Abstract
Background
Oral squamous cell carcinoma (OSCC) is a prevalent form of head and neck cancer characterized by aggressive behavior and a poor prognosis. Conventional therapies have demonstrated limited effectiveness, underscoring the need for innovative strategies that target the molecular mechanisms involved in OSCC progression. Multitargeting agents present a promising approach by simultaneously addressing several key pathways, potentially addressing issues of treatment resistance. Desmostachya bipinnata, a medicinal plant renowned for its anticancer properties, contains bioactive compounds that may serve as effective treatments for OSCC.
Objectives
This study aims to investigate the therapeutic potential of bioactive compounds from Desmostachya bipinnata in treating OSCC. It uses bioinformatics and molecular docking techniques to identify key molecular targets and pathways, evaluate compound binding affinities, and propose novel multitargeting agents for OSCC therapy.
Materials and Methods
This study aimed to explore the therapeutic potential of Desmostachya bipinnata compounds for OSCC using bioinformatics and molecular docking. Six of the 19 compounds screened were excluded due to toxicity, leaving 14 for further analysis. GeneCards, DisGeNet, and Gene Expression Omnibus (GEO) databases identified 3,278 OSCC-related genes, and SwissTargetPrediction predicted 221 targets. Protein–protein interaction and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis pinpointed significant hub genes. Molecular docking of four selected compounds (linoleic acid, kaempferol, daucosterol, stigmasterol-glucoside) with six key targets (MMP2, PTGS2, STAT3, MAPK1, MMP9, AKT1) revealed strong binding affinities, suggesting potential therapeutic efficacy.
Results
This study evaluated potential therapeutic compounds from Desmostachya bipinnata for OSCC through a comprehensive approach. After assessing the toxicity of 19 compounds, six were excluded due to predicted adverse effects, leaving 14 for further analysis. We identified 3,278 OSCC-related genes by integrating data from GeneCards, DisGeNet, and GEO databases. Using SwissTargetPrediction, we narrowed down 221 unique targets for these compounds and identified 95 common targets with OSCC genes. Protein–protein interaction analysis via STRING and Cytoscape, along with Molecular Complex Detection (MCODE), highlighted a significant gene cluster. Expression analysis with Gene Expression Profiling Interactive Analysis (GEPIA) led to the exclusion of low-expressing genes (IL6, MAPK3, ESR1, BCL2), focusing on MMP2, PTGS2, STAT3, MAPK1, MMP9, and AKT1, which are involved in cancer-related pathways. Molecular docking studies showed that linoleic acid, kaempferol, daucosterol, and stigmasterol-glucoside exhibit strong binding affinities to these targets, suggesting their potential as effective therapeutic agents. Activity predictions confirmed their antineoplastic properties, underscoring their potential utility in OSCC treatment.
Conclusion
The findings indicate that Desmostachya bipinnata compounds exhibit promising multitargeting activity against OSCC. The strong binding affinities and interaction profiles of these compounds with key OSCC-related targets support their potential as effective therapeutic agents. Further experimental validation is needed to confirm these results and explore the clinical applicability of these compounds in OSCC treatment.
Keywords
Introduction
Oral squamous cell carcinoma (OSCC) is a major type of head and neck cancer, characterized by its aggressive nature and poor prognosis. 1 Despite advances in surgical techniques, radiotherapy, and chemotherapy, the survival rate for OSCC remains low, primarily due to late diagnosis and the development of resistance to conventional therapies.2, 3 This underscores the urgent need for novel therapeutic strategies that effectively target the molecular mechanisms driving OSCC progression. 4
One promising approach is using multitargeting agents that can simultaneously modulate several critical pathways involved in cancer development and metastasis. 5 Such agents can potentially overcome the limitations of single-target therapies, which often lead to resistance and limited efficacy. 6 In this context, natural compounds have gained attention for their multitargeting capabilities and lower toxicity profiles than synthetic drugs. 7
Desmostachya bipinnata, a medicinal plant known for its wide range of pharmacological properties, has shown potential as a source of bioactive compounds with anticancer activities. 8 Previous studies have highlighted various compounds from Desmostachya bipinnata that exhibit anti-inflammatory, antioxidant, and antiproliferative effects, making it a candidate for cancer therapy. 9
To systematically explore the therapeutic potential of Desmostachya bipinnata in OSCC, this study employs bioinformatics and molecular docking approaches. Bioinformatics tools facilitate the identification of key molecular targets and pathways associated with OSCC, while molecular docking studies allow for the evaluation of binding affinities and interactions between bioactive compounds and these targets.10, 11 This integrated approach aims to uncover the multitarget potential of Desmostachya bipinnata compounds, providing insights into their mechanisms of action and paving the way for future experimental validation.
