Abstract
In China, the majority of ovarian cancer patients (80%–90%) are women who are diagnosed with epithelial ovarian cancer. The SYNPO2 gene has recently been reported to be associated with epithelial ovarian cancer in Europeans. To investigate the association of common variants of SYNPO2 gene with epithelial ovarian cancer in Han Chinese individuals, we designed a case–control study with 719 epithelial ovarian cancer patients and 1568 unrelated healthy controls of Han Chinese descent. A total of 49 tagging single-nucleotide polymorphisms were genotyped; single-single-nucleotide polymorphism association, imputation, and haplotypic association analyses were performed. The single-nucleotide polymorphism rs17329882 was found to be strongly associated with serous epithelial ovarian cancer and with ages ≤49 years, consistent with the pre-menopausal status of analyzed epithelial ovarian cancer cases. Odds ratios and 95% confidence intervals provided evidence of the risk effects of the C allele of the single-nucleotide polymorphism on epithelial ovarian cancer. Imputation analyses also confirmed the results with a similar pattern. Additionally, haplotype analyses indicated that the haplotype block that contained rs17329882 was significantly associated with epithelial ovarian cancer risk, specifically with the serous epithelial ovarian cancer subtype. In conclusion, our results show that SYNPO2 gene plays an important role in the etiology of epithelial ovarian cancer, suggesting that this gene may be a potential genetic modifier for developing epithelial ovarian cancer.
Introduction
Ovarian cancer is the sixth most common cancer among women and results in approximately 140,000 deaths each year. 1 In China, the overall incidence and mortality of ovarian cancer were 7.95/105 and 3.44/105, respectively, in 2009. 2 The majority of ovarian cancer patients (80%–90%) are diagnosed with epithelial ovarian cancer (EOC). Although treatment is initially effective in some patients, there is a considerable risk of recurrence and resistance to therapy. 3 Moreover, due to a lack of detection and screening strategies for early stage and effective therapeutics for late-stage metastases, most patients with EOC are misdiagnosed with widespread intra-abdominal diseases or distant metastases, making EOC the most lethal gynecologic malignancy.4,5 The origin and pathogenesis of EOC are still elusive; EOC has a complicated and heterogeneous histology that cannot be easily explained. Some epidemiological studies have identified several risk factors, such as environmental, growth, and genetic factors, in the development of EOC.6,7 The importance of genetic factors in EOC susceptibility has been recognized by recent studies via association mapping, which is facilitated by genome-wide association studies (GWASs).8–10 In several follow-up studies, large-scale samples have been used by the Ovarian Cancer Association Consortium (OCAC).11–13 For instance, a recent EOC study by OCAC reported a multi-stage GWAS with up to 15,397 cases and 30,816 controls and with six new EOC susceptibility loci (1p34.3, 1p36, 4q26, 6p22.1, 9q34.2, and 17q11.2) identified to reach genome-wide significance. 14 Although GWAS has provided a powerful approach for genetic studies of complex diseases,15–19 the genetic heterogeneity of EOC requires further studies to confirm current GWAS findings and to gain a broader view of different populations.
Podins are a family of proline-rich, actin-binding proteins that play important roles in the processes that occur in both normal and pathological cells and tissues. 20 As the second identified member of the podin family, Synaptopodin-2 (SYNPO2) was encoded by the SYNPO2 gene located at the 4q26. Loss of SYNPO2 expression had been shown to be caused by bladder cancer–associated methylation-dependent epigenetic silencing of gene expression,21,22 which indicates SYNPO2 as a repressor of cancer cell invasion. Although the role of SYNPO2 in cancer cell migration responses has been previously examined by a few researchers,23–25 the relationship between SYNPO2 expression and tumor development remains unclear. A genome-level association between the SYNPO2 gene and EOC in populations of European descent has been reported by a recent GWAS, 14 yet the underlying molecular mechanism of how the SYNPO2 gene contributes to EOC remains unknown. In addition, considering the additive effect of many minor genetic factors to disease occurrence, the exploration of possible associations between this gene and EOC among other (e.g. those who are not of European descent) genetically independent populations may shed light on its fundamental mechanism. In this study, we conducted association analyses in a group of 2287 Han Chinese females to further investigate the susceptibility of the SYNPO2 gene to EOC.
