Abstract
Haploinsufficiency of tumor suppressor genes, wherein the reduced production and activity of proteins results in the inability of the cell to maintain normal cellular function, is one among the various causes of cancer. However the precise molecular mechanisms underlying this condition remain unclear. Here we hypothesize that single nucleotide polymorphisms (SNPs) in the 3′untranslated region (UTR) of mRNAs and microRNA seed sequence (miR-SNPs) may cause haploinsufficiency at the level of proteins through altered binding specificity of microRNAs (miRNAs). Bioinformatics analysis of haploinsufficient genes for variations in their 3′UTR showed that the occurrence of SNPs result in the creation of new binding sites for miRNAs, thereby bringing the respective mRNA variant under the control of more miRNAs. In addition, 19 miR-SNPs were found to result in non-specific binding of microRNAs to tumor suppressors. Networking analysis suggests that the haploinsufficient tumor suppressor genes strongly interact with one another, and any subtle alterations in this network will contribute to tumorigenesis.
Keywords
Introduction
Cancer is a complex genetic disease involving structural and expression abnormalities of both coding and non-coding genes. The search for cancer causing genes has identified two major classes namely the oncogenes and tumor suppressor genes (TSGs). Activation or over expression of a proto-oncogene, such as
In principle, haploinsufficient TSGs are impaired by a 50% reduction in expression or activity. However,
Haploinsufficiency of multiple genes cooperate to promote tumorigenesis, a phenomenon called ‘compound haploinsufficiency'. The 5q deletion syndrome (5q-) is a paradigm of compound haploinsufficiency and demonstrates the importance of combinatorial interactions to elicit specific phenotypes.
12
Experimental evidence has shown that co-suppression of linked 8p TSGs promotes tumor formation more potently than any individual gene.
13
All the available evidence indicates that the functioning of a cell depends on the appropriate expression levels of proteins and, more importantly, that cell signalling pathways involve complex interactions between many proteins. Not only genes, but also microRNAs (miRNAs), a class of small non-coding RNAs, are shown to be haploinsufficient and to cause developmental abnormalities in humans.
14
In addition, genes involved in miRNA biosynthesis pathway, such as
miRNAs act at the post-transcriptional level of gene regulation through RNA-induced silencing complex (RISC) mediated translational inhibition or mRNA cleavage. The miRNA target recognition is mediated through the sequence complementarity between the 2–8 nt at the 5′ end of miRNA (seed sequence) and the 3′-UTR of target mRNA.
18
Based on the degree of complementarity, the mRNA is either guided for inhibition of translation or degradation resulting in the decrease of protein encoded by the target messenger.
19
This has extended the dimensions of haploinsufficiency as miRNAs can lead to the production of insufficient amount of proteins. Bioinformatics prediction and experimental analysis suggested that miRNAs can regulate approximately half of the mammalian genes, with a significant number of important oncogenes and tumor suppressor genes involved.20,21 A recent genome-wide association study has suggested that a gene with more than two miRNA target sites will have higher variability of expression than a gene which is not regulated by a miRNA. The variability is further increased by SNPs in the miRNA target sites.
22
Polymorphisms in the miRNA regulatory pathway are a novel class of functional polymorphisms present in the human genome. The initial demonstration that miRNA binding site variations can result in a phenotype was provided by Abelson et al who identified a mutation in the miR-189 binding site of
Single nucleotide polymorphisms (SNPs) are single base pair changes in DNA that occur with a frequency of about 1 in 12,500 bp in the genome. 27 Several studies have shown that SNPs in microRNA networks moderately increase the risk of cancer incidence. 28 In general, sequence variations in pri-miRNAs, pre-miRNAs, mature miRNAs, and microRNA binding sites potentially affects the processing and/or target selection of miRNAs. As a consequence, aberrant expression of hundreds of genes and pathways greatly affecting miRNA function may occur. 29 SNPs in mature miRNAs and miRNA binding sites function analogously to modulate the miRNA-mRNA interaction and create or destroy miRNA binding sites. Supporting this idea, SNPs within the miRNA binding sites of genes have been implicated in susceptibility to various types of cancer.30–33 Additionally, functional support for individual miR-SNPs implicated in cancer do exist.
In this study we analyzed the role of SNPs that occur at miRNA binding sites and miR-SNPs and their contribution towards haploinsufficiency of tumor suppressor genes.
