Abstract
Esophageal squamous cell carcinoma is one of the most aggressive malignancies worldwide. Special AT-rich sequence binding protein 1 is a nuclear matrix attachment region binding protein which participates in higher order chromatin organization and tissue-specific gene expression. However, the role of special AT-rich sequence binding protein 1 in esophageal squamous cell carcinoma remains unknown. In this study, western blot and quantitative real-time polymerase chain reaction analysis were performed to identify differentially expressed special AT-rich sequence binding protein 1 in a series of esophageal squamous cell carcinoma tissue samples. The effects of special AT-rich sequence binding protein 1 silencing by two short-hairpin RNAs on cell proliferation, migration, and invasion were assessed by the CCK-8 assay and transwell assays in esophageal squamous cell carcinoma in vitro. Special AT-rich sequence binding protein 1 was significantly upregulated in esophageal squamous cell carcinoma tissue samples and cell lines. Silencing of special AT-rich sequence binding protein 1 inhibited the proliferation of KYSE450 and EC9706 cells which have a relatively high level of special AT-rich sequence binding protein 1, and the ability of migration and invasion of KYSE450 and EC9706 cells was distinctly suppressed. Special AT-rich sequence binding protein 1 could be a potential target for the treatment of esophageal squamous cell carcinoma and inhibition of special AT-rich sequence binding protein 1 may provide a new strategy for the prevention of esophageal squamous cell carcinoma invasion and metastasis.
Keywords
Esophageal squamous cell carcinoma (ESCC) is the main type of esophageal cancer in China. Most ESCC cases are presented in an advanced stage.1,2 Though the tremendous advances in diagnosis and treatment, the prognosis of ESCC is still not satisfactory. The 5-year survival rate for patients with ESCC remains poor, which is largely attributable to the high rates of extensive local invasion and regional lymph node metastasis.3,4 Therefore, it is urgent to need better preventive approaches and more effective treatment modalities.
Special AT-rich sequence binding protein 1 (SATB1) has emerged as a new regulator for cancer biology. 5 SATB1 regulates the expression of a number of targeted genes which have potential roles in oncogenic pathways.6,7 Recent studies demonstrated that the dysregulation of SATB1 is involved in multiple types of cancer;8,9 however, the critical role of SATB1 in ESCC remains unclear. The aim of this study was to determine whether SATB1 affects the biological behaviors of ESCC and further elucidate whether this effect works through the epithelial–mesenchymal transition (EMT) pathway.
Materials and methods
Tissue specimens
Ninety fresh tumor tissues were taken immediately after surgery at the First Affiliated Hospital of Sun Yat-Sen University from July 2010 to April 2015 and stored. None of the patients had received radiotherapy or chemotherapy before surgery. Tumor tissues were dissected from the resected specimens, and the normal tissue block was taken from the distal resection margin. The study was approved by the Institutional Review Board of Sun Yat-Sen University. All subjects or their caregivers have written informed consent.
Cell culture
ESCC cell lines YES2, COLO-680N, EC9706, and KYSE450 were from the tumor cell bank of Chinese Academy of Medical Sciences. A normal esophageal epithelial cell (Het-1A) was purchased from Jenniobio Biotechnology (Guangzhou, China). All the cell lines were grown in RPMI 1640 supplemented with 10% fetal bovine serum (FBS), 100 Ag/AL streptomycin, and 100 Ag/AL penicillin (pH 7.2–7.4) in a humidified incubator containing 5% CO2 at 37°C.
Quantitative real-time polymerase chain reaction analysis
Total RNA from cell or tissue was collected and extracted by using TRIzol (Invitrogen, Grand Island, NY, USA). Quantitative real-time polymerase chain reaction (qRT-PCR) analysis was performed to measure the messenger RNA (mRNA) and micro RNA (miRNA) expression. The sequences of the SATB1 primers were as follows: forward 5′-TGCAAA-GGTTGCAGCAACCAAAAGC-3′ and reverse 5′-AACATGGATAATGTGGGGCGGCCT-3′. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as a housekeeping gene for normalization, with GAPDH primers as follows: forward 5′-GTGGACCTGACCTGCCGTCT-3′ and reverse 5′-GGAGGAGTGGGTGTCGCTGT-3′. Blank samples in which the template was omitted were used as controls.
