Abstract
Introduction
Hypertension has emerged as a major public health concern. 1 It seems to be an independent risk factor for cardiovascular diseases, such as heart failure, stroke, renal insufficiency, and coronary artery disease. Various mechanisms for hypertension have been put forward including the dysfunctions of the sympathetic nervous system (SNS) and the renin-angiotensin-aldosterone system (RAAS), and the disorders of gut microbiota and metabolism. 2 A detailed understanding of the molecular mechanisms of hypertension is essential for the development of novel strategies for the control of high blood pressure for individuals. 3
An increasing amount of research suggested that bidirectional communication between the gut microbiota and microRNAs (miRNAs) played an important role in host diseases, including cancer, inflammatory bowel disease, and neurological disorders. 4 In recent years, several studies also indicated that the miRNA-microbiome interaction played a key role in the adjustment of blood pressure.5–7 These previous studies mainly focused on miRNAs (eg, miR-21, miR-122, and miR-129) and gut microbes (eg, Firmicutes/Bacteroidetes ratio). It is well known that fecal samples contain abundant miRNAs. These miRNAs can enter bacteria and regulate bacterial growth, participating in shaping the gut microbiota. 8 Human microbiota is integral in multiple physiological processes of the host. 9 Dysbiosis of gut microbiota causes hypertension via regulating metabolites and modulating inflammatory mediators. 10 Therefore, the therapeutic rationale for targeting the miRNA-microbiome axis may be possible to treat hypertension. 4 Although the structure and composition of the gut microbiota in two-kidney one-clip (2K1C) rats have been characterized by the use of the high throughput sequencing method,11,12 fecal miRNA expression profiles and their relationship with changes in gut microbiota in 2K1C rats remain largely unknown.
A lot of research has been concentrated on traditional folk medicine in modulating the fecal miRNA and the gut microbiota.13,14 Dracocephalum moldavica L., known as Xiang-qing-lan (in Chinese), is an herbaceous annual plant and belongs to the genus Dracocephalum within the family Lamiaceae. 15 It is widely distributed throughout China (eg, Heilongjiang, Liaoning, Jilin, Inner Mongolia, Shanxi, Gansu, Shaanxi, and Qinghai) and thrives in a variety of environments (eg, dry mountains, valleys, and rocky river flats). This plant has a straight and cylindrical root, petiolated lower leaves, and sessile upper leaves with spiny-serrate edges. The corolla of flowers is usually purple-blue, and the fruits are ovoid and tetranucleate. 15 In Mongolian medicine and Uighur medicine, it is used to treat coronary heart disease, myocardial ischemia, hypertension, angina pectoris, etc. 16 In addition, D. moldavica is used to treat neurological diseases, stop vomiting and sweating in Mexican and Iran.17,18 In recent years, more and more researchers have been attracted to investigate biologically active compounds of D. moldavica.19,20 Up to now, approximately 154 compounds have been isolated and identified from aerial parts of D. moldavica, such as flavonoids, steroids, glycosides, saponins, tannins, phenols, and essential oils. 15 Previous research has suggested that D. moldavica extracts (DMEs) demonstrated a diverse range of pharmacological effects, such as antioxidant, anti-inflammatory, antimicrobial, cardiovascular protection and other activities.21–23 In addition, DMEs have therapeutic effects on hypertension.20,24 However, the mechanism of DMEs in treating hypertension remains unclear. Thus, it is essential to elucidate the mechanism of the anti-hypertension effects of DMEs to ensure better treatment of hypertension.
Recently, DMEs were found to be involved in the regulation of miRNA expression.25,26 Our group also reported the crucial role of DMEs in modulating the gut microbiota in an experimental model of chronic colitis or 2K1C renal hypertension.27,28 We, therefore, speculated that it might modulate the miRNA-microbiome axis to achieve its diverse effects. Following this train of thought, the current study was designed to explore the influence of DMEs on the alterations of miRNA expression patterns and the changes in gut microbiota in rats suffering from hypertension. At the same time, we reported the mechanisms of how DMEs ameliorate hypertension through modulating the interaction between miRNAs and gut microbiota for the first time.