Materials and Methods
Datasets
A total of 110 haploinsufficient genes known to have a role in tumor initiation and progression were considered for the study. Among the total list, 63 tumorigenic haploinsufficient genes were obtained from Dang et al, 2008 34 while the remaining 47 genes were compiled using keyword search in PubMed and Online Mendelian Inheritance in Man (OMIM) database. For a comprehensive list of the haploinsufficient tumorigenic genes included in this study with their corresponding reference, see Supplement 1.
Analysis of 3′UTR polymorphisms in mRNA targets and miR seed
The haploinsufficient genes were analyzed for polymorphisms in their 3′UTR by submitting the respective gene ID in ‘Polymorphism in microRNA Target Site’ (PolymiRTS) database, 35 http://compbio.uthsc.edu/miRSNP/, designed to identify SNPs that disrupt the regulation of gene expression by miRNAs in human and mouse (last accessed on June 7th, 2012). The database is organized to provide links between SNPs in miRNA target sites, cis-acting expression quantitative trait loci (eQTLs), and the results of genome-wide association studies (GWAS) of human diseases. We applied filters to identify only the SNPs in the 3′UTR that were known to create binding sites for miRNAs and SNPs occurring in the seed region of miRNA themselves.
Classification and validation of SNPs obtained from polymiRTS
The SNPs predicted by the polymiRTS database were classified based on their effect into the following categories: (i) SNPs that lead to the creation of miRNA binding sites in the mRNA, (ii) SNPs that disrupted the originally existing miRNA binding sites and also resulted in creation of new miRNA binding sites in the mRNA, and (iii) miR-SNPs that altered the binding specificity of miRNAs. After classification, the SNPs were verified by submitting their rs ID in the NCBI's dbSNP Short genetic variation database http://www.ncbi.nlm.nih.gov/projects/SNP/using the batch query option. The output was downloaded in BED format and analyzed. Among the total 158 SNPs submitted, 156 SNPs were validated by dbSNP while 2 miR-SNPs (rs116596918 and rs116838571) were found to be new entries and validated by microRNA-related Single Nucleotide Polymorphism database (http://www.bioguo.org/miRNASNP/miRNA_details.php). 36
Functional network prediction using GeneMANIA Cytoscape plug-in
To identify the functional significance, GeneMANIA Cytoscape plug-in which employs the GeneMANIA algorithm37,38 was utilized with default settings to plot the interactions among the haploinsufficient TSGs. The complete list of tumorigenic haploinsufficient genes were uploaded to Cytoscape 39 using GeneMANIA plug-in (http://www.genemania.org/plugin/). Further, only the genes that were found to have SNPs in their miRNA binding sites and those that are non-specifically targeted by miRNAs harboring miR-SNPs were uploaded individually to study the interactions among this subgroup. The GeneMANIA Cytoscape plug-in integrates association networks from multiple sources into a single composite network using a conjugate gradient optimization algorithm. This algorithm produces networks from the data either directly (as in the case of protein or genetic interactions) or by using an in-house analysis pipeline to convert profiles to functional association networks.
Results and Discussion
Polymorphisms in 3′UTR of mRNA
Our analysis of tumorigenic haploinsufficient genes for genetic variations in the 3′UTR showed that 26% (29 out of 110) of them had at least one 3′UTR SNP resulting in creation of new binding sites for miRNAs thereby bringing the respective mRNA variant under the control of more new miRNAs. In total, 129 SNPs were found to create binding sites for 234 miRNAs in 29 genes (Table 1). Altogether, these SNPs contribute to haploinsufficiency by bringing the polymorphic mRNA under the control of more new microRNAs thereby leading to translational repression or mRNA degradation.
List of SNPs in the haploinsufficient tumor suppressor genes that created microRNA binding sites in their 3′UTR.

For convenience these microRNAs are not indicated in the table, and for details the reader is referred to Supplement 2.
Another 10 SNPs were found to bring 8 different genes under the control of additional new miRNAs due to the presence of the respective SNP (Fig. 1). A single SNP can create miR binding site for several miRNAs as well as delete a miR binding site. For example, the SNP rs186304832 in the 3′UTR of

Effect of SNPs occurring in the 3′UTR of haploinsufficient tumor suppressor genes.
