Abstract
Background and Aims:
The human gastrointestinal system hosts a rich and diverse microbial community, surpassing a staggering 10[14] bacteria, collectively harbouring more than 5,000,000 genes. The aim of this study is to carry out a systematic review of human studies in order to investigate the correlation between the gut microbiota and pancreatic disorders.
Materials and Methods:
To systematically compile relevant research between 2011 and 2022, a thorough exploration of the MEDLINE and EMBASE databases was executed. The initial search results, encompassing all identified studies, were meticulously scrutinised, including a comprehensive review of their reference lists to ensure the completeness of our search. Initially, our search strategy yielded a total of 114 items. Subsequent refinement involved the removal of 58 duplicate articles identified in both searches. Following a meticulous screening process involving title, abstract and keyword assessments, 36 studies progressed to the next phase of evaluation. However, this cohort underwent further scrutiny, leading to the exclusion of 11 studies categorised as reviews or case reports, 5 in languages other than English that have been published and 20 that did not investigate the microbiome. As a result, a focused set of 20 papers emerged for in-depth evaluation for eligibility.
Results:
The demographic profile of the patients revealed that nearly half were men, with an average age of 52 years (interquartile range of 44-56). The interquartile range further ranged from 45% to 52%, reflecting the distribution of ages within the group. Among the studies scrutinising acute pancreatitis, gallstones emerged as the predominant cause in 9 out of 15, while 7 studies focusing on chronic pancreatitis found no evidence of gallstones.
Conclusion:
This comprehensive, systematic investigation has furnished robust evidence substantiating the intricate connection between pancreatitis and the microbiota.
Introduction
The human gastrointestinal system hosts a rich and diverse microbial community, surpassing a staggering 10[14] bacteria, collectively harbouring more than 5,000,000 genes.[1,2] Firmicutes and Bacteroidetes, the predominant bacterial types, constitute 80% to 90% of the gut microbiota, wielding a profound influence on various facets of human physiology. This influence extends to crucial aspects such as metabolism, mucosal immune system modulation, vitamin synthesis, digestion processes and the architectural dynamics of the digestive tract.[3,4] Antigenic effects and systemic metabolism are two parts of the gut microbiome’s complex influence on the immune system. Because the intestinal barrier acts as a defence against the growth of pathogenic flora, maintaining the integrity of the gut mucosa is of the utmost importance. Several gastrointestinal problems, including IBS, IBD, obesity, metabolic syndrome, diabetes, and pancreatic diseases, such as pancreatic cancer, have dysbiosis as an underlying cause.[5,6]
‘Gut microbiome’ encompasses the complex network of bacteria, their genomes and the ambient factors in the immediate vicinity.[7] The microbiome assumes a pivotal role in shaping the mucosal immune system, actively participating in digestion, nutrient storage and vitamin production, especially in a state of optimal health.[8,9] Recent studies illuminate the presence of a diverse microbiome not only in the gastrointestinal tract but also within human blood, underscoring its existence in both healthy and diseased states.[9-18] Contrary to the prevailing belief that a normal pancreas lacks its own microbiome, recent research challenges this notion. Even individuals with ostensibly normal pancreases can experience microenvironmental alterations due to the migration of gut flora into the pancreas. Recent research has shown that pancreatic illness patients often have altered gut flora, which may play a role in the onset of pancreatic cancer, acute pancreatitis (AP) and chronic pancreatitis (CP).[19]
While a definitive causal relationship between gut dysbiosis and pancreatic illnesses remains elusive due to a dearth of knowledge, factors such as genetics, nutrition and medications (e.g., metformin and proton pump inhibitors (PPI)) collectively shape the human microbiome composition. Alterations in the microbiome are implicated in various inflammatory, neoplastic and metabolic diseases, including those affecting the pancreas, as confirmed through extensive research.[20]
The complex aetiology and clinical intricacies of AP and CP pose significant challenges in their diagnosis, characterisation and treatment. Current diagnostic methods struggle to predict individuals prone to developing severe complications. Recognising these challenges underscores the urgent need to enhance our understanding of disease progression in AP and CP. This systematic study seeks to fill in some of the blanks by providing a detailed description of the taxonomic and functional features of the gut and blood microbiomes in patients with pancreatitis at various stages. The main goal is to determine if alterations in these microbiomes occur before pancreatitis develops or as a result of the illness.[21,22]
Understanding microbiome changes associated with pancreatitis holds promise for developing non-invasive predictive tests and innovative therapies capable of altering the natural course of these diseases. Despite the promising prognosis for this field, it is essential to acknowledge that it is still in its infancy, emphasising the need for further studies. A detailed awareness of research constraints is crucial to guide future investigations and direct research efforts. The aim of this study is to carry out a systematic review of human studies in order to investigate the correlation between the gut microbiota and pancreatic disorders.