Western blot
Protein was isolated from cell or tissue using radioimmunoprecipitation assay (RIPA) buffer. Total protein was electrophoresed by 12% sodium dodecyl sulfate (SDS) and transferred to the polyvinylidene difluoride (PVDF) membranes (Millipore, Billerica, MA, USA). After blocking with 10% non-fat milk, the membrane was incubated with rabbit polyclonal anti-SATB1, anti-E-cadherin, anti-vimentin (Abcam Inc., Cambridge, CA, USA), and β-actin (Santa Cruz Biotechnology, CA, USA). Blots were measured by using the electro-chemi-luminescence (ECL) system.
Establishment of stable SATB1-knockdown cell lines
Two short-hairpin RNA (shRNA) sequences were designed based on the SATB1 sequence (NM_002971) identified by shRNA Target Finder (Ambion; Life Technologies, Carlsbad, CA, USA): shRNA1 (2566), 5′-GTCCACCTTGTCTTCTCTC-3′; shRNA2 (2364), 5′-AAGGACAATTCCGGTTTAGAG-3′. The oligoduplexes were cloned into the pSilencer3.1 vector (Ambion; Life Technologies). The resultant constructs were transfected into KYSE450 cells using Lipofectamine 2000 (Invitrogen, Life Technologies) according to the manufacturer’s instructions. Cells transfected with empty vector (pSilencer3.1) were used as controls. Stable shRNA transfectants were selected by culturing with 600 mg/mL of G418 (GibcoBRL) for 3 weeks. Western blot and qRT-PCR were used to confirm the downregulation of SATB1 expression.
CCK-8 assay and colony formation assay
CCK-8 assay was performed to evaluate the cell proliferation according the manufacturer’s instructions. Optical density (OD) was measured at 450 nm by using an enzyme immunoassay instrument (BioRad, Hercules, CA, USA). For colony formation assay, cells were cultured for 7 days. Then cells were fixed with 4% formaldehyde and stained with 1.0% crystal violet.
Transwell migration and invasion assays
Cells were suspended in 200 mL serum-free medium and seeded into the upper chamber of a transwell insert with an 8-mm pore size membrane (Corning Costar Corp, Cambridge, MA, USA). Dulbecco’s modified eagle medium (DMEM) containing 10% FBS was placed in the lower chamber as a chemoattractant. After incubation for 24 h, non-migrated cells in the upper chamber of the transwell insert were removed with a cotton swab, and the migrated cells on the underside of the filter membrane were fixed and stained with 0.1% crystal violet. The number of migrated cells was counted in five randomly selected microscopic fields and photographed. The protocol used for the invasion assay was the same as that used for the migration assay, except that the transwell insert was coated with Matrigel (BD Biosciences, Heidelberg, Germany).
Statistical analysis
Results are presented expressed as mean ± SD (standard deviation). Student’s t-test was performed to measure the difference between two groups, and differences between more than two groups were assessed using one-way analysis of variance (ANOVA). A p-value <0.05 was considered significant.
Results
Expression of SATB1 in primary ESCC tissues
To investigate the clinical significance of SATB1 expression in human ESCC progression, we first analyzed the mRNA and protein expression of SATB1 in a total of 90 pairs of esophageal tumor tissues and distal normal paired tissues by qRT-PCR and western blot. The expression level of SATB1 was significantly elevated in ESCC tissues compared with that in normal paired tissues both at the mRNA and protein levels (Figure 1(a) and (b); p < 0.01).

(a) The expression level of SATB1 mRNA in ESCC tissues; (b) the expression of SATB1 protein in ESCC tissues.
Expression of SATB1 in ESCC cell lines
The mRNA and protein expressions of SATB1 in four ESCC cell lines, YES2, COLO-680N, EC9706, KYSE450, and a normal esophageal epithelial cell line Het-1A are shown in Figure 2(a) and (b). All the four ESCC cell lines expressed higher SATB1 than the normal cell line Het-1A. SATB1 mRNA and protein was highly expressed in KYSE450 and EC9706 cells. So KYSE450 and EC9706 cell lines were chosen for a series of SATB1-knockdown experiments. To investigate the role of SATB1 gene in human ESCC progression, two stable SATB1-knockdown KYSE450 and EC9706 cell lines were established using shRNA. SATB1-shRNA1 and SATB1-shRNA2 KYSE450 and EC9706 cell lines were effective in downregulating SATB1 protein expression (Figure 2(c) and (d)). Protein expression of SATB1 was barely detectable in SATB1-shRNA1 and SATB1-shRNA2 KYSE450 and EC9706 cells.