Ours is the first study to identify the SNP and miRNA mediated control of tumor suppressor dose as well as highlighting the importance of genetic variations in cancer. This mechanism is in agreement with the continuum model for tumor suppression that is related to the level of expression or activity of the TSG rather than to the discrete step-by-step changes in gene copy number. 11
Polymorphisms in microRNA seed sequence
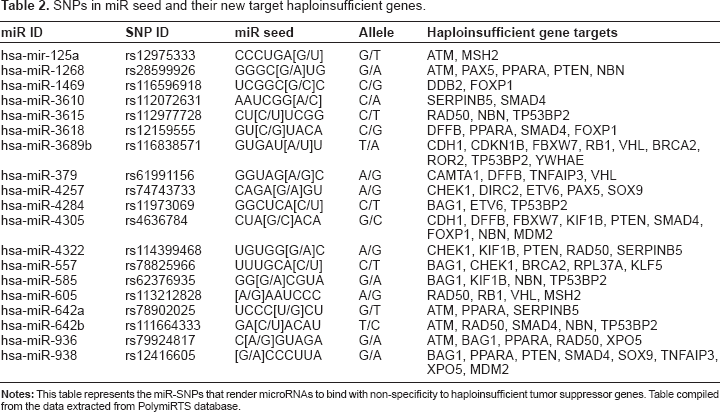
In addition, SNPs that occur in the seed sequence of miRNAs can alter the binding specificity of these miRNAs resulting in the binding to new non-target mRNAs. We analyzed 19 such miR-SNPs that cause miRNAs to bind new haploinsufficient tumor suppressor genes (Table 2). The SNP rs4636784 occurring in the seed of hsa-miR-4305 alters the binding specificity, thereby enabling it to target 6 new haploinsufficient genes (
SNPs in miR seed and their new target haploinsufficient genes.
Networks
Although reports on haploinsufficient genes associated with cancer is limited, analysis of the existing data shows that at least one third of total haploinsufficient genes to be tumorigenic. Tumor suppressors often act as components of complex networks, the overall function of which can be impaired by genetic and epigenetic alterations.41,42 For this reason we analyzed whether the haploinsufficient tumor suppressor genes share a common pathway or fall in a common interacting or functional network. The GeneMANIA algorithm was utilized for this purpose, and results indicate that almost 99% of haploinsufficient genes have a strong interaction, either directly or indirectly.

Interactions among 109 haploinsufficient genes plotted using GeneMANIA plug-in available in Cytoscape.
Next we tested whether the 39 genes with SNPs in miRNA binding sites or targeted by miRNAs with altered specificity due to miR-SNPs fall in to a particular network or pathway. Results of GeneMANIA networking analysis suggested strong interactions between these genes (Fig. 3). DNA repair genes and cell cycle checkpoint genes were enriched in this network and any aberrations in their expression offers selective advantage for the tumors to acquire additional genomic changes or mutations.

Interactions among 39 haploinsufficient genes that either had miRNA binding site SNPs or were targeted non-specifically by miRNAs carrying miR-SNPs.
Conclusion
With the expanding understanding of the roles of miRNAs in various cellular processes and diseases, this is the first report to discuss the role of miRNAs and its contribution towards tumor suppressor gene haploinsufficiency at the protein level via 3′UTR SNPs and miR-SNPs. Further, this could be an alternate means for attaining compound heterozygosity observed in several tumors and the mechanism is in complete agreement with the continuum model of tumor suppression. 11 Our data suggests that 26% of the tumorigenic haploinsufficient genes were brought under the control of new miRNAs due to 3′UTR SNPs. The identification of 10 SNPs that drive haploinsufficiency by bringing the polymorphic mRNA under the control of miRNAs, other than the miRNAs which are deleted, needs experimental validation in tumors. This alteration in the miRNA mediated gene regulation may cause predisposition to cancer initiation and progression.
Evidence for co-operative contribution of oncogenic mutations with tumor suppressor haploinsufficiency also exist. 47 The realization that miR-SNPs play central roles in the aberrant regulation of tumor suppressor genes has provided a new perspective on our understanding of pathophysiologic mechanisms. In addition, networking analysis reveals strong interactions between the haploinsufficient tumor suppressor genes. Any subtle alteration in this network of genes due to SNPs at the 3'UTR and miR-SNPs may contribute to pathogenesis. We suggest that at least a few among these SNPs and their effect on miRNA binding will aid in the diagnosis and/or prognosis of such type of cancers, if experimentally validated. While challenges remain in this regard, the pace of development in this field suggests that new discoveries are forthcoming.