Results
DME Improved the Blood Pressure in 2K1C Rats
The effect of DME on the blood pressure in 2K1C rats was explored. The in vivo experiment scheme is shown in Figure 1. Systolic blood pressure (SBP) levels in the three groups were measured by tail cuff (Supplemental Table S1). No statistically significant differences in the baseline measures were found among all groups. When compared to a sham control, SBP in the 2K1C group was found to be significantly higher after four weeks of operation (p < 0.05). Since treatments were not started yet at the fourth week, SBP was increased in the treatment rats as compared to that of the sham-operated rats (p < 0.05). In contrast, SBP was significantly reduced in the treatment group after 4 weeks of intervention with DME (p < 0.05) (Supplemental Table S1).

In vivo experiment scheme.
Analysis of Chemical Constituents of DME
Through high-performance liquid chromatography (HPLC) analysis, chemical constituents were analyzed in the DME (Supplemental Fig. S1). The chromatograms showed that rosmarinic acid, tilianin, luteolin, apigenin, disometin and an unknown compound were found. The results demonstrated that tilianin, rosmarinic acid and an unknown compound were the main compounds in the DME.
DME Alleviated the Dysregulation of miRNA Expression in 2K1C Rats
We determined the levels of miRNA expression in fecal samples. Sequencing analysis of sRNA yielded 103,266,274, 105,696,459, and 105,696,459 raw reads from the sham, model, and treatment groups, respectively. A total of 98,923,540, 100,229,491, and 95,173,160 clean reads with lengths ranging from 18 to 40 nt were obtained by trimming adapter sequences and filtering low-quality tags, representing 31,981,197, 32,414,441, and 11,400,545 unique small RNAs in the sham, model, and treatment groups, respectively. The conserved miRNAs were identified by comparing the sRNA clean reads with the mature miRNAs in the miRbase. The differential expression of 21 miRNAs between the sham group and the model group was identified, including 17 upregulated and 4 downregulated miRNAs (Figure 2A). However, compared to the model group, 27 differentially expressed miRNAs were screened out in the treatment group (Figure 2B). Specifically, 2K1C considerably caused the dysregulated the expression of let-7a-5p, miR-196b-5p, miR246-5p, miR335-5p, miR-3557-3p, and miR-3596a. These changes were counteracted by DME administration (Figure 2C and D). Then, differentially expressed miRNAs (let-7a-5p and miR-196b-5p) were verified by qPCR, and the results were consistent with that of direct sequencing (Figure 2E and F).

DME alleviated the dysregulation of miRNA expression in 2K1C rats. (A) The volcano map showed the expression of miRNA within the sham and the model groups. (B) The volcano map showed the expression of miRNA within the model and the treatment groups. (C) Bar plot representing log2 Fold change of 6 differentially expressed miRNAs, of which 1 miRNA were down-regulated (green) and 5 up-regulated (red) in the model group relative to the sham group. (D) Bar plot representing log2 Fold change of 6 differentially expressed miRNAs, of which 5 miRNAs were down-regulated (red) and 1 up-regulated (green) in the treatment group relative to the model group. (E) Fold change of the selected miRNA expression in the model group relative to the sham group was determined using PCR analysis. (F) Fold change of the selected miRNA expression in the treatment group relative to the model group was determined using PCR analysis. Values are expressed as the mean ± SEM and analyzed by unpaired t-test analysis, n = 3. * p < 0.05 versus sham group, # p < 0.05 versus model group.