Materials and Methods
To systematically compile relevant research between 2011 and 2022, a thorough exploration of the MEDLINE and EMBASE databases was executed. Pancreatitis and microbiota-related literature were more easily located by making use of Medical Subject Headings (MeSH). Along with the usual title, abstract and keyword parameters, these MeSH phrases were also included in the search. The initial search results, encompassing all identified studies, were meticulously scrutinised, including a comprehensive review of their reference lists to ensure the completeness of our search. Articles falling under the categories of review articles, case studies, animal experiments, or those lacking English translations were deliberately excluded from our analysis.[23] Commencing the search in year 2011 was a strategic decision grounded in the availability of next-generation high-throughput sequencing, which significantly enhanced the economic viability of metagenomic studies. This was a radical change from earlier research that frequently used antiquated methods like fluorescent in situ hybridisation (FISH). Studies excluding individuals under the age of 18 or pertaining solely to pancreatic ductal adenocarcinoma (PDAC), cystic fibrosis, autoimmune pancreatitis, graft pancreatitis, or Helicobacter pylori illness were also removed. This strategic exclusion aimed to streamline the investigation of microbiome changes associated with or caused by pancreatitis, mitigating the potential confounding effects of systemic factors on the microbiota. Notably, Helicobacter pylori-only research was subjected to additional scrutiny, and studies were excluded if the bacteria inadequately represented the gut microbiome. Factors such as current dietary regimen, alcohol usage, metabolic diseases (including diabetes, obesity, or pancreatic exocrine insufficiency), PPI use and antibiotic use were not employed as grounds for excluding individuals from the study. Instead, these aspects were considered variables during the evidence quality review, adhering to a comprehensive grading approach. To ensure a rigorous and unbiased evaluation, two independent reviewers meticulously assessed abstracts and titles (Figure 1).
Risk of assessment bias

Data Extraction
Microsoft Excel (2016 Edition; Microsoft Corp., Redmond, Washington, USA) was used by two separate reviewers to methodically organise the data extracted from the qualifying studies. Discrepancies that arose during this process were thoughtfully resolved through extensive discussions between the reviewers, ensuring a consensus on the extracted information. The data that were extracted covered a wide range of topics, including study details, patient demographics, disease details, methods for microbiome analysis, bacterial abundance, diversity metrics, functional analyses (like KEGG pathway analysis), relative abundance of individual taxa, endotoxin abundance, assessments of intestinal permeability and absorption and evaluations of functional analyses. Patients were considered adequately treated in clinical trials for AP if they exhibited two of the following symptoms: characteristic abdominal pain, biochemical markers of pancreatitis (elevated amylase or lipase levels greater than three times the upper limit of normal) and/or radiographic evidence of pancreatitis on cross-sectional imaging. The standards for severity grading were determined by the Atlanta classification, which consisted of either the 1992 classification or the 2012 version, whichever was published first. Studies delving into CP were considered adequate when participants met specific criteria, including visible pancreatic calcifications on CT or endoscopic ultrasonography, moderate-to-severe pancreatic ductal lesions on endoscopic retrograde or magnetic resonance pancreatography (Cambridge classification), or histological features indicative of pancreatitis on a sufficiently obtained surgical specimen of the pancreas.[24,25]
