Abstract
Background
Breast cancer remains a significant global health challenge despite the emergence of various drug molecules. However, the adverse side effects of several drugs and chemotherapy necessitate the exploration of novel therapeutic strategies. Identifying effective therapeutic proteins specific to breast cancer is complex, and finding potential natural, non-cytotoxic inhibitors presents an even more significant challenge in this field.
Objectives
In this study, we aimed to identify various proteins responsible for the development of breast cancer, as well as explore the potential therapeutic application of various isoflavones as complementary agents for breast cancer management.
Materials and Methods
Analysis of The Cancer Genome Atlas (TCGA) RNA-Seq and protein expression data at the Human Protein Atlas was performed for the identification of proteins. Furthermore, selected proteins were used for molecular docking and dynamics against various isoflavone derivatives. In addition, pharmacokinetic activity was performed for the isoflavone derivatives.
Results
Molecular docking exhibited the most potent binding energy of −9.6 kcal/mol for the CRMP2-genistin complex, closely followed by the HER2-daidzin complex with a binding energy of −9.4 kcal/mol. Subsequent molecular dynamics simulations showed dynamic behavior, structural integrity, stability, and interaction stability of HER2 protein with ligand daidzin. According to ADMET data, most soy isoflavones satisfy the Lipinski, Pfizer, Ghose, and GoldenTriangle criteria, indicating drug-like properties. Immunotoxicity projections indicate daidzein has the least adverse effects, while in silico, cytotoxicity assays indicate minimal overall risk. Glycitin and daidzin have the lowest levels of cytotoxicity. According to the comprehensive ADMET profiles, soy-derived isoflavones can safely complement current breast cancer therapeutics.
Conclusion
Computational analysis revealed that these ligands had inhibitory potential against BC-related HER2 and CRMP2 proteins. These isoflavones could be used to develop nutraceuticals to ensure safe and effective breast cancer management.
Keywords
Introduction
Breast cancer or carcinoma (BC) mostly arises in the epithelial cells that line the ducts (85%) and are known as ductal carcinomas, while sometimes in lobules (15%) in the glandular tissue known as ductal carcinomas of the breast (Feng et al., 2018; Makki, 2015). BC can be non-invasive, which does not go beyond the milk ducts or lobules in the breast, or invasive, which spreads into surrounding breast tissue and then spreads to the nearby lymph nodes or other distant organs in the body. The hallmarks of advanced metastatic BC are stromal invasion and metastasis to regional lymph nodes or distant organs, which become incurable because of metastasis (Makki, 2015; Nwabo Kamdje et al., 2014; Testa et al., 2020). However, BC treatment can be highly effective, especially when the disease is identified early. BC remains a global health concern, exacerbated by issues such as drug resistance and adverse effects from conventional therapies. Studies also emphasize the role of various proteins in BC signaling. For instance, the human epidermal growth factor receptor-2 (HER2), also known as the ERBB2 pathway, is crucial for tumor progression and has been extensively targeted in therapies, although resistance remains a significant challenge (Angus et al., 2023). Several cancer therapeutic studies are focusing on the HER/EGFR/ErbB family of receptor tyrosine kinases (RTKs) and have exhibited promising outcomes, offering an alternative therapy with efficacy comparable to chemotherapy, especially in mutated forms of EGFR (Kjær et al., 2020; Schoeberl et al., 2017). Comprising four homologous RTKs—EGFR/ErbB1/HER1, ErbB2/HER2, ErbB3/HER3, and ErbB4/HER4—the EGFR/ErbB family kinases play a pivotal role in intercellular signaling, cell proliferation, differentiation, and migration (Ding et al., 2020; Green et al., 2016; Kunte et al., 2020; Loibl & Gianni, 2017). These single-pass transmembrane proteins translate extracellular signals, such as ligands or growth factors, into activating specific cell signaling cascades. BC is categorized into molecular subtypes by the presence or absence of estrogen, progesterone hormone receptors, and HER2 (Kunte et al., 2020; Testa et al., 2020; Webster et al., 2009). It has been well-established that estrogen participates in the pathophysiology of BC (Al-Shami et al., 2023). Estrogen receptor 1 (ESR1) or ER alpha (Era) promotes the proliferation of cancer tissues, while estrogen receptor 2 (ESR2) or ER beta (Erb) can protect against the mitogenic effect of estrogen in breast tissue. The expression status of Era and Erb may highly influence the development, treatment, and prognosis of BC (Paterni et al., 2014; Zhou & Liu, 2020). The top protein for target prediction of BC in our study is HER2, a member of the family of RTKs whose overexpression has been reported in numerous cancers, including BC (Galogre et al., 2023; Swain et al., 2023). Research indicates that about 20–30% of BC exhibit an overexpression of HER2, resulting in heightened activation of HER2 and the initiation of multiple downstream pathways that lead to uncontrolled cancer cell proliferation. HER2 binds tightly to other ligand-bound epidermal growth factor (EGF) receptor family members to form a heterodimer, stabilizing ligand binding and enhancing kinase-mediated activation of downstream signaling pathways, such as those involving mitogen-activated protein kinase (MAPK) and phosphatidylinositol-3 kinase (PI3K) (Hsu & Hung, 2016; Mayer & Arteaga, 2016). Drugs targeting these specific proteins or their interactions could offer novel treatment strategies for BC patients. Molecular docking and dynamic simulation studies provide valuable insights into the potential therapeutic interactions between distinct inhibitor drugs and crucial protein targets. In particular, a range of inhibitory drugs, including afatinib, pertuzumab, trastuzumab, ado-trastuzumab emtansine, and ceritinib, demonstrate substantial interactions with these protein targets. Notably, each of these drugs exhibits different inhibitory potentials, reflecting their unique mechanisms of action, and is accompanied by well-known side effects, emphasizing the importance of re-evaluating treatment strategies for BC.