DME Ameliorated Gut Microbiota Dysbiosis in 2K1C Rats
Since DME exerts a powerful influence on intestinal miRNA expression in 2K1C rats, we performed bacterial 16S rRNA sequencing to measure the influence of DME on the fecal microbial community. The composition of intestinal flora across the different groups was compared. The results revealed that Firmicutes and Bacteroidetes were the dominant phyla of feces. (Supplemental Fig. S2). At the order level, 2K1C rats showed a reduced content of Lactobacillus and an increased content of Bacteroides. However, the alterations of Lactobacillus and Bacteroidetes were ameliorated by DME treatment (Figure 3A). Additionally, Lactobacillus was the dominant genus in the three groups (Figure 3B). 2K1C rats showed a decreased abundance of Lactobacillus. In contrast, the abundance of this genus was reversed by DME. In addition, the differences in fecal microbiota across diverse groups were compared using the Metastatic method with Fisher's exact test. Compared with the sham group, two genera (Papillibacter and Desulfovibrio, the members of the phyla Firmicutes and Proteobacteria, respectively) displayed an increase in the model group (Supplemental Fig. S3). In contrast, DME treatment attenuated the 2K1C-mediated upregulation of these two genera.

The DME treatment modulated the imbalance of gut microbiota in 2K1C rats. (A) Composition of the gut microbiota at the order level. (B) Composition of the gut microbiota at the genus level.
Associations Between the Differentially Expressed miRNAs and Gut Microbiota
We also analyzed the relationship between the differentially expressed miRNAs and the 3 most significantly altered gut microbes. As shown in Figure 4, let-7a-5p showed a positive relationship with Lactobacillus and a negative relationship with Papillibacter and Desulfovibrio. Furthermore, miR-196b-5p, miR246-5p, miR335-5p, miR-3557-3p, and miR-3596a showed a positive correlation with Papillibacter and Desulfovibrio and a negative correlation with Lactobacillus. Taken together, these miRNAs might affect the gut microbiota in rats with hypertension.
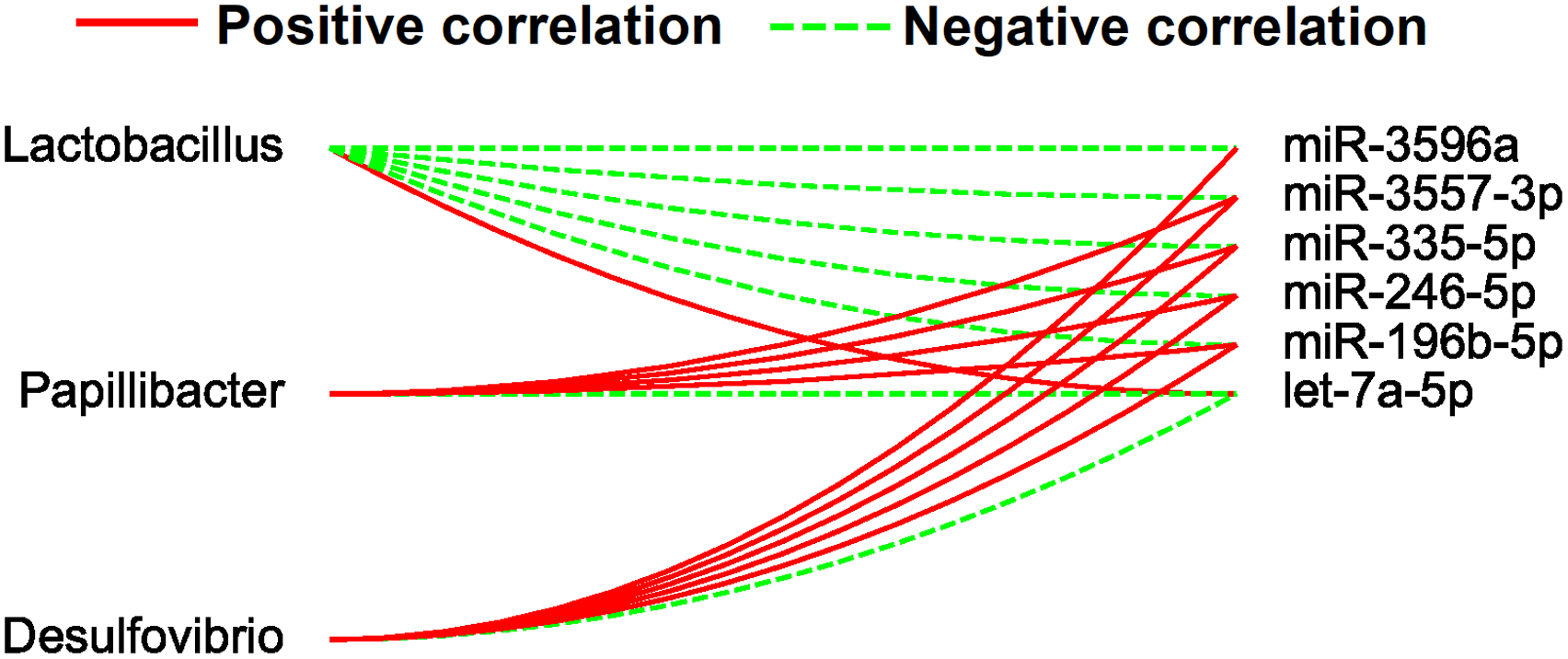
Associations between the 3 most significantly altered gut microbes and the differentially expressed miRNAs (DME significantly affect). Associations with the miRNAs are shown for each genus microbiota with the direction of correlation indicated by red (positive) or green (negative) lines.
Relationships Between Circulating Metabolites and Gut Microbiota
