Abstract
Background/Objectives
Diabetes and cancer are common diseases, and epidemiologic evidence suggests that people with diabetes are at significantly higher risk for many forms of cancer. Both these diseases are among the highly complex, heterogeneous, and robust. Transcriptional dysregulation and abnormal signaling are major contributors to disease heterogeneity, posing challenges for therapeutic development. These days, herbal compounds are also used for the diagnostic purpose for many diseases, including diabetes and cancer.
Materials and Methods
Here, to understand the common genes and the commonly enriched functions (pathways) between the two diseases and to unravel the potential herbal targets, we have studied the publicly available gene expression datasets (diabetes and cancer) and performed docking for the protein structure of the common 31 genes against the herbal compound of our interest—apigenin, quercetin, and resveratrol.
Results
We identified 31 common differentially expressed genes and 14 enriched pathways, including insulin and phosphatidylinositol 3-kinase-protein kinase B signaling, critical to both diseases.
Conclusion
Molecular docking revealed eukaryotic translation initiation factor 5B, DEAH-box helicase 16, integrin beta 7, and O-GlcNAc transferase as high-affinity targets for apigenin, quercetin, and resveratrol, with favorable absorption, distribution, metabolism, and excretion properties (e.g., no Lipinski violations).
Introduction
Among the most common and complex diseases, diabetes and cancer are common diseases, and epidemiologic evidence suggests that people with diabetes are at significantly higher risk for many forms of cancer.1–3 These diseases are considered very hard in terms of diagnosis due to their highly complex and heterogeneous nature. The complexity arises from transcriptional dysregulation and pathway-level aberrations, which hinder therapeutic strategies.1, 4–6 Recent studies highlight shared insulin/phosphatidylinositol 3-kinase-protein kinase B (PI3K-AKT) signaling in cancer and diabetes. 7
In a simplified way, it can also be said that cancer and/or diabetes arise due to failure at multiple levels in multicellular organisms, including genetic lesions, abnormal signaling, and metabolic disorders. After such changes, the cell appears distinct from the normal cell, referred to as a cancer cell. Such cells appear to be different in their physiological behavior and morphology compared to the respective normal cells. The incidence of breast cancer and diabetes mellitus in females in Saudi Arabia is nearly 21% and 21.5%, respectively. The studies revealed that approximately 27.9% of the females were not aware of having diabetes mellitus despite the availability of advanced diagnostic facilities, and these subsets of the female population have a high susceptibility to the development of breast cancer at a later age. 8 To no surprise, diabetes mellitus in the region of Ha’il accounts for the highest percentage, and diabetes patients in the Ha’il region have a strong family history of breast cancer. 9
These days, herbal compounds are also used for diagnostic purposes in the treatment of various diseases, including diabetes and cancer.2–4 There is a large number of natural products that have pharmacological or biological activities and are considered to have therapeutic benefits in treating diseases. They are also considered an important source of inspiration for the development of potential novel drugs. 2 With advancements in technologies, natural products and their derivatives are of potential interest for research to unravel therapeutic agents.10–13
Here, to understand the common genes and the commonly enriched functions (pathways) between the two diseases and to unravel the potential herbal targets, we have studied publicly available gene expression datasets (diabetes and cancer) and performed docking for the protein structure of the standard 31 genes against the herbal compounds of our interest—apigenin, quercetin, and resveratrol.14–16
Materials and Methods
Data Acquisition and Normalization
In the initial step, we selected the raw expression datasets of interest, namely GSE10797 (cancer) 17 and GSE13760 (diabetes), 18 and processed them through normalization, which was performed using the Empirical Bayes (EB) method, obtaining log2 values for all mapped genes as illustrated in the workflow (Figure 1a).
Comparative Gene Expression Profiling. (a) Workflow, (b) and (c) Venn Diagram for Epithelial and Stromal Breast Cancer Differentially Expressed Genes (DEGs) and Enriched Pathways, (d) and (e) Venn Diagram for Epithelial, Stromal Breast Cancer, and Diabetes DEGs and Enriched Pathways, (f) and (g) Common Genes and Pathways Between Cancer and Diabetes.