Isoflavones, particularly those derived from soy, have garnered attention for their potential therapeutic effects. In this work, we use an in silico approach to predict the binding interactions between the different isoflavones and targeted proteins involved in the development of BC in order to investigate the anti-cancer potential of the isoflavones derived from soy. Furthermore, molecular dynamics simulation (MDS) analysis was carried out to verify the protein and ligand complexes’ stability. Additionally, ADMET and computational pharmacokinetic study of these isoflavones was conducted to understand their physicochemical properties.
Materials and Methods
Analysis of The Cancer Genome Atlas (TCGA) RNA-Seq and Protein Expression Data at the Human Protein Atlas (HPA)
For this metadata analysis, we used data from TCGA, which offered transcriptomics information for 1,075 patients. There were 12 men and 1,063 women in this sample. It categorizes them into different groups, including “cancer tissue enriched,” “cancer group enriched,” “cancer tissue enhanced,” “expressed in all,” “mixed,” and “undetected.” Analysis of RNA expression data for five putative BC-associated proteins was obtained from TCGA and is accessible on the Protein Atlas website (
Intra-species Protein–Protein Interaction (PPI) Network Construction
The PPI was conducted in the EMBL-EBI IntAct database (
Prediction and Selection of Diagnostic Marker Protein Target for BC
This modeling study selected five proteins based on cancer specificity, tau specificity score (RNA), and tissue specificity to predict diagnostic marker proteins associated with BC. A critical protein is HER2, well known for its overexpression in specific subtypes of BC, and it is upregulated in blood from such BC patients (Magis et al., 2020).
Retrieval of Protein Structures
After TCGA RNA-Seq data, HPA, PPI, and pathway analysis, we identified HER, CRMP2, CA15.3, ESR1 and ESR2 as promising drug targets for BC. The crystal structure of the human HER2 (PDB ID- 7PCD), CRMP2 (PDB ID- 5LXX), CA15.3 (PDB ID- 1Y8X), MUC1 (PDB ID- 5T6P), ESR1 (PDB ID- 6CHZ), and ESR2 (PDB ID- 5TOA) was obtained from the RCSB-Protein Data Bank (Rutgers University, NJ, USA).
Retrieval of Molecules or Compounds
Soybean-derived six prominently known isoflavonoid compounds, daidzein (CID: 5281708), daidzin (CID: 107971), genistein (CID: 5280961), genistin (CID: 5281377), glycitein (CID: 5317750), and glycitin (CID: 187808), were retrieved from NCBI-PubChem. Meanwhile, known inhibitors and drugs of HER2 were downloaded from EMBL-CHEBI and DrugBank and used as positive controls for molecular docking.
Molecular Docking
This computational method employed AutoDock Vina (version 4.2) to predict the molecular-level interactions between two entities: anti-cancer drugs or inhibitors and target proteins (Trott & Olson, 2009). We performed molecular docking to determine the best binding configuration and energy (kcal/mol) between these compounds and target proteins. First, we assigned Gasteiger-Marsili partial charges to the ligand molecules in PDB format and converted them to PDBQT format. Next, we processed the target protein by removing water molecules and adding polar hydrogen atoms. We then introduced partial atomic Kollman charges and distributed the charge deficit. Finally, we converted the PDB file to PDBQT format (Ashraf et al., 2021; Elkhalifa et al., 2023).
MDS Trajectory Analysis
MDS was performed on a Linux operating system (LOS)-based machine using the GROMACS 2019.2 program and the GROMOS 43a1 force field (Abraham et al., 2015; Páll et al., 2015). A 50 ns MDS production run was conducted for individual systems (unbound and docked proteins) at a constant temperature of 300 K and pressure of 1 bar. Following the completion of MDSs, individual atomic coordinates within the overall MD trajectory for both the protein and the docked complex were subject to thorough analysis. Various GROMACS tools, namely, g_rmsd, g_rmsf, g_gyrate, g_sasa, and g_hbond, were employed for trajectory analyses. The resulting trajectory data were visualized using xmgrace 5.0.5 software, and the generated plots for root mean square deviation (RMSD), root mean square fluctuation (RMSF), radius of gyration (Rg), solvent-accessible surface area (SASA), and the number of hydrogen bonds, presented in .xvg files, were further illustrated using Microsoft 365 Excel Worksheet.