Materials and methods
Subjects
The study subjects consisted of 719 EOC patients and 1568 healthy female controls. All cases were recruited from three hospitals (The First Affiliated Hospital of Xi’an Jiaotong University, The Second Affiliated Hospital of Xi’an Jiaotong University, and Northwest Women and Children’s Hospital) between April 2012 and July 2015. Moreover, all cases were newly diagnosed and untreated EOC patients without a history of previous cancer or metastasized cancer from any other organs. If any of their relatives have been diagnosed of ovarian or breast cancer, the patients were excluded from our study. Based on independent reports from two senior pathologists, all tumors were histopathologically confirmed as primary epithelial ovarian carcinoma according to criteria from the World Health Organization. The age-matched controls, with the selection criteria of no personal or family history of ovarian or breast cancer, were recruited from healthy women at the health check-up centers of the same three hospitals. All subjects were recruited from the city of Xi’an in Shaanxi Province, China. Based on self-reports and medical records regarding their own and their paternal grandparents’ places of birth, we excluded those who were not born in Xi’an or did not have three generations of family who had originated from Xi’an. Moreover, given the fact most Chinese women are non-smokers and non-drinkers, our subjects only included women who did not smoke cigarettes or drink alcohol. For the EOC patients, detailed clinical information on the cancer diagnosis, disease FIGO (International Federation of Gynecology and Obstetrics, 2009) stage, tumor histopathology, and cell differentiation were extracted from their medical records. This study was performed in accordance with ethical guidelines from the Declaration of Helsinki (version 2002) and was approved by the Xi’an Jiaotong University Ethics Committee. All participants had written (signed) informed consent forms.
Single-nucleotide polymorphism selection and genotyping
We searched for single-nucleotide polymorphisms (SNPs) with minor allele frequencies (MAFs) over 0.01 within the genomic region of the SYNPO2 gene in the HapMap CHB database and found 118 SNPs. Afterward, MAF ⩾0.01 with pair-wise tagging and r2 ⩾0.6 were used as the cutoff criteria for tag SNPs selection, which resulted in 49 tag SNPs covering the region of the SYNPO2 gene for the study (Supplemental Table S1). Peripheral venous blood samples were collected in plain tubes. Genomic DNA was isolated from peripheral blood leukocytes according to the manufacturer’s protocol (Genomic DNA kit; Axygen Scientific Inc., Union City, CA, USA) and stored at −80°C for SNP analysis. SNPs genotyping was performed at the Sequenom MassARRAY platform with the iPLEX Gold chemistry (Sequenom, San Diego, CA, USA) based on the manufacturer’s protocols. Polymerase chain reaction (PCR) primers and locus-specific extension primers were designed with MassARRAY Assay Design software package (v3.1). A total of 50 ng of DNA was used in each multiplexed PCR well as a template. The PCR products were treated with shrimp alkaline phosphatase (USB, Cleveland, OH, USA) before the iPLEX Gold primer extension reaction. The single base extension products were desalted with SpectroCLEAN resin (Sequenom), and then an aliquot of 10 nL of the desalted product was spotted onto a 384-format SpectroCHIP with the MassARRAY Nanodispenser. Matrix-assisted laser desorption/ionization-time of flight (MALDI-TOF) mass spectrometer was employed for determination and MassARRAY Typer 4.0 software was used for data acquisition. For quality control, the statuses of both case and control samples were blind during all genotyping processes. Meanwhile, 5% of random samples were repeated and the results were 100% concordant.