Our approach has currently focused only on analyzing 3′UTRs, although a small subset of miRNAs can target 5′UTRs, coding regions, and gene promoters. miR-SNPs leading to non-specific binding of miRNAs to tumor suppressors may also result in the loss of binding to their original targets. If the target is an oncogene, it will be over expressed and will result in tumorigenesis. This has not been focused on in this study. Since the rules for miRNA binding to its target changes constantly, 48 the databases need constant updating. The 110 tumor suppressor genes chosen for our study were already proven to be haploinsufficient in several cancers through experimental evidence. This list may likely grow due to continuous identification of haploinsufficient tumor suppressors.
Funding Sources
This research was supported in part by a grant from the Department of Biotechnology (DBT), New Delhi (Grant Number BT/PR10023/AGR/36/27/2007) to AKM. MM is a recipient of research fellowship from Council of Scientific and Industrial Research (CSIR), New Delhi. We also thank DST-FIST and UGC-SAP for the infrastructure provided by grants to the department.
Author Contributions
AKM conceived and designed the study. MM and GR performed the experiments. AKM, MM, and GR analyzed the data. MM and AKM prepared the manuscript. All authors read and approved the final manuscript.
Competing Interests
Author(s) disclose no potential conflicts of interest.
Disclosures and Ethics
As a requirement of publication author(s) have provided to the publisher signed confirmation of compliance with legal and ethical obligations including but not limited to the following: authorship and contributorship, conflicts of interest, privacy and confidentiality and (where applicable) protection of human and animal research subjects. The authors have read and confirmed their agreement with the ICMJE authorship and conflict of interest criteria. The authors have also confirmed that this article is unique and not under consideration or published in any other publication, and that they have permission from rights holders to reproduce any copyrighted material. Any disclosures are made in this section. The external blind peer reviewers report no conflicts of interest.
Footnotes
List of Haploinsufficient genes known to be involved in tumorigenesis

| Haploinsufficient genes | References |
|---|---|
| ANX7, APC, ARF, ATM, BAG1, BECN1, BRCA1, BRCA2, BUBR1, CAMTA1, CAV1, CDKN1B, CDKN1C, CDKN2B, CSN5, CTCF, DFFB, DIRC2, EEF1E1, ETV6, FBXW7, FOXP1, H2 AX, INI1, INK4C, KLF6, LIG4, LIS1, LKB1, MEL-18, MYH, NHERF1, NKX3-1, NPM1, P53, PAX5, PLK4, PPARA, PPARG, PRKAR1 A, PTCH1, PTEN, RAD50, RASSF1 A, ROR2, RPRM, SAM68, SERCA2, SMAD5, SOCS1, SOX9, SPRED1, ST7, TGFB1, TP53BP2, TP73, TRAPPC2, TSC1, TSC2, VHL, WT1, WWOX, YWHAE | 11 |
| ATR | 18 |