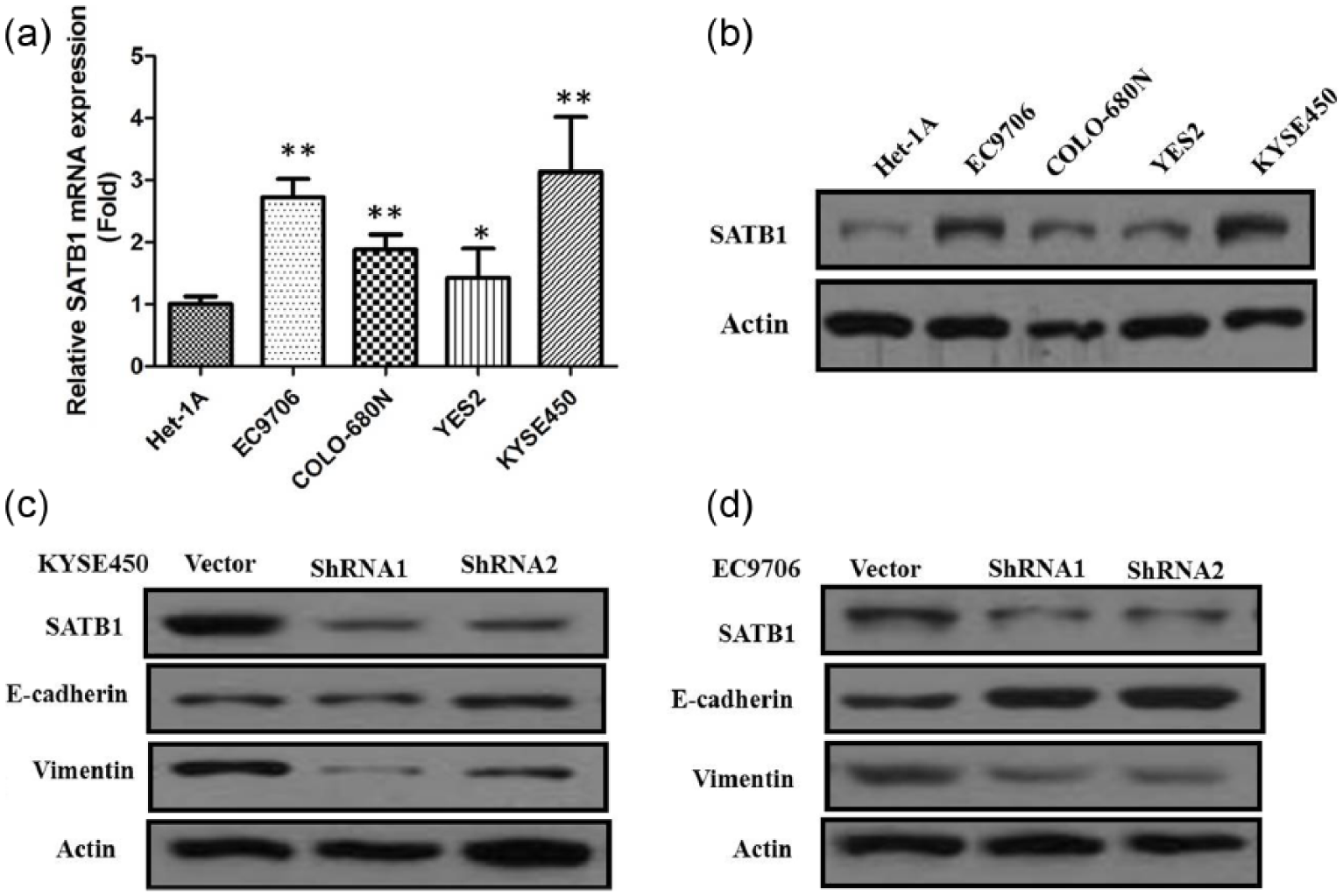
(a) qRT-PCR showing expression level of SATB1 mRNA in ESCC cell lines; (b) western blots showing expression of SATB1 in ESCC cell lines; (c) western blots showing that shRNA treatment of SATB1 markedly decreased SATB1 levels in KYSE450 cells; (d) western blots showing that shRNA treatment of SATB1 markedly decreased SATB1 levels in EC9706 cells.