Statistical analyses
A chi-square test and t-test were used to identify associations of demographic variables (age, age at menarche, and menopausal status) between cases and controls. The Hardy–Weinberg equilibrium (HWE) for each SNP was estimated using Haploview v4.2. Allelic and genotypic association tests were performed with the genetic analysis software PLINK v1.9 and a logistic model was implemented for each SNP to investigate the association between genetic polymorphisms and EOC risk. A Bonferroni correction was applied to address potential multiple comparison problems, and the strength of association between the SYNPO2 gene and EOC risk was assessed by odds ratios (ORs) and 95% confidence intervals (CIs) from the logistic model. The associations in the different subgroups of our subjects were also evaluated. In addition to the single-SNP association analyses, we also conducted haplotype-based analyses. The inter-marker relationship was determined by performing a pair-wise linkage disequilibrium (LD) with Haploview v4.2. The haplotype frequencies were estimated using GENECOUNTING v2.2. In addition, the haplotypic association analyses included a likelihood ratio test that was followed by permutation testing that was performed for common haplotypes (frequency >0.01).26,27 Imputations based on genotype data were implemented through IMPUTE2 software with 1000 Genomes CHB data as the reference panel. Subsequently, association tests based on the imputed dosage data were conducted with the software SNPTEST v2. Notably, the parameter “average certainty” calculated in IMPUTE2 was employed as the main indicator of imputation quality. 28 The threshold of indicator was chosen by exploring the patterns of Q–Q plots based on the p-values from association analyses of multiple marker sets obtained with different certainty thresholds. A p-value of 0.05 was selected as a threshold for statistical significance in all analyses.
Results
Baseline characteristics of the subjects
As shown in Table 1, 719 EOC cases and 1568 healthy female controls were matched by age with the mean age of 54.74 and 54.57 years, respectively. There were no significant differences with regard to age (p = 0.548) and menopausal status (p = 0.436) between cases and controls. Compared with the controls, the EOC cases were more likely to be of younger age at menarche (p < 0.001). Therefore, age at menarche was adjusted for any residual confounding effect in subsequent multivariate logistic regression analyses. To determine the effect of genetic polymorphisms in the SYNPO2 gene on the onset of sporadic cancer, EOC cases were divided into the following three subgroups: 40–49 years (23.1%), 50–60 years (59.5%), and 61–70 years (18.4%; Table 1). Moreover, to evaluate the potential effect of these polymorphisms on pathological characteristics, EOC cases were also divided into subgroups according to their FIGO stage, histopathological type, and histopathological differentiation grade (Table 1). A comparison among FIGO Stage IV (14.6%), FIGO Stage II (10.8%), and FIGO Stage I (8.3%) indicated that the highest numbers of EOC cases were in FIGO Stage III (66.2%). Furthermore, among the five histopathological types, the highest numbers of EOC cases were serous (44.5%) when compared to mucinous (37.3%), endometrioid (7.4%), clear cell (6.5%), and unclassifiable (4.3%) types. According to the histopathological differentiation grade, the highest numbers of EOC cases were moderate differentiation (63.7%) when compared to high differentiation (27.7 %) and low differentiation (8.6%).
Distributions of selected variables in epithelial ovarian cancer cases and healthy controls.

SD: standard deviation; FIGO: International Federation of Gynecology and Obstetrics.
Allelic and genotypic association of SNPs with EOC risk
The results of single-SNP-based association analyses of 49 SNPs, as well as the HWE test, are summarized in Table 2 and Supplemental Table S2. All genotype distributions among the subjects were in agreement with HWE (Table 2 and Supplemental Table S2). A significant association for the SNP rs17329882 (adjusted p = 0.000277) was observed, indicating that the C allele was an EOC risk allele, and the association with EOC risk still remained after Bonferroni correction (Table 2). A similar positive effect of SNP on EOC risk was further revealed by logistic regression with regard to the ORs and 95% CIs (OR = 1.37, 95% CI: 1.15–1.61; Table 2) after adjustments for age, age of menarche, and menopausal status. Moreover, genotypic association analyses also confirmed the conclusion with a similar pattern (adjusted p = 0.000976). The other 48 SNPs did not significantly differ in either genotype or allele distribution (Supplemental Table S2). In stratification analyses, an increased trend of EOC associated with rs17329882 was even significant for women younger than 49 years (p = 0.017), with an older age of menarche (>15 years, p = 0.008) and pre-menopausal status (p = 0.004), as well as in the subgroups of FIGO Stage IV, serous EOC, and high differentiation grade (p = 0.007, 0.002, and 0.002, respectively; Table 3). Furthermore, we performed power calculations for our study using PGA v2.0. Our sample size can detect SNP association with 86% power at a false-positive rate of 5% and the associated SNP (rs17329882) OR of 1.37.