Distinguishing between research on the gut microbiome and studies examining the blood microbiome, the former relied on samples extracted from the gastrointestinal tract (pancreas, biliary tract, or the alimentary tract), while the latter used peripheral blood. The microbiological community evaluation assays were classified into two groups: those that relied on culture-dependent methods and those that did not. The latter group was further divided into limited detection methods (such as FISH, denaturing gradient gel electrophoresis, or focused sequencing) and high-throughput approaches (such as unbiased sequencing of 16S rRNA gene amplicons). The 16S rRNA gene was used as a marker sequence in all culture-independent approaches; however, whole-genome shotgun sequencing was not included. Studies that included diversity analysis, which is a formal evaluation of α or β diversity, were categorised as carrying out a thorough microbiome analysis. The term ‘diversity’ was understood in the context of the variety of life in a given area, encompassing the absolute number of species present and their distribution within the area. Beta diversity, which delineates diversity between groups, was determined by the extent of shared species. Given the methodological variations in diversity analysis among studies, categorical data were retrieved for each variable, indicating changes such as increased, decreased, or not substantially different for α diversity, and considerably or not significantly different for β diversity.[26,27]
Data Extraction (Risk of Bias/Quality of Evidence)
Two separate reviewers meticulously examined all qualifying journal articles in an effort to be objective and provide a thorough evaluation of evidence quality. The research was evaluated for potential bias using the Cochrane risk of bias instrument (RoB 2) and the quality assessment tool for diagnostic accuracy studies (QUADAS-2). Both observational studies and randomised controlled trials (RCTs) were included in this comprehensive review, and they were both evaluated using different criteria to guarantee a solid evaluation of the quality of the evidence.[28,29] A GRADE system was applied to RCTs, with a baseline grade of 4, signifying high-quality evidence. However, this score could be subject to adjustments based on factors such as publishing bias, indirectness, inconsistency and the risk of prejudice. A score of 3 denoted moderate quality, 2 indicated low quality and 1 signalled extremely low quality. The resultant GRADE scores for all RCTs, alongside the RoB 2 results for each trial, were systematically presented in tabular form, offering a comprehensive overview of evidence quality. Observational research, on the other hand, underwent evaluation using the Newcastle-Ottawa Scale (NOS). The QUADAS-2 and RoB 2 tools employed a set of signalling questions across various domains, with responses of ‘yes’ indicating a low likelihood of bias, while ‘no’ and ‘unclear’ suggested higher potential bias. A domain was categorised as having a ‘low’ bias risk if all signalling questions within it received a ‘yes’ response. Conversely, multiple ‘no’ responses indicated a ‘high’ probability of bias, and any other combinations were designated as ‘unclear’. Rather than relying on a single overall score, both QUADAS-2 and NOS analyses presented scores for each component in tabular or graphical formats, avoiding the pitfalls associated with summary scores.[30] This comprehensive approach aimed to provide a nuanced understanding of evidence quality, taking into account various factors and domains crucial to the evaluation of bias and study quality.[31,32]
Results
Search Result and Study Selection
Initially, our search strategy yielded a total of 114 items. Subsequent refinement involved the removal of 58 duplicate articles identified in both searches. Following a meticulous screening process involving title, abstract and keyword assessments, 36 studies progressed to the next phase of evaluation. However, this cohort underwent further scrutiny, leading to the exclusion of 11 studies categorised as reviews or case reports, 5 published in languages other than English, and 20 that did not investigate the microbiome.
As a result, a focused set of 20 papers emerged for in-depth evaluation for eligibility. During this thorough assessment, one publication was excluded due to various reasons: (a) It was initially abstracted and subsequently released as a full text (with the entire publication analysed); (b) it failed to specify whether patients suffered from acute or CP; (c) it lacked detailed methodologies for microbiome analysis; and (d) it did not permit the extraction of data specific to pancreatitis patients. Consequently, a final set of 15 publications was deemed suitable for inclusion in the qualitative synthesis, ensuring that the selected studies met rigorous criteria for relevance and methodological clarity.