Non-targeted metabolomics methods were utilized to characterize the serum metabolites and to analyze the relationship between the intestinal microbiota and the circulating metabolites identified. Serum metabolomics results showed that 31 metabolites were significantly different in the model rats compared with the sham group (Figure 5A), 21 metabolites were raised and 10 metabolites were reduced (Supplemental Table S2). It was noted that only 14 metabolites were known. Moreover, 72 metabolites were marked as differential metabolite candidates in the comparison between the treatment and model groups (Figure 5B). Among these metabolites, 22 were known (Supplemental Table S3). Especially, the disorders of 14 serum metabolites caused by 2K1C were partially ameliorated by treatment with DME. As shown in Figure 5C to G, the serum levels of arachidonic acid, hippuric acid 2, fumaric acid, 2, 6-biphosphate degr prod 2, and xylose 1 were elevated in the model group. Those alterations were attenuated by treatment with DME.

The changes in gut microbiota by the DME influenced the circulating metabolites composition. (A) Volcano plots of differential metabolites between the sham and model groups. (B) Volcano plots of differential metabolites between the model and treatment groups. (C-G) Concentrations of arachidonic acid, hippuric acid 2, fumaric acid, 2, 6-biphosphate degr prod 2, and xylose 1, respectively. (H) Correlation heatmap between metabolites and gut microbiota at the genus level.
The relationship between the metabolites and gut microbiota was further analyzed. Fructose 2, 6-biphosphate degr prod 2 and xylose 1, which were elevated in the 2K1C rats, showed a positive correlation with Desulfovibrio and a negative correlation with Lactobacillus, demonstrating that the reduced counts of Lactobacillus and enhanced counts of Desulfovibrio were correlated with higher serum levels of fructose 2, 6-biphosphate degr prod 2 and xylose 1 in 2K1C rats (Figure 5H). When the abundances of Lactobacillus and Desulfovibrio were changed by the DME intervention, serum levels of fructose 2, 6-biphosphate degr prod 2 and xylose 1 were decreased in 2K1C rats (Figure 5H).
Altered Metabolic Pathways Induced by the Differential Metabolites
The radar plot in Supplemental Fig. S4 showed ten metabolites, illustrating how these metabolites distinguish the model group from the sham group, and distinguish the treatment group from the model group, respectively. We found that these differential metabolites participated in regulating diverse signaling pathways, including the Citrate cycle (TCA cycle), Alanine, aspartate and glutamate metabolism, Arachidonic acid metabolism, Pyrimidine metabolism, Biosynthesis of unsaturated fatty acids, Tyrosine metabolism, and Arginine and proline metabolism (Supplemental Fig. S4). These differential metabolites and corresponding pathways of change were summarized in Supplemental Tables S4 and S5, respectively. Treemap of the KEGG pathway showed that the differentially expressed metabolites in the three groups were mainly involved in the Arachidonic acid metabolism, TCA cycle, and Alanine, aspartate and glutamate metabolism (Supplemental Fig. S4). Especially, arachidonic acid and fumaric acid, which were increased in the 2K1C rats, might involve in regulating these three pathways.