Materials and Methods
Compound Selection and Toxicity Analysis
Bioactive compounds from Desmostachya bipinnata were initially retrieved from existing literature, resulting in a list of 19 compounds. These compounds underwent rigorous toxicity analysis using ProTox II webserver
12
(
Target Identification for OSCC
A comprehensive gene search was conducted across multiple databases to identify genes associated with OSCC. Three different databases were used: GeneCards, DisGeNet, and Gene Expression Omnibus (GEO).13, 14 Using GeneCards, a keyword search for “OSCC” yielded 2,304 genes. In the DisGeNET database, a search for “Oral cancer” identified 734 genes. Additionally, the GEO database (GSE37991) was analyzed to identify differentially expressed genes in OSCC samples. This analysis included 40 OSCC samples and 40 standard samples, resulting in identifying 999 upregulated genes with a log fold change (LogFC) greater than one and an adjusted p value of less than .05.
Compound–Target Prediction
The SwissTargetPrediction tool was utilized to predict potential targets for the selected 14 nontoxic compounds (
Protein–Protein Interaction (PPI) and Expression Analysis
To understand the interactions and pathways involved, a PPI network was constructed using the STRING database (version 12.0) with a confidence score of 0.700 (
KEGG Pathway Enrichment
Significant hub genes with high expression profiles were analyzed to determine their involvement in various Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways. This analysis was conducted using the ShinyGo 0.76 web server, a tool designed for pathway enrichment analysis (
Molecular Docking Studies
The three-dimensional structure of the protein encoded by these genes was downloaded from Protein Data Bank (PDB) (
Docking studies were conducted using PyRx software. Initially, the proteins were prepared by removing water molecules and heteroatoms, and hydrogen atoms were added. The one with the best binding affinity was selected for each compound among the different conformations generated. The docking results were visualized using Discovery Studio Visualizer.
The docking interactions were assessed based on binding affinity (kcal/mol) and interacting amino acids. The results demonstrated the potential efficacy of the bioactive compounds from Desmostachya bipinnata in binding effectively with these key targets, indicating their promise as therapeutic agents against OSCC.
Activity Prediction
To predict the biological activities associated with the identified leads, the PASS (Prediction of Activity Spectra for Substances) webserver was utilized (
Results
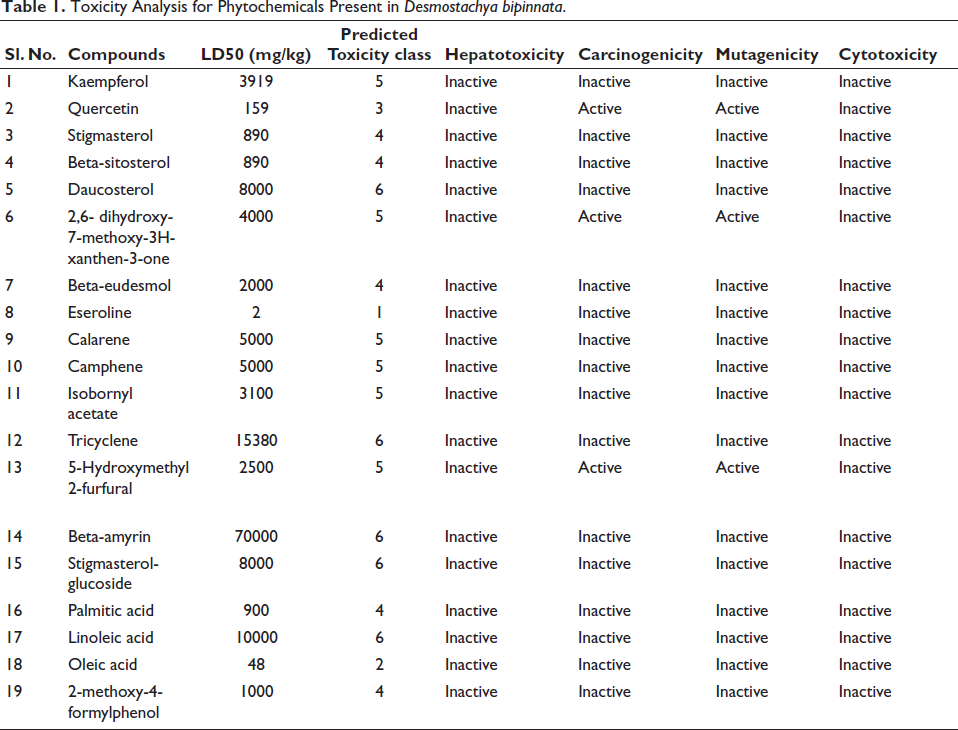
During the toxicity analysis, it was observed that out of 19 compounds, 6 compounds—quercetin, 2,6-dihydroxy-7-methoxy-3H-xanthen-3-one, eseroline, 5-hydroxymethyl-2-furfural, and oleic acid—were predicted to have adverse toxic effects. Consequently, these six compounds were excluded from further analysis to ensure the safety and efficacy of the potential leads. The remaining 14 compounds did not show significant toxicity and were selected for compound target prediction (Table 1). This step aimed to identify potential molecular targets and pathways associated with these compounds, facilitating the selection of promising candidates for further experimental validation and development.