Study Characteristics
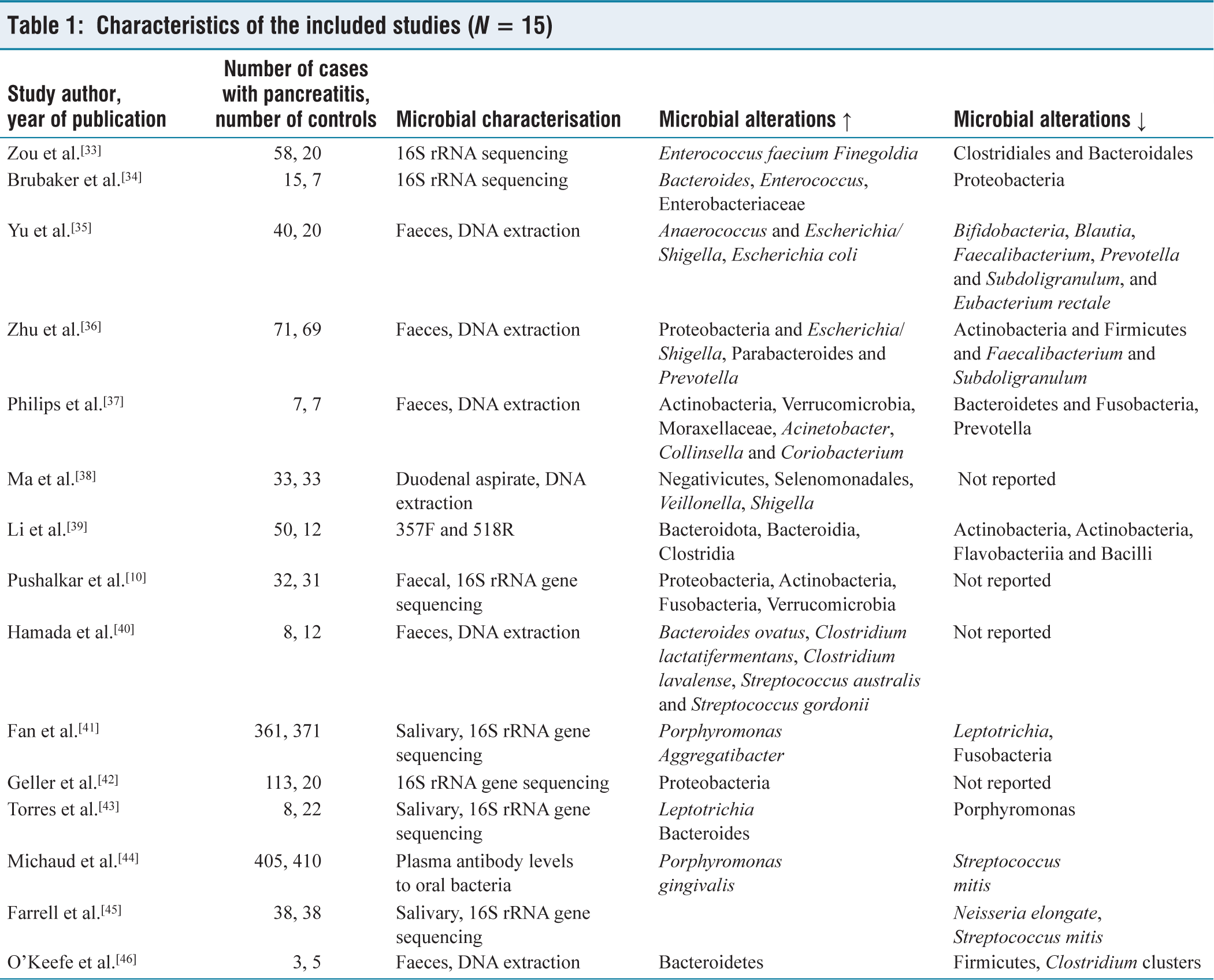
Table 1 presents a comprehensive overview of the 15 papers under consideration for this study. The demographic profile of the patients revealed that nearly half were men, with an average age of 52 years (interquartile range of 44 to 56). The interquartile range further ranged from 45% to 52%, reflecting the distribution of ages within the group. Among the studies scrutinising AP, gallstones emerged as the predominant cause in 9 out of 15, while the 7 studies focusing on CP found no evidence of gallstones. The intricate link between oral microbial compositions and pancreatic illnesses was explored in studies, such as those conducted by Fan et al.[41] and Michaud et al.,[44] revealing higher levels of Porphyromonas gingivalis in pancreatic cancer patients compared to healthy controls. Notably, this bacterium was associated with a twofold increased risk of pancreatic cancer. Farrell et al.[45] delved into differences in the oral microbiome among pancreatic cancer patients, those with CP, and healthy controls, although the causal or consequential nature of these microbial alterations remains uncertain, given the case-control nature of all investigations.[46]
Characteristics of the included studies (N = 15)
Recent research has unveiled the presence of bacteria in the tumour microenvironment of various cancers, including pancreatic cancers in humans. The bacterial genera found in pancreatic tumours mirror those commonly present in the human digestive tract, particularly Proteobacteria. Studies using genetically modified mice models, such as the Ptf1aCre, LSL-KrasG12D (KC) and Ptf1aCre, LSL-KrasG12D, Trp53R172H (KPC) models, revealed the impact of gut bacteria on pancreatic tumour development. Chang et al. demonstrated that oral antibiotics impeded neoplasm progression, while replenishing germ-free mice with specific bacteria accelerated the disease.