| BIM (BCL2 L11) | 17 |
| BLM | 21 |
| BUB3 | 30 |
| CDH1 | 44 |
| CDKN1 A | 28 |
| CHK1 (CHEK1) | 33 |
| CLU | 9 |
| CREBBP (CBP) | 49, 47, 5 |
| DDB2 | 26 |
| DGCR8 | 42 |
| DICER1 | 32, 2, 41 |
| DLL4 | 19, 23 |
| DMP1 | 25, 51, 34 |
| DOK2 | 4 |
| FBXO11 | 14 |
| FEN1 | 31 |
| IKZF1 (IKAROS) | 39, 29, 52 |
| KIF1Bβ | 40 |
| KLF5 | 10 |
| MAD2 L1 | 38 |
| MDM2 and MDM4 | 50 |
| MLH1 | 48 |
| MSH2 | 12, 13, 6 |
| MUS81 | 36 |
| NBN | 15 |
| NF1 | 27, 59 |
| NF2 | 35 |
| PINX1 | 58 |
| RAD17 | 8 |
| RB1 | 53, 57 |
| Ribosomal proteins (RPL35, RPL37 A, RPS19 and RPS8) | 1 |
| RPS14 | 16, 3 |
| RUNX1 | 45 |
| RXRA | 24 |
| SERPINB5 (Maspin) | 43 |
| SMAD4/DPC4 | 56 |
| TARBP2 | 37 |
| TIP60 | 20 |
| TNFAIP3 | 55, 7 |
| TP53BP1 | 54 |
| TP63 (TAp63) | 46 |
| XPO5 | 22 |
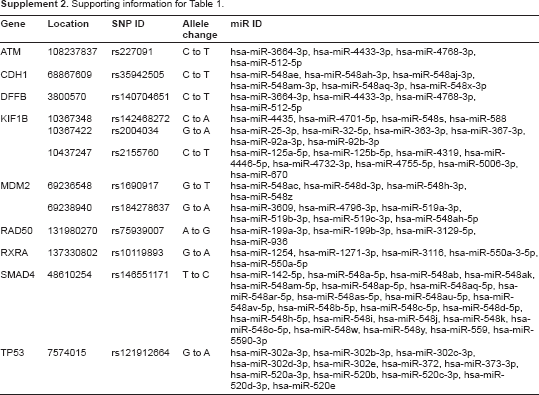
Supporting information for Table 1
| Gene | Location | SNP ID | Allele change | miR ID |
|---|---|---|---|---|
| ATM | 108237837 | rs227091 | C to T | hsa-miR-3664-3p, hsa-miR-4433-3p, hsa-miR-4768-3p, hsa-miR-512-5p |
| CDH1 | 68867609 | rs35942505 | C to T | hsa-miR-548ae, hsa-miR-548ah-3p, hsa-miR-548aj-3p, hsa-miR-548am-3p, hsa-miR-548aq-3p, hsa-miR-548x-3p |
| DFFB | 3800570 | rs140704651 | C to T | hsa-miR-3664-3p, hsa-miR-4433-3p, hsa-miR-4768-3p, hsa-miR-512-5p |
| KIF1B | 10367348 | rs142468272 | C to A | hsa-miR-4435, hsa-miR-4701-5p, hsa-miR-548s, hsa-miR-588 |
| 10367422 | rs2004034 | G to A | hsa-miR-25-3p, hsa-miR-32-5p, hsa-miR-363-3p, hsa-miR-367-3p, hsa-miR-92a-3p, hsa-miR-92b-3p | |
| 10437247 | rs2155760 | C to T | hsa-miR-125a-5p, hsa-miR-125b-5p, hsa-miR-4319, hsa-miR-4446-5p, hsa-miR-4732-3p, hsa-miR-4755-5p, hsa-miR-5006-3p, hsa-miR-670 | |
| MDM2 | 69236548 | rs1690917 | G to T | hsa-miR-548ac, hsa-miR-548d-3p, hsa-miR-548h-3p, hsa-miR-548z |
| 69238940 | rs184278637 | G to A | hsa-miR-3609, hsa-miR-4796-3p, hsa-miR-519a-3p, hsa-miR-519b-3p, hsa-miR-519c-3p, hsa-miR-548ah-5p | |
| RAD50 | 131980270 | rs75939007 | A to G | hsa-miR-199a-3p, hsa-miR-199b-3p, hsa-miR-3129-5p, hsa-miR-936 |
| RXRA | 137330802 | rs10119893 | G to A | hsa-miR-1254, hsa-miR-1271-3p, hsa-miR-3116, hsa-miR-550a-3-5p, hsa-miR-550a-5p |
| SMAD4 | 48610254 | rs146551171 | T to C | hsa-miR-142-5p, hsa-miR-548a-5p, hsa-miR-548ab, hsa-miR-548ak, hsa-miR-548am-5p, hsa-miR-548ap-5p, hsa-miR-548aq-5p, hsa-miR-548ar-5p, hsa-miR-548as-5p, hsa-miR-548au-5p, hsa-miR-548av-5p, hsa-miR-548b-5p, hsa-miR-548c-5p, hsa-miR-548d-5p, hsa-miR-548h-5p, hsa-miR-548i, hsa-miR-548j, hsa-miR-548k, hsa-miR-548o-5p, hsa-miR-548w, hsa-miR-548y, hsa-miR-559, hsa-miR-5590-3p |
| TP53 | 7574015 | rs121912664 | G to A | hsa-miR-302a-3p, hsa-miR-302b-3p, hsa-miR-302c-3p, hsa-miR-302d-3p, hsa-miR-302e, hsa-miR-372, hsa-miR-373-3p, hsa-miR-520a-3p, hsa-miR-520b, hsa-miR-520c-3p, hsa-miR-520d-3p, hsa-miR-520e |