Toxicity Analysis for Phytochemicals Present in Desmostachya bipinnata.
OSCC Targets
Upon analysis, 2,304 genes were identified using GeneCards, and 734 genes were obtained from DisGeNet. Gene expression analysis was also performed on 40 OSCC samples and 40 normal samples from the GEO database. The criteria for upregulated genes were set with a LogFC (log fold change) > 1 and an adjusted p value < .05. This analysis resulted in the identification of 999 upregulated genes in GEO.
Combining the results from all three databases—GeneCards, DisGeNet, and GEO—and removing duplicate entries, a total of 3,278 genes related to OSCC were obtained. This comprehensive gene list serves as a valuable resource for further investigations into the molecular mechanisms and potential therapeutic targets for OSCC.
Compounds Target
The targets of the 14 compounds were predicted using the SwissTargetPrediction tool, with a probability score threshold set at >0.06 (Supplementary Figure S1). This analysis initially identified 383 potential targets. After consolidating the data and removing duplicates, 221 unique targets were obtained. These unique targets provide a focused list for further exploration of the compounds’ potential interactions and therapeutic applications. A Venn diagram was constructed using the interactive Venn online web server (
Heatmap Visualizing the Relationship Between Compound Targets and OSCC Targets.
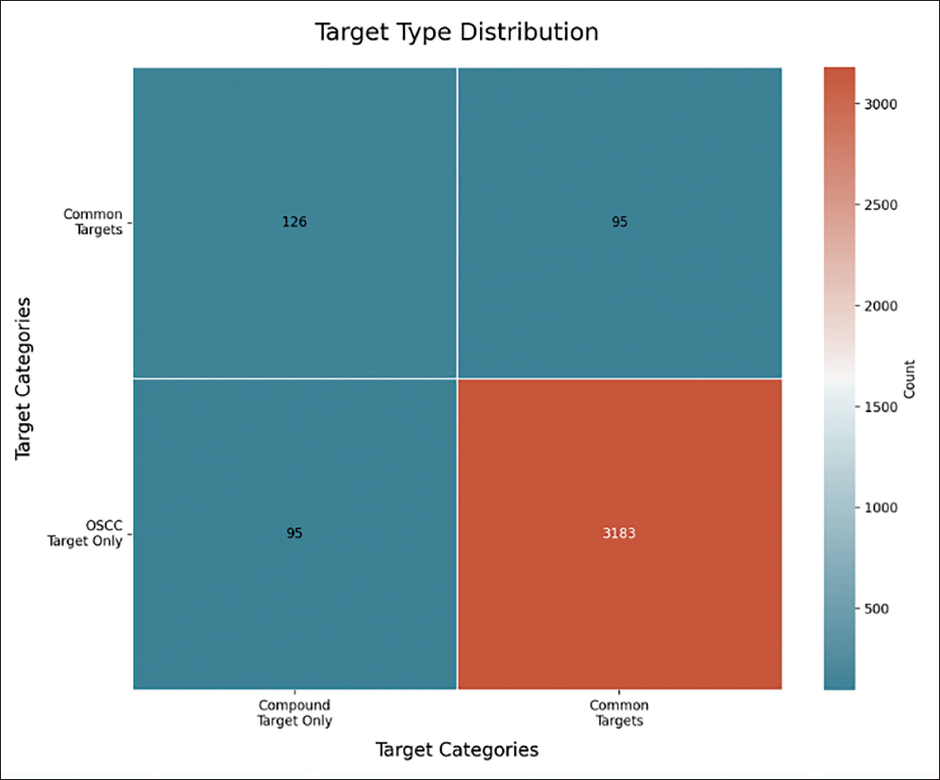
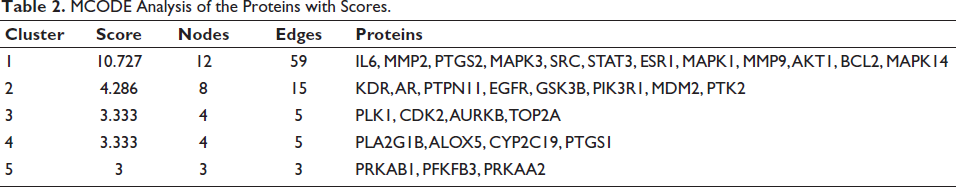
The 95 common genes identified as potential OSCC targets were analyzed using the STRING web server to examine their PPI. The results of this PPI analysis were visualized using Cytoscape software (Figure 2a and b) (Supplementary Figure S2). To identify highly interconnected genes, MCODE analysis was performed. The highest MCODE score obtained was 10.727, indicating a highly significant cluster (Table 2). The genes within this high-scoring cluster included IL6, MMP2, PTGS2, MAPK3, SRC, STAT3, ESR1, MAPK1, MMP9, AKT1, BCL2, and MAPK14.