[47] In addition, the immune cell composition inside the tumour microenvironment was changed by microbial ablation. The ratio of tumour-protective M1-like Tumor-Associated Macrophages (TAMs) to immunosuppressive CD206+ M2-like TAMs increased, and the balance shifted from these two groups. Repopulating the microbiota had the interesting effect of reversing these changes. A further study that utilised various mice models discovered that when the gut microbial population was reduced through antibiotic-induced depletion, there was an upregulation of Th1 and Tc1 cells inside the tumour microenvironment, and the pancreatic tumour burden was decreased.[48] Altogether, these results shed light on the complex interplay between the gut microbiota and pancreatic diseases, which may lead to new approaches to treating these conditions.
Discussion
The intricate relationship between the microbiota and pancreatic health and disease has been the subject of numerous scholarly reviews. The uniqueness of this investigation lies in the fact that it is the first of its type to examine pancreatitis in all its complexity. Memba et al. (2017)[49] conducted a systematic review, although they only included five pancreatitis-related articles that met their criteria at that time. This suggests that their primary focus was on pancreatic ductal adenocarcinoma.[49] Nevertheless, our present systematic review examines pancreatitis from a more comprehensive angle, examining 10 papers and exploring the substantial evidence around changes in microbial composition using a qualitative analysis. When we look at the data from Poudel et al., we find an intriguing finding: the levels of Firmicutes are even lower in patients with PDAC compared to those with AP.[50] This discovery adds another layer of complexity to our understanding of the microbial dynamics involved in many pancreatic disorders. The study focuses on Faecalibacterium prausnitzii, a commensal microbe that is highly prevalent in the gut and belongs to the Firmicutes phylum. This bacterium may aid in lowering Firmicutes bacterial populations in pancreatic ductal adenocarcinoma and AP. To completely comprehend the potential involvement of F. prausnitzii in various diseases, further species-level microbiome studies are required.