Knockdown of SATB1 inhibited ESCC cell proliferation
We investigated the proliferation activity by CCK-8 assay in vitro. The results showed that the growth rate of all SATB1-shRNA KYSE450 and EC9706 cells were slower than that of the empty vector control cells (p < 0.01, Figure 3(a) and (b)). Similarly, the results of colony-formation assays revealed that clonogenic survival was significantly decreased following inhibition of SATB1 in KYSE450 and EC9706 cells by SATB1-shRNA (Figure 3(c) and (d)). These data suggest that SATB1 may act as an oncogene involved in the promotion of ESCC cell proliferation.

(a) SATB1 knockdown inhibited cell proliferation of KYSE450 cells. Cell number was measured by CCK-8 assay; (b) SATB1 knockdown inhibited cell proliferation of EC9706 cells; (c) knockdown of SATB1 expression decreased the colony formation of KYSE450 cells; (d) knockdown of SATB1 expression decreased the colony formation of EC9706 cells.
Inhibition of SATB1 could reverse EMT and suppress the migration and invasion of ESCC cells
Furthermore, the transwell assay also showed knockdown of SATB1 could significantly suppress the migratory and invasion ability of KYSE450 and EC9706 cells. In addition, the number of cells invading the rear side of the membrane in the SATB1-shRNA groups were markedly decreased in KYSE450 and EC9706 cells compared with the empty vector control cells. The migratory and invasion capability of the cells of SATB1-shRNA groups in KYSE450 and EC9706 cells were significantly reduced compared with that of the empty vector control cells (Figure 4(a) and (b); p < 0.01). To study whether SATB1 downregulation could induce EMT, western blotting was performed to show expression of EMT-relevant markers, and the results indicated that E-cadherin expression was significantly increased, while expression of vimentin was greatly reduced with SATB1 knockdown in all SATB1-shRNA KYSE450 and EC9706 cells, indicating that SATB1 is involved in the EMT process in ESCC (Figure 2).
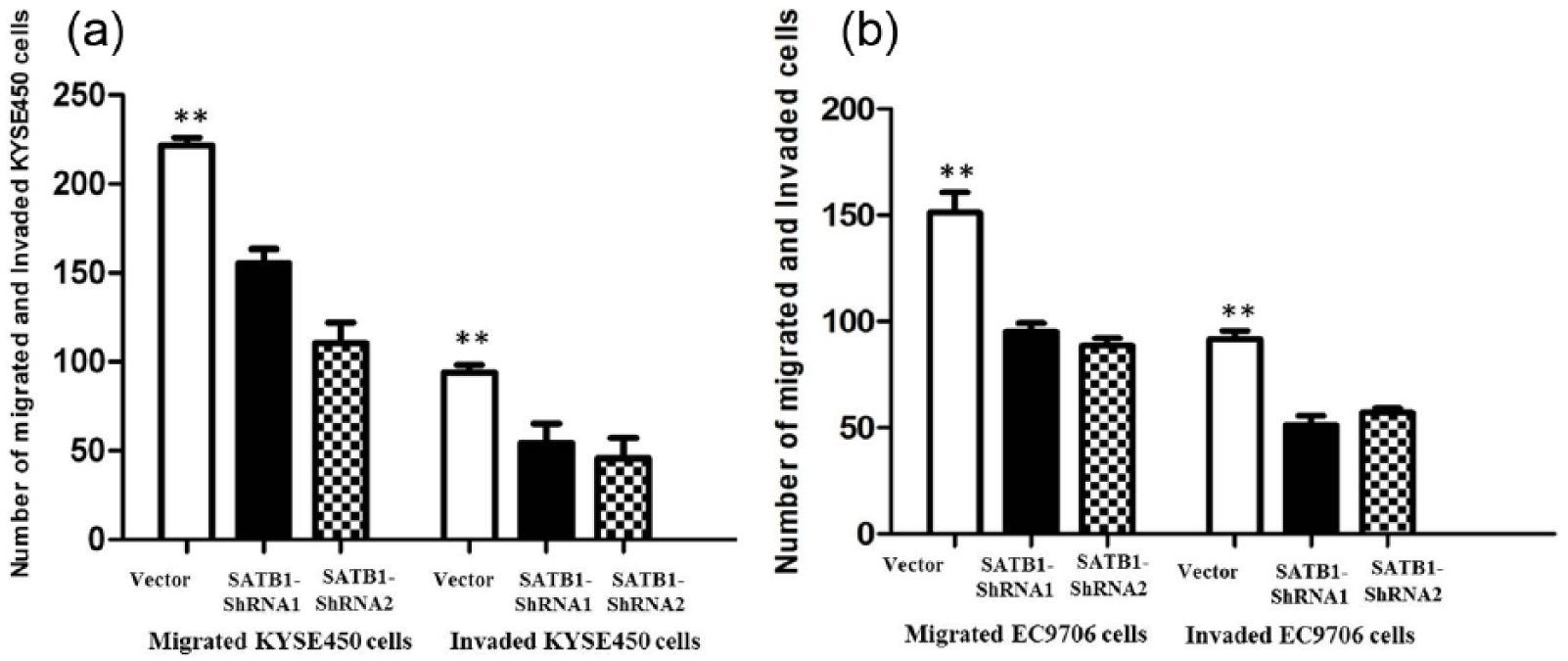
(a) Inhibition of invasion and migration of KYSE450 cells by SATB1-shRNA; (b) inhibition of invasion and migration of EC9706 cells by SATB1-shRNA.
Discussion
ESCC presents symptomatically late in the course of the disease and, despite currently available therapies, carries a poor survival rate. Although great advances have been achieved in the treatment of ESCC over the past few decades, the prognosis of ESCC remains poor, and the overall 5-year survival is less than 15%.10,11 The development of ESCC is also believed as a multiple-step process caused by the accumulation of activation of oncogene; 12 however, the molecular mechanisms underlying ESCC progression remain largely unknown. It is urgent to understand the molecular mechanisms of ESCC, and it is of crucial significance for the development of therapeutic strategies for advanced ESCC patients.