(a) PPI Visualized in Cytoscape.
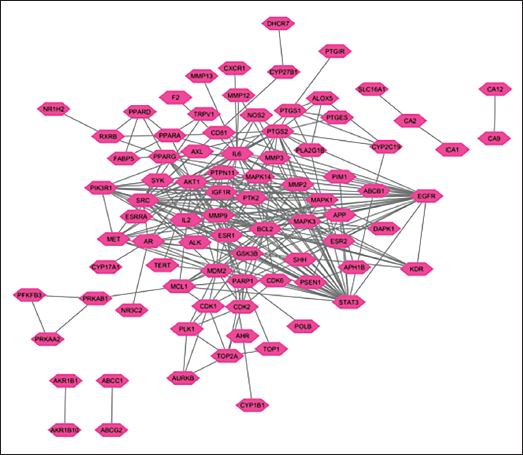
(b) MCODE Analysis with the Highest Score.
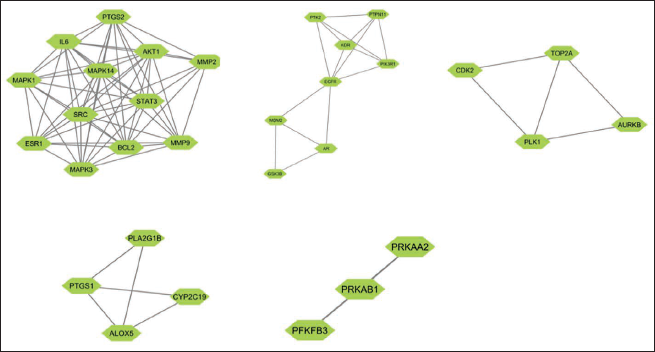
MCODE Analysis of the Proteins with Scores.
Expression Analysis
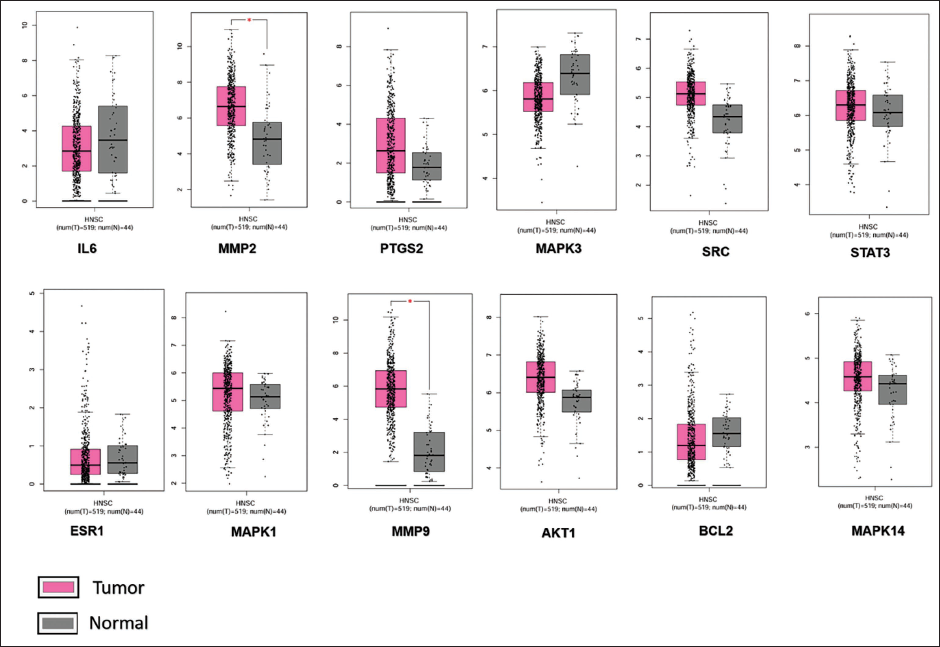
Expression profile analysis for the 12 identified genes was conducted using the GEPIA database. This analysis revealed that IL6, MAPK3, ESR1, and BCL2 had low expression levels in HNSC (Figure 3). Consequently, these four genes were excluded from further analysis due to their low expression in the relevant cancer type. This step ensured that the focus remained on genes with significant expression in HNSC.
Expression Profile of Hub Genes.