Assessment of ADMET and Computational Pharmacokinetics
The SwissADME (release 2017) (Daina & Zoete, 2016; Daina et al., 2017) and ADMETlab 3.0 (Fu et al., 2024) are freely accessible online resources that assess small compounds’ pharmacokinetics, drug-likeness, and medicinal chemistry using comprehensive experimental datasets. The SMILES representation of the compounds was inputted, and 2D structure files were produced using SwissADME and ADMETlab 3.0 (Ghose et al., 1999; Lipinski et al., 2001; Martin, 2005; Muegge et al., 2001; Veber et al., 2002).
Results
Evaluation of RNA-Seq Information from TCGA
The RNA tissue category classification is based on analyzing mRNA expression levels of CDK4 and CDK6 in 17 different cancer tissues, using data from TCGA. This data is presented as the median fragments per kilobase of exon per million (FPKM) reads and provides valuable insights into mRNA expression across these cancer tissues. We used the HPA database to confirm the protein expression, which indicates that HER2, CRMP2, CA15.3, and ESR1 are strongly associated with BC (Figure 1A–B). This information makes understanding the gene expression patterns and possible prognostic value within different cancer types easier.
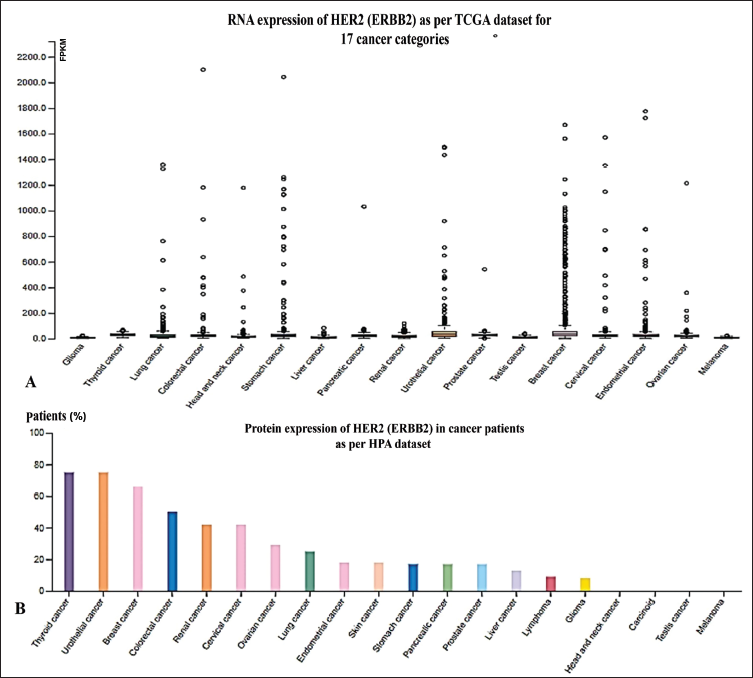
Most cancer tissues showed moderate to strong cytoplasmic immunoreactivity for CA15.3 or MUC1, especially for lung, breast, and ovarian cancer types. Nuclear positivity was observed in a few cases. TCGA RNA expression data of ESR2 categorizes it as undetected in any cancer category. In contrast, protein expression of ESR2 showed weak to moderate membranous positivity in cervical and colorectal cancer only. TCGA data (cancer tissue) suggest low cancer and low tissue specificity for HER2 gene transcript, located intracellularly, and has 11 different gene transcripts. Cell line expression analysis and group-enriched cell line specificity at HPA show that it is BC-specific, with a tau specificity score of 0.44. The tau specificity score ranging from 0 to 1 serves as a quantitative, graded scalar assessment of the specificity of gene expression. It is also known as a gene characterization score (Lüleci & Yılmaz, 2022; Fredholm et al., 2017). The other top four predicted proteins as diagnostic markers of BC in our model were CRMP2, CA15.3, ESR1, and ESR2, showing strong involvement in BC (Table S1 in the supplementary material). Table S1 in the supplementary material presents a comprehensive list of cancer-related human proteins that serve as Food and Drug Administration (FDA)-approved drug targets for various cancer types.
