Abstract
Hepatocellular carcinoma is one of the leading causes for cancer-related mortality worldwide. SIRT3 may function as either oncogene or tumor suppressor in a panel of cancers; however, the role of SIRT3 in hepatocellular carcinoma remains unclear. In this study, we assayed the expression level of SIRT3 in hepatocellular carcinoma tissues by quantitative reverse transcriptase-polymerase chain reaction and immunohistochemistry. A loss-of-function approach was used to examine the effects of SIRT3 on biological activity, including cell proliferative activity and invasive potential. The results demonstrated that the expression levels of SIRT3 protein in hepatocellular carcinoma tissues were significantly downregulated compared with those in adjacent non-cancerous tissues. Furthermore, SIRT3 could decrease cell proliferation and inhibit cell migration/invasion in hepatocellular carcinoma cell line. Taken together, these results elucidated the function of SIRT3 in hepatocellular carcinoma development and suggested that SIRT3 might function as tumor suppressor in hepatocellular carcinoma by targeting PI3K/Akt pathway.
Hepatocellular carcinoma (HCC) is the fifth most common cancer with the worldwide incidence of more than 700,000 new cases annually and is currently the third leading cause of cancer-related mortality worldwide.1,2 The incidence of HCC is dramatically increased in recent years. Hepatic resection remains a potentially curative treatment for HCC, but the 5-year survival of rate is low.3–5 However, the molecular mechanisms underlying HCC development remain largely unclear. Identifying novel molecules of HCC development and elucidating molecular mechanism responsible for post-surgical recurrence and metastasis will contribute to improving overall survival of HCC patients.
Mitochondrial deacetylase sirtuin 3 (SIRT3), one member of the NAD+-dependent class III deacetylase sirtuin family, plays pivotal roles in preventing cell aging and transformation via regulation of mitochondrial metabolic homeostasis.6,7 Emerging evidences demonstrate that SIRT3 may function as either oncogene or tumor suppressor in a panel of cancers, suggesting that it is one of the important regulators of gene expression by participating in tumorigenesis.8–11 SIRT3 has been identified to be differentially expressed in many human cancers including breast cancer and prostate cancer. 12 However, the biological function of SIRT3 in HCC has been poorly characterized. The aim of this study is to elucidate the mechanism for the effect of SIRT3 on HCC. In this study, we first examined the expression of SIRT3 in HCC tissues. Furthermore, the effects of SIRT3 on cell growth, invasion, migration of HCC cell lines, and related pathway were evaluated.
Materials and methods
Human tissue specimens
A total of 92 paired HCC samples and normal pericarcinomatous tissues (located >5 cm from the tumor) were collected from 92 patients receiving surgery at the People’s Hospital of Guizhou Province. These tissue samples were conserved in liquid nitrogen after collection or prepared in paraffin sections. No systemic or local treatment had been received before operation. Both non-tumor and tumor tissues were histologically confirmed. Informed consent was obtained from each patient and was approved by the Institute Research Ethics Committee at Guizhou Medical University.
Cell culture
Human liver cancer cell line SMMC-7721, Bel-7404, Huh7, HepG2, and normal liver cell HL-7702 cells were from American Type Culture Collection (Manassas, VA, USA) and cultured in Dulbecco’s Modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum, streptomycin (100 mg/mL), and penicillin (100 U/mL).
Immunohistochemistry
Immunohistochemistry (IHC) was performed on formalin-fixed, paraffin-embedded primary HCC and adjacent normal tissues. Briefly, the slides were subjected to antigen retrieval in Target Retrieval Solution, pH 9 (DAKO) with PT Link (DAKO). Tissues were incubated in a rabbit monoclonal antibody against SIRT3 (1:100, Abcam, Cambridge, MA, USA). Negative controls were incubated with mouse or rabbit IgGs (DAKO). The percentage of positive cells was divided into four grades (percentage cores): <1% (0), 1%–25% (1), 26%–50% (2), 51%–75% (3), and >75% (4). The intensity of staining was divided into four grades (intensity scores): negative (0), weak (1), moderate (2), and strong (3). The histological score (H-score) was determined using the following formula: overall scores = percentage score × intensity score. The final score was obtained by multiplying the scores for IHC intensity by the percentage of positive cells. A final score of <3 was considered as negative, and a final score of 4–9 was considered as positive. Two histopathologists were blindly assigned to review the slides and score the staining.