SATB1 is a nuclear matrix attachment region binding protein (MBP) which participates in many pathophysiological processes, including cancer. 13 SATB1 has recently been emphasized in various malignant tumor types.14,15 Recent studies have shown that SATB1 expression promoted the growth and metastasis of breast cancer, and that it might serve as an independent prognostic marker for predicting outcome.16,17 Nevertheless, the clinical significance of SATB1 protein in patients with ESCC remains unknown. Our study showed that the mRNA and protein expression of SATB1 were obviously increased in ESCC tissues compared with paired adjacent normal mammary tissues, which was coincident with the status of SATB1 in other cancers. Then, we used two shRNA sequences to knockdown the SATB1 expression in KYSE450 and EC9706 cells which express higher levels of SATB1. Our results showed that a reduction in SATB1 expression resulted in inhibition of proliferation of ESCC cells, and depletion of SATB1 significantly decreased the colony formation rate and colony size. Furthermore, there are potentially certain correlations between SATB1 expression and EMT markers in various cancers. 18 This is due to downregulation of Snail, ZEB2, and Slug, and upregulation of E-cadherin. Consistent with previous studies, we observed that SATB1 depletion is associated with a significant increase of E-cadherin and reduced vimentin expression in the established SATB1-shRNA KYSE450 and EC9706 cells. Then, we investigated whether SATB1 is required for the invasive phenotype of ESCC cells. In agreement with the loss of mesenchymal features and gain of epithelial features in established SATB1-shRNA KYSE450 and EC9706 cells, SATB1 knockdown decreased the migration and invasion capabilities of ESCC cells compared with the control shRNA cells.
In summary, our study showed that SATB1 expression was significantly increased in ESCC clinical samples. Moreover, inhibition of SATB1 may provide a new strategy for the prevention of ESCC invasion and metastasis. In the future, more studies are needed to verify the role of SATB1 as a reliable clinical predictor of outcome for ESCC patients.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