KEGG Enrichment Analysis
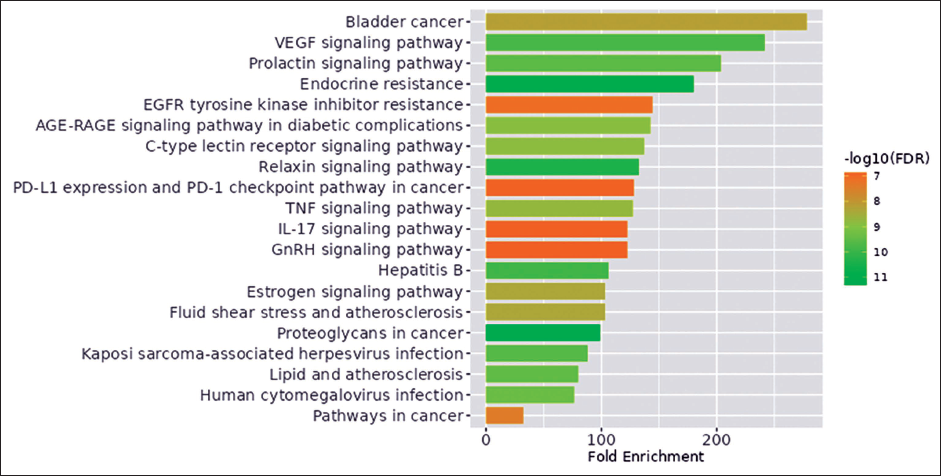
KEGG enrichment analysis of the remaining eight genes was performed using the ShinyGO web server. The analysis indicated that these genes are enriched in various metabolic pathways. For the purposes of this study, genes involved in cancer-related pathways were specifically selected. The genes identified as being involved in pathways in cancer were MMP2, PTGS2, STAT3, MAPK1, MMP9, and AKT1 (Figure 4) (Supplementary Figure S3). These six genes were chosen for further investigation due to their potential roles in cancer biology and their relevance as therapeutic targets.
In their investigation, Zhang et al. scrutinized the KEGG enrichment for 27 target genes associated with stemazole, revealing their enrichment across 53 distinct KEGG pathways. This comprehensive analysis highlights the diverse roles these genes play in various biological contexts within their study. We derived inspiration from Zhang et al.’s study and conducted a parallel KEGG enrichment analysis for our three genes. The objective pursued by Zhang et al. resonates with our study, emphasizing the importance of understanding the significance of target genes within different pathways. 20
Hub Genes Enrichment in Pathways in Cancer.
Molecular Docking