Quantitative real-time reverse transcriptase-polymerase chain reaction
Total RNA was extracted by RNeasy Plus Mini Kit (QIAGEN), according to manufacturer’s recommendations. RNA concentration was determined spectrophotometrically at 260 and 280 nm and RNA samples were stored at −80°C until use. One-step reverse transcriptase-polymerase chain reaction (RT-PCR) was performed using the QuantiTect SYBR Green RT-PCR kit (QIAGEN GmbH, Hilden, Germany) following manufacturer’s protocol. Expression levels of SIRT3gene were measured by quantitative real-time RT-PCR using the LightCycler thermal cycler system (Roche Diagnostics, Indianapolis, IN, USA). Glyceraldehyde phosphate dehydrogenase (GAPDH) was used as housekeeping gene. The primers for polymerase chain reaction (PCR) are as follows: SIRT3, forward 5′-CCCCAAGCCCTTTTTCACTTT-3′ and reverse 5′-CG ACACTCTCTC AAGCCCA-3′; GAPDH, forward 5′-GAGTCAACGGATTTGGTCG T-3′ and reverse 5′-AATGAAGGGGTCATTGATGG-3′. Results were normalized to that of GAPDH and quantification of relative expression was determined by the 2−ΔΔCt method.
Western blot analysis
Protein lysates from tissues and cells were subjected to 10% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and target proteins were detected with primary antibodies anti-SIRT3, anti-PI3K, antiphospho-PI3K (Y607), anti-AKT, antiphospho-AKT (S473), and β-actin (Abcam, Cambridge, MA, USA), respectively. After incubation with appropriate horseradish peroxi-dase (HRP)-conjugated secondary antibodies (Jackson ImmunoResearch, West Grove, PA, USA), protein bands were visualized using enhanced chemiluminescence (ECL) plus Western blotting detection reagents followed by exposure to Hyperfilms (Amersham, Buckinghamshire, UK).
Recombinant adenovirus construction and tumor cell infection
The SIRT3-recombined adenoviral expression vector and the control vector were constructed by the rapid BP/LR reaction in the Gateway cloning system (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. Two SIRT3 siRNA sequences were designed using the Oligoengine software and verified by nucleotide BLAST searches. Cells grown to 50%–60% confluence in 10-cm dishes were transiently transfected with adenoviral vectors, siRNA sequences, or their corresponding mock sequences using Lipofectamine2000 (Invitrogen). After transfection for 48 h, the cells were harvested and the assays were performed including quantitative reverse transcriptase-polymerase chain reaction (qRT-PCR) and Western blot.
Cell Counting Kit-8 cell viability assay
After transfection, the cells were mixed with 10 mL of Cell Counting Kit-8 (CCK-8) solutions per well and incubated for further 1 h at 37°C. The amount of formazan dye generated by cellular dehydrogenase activity was measured for absorbance at 450 nm by a microplate reader (Molecular Devices, Sunnyvale, CA, USA). The optical density values of each well represented the survival/proliferation of HCC cells.
Colony formation assay
Cell clone formation was assessed by colony formation assay. To be brief, tumor cells (1.0 × 103 cells/60 mm culture dish) were seeded in triplicate and incubated at 37°C for 2 weeks to form clones. The cells were washed with phosphate-buffered saline (PBS), fixed with 4% paraformaldehyde for 15 min, and stained with crystal violet (1% paraformaldehyde, 0.5% crystal violet, and 20% methanol in PBS) for 30 min. The clone number on each plate was counted to measure cell survival ability.
Migration assay
In vitro migration was assessed using the Boyden chamber assay (Neuro Probe, Gaithersburg, MD, USA). Cells in serum-free culture medium were added to the upper chamber of the device, and the lower chamber was filled with 10% fetal bovine serum (FBS) culture medium. After the indicated period of incubation, the upper surface of the filter was carefully removed with a cotton swab. The filter was then fixed, stained, and photographed. Migrated cells were quantified by counting the cells in three random fields/filter. The wound-healing assay was also assessed using a culture insert (ibidi, Martinsried, Germany), which was transferred into a plate. The cells were seeded at a density of 2 × 105 cells/well and allowed to attach. After incubation, the culture insert was removed with sterile tweezers and washed with PBS. The plate was filled with culture medium containing 2% serum for cell migration. Cells were photographed for quantification according to the denuded area closure.