Discussion
Intestinal miRNAs can interact with host microbiota and influence the pathogenesis of hypertension. 6 Several studies have assessed the effects of D. moldavica on the gut microbiota and miRNA expression profiles, respectively.25,26,27,28 Previous reports also showed that DMEs possessed significant medicinal values to treat hypertension.20,24 However, the influence of DME on the fecal miRNA expression pattern and the miRNA-microbiome axis in 2K1C rats remains to be elucidated. In this context, the present study brings novelty and originality by aiming to provide scientific arguments for the anti-hypertension mechanism of DME in 2K1C rats based on the miRNA-microbiome axis.
To better understand the mechanism of the anti-hypertensive effects of DME, a chemical component analysis was performed. We found that tilianin, rosmarinic acid and an unknown compound were the main compounds in the DME. These results are consistent with our earlier findings. 23 Previous studies have also shown that tilianin and rosmarinic acid were effective against hypertension.29,30 Additionally, the presence of rosmarinic acid in DMEs has antioxidant potential. 31 Tilianin from DMEs was proven to display anti-inflammatory activity. 32 Taken together, these results suggested that rosmarinic acid and tannins might partially mediate the effects of DMEs on blood pressure regulation. However, further in-depth research is needed on the structural information and biological functions of an unknown compound.
Although previous study has proven that DME can ameliorate intestinal inflammation and hypertension by adjusting the profile of gut microbiota in 2K1C rats, 28 the molecular mechanism of such gut microbiota response remains to be further studied. In this regard, we investigated the regulatory mechanism of DME on gut microbiota in rats with hypertension.
Based on recent evidence, nutrients disturb miRNA expressions that could potentially contribute to the pathogenesis of hypertension. 7 Emerging evidence suggests that DME and its active compounds modified miRNA signatures during vascular dementia.25,33 Previous studies also reported that compounds tannins and rosmarinic acid modulate miRNA expression.34,35 Thus, we speculated that the protective benefit of DME in hypertension might be related to the regulation of miRNAs. To verify this hypothesis, RNA sequencing and analysis were performed. The results revealed that 2K1C induced the dysregulated expression of miRNAs, such as let-7a-5p, miR-196b-5p, miR246-5p, miR335-5p, miR-3557-3p, and miR-3596a. Importantly, DME treatment recovered the expression of these miRNAs in 2K1C rats. In particular, let-7a-5p, miR-196b-5p, and miR-335-5p have been reported to be associated with hypertension,36,37 which is in agreement with our study. However, as far as we know, the effects of miR-246-5p, miR-3557-3p, and miR-3596a on hypertension have not been reported yet. Our findings underscore the critical roles of DME that mediates miRNA expressions.