Various studies have demonstrated that as CP advances, the microbial composition changes more visibly and the relative quantity of commensal organisms declines, regardless of the underlying aetiology of the illness.[42] There is mounting evidence that alterations in microbial composition may be an outcome of CP rather than a cause of the disease. Our research provides new insight into the role of ecoimmunonutrition and fibre as treatment modalities for AP, which have been associated with improved functional and clinical results. Due to the lack of thorough diversity evaluations, it is impossible to say with certainty whether these benefits are due to the normalisation of the gut microbiota or other known effects of fibre/probiotics. It would be a mistake to assume anything just because these results are promising; for instance, RCTs have demonstrated contradictory outcomes when employing specific agents; for instance, Lactobacillus + Bifidobacterium + fibre had detrimental effects, while Lactobacillus plantarum + fibre had beneficial effects. Due to the lack of evaluation of the effects of these medicines on gut flora and intestinal permeability, alternative explanations for the observed outcomes may exist.[51-53]
In AP, the number of Bifidobacteria is lower than in healthy controls, and in CP, the number of Bifidobacteria is inversely proportional to the severity of the condition. Bifidobacterium strains were surprisingly absent from all of the trials that made up this investigation. We need more research on species-level changes in the microbiome to support commensal microbial recovery in pancreatitis patients. Treatment options like narrow-spectrum antibiotics and probiotics would not be addressed until that time has passed.[54] Although this is the most comprehensive systematic review on the subject to date, it is crucial to recognise a number of limitations. One major drawback is that different studies use different approaches to assess the microbiome. Discordances persisted, likely as a result of variations in the regions of the 16S rRNA gene that were amplified, despite the predominant use of high-throughput sequencing techniques. It was also difficult to draw definite conclusions regarding the composition of microbial communities due to inconsistent reporting of data and methodologies and the fact that some studies did not perform a systematic diversity study.
Heterogeneity also impacted the reporting of intestinal permeability, a crucial component when considering changes in microbial composition. It is important to recognise that intestinal barrier disruption is just one of many pathogenic routes that can lead to complications of AP. Autodigestion and microcirculatory dysfunction are two other important pathways. When studying the relationship between shifts in microbial composition and disease prognosis, it is crucial to keep these various pathogenic pathways in mind.[55]
Specific restrictions about the applicability and credibility of the data were present in the research that were taken into consideration. The results may not be applicable to those with quiescent CP or acute exacerbations of CP because several studies that looked at the gut microbiota used samples that were not from faeces. Antibiotic and PPI use, metabolic diseases, diet and alcohol use and other variables known to impact the gut microbiome were either not controlled for or not appropriately considered in the majority of research. By carefully controlling for variables like food habits, medication use, diabetes and pancreatic exocrine insufficiency, this study provides a rare chance to learn more about the interdependent relationship between the microbiota and pancreatic disorders.
Conclusion
The intricate pathophysiology underlying pancreatic diseases has been consistently associated with gut microbial dysbiosis, as evidenced by a substantial body of research primarily derived from animal models. The continued relevance of these models in testing hypotheses and enhancing our comprehension of the intricate relationship between the gut microbiota and pancreatic health ensures their continued demand in future investigations. Despite significant strides in unravelling this intricate connection, there persists a considerable amount that remains unknown, particularly regarding the origins and consequences of gut microbial dysbiosis, including alterations in the abundance of different microbial species.
In the quest for a more profound understanding of the human microbiome, numerous large-scale research endeavours are diligently working towards a comprehensive characterisation. These efforts specifically target the metabolic effects arising from the intricate interplay between the microbiome and factors such as dietary patterns, genetic predispositions, pathological conditions and medicinal interventions. To gain insights into how the microbiome intricately influences the spectrum of pancreatic diseases in humans, further exploration of these complex interactions is imperative. Prospective studies evaluating the gut microbiota of individuals at risk of pancreatic diseases are essential to fill existing knowledge gaps. Such research endeavours hold the key to enhancing our understanding of how gut dysbiosis contributes to the aetiology of pancreatic illnesses based on human physiology. By proactively pursuing these initiatives, we stand to unravel more about the intricate links between microbial shifts and the onset of pancreatic diseases, paving the way for the development of more personalised therapeutic interventions.
In conclusion, this comprehensive, systematic investigation has furnished robust evidence substantiating the intricate connection between pancreatitis and the microbiota. While notable progress has been made, the evolving nature of this field demands future advancements characterised by standardised methodology, larger sample sizes and more meticulously controlled interventions. Embracing these changes is crucial to deepening our insights into the complex relationship between the gut microbiota and pancreatic health, thereby enhancing our diagnostic and therapeutic capabilities for pancreatic diseases.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The authors received no financial support for the research, authorship and/or publication of this article.
Credit author statement
All authors contributed equally in the manuscript.
Data availability
Data is included in the manuscript.
Use of artificial intelligence
Nil.