Allele and genotype frequency of single-SNP association analysis.

SNP: single-nucleotide polymorphism; HWE: Hardy–Weinberg equilibrium; OR: risk allele odds ratio in cases and controls; CI: confidence interval; EOC: epithelial ovarian cancer; CTR: control.
Risk allele and significant p-values are in italics, and corrected p-values are in bold after Bonferroni correction.
Obtained in logistic regression models with adjustment for age, age at menarche, and menopausal status.
Stratification analysis for association between rs17329882 genotypes and epithelial ovarian cancer risk in Chinese women.
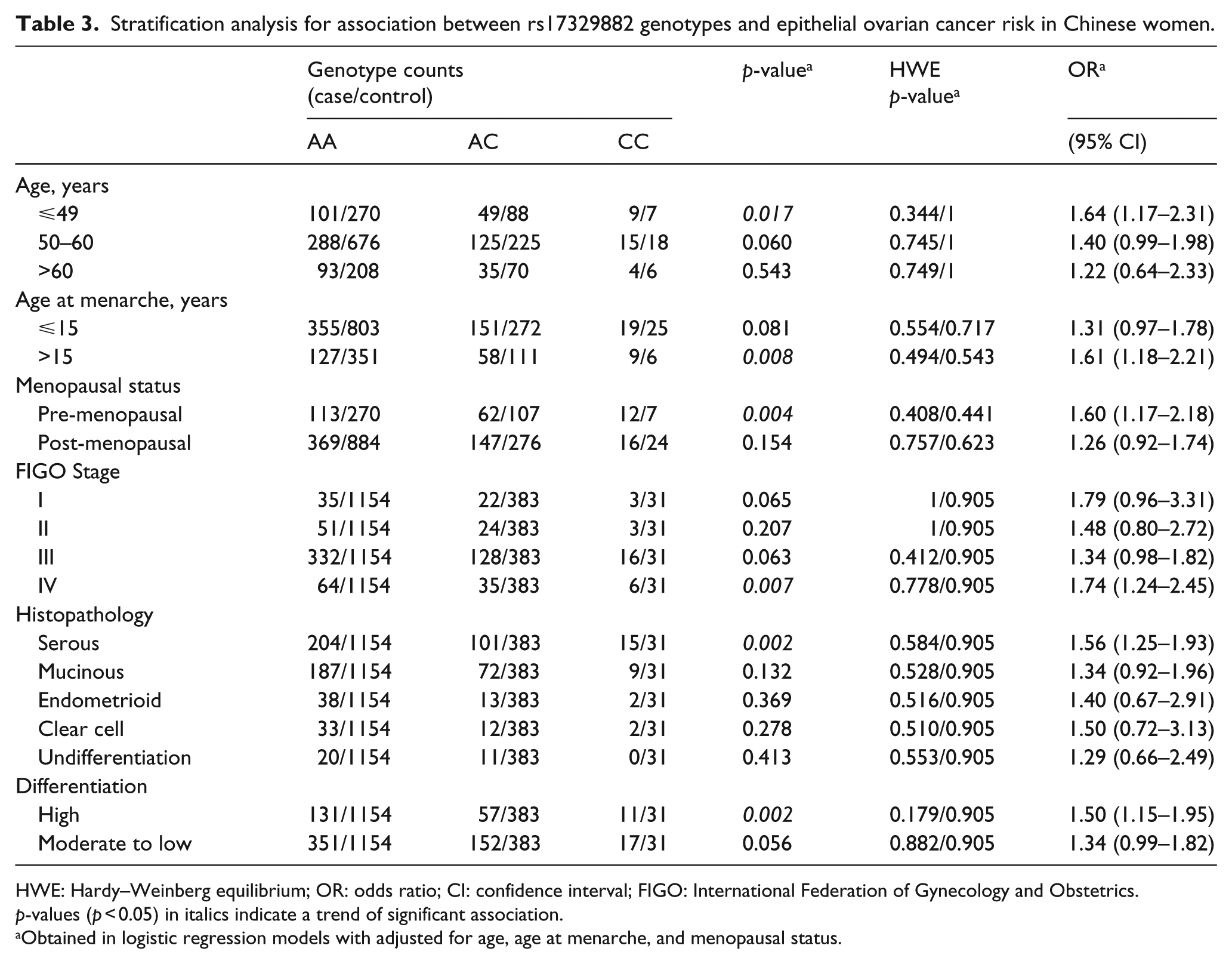
HWE: Hardy–Weinberg equilibrium; OR: odds ratio; CI: confidence interval; FIGO: International Federation of Gynecology and Obstetrics.
p-values (p < 0.05) in italics indicate a trend of significant association.