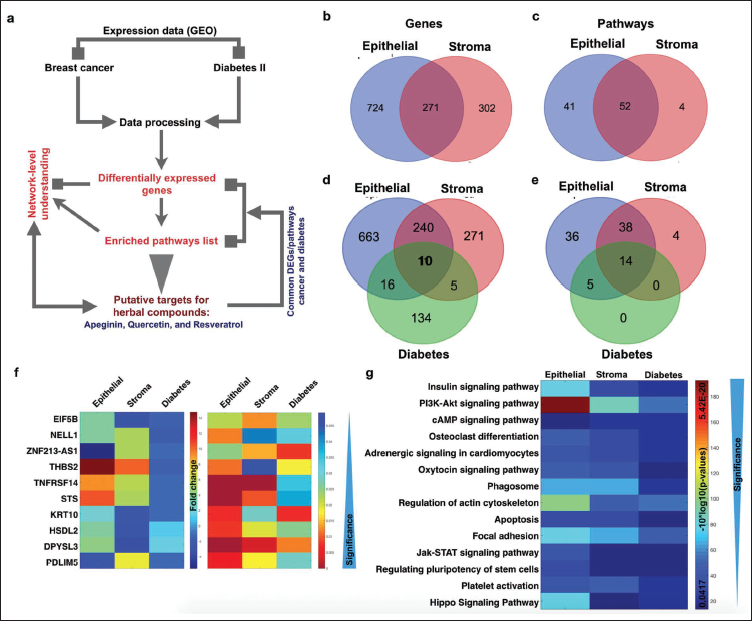
Differential Expression Analysis
The GSE10797 dataset comprised 66 samples, including 5 normal epithelial, 5 normal stroma, 28 cancer stroma, and 28 cancer epithelial samples from the U133A 2.0 platform. The GSE13760 dataset consisted of 21 samples from individuals with type 2 diabetes and 10 control samples, totaling 31 samples for comparison. For the analysis of differential gene expression, we conducted comparisons between tumor and standard samples within their respective datasets, resulting in four lists of differentially expressed genes (DEGs). In summary, the fundamental steps encompass raw file processing, intensity calculation, and normalization. For normalization, the most commonly employed approaches include GC robust multi-array average (GCRMA), robust multi-array average (RMA), and EB.19–26 In this study, we utilized the EB method for raw intensity normalization. Following normalization, our objective was to comprehend gene expression patterns and their inferred functions.27, 28 In the prediction of differential gene expression and statistical analysis, MATLAB functions such as the MAT test were employed.
Pathway and Network Analysis
For pathway analysis, we referred to the Kyoto Encyclopedia of Genes and Genomes (KEGG)
29
database and implemented our code designed for pathway and network analysis. A further STRING database was also used to characterize the common pathways associated with cancer and diabetes. The list of 31 common genes between diabetes and cancer was uploaded to the STRING database (
For generating DEG networks, FunCoup2.034 was employed for all networks throughout the study, and Cytoscape 3.1 was used for network visualization. MATLAB was the primary tool for coding and calculations in most instances. FunCoup predicts four different classes of functional coupling or associations, including protein complexes, protein–protein physical interactions, metabolic, and signaling pathways. 30
Molecular Docking (MD)
For docking analyses, SwissDock was utilized, and the docking was performed using the web interface against all three selected herbal compounds for the common DEGs between diabetes and cancer. 31
Absorption, Distribution, Metabolism, and Excretion (ADME) Analysis
QikProp predicted pharmacokinetic properties (e.g., molecular weight, donor/acceptor H-bonds) to assess drug-likeness (Lipinski’s Rule of Five).