Statistical analysis
Data were analyzed using SPSS 13.0 software and plotted as mean ± standard deviation. Statistically significant differences were analyzed by one-way analysis of variance. p < 0.05 was considered statistically significant.
Results
SIRT3 is significantly downregulated in HCC tissues and cell lines
We evaluated the expression of SIRT3 messenger RNA (mRNA) and protein in 92 pairs of HCC and non-tumor tissues by qRT-PCR and Western blot. As shown in Figure 1(a) and (b), we found lower expression levels of SIRT3 mRNA in the HCC tissues compared to non-tumor tissues. Similar results were obtained by Western blot analysis of SIRT3 protein expression in HCC and adjacent non-cancerous tissues. Next, we investigated the SIRT3 protein levels in the human HCC and non-tumor tissues by IHC. The positive rates of SIRT3 protein detection were 65.22% (60/92) in non-tumor tissues and 28.26% (26/92) in tumor tissues, and there exists a significant difference (Figure 1(c) and (d),

(a) SIRT3 mRNA expression levels in 92 paired HCC tissues and corresponding non-neoplastic liver tissues (**p < 0.01), (b) Western blot assay showing expression level of SIRT3 in 92 paired HCC tissues and corresponding non-neoplastic liver tissues, (c) negative expression of SIRT3 in HCC tissues, and (d) positive expression of SIRT3 in corresponding non-neoplastic liver tissues.

(a) SIRT3 mRNA expression levels in a series of human HCC cell lines (**p < 0.01) and (b) SIRT3 protein expression in a series of human HCC cell lines.
SIRT3 inhibited the proliferation of HCC cells
Given that SIRT3 is markedly downregulated in HCC tissues and cell lines, we hypothesized that SIRT3 may function as a tumor suppressor. To investigate whether SIRT3 regulates HCC cell proliferation, we performed in vitro gain-of-function analyses by overexpressing SIRT3 with a lentiviral vector encoding SIRT3 in HepG2 cells. Conversely, Huh7 cells were transfected with lentiviral vectors encoding SIRT3 siRNA or control siRNA. The levels of SATB1 in the resultant cell lines with forced SIRT3 expression (designated as HepG2-SIRT3) and silenced SIRT3 expression (designated as Huh7-siSIRT3-1 and Huh7-siSIRT3-2) were verified by qRT-PCR and Western blot (Figure 3). Therefore, we investigated whether SIRT3 would affect human HCC cell proliferation. The CCK-8 cell viability assay and colony formation assay showed that SIRT3 overexpression significantly inhibited the proliferation ability and colony formation numbers of HepG2 cells (Figure 4(a) and (b)); in contrast, Huh7-siSIRT3 cells had higher rates of cell proliferation after SIRT3 knockdown (Figure 4(c) and (d)).

(a) qRT-PCR showing expression level of SIRT3 mRNA in HepG2-SIRT3 cells was significantly increased compared with control cells (**p < 0.01), (b) Western blots showing the expression of SIRT3 protein in HepG2-SIRT3 cells was significantly increased compared with control cells, (c) qRT-PCR showing expression level of SIRT3 mRNA in Huh7-siSIRT3 cells was significantly decreased compared with control cells (**p < 0.01), and (d) Western blots showing the expression of SIRT3 protein in Huh7-siSIRT3 cells was significantly decreased compared with control cells.