Obtained in logistic regression models with adjusted for age, age at menarche, and menopausal status.
Imputation and haplotypic association analyses
The imputation was implemented for a 5-Mb genomic region that included the SYNPO2 gene. An average certainty threshold of 0.7 was implemented to exclude imputed SNPs with low accuracy. This threshold was chosen by the pattern of Q–Q plots based on the p-values of the association analyses of multiple marker sets obtained at different certainty thresholds (Supplemental Figure S1). Moreover, MAF ⩾0.01 was employed as the second filter criterion because this work focused on common SNPs. With implement of those filters, 1784 SNPs were imputed with the reference panel and then tested for associations. We present 92 significant SNPs imputed with the reference panel (p < 0.05) in Supplemental Table S3, and the results of the association analyses based on the imputations are also shown in Figure 1.

Regional association plots based on imputed region covering the SYNPO2 gene. Imputed SNPs are indicated by circles, and genotyped SNPs are indicated by squares.
To perform haplotype-based association analyses, we examined the LD structure of 49 SNPs in genotype data and identified 16 LD blocks, among which included rs17329882 (rs17263971–rs17329882–rs2045547; Figure 2). Subsequent haplotypic association analyses were used to test the LD block (Table 4). A significant global p-value (p < 0.001) was obtained (presented in Table 4), and a haplotype in the LD block was positively associated with the EOC risk. For example, HAP4 (A-C-G) was significantly associated with EOC risk as a risk haplotype and its frequency increased almost sixfold in EOC cases (p < 0.001). Other results with a similar trend also indicated a positive genetic effect on serous EOC susceptibility (Table 4).

LD structure based on the genotype data. LD blocks are indicated by the shaded matrices, and the associated SNP rs17329882 in a red circle is located in LD Block 13.
Haplotypes’ frequency and association analyses.

EOC: epithelial ovarian cancer; CTR: control; SNP: single-nucleotide polymorphism.
Haplotypes in italics are the significant ones in the study. Rare haplotypes are not shown, if the frequency is less than 1%.
Based on 10,000 permutations.
Based on comparison of frequency distribution of all haplotypes for the combination of SNPs.
Discussion
In the current case–control study, we found that the C allele of rs17329882 in the SYNPO2 gene was significantly associated with EOC risk. Specifically, in the serous EOC subgroup, individuals harboring the CC and AC genotypes of rs17329882 exhibited higher risk for EOC than individuals with the AA genotype. ORs and 95% CI also reflected the positive effects of the C allele of the SNP on EOC. Recently, a GWAS reported that rs17329882 was significantly associated with the increased risk of EOC in populations of European descent, 14 specifically those with the serous EOC subtype. A comparison between our results and previous GWAS results indicated that the C allele was a risk effect allele in both studies. Furthermore, the OR of rs17329882 was 1.37 in our study and 1.09 in the GWAS report, while the risk allele frequency was much higher in European populations (0.16 in our samples; 0.24 in the GWAS samples). Despite the similarities in general association patterns between our study and the GWAS, there were apparent discrepancies that were potentially caused by ethnicity differences, genetic heterogeneity, sample size, and various levels of data quality. Although no additional evidence of potential interactions between covariates was involved in our study, an increased trend of EOC risk for rs17329882 C allele carriers was observed in pre-menopausal women of older age at menarche, as well as in subgroups of FIGO Stage IV and high differentiation. Although age at menarche was not significantly related to the risk of ovarian cancer overall, later age at menarche was associated with worse EOC survival.29,30 It is still unclear whether an older age of menarche is associated with EOC considering the lifetime ovulation factor in OC etiology. Further studies are necessary to confirm possible interactions in larger independent populations. Indeed, analyses of only several SNPs are insufficient to draw a conclusion.31 –35 Our haplotype analyses also indicated that the rs17329882 was significantly associated with EOC, especially for serous EOC. The SNP of rs17329882 is located in the intron region of the SYNPO2 gene. Certain type spacer sequences in the intron regions may contain functional elements with their particular functions unknown, such as transcription factor binding sites for uncharacterized transcription factors or other structural features. 36 Given that the SNP of rs17329882 may coexist in LD with other undiscovered variants or may be involved with certain unpredicted regulation machinery to confer a risk for EOC, the SNP rs17329882 will be a proper subject for future studies.