Results
Based on our analysis, we conclude that there are 31 DEGs and 14 enriched pathways. Among the 14 pathways, insulin, PI3K-AKT, osteoclast, adrenergic, and oxytocin are the pathways known to play critical roles in cancer and diabetes. Eukaryotic translation initiation factor 5B (EIF5B), DEAH-box helicase 16 (DHX16), integrin beta 7 (ITGB7), and O-GlcNAc transferase (OGT) display the lowest full fitness and delta G for all the three selected herbal products—apigenin, quercetin, and resveratrol.15, 16
A comparative analysis of GSE10797 (cancer) and GSE13760 (diabetes) identified 31 common DEGs (Figure 1a). Pathway enrichment revealed 14 shared pathways, including insulin and PI3K-AKT signaling (Figure 2).
Network-level Understanding of the Differentially Expressed Genes (DEGs) and Their Potential Impact. (a) Common DEGs Between Cancer and Diabetes and Their Known Association Based on Network Database, (b)–(h) Selected Genes from Common DEGs and Their Associated DEGs from Over DEGs List (Cancer and Diabetes Both).
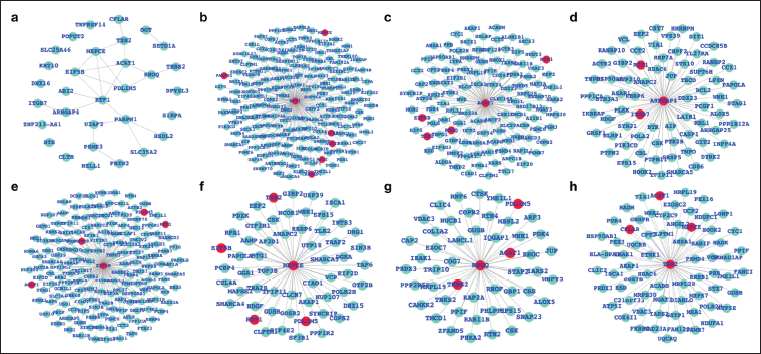
Combined and comparative gene expression profiling for breast cancer and diabetes: In this work, our primary focus has been on the combined study of gene expression patterns in breast cancer and diabetes, aiming to identify potential targets for herbal extracts shared between the two conditions.
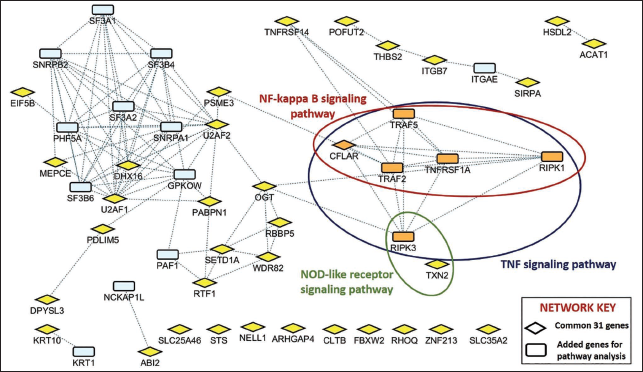
Insulin, AKT, osteoclast, and oxytocin signaling pathways, and among the commonly enriched pathways: Given our main objective to comprehend the potential list of genes and functions crucial in cancer and diabetes as putative markers and targets, we have concentrated on the overall common DEGs and the commonly enriched pathways between diabetes and cancer (Figure 2). The connectivity of genes shared between cancer and diabetes indicates that out of the 31 genes, 20 have a substantial number of links, and 6 genes have more than 50 links with the overall DEGs. This implies that genes shared between cancer and diabetes play a dominant role in controlling functions, with EIF5B, neural EGFL like 1 (NELL1), and ZNF213 antisense RNA 1 (ZNF213-AS1) being connected to more than 100 DEGs in both cancer and diabetes (Figure 3). Furthermore, pathways such as the spliceosome, tumor necrosis factor (TNF) signaling pathway, necroptosis, nuclear factor kappa B (NF-κB) signaling pathway, nucleotide-binding oligomerization domain-like receptor (NLR) signaling pathway, and apoptosis were found to be significantly enriched (Figure 4). However, an intensive literature search revealed that only TNF signaling, NF-κB signaling pathway, and NLR signaling pathway were involved in the pathogenesis of both cancer and diabetes. Recent studies have shown that TNF-α plays anti-neoplastic roles, and it exerts its effect through activating distinct signaling pathways such as NF-κB. Furthermore, TNF-α is also directly known to interfere with the signaling of insulin through its receptor and consequently blocks the biological actions of insulin. Similarly, these immune receptors (toll-like receptors (TLRs) and NLRs) are reported to be involved in obesity-mediated insulin resistance and are upregulated in monocytes from patients with type 2 diabetes compared with healthy individuals. The role of NLRs is also known in mediating inflammation-associated cancer progression, offering the opportunity to target these pathways.