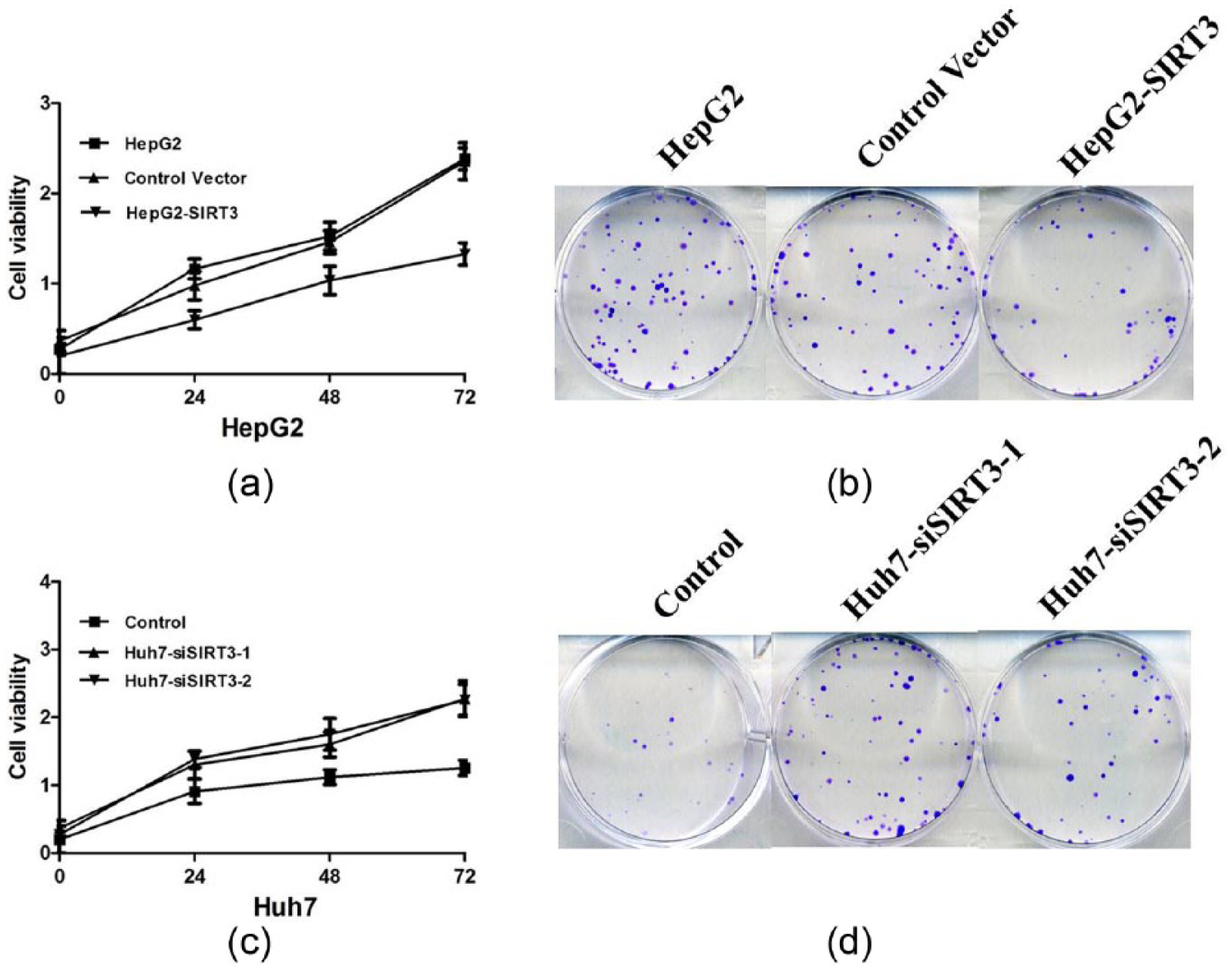
Cell proliferation was measured by CCK-8 cell viability assay; colony formation was analyzed by colony formation assay. (a) and (b) Overexpression of SIRT3 inhibits cell viability and colony formation of HepG2 cells; (c) and (d) SIRT3 knockdown promotes the cell viability and colony formation of Huh7 cells.
SIRT3 suppresses HCC cell migration and invasion in vitro
We performed transwell analyses to investigate whether SIRT3 regulates HCC cell invasion and migration. Our results showed that ectopic expression of SIRT3 significantly suppressed the invasion and migration of HepG2 cells (Figure 5(a) and (b)). In contrast, the migration and invasion of Huh7 cells sharply increased when endogenous SIRT3 was silenced by siRNA (Figure 5(c) and (d)), suggesting that SIRT3 suppresses tumor metastasis in HCC in vitro.

(a) and (b) Overexpression of SIRT3 inhibits the ability of migration and invasion of HepG2 cells; (c) and (d) SIRT3 knockdown promotes the ability of migration and invasion of Huh7 cells.