Analysis of Intra-species PPI Network
The intra-PPI data of five shortlisted gene transcripts were filtered to focus on their effective interactor proteins. The IntAct circular layout visual representation of the PPI network of HER2, a gene with the Ensembl ID ENSG00000141736, showed the interactions between HER2 and 860 other human proteins. Based on stringent criteria, this result was further enriched with a high-confidence MI score, robust experimental evidence, their interaction types, and associated interactions. Twenty-four proteins were selected that were interacting with HER2 by colocalization, phosphorylation, and dephosphorylation. ERBB2 engages in colocalization with STAT3, supported by a confidence MI score of 0.78. Similarly, LRIG1 colocalizes with ERBB2, indicated by interaction and an MI score of 0.75. ERBB2 and PTPN12 participate in dephosphorylation interactions and have a confidence MI score of 0.59. Additionally, ERBB2 colocalizes with EEA1, supported by an MI score of 0.57. The analysis was further refined, and the network was recreated using HPA data. It identified 57, 49, 7, 6, and 4 human proteins interacting with ESR1, HER2, DPYSL2, ESR2, and MUC1, respectively (Figure 2A–E). The edges and nodes in this network are color-coded, with green edges representing direct interactions between proteins and black edges indicating physical associations. The PPI network implies that ERBB2 and ESR1 proteins are strongly regulatory within the cellular context.
The Intra-Protein–Protein Interaction (PPI) Network Demonstrates (A) HER2, (B) MUC1, (C) ESR1, (D) DPYSL2, and (E) ESR2 and Their Respective Interacting Partners (Derived from Data in the Human Protein Atlas). The Edges’ and Nodes’ Colors Signify Proteins’ Interaction Type and Subcellular Localization. The Edges’ Colors Distinguish Direct Interactions (Green) and Physical Associations (Black) Among the Chosen Protein and Interacting Partners.
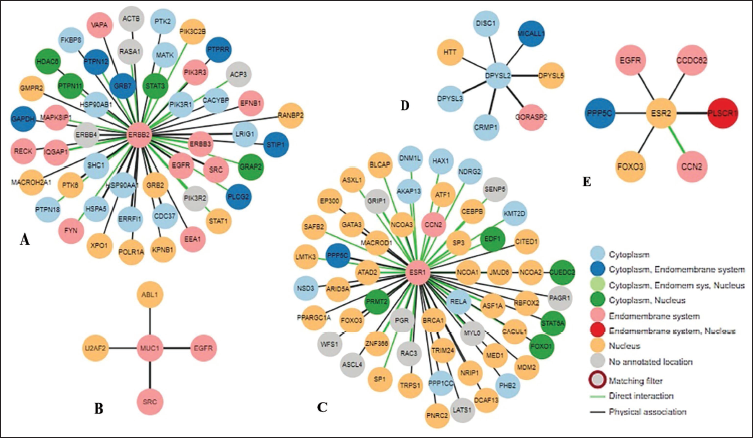
The PPI results indicate a network of protein interactions with associated interaction types, confidence levels, and MI scores. Notably, HER2 directly interacts with ERBB3 with a high confidence level, supported by an MI score of 0.97. ESR1 demonstrates a highly interactive network with NCOA1, showing a high-confidence physical association with an MI score of 0.84, contrary RELA and BRCA1 exhibit direct interactions with high confidence, featuring MI scores of 0.83 and 0.81, respectively.
Molecular Docking
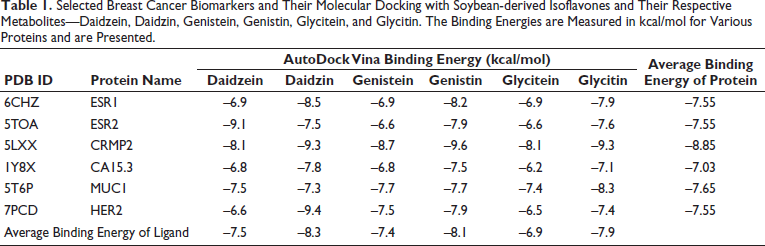
By applying AutoDock Vina, molecular binding energies (kcal/mol) for various proteins against these isoflavones (daidzein, daidzin, genistein, genistin, glycitein, and glycitin) were presented in Table 1. ESR2’s binding energies range from –6.6 to –9.1 kcal/mol. Meanwhile, ESR1 demonstrates binding energies varying from –6.9 to –8.5 kcal/mol. CRMP2 exhibits binding energies between –8.1 and –9.6 kcal/mol. CA15.3 shows binding energies spanning from –6.2 to –7.8 kcal/mol. MUC1 displays binding energies ranging from –7.3 to –8.3 kcal/mol. Finally, for HER2, the binding energies range from –6.5 to –9.4 kcal/mol. Daidzein exhibits the highest affinity of –9.1 kcal/mol with Erβ, whereas it has an exceptionally low affinity to HER2 (–6.6 kcal/mol). Daidzin exhibits the highest binding energy of –9.4 kcal/mol with HER2. Genistein binds to CRMP2 with the highest affinity of –9.6 kcal/mol. The average binding energy of these six compounds was highest (–8.85 kcal/mol) for CRMP2 while lowest (–7.03 kcal/mol) for cancer antigen 15-3. This comparative analysis reveals variations in binding energies across different proteins and isoflavones.