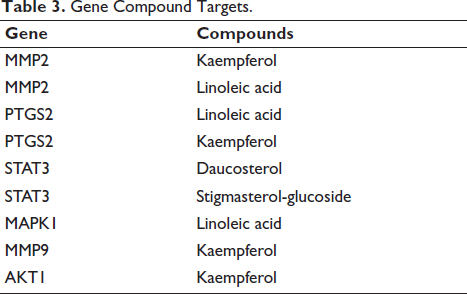
From compound target analysis, the four compounds, namely, linoleic acid, kaempferol, daucosterol, and stigmasterol-glucoside, would target these six genes in OSCC (Table 3). Docking analysis revealed that all compounds exhibited good binding affinities towards all six target proteins. The binding affinities ranged from –4.4 to –10.6 kcal/mol, indicating strong interactions (Table 4, Figure 5). These affinities were comparable to those of the control drug, Carboplatin, suggesting that the compounds could potentially serve as effective therapeutic agents.
Gene Compound Targets.
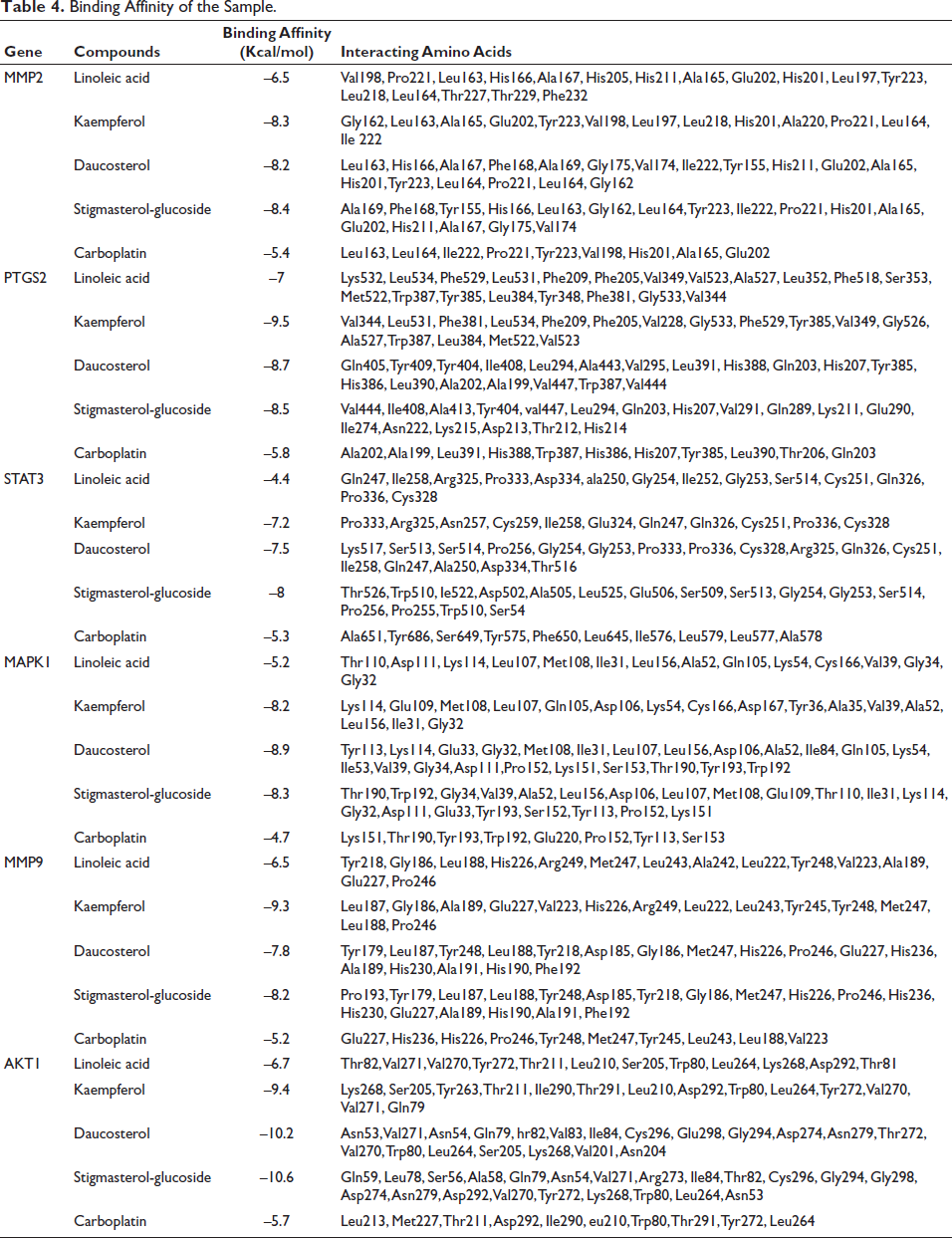
Binding Affinity of the Sample.
Docked Structure of Compounds and Targets.
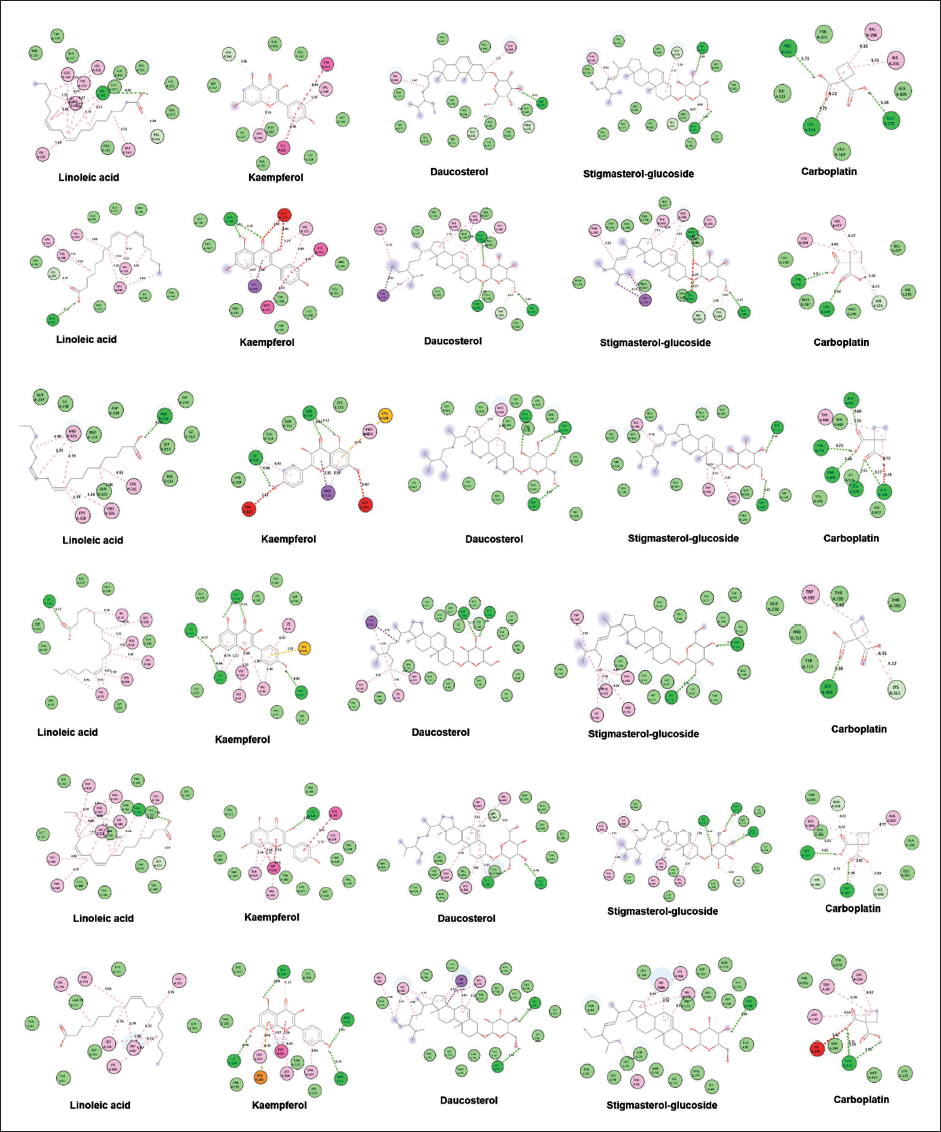
The docking studies showed that the interacting amino acids of the compounds were located within the active sites of the target proteins, which support their potential efficacy. Detailed information, including the gene and its corresponding protein PDB ID, grid coordinates, and the amino acids involved in the active site interactions, is presented in Table 5. This comprehensive data provides valuable insights into the molecular interactions and supports the potential of these compounds for further development as cancer therapeutics.