It has been well-documented that fecal miRNAs shape the gut microbiota. 38 More importantly, several studies also underscored the relationship between gut microbiota and hypertension.39,40 This prompted us to investigate whether DME ameliorated hypertension through its modulatory effects on the gut microbiota mediated by miRNAs. Firstly, the effects of DME on the profile of gut microbiota were determined. Our results showed a decrease in the relative abundance of Lactobacillus and an enhancement in the relative abundance of Desulfovibrio were detected in 2K1C rats when compared with the controls. Lactobacillus has been widely used for balancing blood pressure. 41 According to the literature, Desulfovibrio species level was increased in hypertensive rats. 42 Our findings aligned with these reports. LEfSe analysis was also performed to identify differentially abundant taxa. We found that 2K1C rats were characterized by a higher enrichment of Anaerobiospirillum at each level from phylum to genus. It has been reported that the relative proportion of Anaerobiospirillum was significantly associated with gut inflammation in rats with Alzheimer's disease. 43 Thus, alteration of the Anaerobiospirillum level might contribute to an inflammatory response in 2K1C rats. Interestingly, the treatment of the DME caused a decrease in the counts of Desulfovibrio and Anaerobiospirillum as well as an enhancement in the count of Lactobacillus, implying its potential application in shaping the gut microbiota. This was similar to our previous study in which DME attenuated gut microbiota dysbiosis in the 2K1C rats by increasing the abundance of the genus Lactobacillus and decreasing the abundance of Desulfovibrio and Anaerobiospirillum genera. 28 Secondly, the correlation between the differentially expressed miRNAs and gut microbiota was explored. As expected, the correlation analysis illustrated that the key microbiota (Lactobacillus, Papillibacter, and Desulfovibrio) had a close relationship with the 6 differentially expressed miRNAs (let-7a-5p, miR-196b-5p, miR246-5p, miR335-5p, miR-3557-3p, and miR-3596a). Taken together, we confirmed that DME participated in the regulation of gut microbiota by miRNAs. This study adds to our understanding of the interaction between miRNAs and gut microbiota in 2K1C hypertensive rats.
Evidence indicated that the reshaping of gut microbiota was related to antihypertensive therapy. 44 The gut microbiota can manipulate host metabolism by modulating metabolic pathways. To elucidate the role of gut microbiota in hypertension, it was necessary to determine the possible impact of gut microbiota on the host metabolites. Thus, we used metabolomics analysis of serum samples to gain insight into any metabolic deviation associated with the gut microbiota of 2K1C rats. Our results showed that DME ameliorated the induction in the levels of arachidonic acid, hippuric acid 2, and fumaric acid in this study. A previous study showed that the metabolic abnormality of arachidonic acid is closely related to intestinal flora disturbance in mice suffering from atherosclerosis and inflammation. 45 This metabolite is the signaling molecule that regulates arachidonic acid metabolism and biosynthesis of unsaturated fatty acids, which initiates the inflammatory response. 46 In addition, hippuric acid 2 involved various pathways such as tryptophan, purine, kynurenine, and phenylalanine metabolism. These metabolic disorders could be associated with oxidative damage and inflammation. 47 Furthermore, the untargeted metabolomics method revealed that the level of fumaric acid was increased in rats with salt-sensitive hypertension, and might participate in blood pressure regulation. 48 Considering the potential effect of these metabolites on oxidative stress, inflammation and blood pressure, we speculated that the regulatory effect of DME on hypertension was associated with the modulation of host metabolism through regulating the gut microbiota of 2K1C rats.
To further confirm the correlation between relative quantities of metabolites and relative abundance of intestinal bacteria, Spearman correlation analysis was performed. We observed that metabolites 2, 6-biphosphate degr prod 2 and xylose 1 were increased in 2K1C rats. The levels of these metabolites were recovered to different degrees after the DME administration. A significant correlation between serum levels of these two metabolites and 7 bacterial genera reveals that 2, 6-biphosphate degr prod 2 and xylose 1 might be other major metabolites that interact with gut microbiota. In this study, the contents of 2, 6-biphosphate degr prod 2 and xylose 1 had a positive correlation with the population of Desulfovibrio and a negative correlation with the population of Lactobacillus. DME treatment decreased serum levels of these two metabolites in 2K1C rats, which was directly correlated with an enhancement in the population of Lactobacillus and a reduction in the population of Desulfovibrio. However, the correlation between 2 differentially expressed metabolites (2, 6-biphosphate degr prod 2 and xylose 1) and hypertension was unclear. According to previous literature reports, we just found evidence that xylose 1 affects the transport disorder of sugar. 49 Further studies are necessary to identify the accurate mechanism by which 2, 6-biphosphate degr prod 2 and xylose 1 affect blood pressure in 2K1C rats.
