective nature of our study might have reduced the prognostic value of RBM5. Further large-scale studies of patients with advanced gastric cancer may clarify the etiological significance of RBM5. Meanwhile, we found that RBM5 levels were associated with the depth of tumor infiltration and lymph node metastasis concerning overall survival. Most patients with gastric cancer and lymph node metastasis have poor prognoses even after curative resection. 35 Although the biological role of RBM5 in gastric cancer metastasis has not been fully addressed yet, decreased RBM5 expression is part of a 17-gene expression signature predictive of metastasis and poor clinical outcome for various types of human cancers. 36 Oh et al. 37 demonstrated that RBM5 alters the expression of genes involved in metastasis, suggesting that the loss of RBM5 function may increase the metastatic potential of lung cancer.
To elucidate the molecular basis of the tumor-suppressing role of RBM5 regarding gastric cancer progression, we investigated the effect of RBM5 silencing on gastric cancer cell proliferation. RBM5 silencing stimulated gastric cancer cell growth. Contrarily, restoring RBM5 expression reduced their proliferation. It appears that RBM5 plays an inhibitory role in the regulation of cell proliferation. However, the molecular mechanisms by which RBM5 exerts antitumor effects in gastric cancer remain to be investigated. We previously demonstrated that RBM5 regulates p53 transcriptional activity in various cancer cells and that RBM5-mediated growth suppression and apoptosis are linked to p53 activity.
21
RBM5 silencing decreased p53 protein expression and transcriptional activity. Conversely, restoration of RBM5 expression recovered p53 transactivation in RBM5-silenced cells. Furthermore, RBM5 silencing reduced
Recurrence and metastasis after curative operation occur in some patients with advanced tumors. Sometimes, patients with the same TNM stage display considerable variability in tumor recurrence and clinical outcome; hence, we need specific biomarkers for high-risk patients with gastric cancer to facilitate the development of individualized treatment. 42 For example, surveillance of early recurrence and aggressive postoperative systemic chemotherapy could be selected for patients with reduced RBM5 expression in endoscopic biopsy or surgical specimens. In addition, we previously revealed that depletion of endogenous RBM5 conferred resistance to antitumor drugs and, contrarily, RBM5 overexpression increased sensitivity to antitumor drugs. 21 Similarly, Li et al. 43 found that RBM5 re-sensitized cisplatin-resistant lung cancer cells to cisplatin and that RBM5 could predict response to cisplatin in lung cancer cells. These data suggest that the suppression or absence of RBM5 is associated with resistance to various chemotherapeutic agents. Thus, enhancing RBM5 function may augment responses to chemotherapy in the treatment of advanced gastric cancer. Furthermore, a better understanding of the expression patterns of RBM5 may provide novel therapeutic approaches for advanced gastric cancer in the future.
In conclusion, we demonstrated for the first time that RBM5 expression is decreased in gastric cancer tissue according to tumor progression and that the protein plays a tumor-suppressive role in vitro, suggesting that RBM5 downregulation is pivotal to the acquisition of malignant potential in gastric cancer cells and that the loss of RBM5 function contributes to gastric cancer development. Further research is required to validate the biological regulation of RBM5 in the progression of gastric cancer.11,12
Footnotes
Acknowledgements
We thank Ms Hatsumi Ueda and Ms Terumi Hatakeyama for their research assistance.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. Participants provided written informed consent for their tissues to be used for research purposes.
Funding
This work was supported by JSPS KAKENHI Grant number 25460921.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