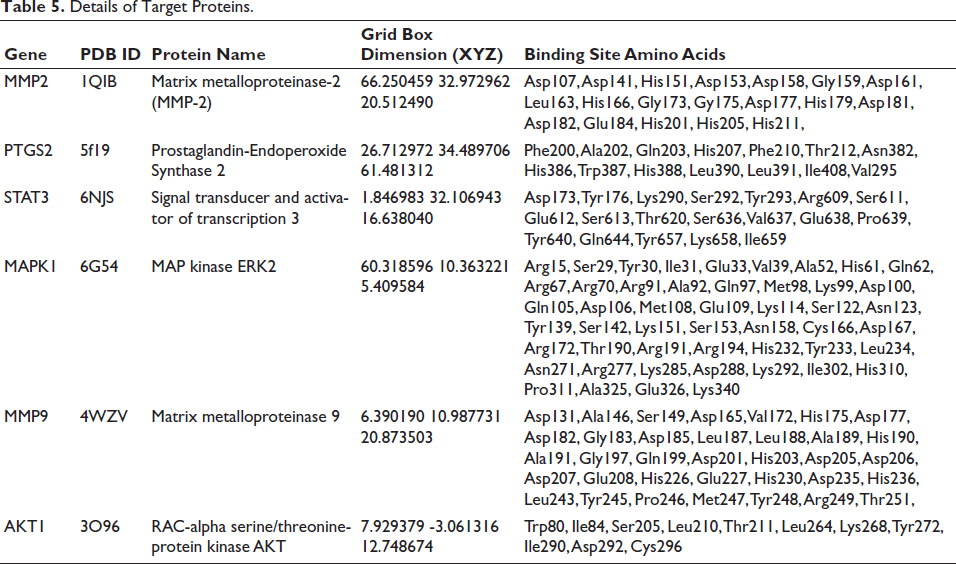
Details of Target Proteins.
These results also suggest that all four compounds can effectively interact with all six target proteins in OSCC, highlighting their potential multitargeting activity. This indicates that each compound has the capability to bind and potentially modulate multiple key proteins involved in OSCC pathways, which could enhance their therapeutic efficacy. The ability of these compounds to interact with multiple targets suggests a promising strategy for developing multifaceted treatments for OSCC, potentially leading to better clinical outcomes by simultaneously influencing various molecular mechanisms of the disease.
Known for its antioxidant and anti-inflammatory properties, kaempferol demonstrated strong binding to STAT3 and AKT1. By inhibiting these proteins, kaempferol could potentially reduce tumor growth and metastasis. Linoleic acid showed significant interactions with MMP2 and PTGS2. Linoleic acid’s anti-inflammatory properties might help in reducing OSCC-related inflammation. Daucosterol exhibited strong binding to MAPK1 and MMP9. Daucosterol could play a role in inhibiting cell proliferation and metastasis through these interactions. Stigmasterol-glucoside demonstrated a high affinity for MMP9 and PTGS2, suggesting its potential to modulate inflammation and prevent tumor invasion.
In a parallel study, Alamri investigated the interaction between five phytochemicals sourced from indigenous plants and targets associated with hepatocellular carcinoma (HCC). Their findings revealed binding affinities ranging from –8 to –9.5 kcal/mol. 21 Similarly, in our study, we conducted docking analyses between three fatty acids and three targets linked to oral cancer. Our results exhibited comparable variations in binding affinities. This congruence across studies validates the consistency of our findings and underscores the potential therapeutic relevance of these bioactive compounds in targeting specific disease pathways.