It has been well-established that hypertension is closely related to metabolic dysfunction. In our study, we further identified the key pathways based on the metabolite enrichment analysis. The results indicated that three metabolic pathways (Arachidonic acid metabolism, TCA cycle, and Alanine, aspartate and glutamate metabolism) were regarded as important factors influencing hypertension, which is consistent with the prior reports.50,51
Limitations
Although we found rosmarinic acid, tilianin, luteolin, apigenin, disometin and an unknown compound in DME using HPLC analysis, the characteristic details of the chemical constituents of DME will be needed to study further, which is helpful to clarify the molecular mechanisms underlying DME-mediated impact on the miRNA-microbiome axis in 2K1C rats. Another limitation was the absence of experimental verification of the altered metabolic pathways induced by the differential metabolites.
Conclusion
In summary, DME protects against hypertension. Several compounds were identified in HPLC analysis that may be involved in the blood pressure regulation of DME. Additionally, our study provides a novel antihypertension mechanism: DME restores host metabolism through the interplay between miRNAs and gut microbiota. The presented results imply that the use of drugs to target miRNAs and gut microbiota could be a potential therapeutic strategy against hypertension. More importantly and practically, further investigations into the antihypertension mechanisms of DME in the future may facilitate the development of personalized therapeutics for patients with hypertension.
Materials and Methods
Collection of Plant Material, Preparation of Total Extracts, and Analysis of Physicochemical Properties
The plant samples were harvested from the Tongliao district of the Inner Mongolia Autonomous Region localized in Northern China. The aerial parts of D. moldavica were air-dried before being into a fine powder. Then, DMEs were prepared as described. 22 In brief, the plant powder was extracted twice with 65% ethanol for 120 min at a temperature of 60 °C. After removing the ethanol by vacuum distillation, the obtained extracts were separated with ethyl acetate (EtOAc) using separating funnels. Then, the resultant extracts were subjected to HPLC analysis for the simultaneous quantification of 5 compounds, including rosmarinic acid, tallianine, luteolin, diosmetin, and apigenin. 22
Animal Experiments
All procedures were approved by the Animal Care and Use Committee of Baotou Medical College. Male Wistar rats weighing 160 to 180 g were purchased from Beijing Vital River Laboratory Animal Technology (Beijing, China). All rats were kept in individual cages on a 12-h light/dark cycle with free access to rodent chow and water under a temperature-controlled pathogen-free environment. To induce a 2K1C model, animals were anesthetized with pentobarbitone sodium (50 mg/kg by intraperitoneal injection). The left renal artery was constricted with a silver clip as the previous report to produce hypertensive rats. 28 The sham-operated animals underwent the same operation but without artery ligation. Weekly, the blood pressure was monitored and recorded using a non-invasive blood pressure analyzer provided by Chengdu Instrument Factory (Sichuan, China). The rats were divided into the sham group, model group, and treatment group, respectively. The animals were administrated either DME (50 mg/kg/day) or an equal volume of saline for 4 weeks by oral gavage. Upon termination of the experiments, the investigated animals were anesthetized with an anesthetic overdose (4% pentobarbital sodium), before being euthanized.
Sample Collection
Fresh fecal pellets were collected from the rat's cecum and frozen at −80 °C for analysis of gut microflora and miRNAs. Arterial blood was obtained from the abdominal aorta and serum was separated for serum metabolomics analysis.
Fecal DNA and RNA Extraction
Bacterial genomic DNA was isolated from fecal pellets using the MoBio Power Fecal DNA Isolation kit (Mo BioLaboratories, Carlsbad, USA). Total RNA was extracted with TRIzol reagent (Invitrogen, Carlsbad, USA). The obtained DNA and RNA were evaluated by electrophoresis on a denaturing agarose gel (Solarbio, Beijing, China), and the ratio of OD 260/280 was estimated by a spectrophotometer.