Common Breast Cancer and Diabetic Herbal Putative Target Genes and the Inferred Pathways. (a) Connectivity of the Cancer and Diabetic Genes with All the Differentially Expressed Genes. (b) Protein Classification, (c) Structure of the Three Herbal Drugs Used for Identification of In Silico Drug Targets, and (d) SwissDock Parameters for the Common Drug Targets.
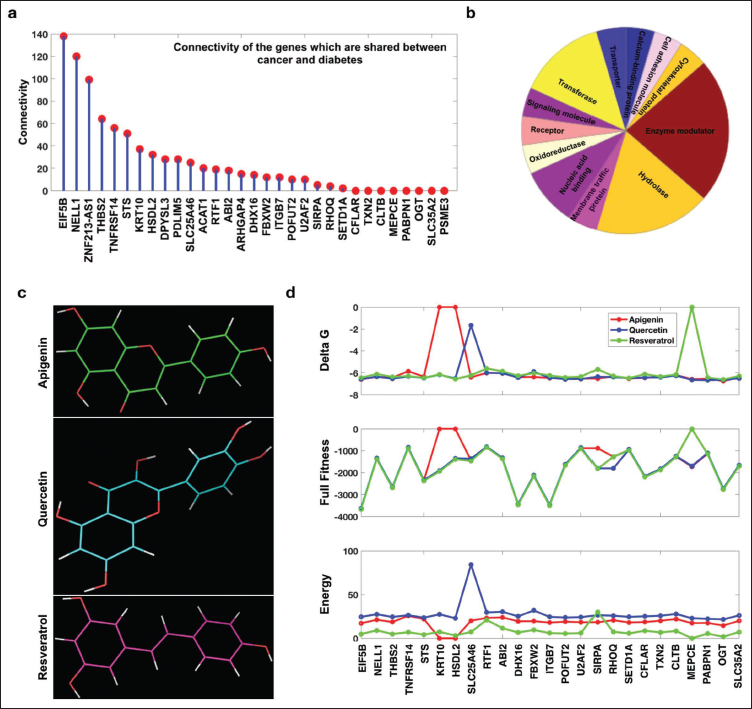
Network Analysis of Common 31 Genes Depicting Shared Pathways Between Diabetes and Cancer.
Herbal targets from common DEGs: Prior to exploring the docking interactions with our herbal compounds, we analyzed the overall connectivity of the 31 shared genes (between diabetes and cancer) (Figure 3a) and additionally examined the protein classes using Panther (Figure 3b). 32 Among these 31 genes, it is observed that enzyme modulators, hydrolases, and nucleic acid-binding proteins are in the majority (Figure 3b). Our focus then shifted to potential markers for the herbal compounds apigenin, quercetin, and resveratrol (Figure 3c). We generated 3D structures for all common DEGs and performed MD. Utilizing the SwissDock server, 31 which takes the 3D structure of proteins and herbal compounds, we obtained results with various parameters. In this study, we emphasized shared genes and pathways, conducting docking specifically for these shared genes. Out of the 31 genes, we obtained SwissDock output for 25 genes. Our MD results indicate that there is no substantial difference among the three herbal compounds; they are very close to each other in terms of total fitness and delta G, with few exceptions among the 25 proteins. Furthermore, ADME analysis using QikProp (a tool for predicting pharmacokinetic and physicochemical properties) also reveals drug-like properties of all the three compounds (Table 1), with no violations of Lipinski’s Rule of Five. However, there is a notable difference in terms of energy (Figure 3d). The docking profile reveals that EIF5B, DHX16, ITGB7, and OGT could be potential herbal targets for both diabetes and cancer.