Imputation is a powerful tool for fine-mapping untyped variants in candidate gene-based association studies. The quality of imputation may affect subsequent association analyses; hence, it is necessary to exclude poorly imputed SNPs. The strict filtering criteria (certainty >0.7 and MAF ⩾0.01) of this study ensured the exclusion of inaccurately imputed markers. The information from these imputed SNPs provided us with a novel approach for scrutinizing our genetic data. Using significant imputed SNPs as the reference (Figure 1 and Table S3), a careful check through regional association plots identified a clustering pattern of a few SNPs in the SYNPO2 gene. Imputation data provided further evidence to support the association of the SYNPO2 gene with EOC risk in the Han Chinese population.
SYNPO2 is expressed in cardiac, smooth, and skeletal muscle and regulates cell shape and motility. 23 Early research has shown that the deletions of SYNPO2 gene at 4q26 and losses of myopodin protein expression seem to be associated with invasive tumor development by affecting the Rho signaling pathways, actin cytoskeleton dynamics, and cell motility.23,37 Moreover, the critical role that myopodin plays in tumor cell migration by inducing changes in actin cytoskeleton networks has also been confirmed;38,39 SYNPO2 expression, for example, is found to inhibit proliferation and invasiveness of tumor growth such as in prostate cancer cells.40,41 In a complex tumor microenvironment, SYNPO2 expression was found to inhibit the migration of neoplastic epithelial cells, to promote migration and cellular interactions of basal cells and to maintain the integrity of the basal layer, which leads to inhibition of tumor cell invasion. 20 SYNPO2 gene methylation and diminished SYNPO2 expression have also been found to have a strong relationship with bladder cancer, colon cancer and melanoma.20,21,42 Therefore, it is reasonable to hypothesize that the SNP rs17329882 within the SYNPO2 gene may be involved in the regulation of SYNPO2 expression. Further study is necessary to determine the role of the SNP rs17329882 (a functional variant or a tagging one) and the exact mechanism that underlies its association with EOC risk.
There are several challenges in our current work. First, our results need to be confirmed in other ethnic groups because SNPs and haplotype structures may vary significantly among different ethnic groups. Second, it is quite difficult to conclusively rule out other potential confounding effects such as population stratification, lifestyle, and diet. Third, future functional experiments are necessary to clarify the underlying molecular mechanisms of the association signal between the SNP rs17329882 within SYNPO2 gene and EOC. Furthermore, the majority of EOC patients had an advanced cancer (FIGO Stage III–IV) in our study. Given that the SYNPO2 gene is thought to be linked to cancer cell migration/invasion, it would be useful to consider a correlation between the SYNPO2 genetic variations and the metastatic character of EOC. However, we did not perform these analyses because of the lack of sufficient information regarding metastatic character of EOC. Finally, the study is also limited by the absence of the data related to genotype-specific expression levels. Thus, some unmeasured factors need to be taken into account in future studies.
In summary, our case–control study indicates that SYNPO2 is a susceptible gene for EOC risk in the Han Chinese population. A strong association between SNP rs17329882 and EOC risk was revealed through several sets of analyses. Further understanding of the pathological mechanisms and functional roles of the SYNPO2 gene in EOC will facilitate potential clinical applications.
Footnotes
Acknowledgements
L.M. and W.Z. contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was totally supported by the National Natural Science Foundation of China (Nos 81201925 and 81070536). The funding sources had no role in the design of this study; the collection, analysis, and interpretation of data; the writing of the report; or the decision to submit the paper for publication.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