RNA Sequencing and Analysis
The mRNA was extracted with the NEBNext Poly(A) mRNA Magnetic Isolation Module (New England Biolabs, Inc., Ipswich, MA, United States), and the RNA-seq library was then generated using a KAPA Stranded RNA-Seq Library Preparation Kit (Illumina, San Diego, CA, USA). The quality of the libraries was evaluated on an Agilent 2100 bioanalyzer system. The prepared libraries were sequenced on Illumina HiSeq 2500 (San Diego, CA, USA). The raw sequencing data quality was checked using FastQC (v0.11.5). Quality trimmed raw reads were then aligned to reference genome sequence using HISAT (version 0.1.4). The data was further analyzed as described previously. 52
High-Throughput 16S Ribosomal RNA (rRNA) Gene Sequencing
The V3–V4 of the bacteria 16S rRNA genes were amplified using barcoded primer sets 341F (5’-ACT CCT ACG GGA GGC AGC AG-3’) and 806R (5’- GGA CTA CHV GGG TWT CTA AT-3’). 53 PCR products were purified using the AmpureXP beads (Beckman Coulter, Fullerton, CA, USA) and quantified using the Agilent 2100 bioanalyzer system. Then the amplicon library was sequenced on the Illumina MiSeq at the BGI-Shenzhen Co. Ltd (Shenzhen, China). The sequences were then clustered into operational taxonomic units (OTUs) with 97% similarity by UPARSE (version 7.0.1090). Alpha diversity was determined with the MOTHUR (version 1.31.2), while beta diversity was determined with the QIIME (version 1.8.0). LDA (Linear Discriminant Analysis) effect size (LEfSe) software was used for LEfSe analysis.
Metabolomics Analysis
After thawing at room temperature, metabolites in the serum were isolated. In brief, 100 μL of the sample was taken, and precooled extract (methanol: acetonitrile volume ratio: 2:1) was added. The mixture was then vortexed for 1 min and incubated at −20 °C for overnight. After centrifugation at 4000 g for 20 min at 4 °C, the supernatant was evaporated to dryness under vacuum. The residue obtained was redissolved in 50% methanol solution until quantification using ultra-performance liquid chromatography-tandem mass spectrometry (Q Exactive, Thermo Fisher Scientific, USA). The separation was performed on a Waters ACQUITY UPLC BEH C18 column (100 mm × 2.1 mm, 1.7 μm, Waters, USA) as described previously. 52 Processing of original chromatographic peak data and functional annotations were performed using the metabolomics R package metaX. 54
Correlation Analysis
Spearman correlation analysis was adopted to explore the correlation between two variables (such as the differentially expressed miRNAs and gut microbiota, and gut microbiota and metabolites), using p-value < 0.05 and R-value > 0.5 as the thresholds.
Statistical Analysis
SPSS statistical software version 21.0 was used for data analysis. The significant differences were assessed by one-way ANOVA or unpaired student's t-test. Data are expressed as means ± SEM, and a value of P < 0.05 indicates statistical significance.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X241261020 - Supplemental material for Dracocephalum moldavica L. Extract Ameliorates Hypertension Through Modulating the Interaction Between miRNAs and Gut Microbiota in 2K1C Rats
Supplemental material, sj-docx-1-npx-10.1177_1934578X241261020 for Dracocephalum moldavica L. Extract Ameliorates Hypertension Through Modulating the Interaction Between miRNAs and Gut Microbiota in 2K1C Rats by Hui Yu, Zhangjun Chen, Huixia Chen and Zhanli Wang in Natural Product Communications
Footnotes
Abbreviations
Authors’ Contributions
HY and ZC conducted the study, performed analysis of the data, and prepared the manuscript. HC performed experiments. ZW and HY designed and supported the study, and edited the manuscript. All authors approved the final version of the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Natural Science Foundation of China (82170296, 82060084, and 82260092).
Ethical Approval
The Ethics Committee of Experimental Animals of Baotou Medical College (Baotou, China) approved the present study (approval no. 2021040).
Statement of Human and Animal Rights
All procedures performed on rats were approved by the Animal Care and Use Committee of Baotou Medical College.
Statement of Informed Consent
NA
Availability of Data
All data either generated or analyzed during this study are included in this published article or available from the corresponding author upon reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