SIRT3 may regulate PI3K/AKT signaling in HCC cells
To explore the mechanisms underlying the effects of SIRT3, we investigated the effect of SIRT3 knockdown on PI3K/AKT signaling, which plays an important role in cell proliferation and survival in HCC. We observed that ectopic expression of SIRT3 significantly reduced the levels of p-PI3K and p-AKT in HepG2 cells, whereas knockdown of SIRT3 resulted in upregulation of expression of p-PI3K and p-AKT in Huh7 cells (Figure 6(a) and (b)), suggesting that SIRT3 may be involved in PI3K/AKT pathway in HCC.

(a) Ectopic expression of SIRT3 significantly reduced the levels of p-PI3K and p-AKT in HepG2 cells and (b) knockdown of SIRT3 resulted in upregulation of expression of p-PI3K and p-AKT in Huh7 cells.
Discussion
HCC is one of the most prevalent malignancies and the third leading cause of cancer-related deaths worldwide. 13 Despite advances in diagnostic and treatment modalities, the prognosis for HCC remains poor. 14 Tumor relapse after surgical resection and distant metastasis are the major causes of the high mortality. 15 Therefore, it is urgent to explore the molecular mechanisms of HCC progress.
SIRT3 was discovered to be associated with regulating multiple cellular and physiological processes in various cancers.16,17 Accumulating studies demonstrated that many types of cancers exhibited aberrant expression or deletion of SIRT3.18–20 However, the function of SIRT3 in HCC development remains enigmatic. In our study, significantly reduced expression of SIRT3 mRNA and protein were observed in HCC tissues, in comparison with normal adjacent tissues by qRT-PCR, IHC, and Western blot analysis. Consistent with our result, Zhang et al. 21 have reported that Sirt3 was downregulated significantly in human HCC tissue compared with peritumoral normal tissue, suggesting the loss of Sirt3 may participate in the genesis and development of HCC.
Then, we used different HCC cell lines to explore possible regulatory mechanism of SIRT3 in the tumorigenesis of HCC. We also showed that SIRT3 expression was downregulated in a panel of HCC cell lines compared with a normal liver cell line HL-7702. Then, we performed gain and loss-of-function experiments in HepG2 and Huh7 to determine the possibility of SIRT3 as a therapeutic target of HCC. We employed the siRNA technique to knockdown of the SIRT3 expression in Huh7 cells in which SIRT3 expression is relatively high; and we overexpressed the SIRT3 levels by viral transduction in HepG2 cells in which SIRT3 expression is relatively low.
SIRT3 can act as a tumor suppressor gene or oncogene in the initiation and development of tumors. Our results found that overexpression of SIRT3 expression had an antitumorigenic effect on HepG2 cells; however, knockdown of SIRT3 in Huh7 cells could result in increased cell proliferation. Moreover, SIRT3 could significantly reduce the colony formation activity of HepG2 and Huh7 cells, in comparison with control. Combining with the previous reports, these observations further confirmed that SIRT3 has key roles in suppressing proliferation of HCC cells. Next, we examined the effect of SIRT3 in HCC cells on migration and invasion by transwell assay. Our transwell migration/invasion systems suggested that overexpression of SIRT3 markedly attenuated the migratory and invasive potency of HepG2 cells in vitro. On the contrary, silencing of SIRT3 significantly promoted Huh7 cell migration and invasion as determined by transwell assay.
To understand the molecular mechanism of SIRT3 in HCC, we focused on specific oncoproteins which could be regulated by SIRT3. Activation of the survival signal PI3K/Akt pathway is closely associated with proliferation and migration of cancer cells. As an oncoprotein, AKT has been reported to maintain tumorigenicity and accelerate tumor growth in different cancer types. In this study, we observed that ectopic expression of SIRT3 significantly reduced the levels of p-PI3K and p-AKT in HepG2 cells, whereas knockdown of SIRT3 resulted in upregulation of expression of p-PI3K and p-AKT in Huh7 cells, indicating that SIRT3 inhibited cell growth via suppressing PI3K/AKT signaling.
In summary, this is a preliminary study to investigate the role of SIRT3 in HCC. We found that SIRT3 has tumor-suppressive roles in HCC via inhibition of cell growth, migration, and invasion. We have also shown that SIRT3 suppresses HCC by inhibiting the activation of PI3K/Akt. Thus, SIRT3 may be a potential biomarker and therapeutic target for HCC.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This research was supported, in part, by the youth fund of The People’s Hospital of Guizhou Province (GZSYQN [2015] 01) and the United Fund of Science and Technology Agency of Guizhou Province (LH2014-7013).