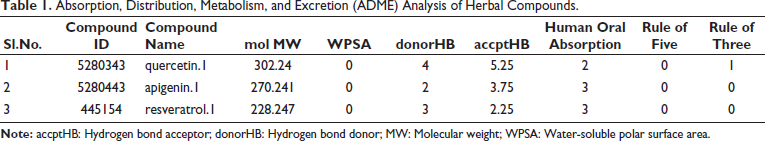
Absorption, Distribution, Metabolism, and Excretion (ADME) Analysis of Herbal Compounds.
Discussion
Recent technological advances addressing challenges in drug discovery, combined with unrealized expectations from current lead-generation strategies, have renewed interest in natural products. Substances derived from animals, plants, and microbes have been used in medicine since its inception. Herbal products are often sought by patients with various diseases, including cancer, as numerous plants and their extracts are considered potential sources for treating different conditions.33–36 In contrast to previous work, our study involves a combined analysis of gene expression profiling for publicly available datasets. We analyzed common DEGs and enriched pathways and conducted a straightforward docking study using SwissDock. 17 Our study suggests major putative drug targets for diabetes and cancer. We selected three herbal compounds—apigenin, quercetin, and resveratrol—to identify potential targets common to both diabetes and cancer. Apigenin, a trihydroxyflavone, induces autophagy in cancer cells and functions as a metabolite and an anti-neoplastic agent. Quercetin, a polyphenolic flavonoid, exhibits chemopreventive, anti-inflammatory, and anti-allergy effects. Resveratrol, a plant polyphenol found in red grapes, is considered a potential herbal drug for treating hyperlipidemia, preventing fatty liver, diabetes, atherosclerosis, and aging. In summary, we progressed from gene expression profiling to the identification of drug targets. Gene expression profiling identified common DEGs between epithelial breast cancer, stromal breast cancer, and diabetes, along with commonly enriched pathways (Figure 1). For the common DEGs, we explored networks and connectivities, observing that EIF5B, NELL1, and ZNF213-AS1 have more than 90 connections with other DEGs. Several genes show higher connections, indicating that these common genes directly or indirectly influence a large number of genes in the overall DEGs list.
Conclusion
Our analysis concludes that there are 31 common DEGs and 14 commonly enriched pathways. Notably, insulin, PI3K-AKT, osteoclast, adrenergic, and oxytocin are pathways known to play critical roles in cancer and diabetes. MD results reveal that EIF5B, DHX16, ITGB7, and OGT consistently exhibit the lowest full fitness and delta G for all three herbal compounds—apigenin, quercetin, and resveratrol.
Footnotes
Abbreviations
accptHB: Hydrogen bond acceptor; ADME: Absorption, distribution, metabolism, and excretion; DEGs: Differentially expressed genes; DHX16: DEAH-box helicase 16; donorHB: Hydrogen bond donor; EB: Empirical Bayes; EIF5B: Eukaryotic translation initiation factor 5B; GCRMA: GC robust multi-array average; ITGB7: Integrin beta 7; KEGG: Kyoto Encyclopedia of Genes and Genomes; MD: Molecular docking; MW: Molecular weight; NELL1: Neural EGFL like 1; NF-κB: Nuclear factor kappa B; NLRs: NOD-like receptors; NOD: Nucleotide-binding oligomerization domain; OGT: O-GlcNAc transferase; PI3K-AKT: Phosphatidylinositol 3-kinase-protein kinase B; RMA: Robust multi-array average; TLRs: Toll-like receptors; TNF: Tumor necrosis factor; WPSA: Water-soluble polar surface area; ZNF213-AS1: ZNF213 antisense RNA 1.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval and Informed Consent
Not applicable.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research has been funded by the Scientific Research Deanship at the University of Ha’il, Saudi Arabia, through project number RG-21 160.