Selected Breast Cancer Biomarkers and Their Molecular Docking with Soybean-derived Isoflavones and Their Respective Metabolites—Daidzein, Daidzin, Genistein, Genistin, Glycitein, and Glycitin. The Binding Energies are Measured in kcal/mol for Various Proteins and are Presented.
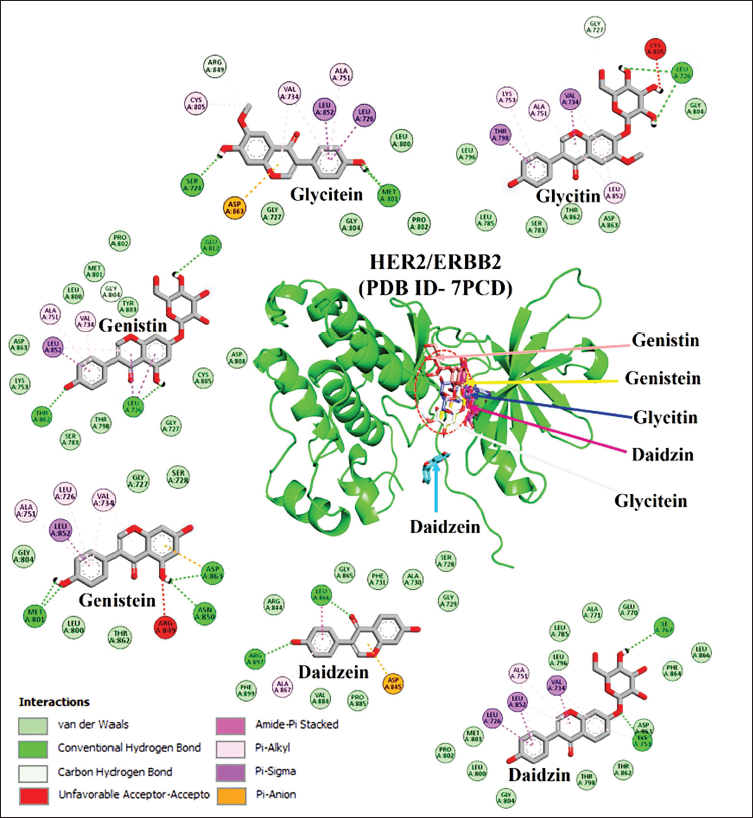
Binding Affinity and Interaction Profile of Isoflavones with HER2
The molecular docking of isoflavones with HER2 is visually represented using PyMol in 3D. This visualization illustrates the interactions between isoflavones and the active site residues of the HER2 protein (Figure 3). Further, the 2D analysis delves into specific interactions for each isoflavone. Daidzein and daidzin, for instance, form two hydrogen bonds each. However, daidzin exhibits stronger interactions due to the formation of 12 van der Waals (VDW), 3 pi–sigma, and 1 pi–alkyl interaction. Genistein establishes four hydrogen bonds, five VDW interactions, three pi–alkyl, one pi–sigma, and one unfavorable acceptor–acceptor bond with HER2 binding cavity residues. Conversely, genistin forms 3 hydrogen bonds, 11 VDW interactions, 2 pi–alkyl, 1 pi–sigma, and 1 carbon–hydrogen bond. Glycitein and glycitin also display distinct interaction patterns with HER2 binding cavity residues, encompassing hydrogen bonds, VDW interactions, pi–alkyl, pi–sigma, and other specific bonds. This comprehensive analysis provides a nuanced understanding of how each isoflavone interacts with the HER2 protein at the molecular level, offering valuable insights for further research in the context of HER2-related conditions.
The Molecular Docking of Isoflavones with HER2 (PDB ID-7PCD). The PyMol 3D Visualization Highlights the Interactions Between Isoflavones and Active Site Residues. The Remaining Five Molecules are Docked Within a Single Binding Cavity Apart from Daidzein. The 2D Discovery Studio Illustration Provides Insights into Eight Types of Interactions: van der Waals (VDW) (Depicted in Green), Conventional H-bond (Dark Green), Carbon–Hydrogen Bond (Light Green), Unfavorable Acceptor–Acceptor (Red), Amide–Pi Stacked (Dark Pink), Pi–Alkyl (Pink), Pi–Sigma (Purple), and Pi–Anion (Orange). This 2D Analysis Offers a Detailed Perspective on the Molecular-level Interactions Occurring Within the Binding Pocket Residues of HER2. Daidzein (LEU866, ARG897) and Daidzin (LYS753, ILE767) Each Form Two Hydrogen Bonds, with Daidzin Displaying Stronger Interactions Than Daidzein Through The Formation of 12 VDW, 3 Pi–Sigma, and 1 Pi–Alkyl Interaction.