Activity Prediction
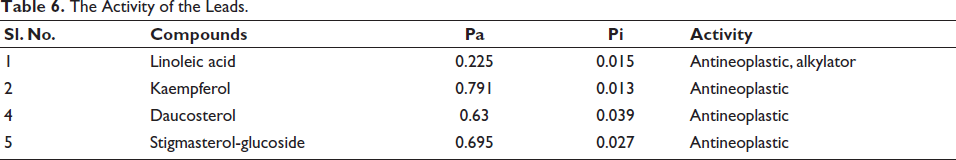
Activity prediction using the PASS web server revealed that all four compounds possess antineoplastic activity, meaning they have the potential to inhibit or prevent the growth of tumors. The probability scores for active antineoplastic properties were higher than the probability scores for inactive properties for each compound (Table 6). This finding underscores the potential of these compounds as effective agents in cancer treatment, particularly in targeting OSCC. The high probability scores further validate the compounds’ potential utility in therapeutic applications and warrant further investigation into their efficacy and mechanisms of action.
The Activity of the Leads.
In this study, we sought to identify and evaluate potential therapeutic compounds from Desmostachya bipinnata for OSCC through a multi-step approach. Initially, we identified 3,278 OSCC-related genes by integrating data from GeneCards, DisGeNet, and GEO databases. Using SwissTargetPrediction, we filtered 221 unique targets for 14 selected compounds and identified 95 common targets with OSCC-related genes. PPI analysis using STRING, Cytoscape, and MCODE highlighted a critical cluster of genes. Subsequent GEPIA analysis led to the exclusion of low-expressing genes (IL6, MAPK3, ESR1, and BCL2), focusing our study on MMP2, PTGS2, STAT3, MAPK1, MMP9, and AKT1, which are involved in cancer-related pathways according to KEGG enrichment analysis via ShinyGO.
Molecular docking studies revealed that the four compounds—linoleic acid, kaempferol, daucosterol, and stigmasterol-glucoside—exhibited strong binding affinities to the six key target proteins MMP2, PTGS2, STAT3, MAPK1, MMP9, and AKT1, with binding affinities comparable to Carboplatin suggesting their potential as effective multitarget agents against OSCC. The interacting amino acids were located in the active sites of the target proteins, indicating potential efficacy. PASS web server predictions confirmed the antineoplastic activity of these compounds. These findings suggest their promise as multitargeting agents for OSCC treatment.
Discussion
The study investigates the potential of compounds derived from Desmostachya bipinnata as novel therapeutic agents for OSCC. By employing a comprehensive approach encompassing compound screening, toxicity assessment, and in silico analysis, several key compounds have been identified, showing promising anticancer properties. Notably, linoleic acid, kaempferol, daucosterol, and stigmasterol-glucoside have emerged as significant candidates for further investigation. Utilizing the SwissTargetPrediction tool, these compounds have been mapped to their respective gene targets, elucidating their potential roles in OSCC treatment.
The findings are validated through PPI networks analyzed via the STRING database and visualized using Cytoscape. This network analysis highlights key interactions between the identified compounds and crucial proteins involved in the progression of OSCC. Subsequent gene expression profiles obtained from GEPIA corroborate these interactions, suggesting that these compounds might influence key pathways associated with OSCC.
Insights from molecular docking studies provide valuable information on the binding affinities and mechanisms through which these compounds exert their effects. The interaction of linoleic acid, kaempferol, daucosterol, and stigmasterol-glucoside with targeted proteins suggests that these compounds could disrupt critical signaling pathways implicated in OSCC.
Finally, the integration of compound screening, toxicological evaluations, and bioinformatics analyses underscores the potential of Desmostachya bipinnata compounds as multitargeting agents for OSCC. These findings lay a promising foundation for further preclinical and clinical investigations, offering the potential to enhance therapeutic strategies for OSCC and improve patient outcomes.
Conclusion
This study underscores the potential of Desmostachya bipinnata compounds as multifaceted therapeutic agents in OSCC treatment. The use of advanced in silico methodologies has provided a robust framework for identifying and validating the efficacy of these bioactive compounds. The significant interactions observed in molecular docking studies highlight their potential as multitarget agents, comparable to established chemotherapeutic drugs like carboplatin. The findings of this study suggest that linoleic acid, kaempferol, daucosterol, and stigmasterol-glucoside have significant potential as multitargeting agents against OSCC. Their strong binding affinities to key cancer-related proteins and predicted antineoplastic activities highlight their promise as therapeutic candidates. These findings pave the way for further experimental validation and clinical exploration, offering hope for more effective and less toxic treatments for OSCC.
Footnotes
Abbreviations
Acknowledgments
The authors wish to express their sincere gratitude to the Department of Health Research under the Human Resource Development Scheme and Saveetha Dental College for their valuable support in conducting this research.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval and Informed Consent
Not applicable.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research is supported by the Department of Health Research under the Human Resource Development Scheme (Women Scientist 2022-1188), India.
Supplemental Material
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