Elucidation of MDS of Daidzin with HER2
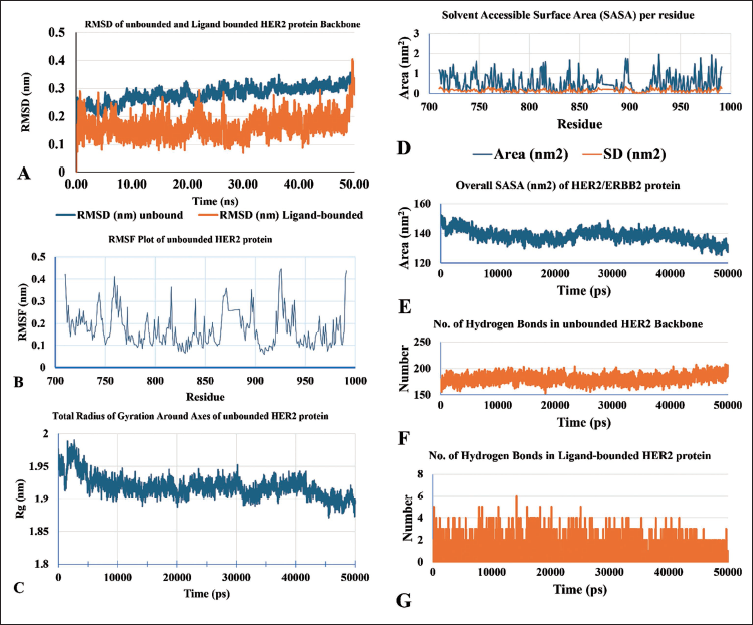
The average binding energy of the ligand daidzin is notably negative across all six proteins studied, with an exceptionally high value of –8.3 kcal/mol. Notably, for the HER2 protein, the binding energy reaches an exceptionally high level of –9.4 kcal/mol. Examination of the RMSD graph for the unbound HER2 protein backbone reveals a deviation ranging from 0.2 to 0.35 nm throughout the 50 ns simulation. In contrast, the ligand-bound HER2 protein displays a deviation between 0.1 and 0.4 nm over the same simulation time. Interestingly, the RMSD plots for the unbound and ligand-bound complexes exhibit numerous overlapped regions, suggesting stability throughout the simulation. After the 50 ns simulation, the unbound and ligand-bound complexes demonstrate equivalent RMSD values, affirming their stability and suitability for further analysis (Figure 4A). Considering that the HER2 protein’s binding domain encompasses 281 residues (amino acids 710–991), the elevated RMSF values observed for specific Cα-backbone residues (ranging from 0.3 to 0.45 nm) signify a heightened level of flexibility, particularly in a distinct region characterized by increased flexibility and accessibility. Conversely, many residues exhibit low RMSF values (ranging from 0.1 to 0.25 nm), indicating constrained movement and a region of reduced flexibility within the HER2 protein structure (Figure 4B). These variations in RMSF values suggest a potential correlation between flexibility and the diverse binding affinities of the residues. The Rg graph illustrates an overall decrease in Rg values from 1.98 to 1.88 nm for the unbound HER2 protein structure, suggesting a compactly packed state and stable folding of the HER2 protein (Figure 4C). The calculation of local SASA per residue revealed a range of 0 to 2.0 nm2, indicating varying exposure of HER2 protein residues to the solvent. Higher SASA values suggest increased surface accessibility for solvent or ligand interactions, potentially leading to more conformational changes (Figure 4D). Conversely, the overall global SASA of the unbounded HER2 protein per residue decreased after 50 ns of MDS, ranging from 150 to 125 nm2. Throughout the 50 ns MDSs, hydrogen bonds between the atoms of unbounded and ligand-bounded HER2 proteins were quantified individually. Figure 4F illustrates the fluctuation in the number of hydrogen bonds among unbounded HER2 backbone residues, ranging from 152 to 208. In contrast, the number of hydrogen bonds between the ligand (daidzin) and HER2 protein varies from 1 to 6 per residue, with an average of 3 hydrogen bonds forming per residue (Figure 4E–G). This analysis provides insights into the dynamic interactions and stability of the protein and its complex with the ligand during the simulation.
This Figure Illustrates the Plots Derived from the 50 ns (50,000 ps) Molecular Dynamics Simulations of the Unbounded and Ligand-bound (Daidzin) HER2 Protein Backbone. (A) Root Mean Square Deviation (RMSD), (B) Root Mean Square Fluctuation (RMSF), and (C) Radius of Gyration (Rg). (D) Local Solvent-accessible Surface Area (SASA) per Residue, (E) Overall Global SASA, (F) the Number of Hydrogen Bonds in HER2 Backbone Residues, and (G) the Number of Hydrogen Bonds Between the Ligand (Daidzin) and HER2 Residues. These Representations Show the Protein’s Structural Stability, Flexibility, and Compactness During the Simulation Period, Showing the Dynamic Behavior of HER2 in Both Unbounded and Ligand-bound States.
ADMET Analysis
ADMET analysis assessed the drug-like properties of six isoflavones derived from soy: daidzein, daidzin, genistein, genistin, glycitein, and glycitin. The evaluation of these compounds revealed that they get efficiently absorbed, distributed, metabolized, and excreted (ADMET) within the body. ADMET analysis assesses different molecular properties, including the count of hydrogen bond acceptors (nHA), the count of hydrogen bond donors (nHD), the count of rotatable bonds (nRot), the count of rings (nRing), the maximum count of atoms in the most extensive ring (MaxRing), the count of rigid bonds (nRig), the count of heteroatoms (nHet), and the formal charge (Fchar). Most of the examined isoflavones complied with the Lipinski, Pfizer, Ghose, and GoldenTriangle criteria, suggesting positive ADMET characteristics (Table S2 in the supplementary material). The Boiled-Egg plot depicts the blood–brain barrier (BBB) penetration capacity of daidzein (1) and glycitein (2); meanwhile, the human intestinal absorption (HIA) tendency of genistein (3) was higher than daidzein (4), glycitin (5), and genistin (6) (Table S2 in the supplementary material). The comprehensive analysis of absorption attributes for the compounds reveals distinct predicted values across various parameters. Daidzin and glycitin (glycosides) have significantly higher HIA values than their aglycone counterparts (daidzein and genistein). Daidzin and glycitin again outperform daidzein and genistein across all bioavailability levels, supporting the potential benefit of glycosylation (Table S3 in the supplementary material).
Discussion
The findings of our study provide a comprehensive analysis of the mRNA and protein expression levels of four key genes—ESR1, CA15.3, HER2, and CRMP2, shedding light on their significant relationships with BC (Chen et al., 2003; Ding et al., 2020; Galogre et al., 2023; Schoeberl et al., 2017; Testa et al., 2020). Their distinct expression patterns provide valuable insights into the potential diagnostic significance of these proteins against BC. CRMP2 exhibited moderate to strong cytoplasmic positivity, predominantly in tumor stroma and a subset of tumor cells in glioma and testis cancer (Barnard et al., 2015). Conversely, colorectal, cervical, and endometrial cancers were negative for CRMP2 expression. CA15.3 showed moderate to strong cytoplasmic immunoreactivity, especially in lung, breast, and ovarian cancers (Goodwin et al., 2021; Li et al., 2022). RNA expression analysis of ESR1, as per TCGA cancer enhanced dataset, showed strong nuclear positivity in breast, endometrial, and ovarian cancers. Other cancer categories were negative for ESR1 RNA and protein expression. ESR2, based on TCGA RNA expression data, was not detected in any cancer category. However, protein expression showed weak to moderate membranous positivity in cervical and colorectal cancer. HER2 receptors command attention, particularly in BC. Approximately 20–30% of BC exhibit HER2 overexpression, fueling uncontrolled cell proliferation (Galogre et al., 2023; Kunte et al., 2020; Schoeberl et al., 2017). As a therapeutic target, trastuzumab emerges as an effective intervention, highlighting its diagnostic significance in BC. With a tau specificity score of 0.44, HER2 demonstrates low tissue specificity, featuring cytoplasmic and membranous expression in various tissues and interactions with 49 proteins (Figure 1A). Glycoprotein mucin 1 (MUC1) or cancer antigen 15-3 (CA 15-3) navigates a complex landscape, a transmembrane glycoprotein overexpressed in BC cells. CA 15-3, an associated glycoprotein, aids in monitoring disease progression and treatment response. Expressing primarily on the apical surface of epithelial cells, especially in the breast, and uterus, it interacts with four proteins, including EGFR, SRC, U2AF2, and ABL1 (Table S1 in the supplementary material). ESR1, also known as Era, emerges as a pivotal biomarker in BC, dictating treatment strategies and prognostication (Fredholm et al., 2017). Its prevalence or absence significantly influences therapeutic responses in hormone receptor-positive BC. ER1 proves particularly prominent in BC, ovarian, and prostate cancers, earning the distinction as a prognostic and diagnostic marker, with a tau specificity score of 0.67 (Table S1 in the supplementary material).
The investigation into the intra-species PPI network landscape, specifically its association with BC progression, delved into a meticulous analysis refined with data from the HPA (Digre & Lindskog, 2023; Thul & Lindskog, 2018). The outcome was a discerning identification of interacting proteins associated with key biomarkers, namely, ESR1, HER2, DPYSL2, ESR2, and MUC1. This network, vividly depicted in Figure 2A–E, unfolds a complex web of 57, 49, 7, 6, and 4 human proteins intricately connected with ESR1, ERBB2, DPYSL2, ESR2, and MUC1. The PPI network’s visual representation underscores the robust regulatory role of ERBB2 and ESR1 proteins within the cellular context, as evidenced by their high number of interacting partners and network topology (Figure 2A and C). This points toward their potential importance in BC development and progression. The subcellular localization of the PPI network reveals that ERBB2, ESR1, and MUC1 express in the endomembrane system, ensuring the inclusion of interactions specific to Homo sapiens in the datasets. A notable finding within this intricate network is the direct interaction between HER2 and ERBB3, supported by a high confidence level and an MI score of 0.97. These findings contribute significantly to our understanding of the molecular landscape associated with BC progression, opening avenues for further exploration and targeted therapeutic interventions. This overexpression, in turn, triggers the activation of the PI3K/AKT signaling pathway, a crucial pathway in cell proliferation and survival, known to be dysregulated in many types of cancers (Mayer & Arteaga, 2016; Testa et al., 2020). The network pinpoints a specific cascade where ERBB2 upregulates ERBB4, activating PIK3CA, a key enzyme in the PI3K/AKT pathway (Figure 2).
In the current study, the therapeutic potential of six soybean-derived bioactive isoflavones and their metabolites (daidzein, daidzin, genistein, genistin, glycitein, and glycitin) was docked with six promising protein targets in BC: ESR1, ESR2, CRMP2, CA15.3, MUC1, and HER2 (Cameron et al., 2017; Shimada et al., 2014). Interestingly, daidzin exhibited the most favorable binding energy and interactions compared to other isoflavones across all studied protein targets (Table 1 and Figure 3). The observed prominence of daidzin in binding energy and interactions compared to daidzein, genistein, genistin, glycitein, and glycitin underscores its potential as a promising candidate for further investigation in BC therapy. On average, these findings contribute valuable information indicating the differential affinities of soybean-derived compounds toward specific protein targets and encouraging further exploration of daidzin as a potential therapeutic agent for BC treatment (Elkhalifa et al., 2023). The study lays the groundwork for future research to understand the molecular mechanisms underlying the observed interactions and potential translational applications in BC therapeutics. Several research studies suggest that they can modulate BC signaling pathways by targeting specific human proteins like HER2, CRMP2, CA15.3, ESR1, and ESR2. Soy isoflavone genistein has demonstrated anti-cancer properties, including suppressing HER2 and ERα expression. HER2 is a RTK overexpressed in specific BC (HER2+). Genistein exhibits a variety of biological functions that contribute to its BC preventive effects. It competes with estrogens for binding to estrogen receptors (ERs) and regulates estrogen-dependent gene expression (Markiewicz et al., 1993). Additionally, genistein acts as a protein tyrosine kinase inhibitor (Shao et al., 1998), suppresses angiogenesis (Fotsis et al., 1995), inhibits DNA topoisomerase II activity (Markovits et al., 1989), induces apoptosis of BC cells (Katdare et al., 2002; Pagliacci et al., 1994), and downregulates HER2 (Katdare et al., 2002; Li et al., 1999) and ERα expression (Chen et al., 2003; Mai et al., 2007). Therefore, the supplementation of genistein suppresses HER2-overexpressing BC cells (Bhat et al., 2021).
The MDS results of the daidzin and HER2 complex provide a comprehensive understanding of the structural dynamics and stability of the binding interaction over a 50 ns simulation period. The RMSD analysis of the unbound HER2 protein backbone reveals a deviation ranging from 0.2 to 0.35 nm, indicating a stable and compactly packed state throughout the simulation. Interestingly, the ligand-bound HER2 protein exhibits a slightly higher deviation (0.1–0.4 nm), suggesting that daidzin induces subtle structural changes, a common phenomenon in ligand binding (Figure 4A). While slight deviations exist, the overlap between unbound and bound RMSD plots suggests overall stability. Specific regions of the HER2 protein exhibit higher flexibility (RMSF values 0.3–0.45 nm), particularly in the binding domain (residues 710-991), indicating potential interaction points. Other regions remain rigid (RMSF 0.1–0.25 nm), suggesting key structural elements (Figure 4B). This flexibility variation might correlate with binding affinities of different residues. The Rg analysis shows a decrease in Rg values from 1.98 to 1.88 nm for the unbound HER2 protein, indicating a compactly packed and stably folded structure (Figure 4C). The analysis of local SASA per residue reveals varying exposure of HER2 protein residues to the solvent, with higher SASA values suggesting increased surface accessibility for solvent or ligand interactions (Figure 4D). The overall global SASA of the unbound HER2 protein per residue exhibits a decrease, indicating a stable protein structure with reduced solvent exposure (Figure 4E).
The diminished SASA values imply a lower content of exposed surface amino acids or higher hydrophobicity of the HER2 protein. Previous studies on ERBb kinases have shown hydrophobic sub-regions, including the C-spine, R-spine, hydrophobic core, and αC-β4 regions. Notably, active conformations minimize SASA compared to inactive forms, and the transition from inactive to active states involves an increase in hydrophobicity, settling into a hydrophobically favorable region indicative of a well-formed C-spine in active conformations (Shi et al., 2011; Telesco & Radhakrishnan, 2009). The evaluation of hydrogen bonds within the unbound HER2 protein backbone demonstrates fluctuations in the number of hydrogen bonds formed among residues (Figure 4F). In contrast, the ligand-bound HER2 complex exhibits varying hydrogen bonds between the ligand (daidzin) and HER2 protein, ranging from 1 to 6 per residue, with an average of 3 hydrogen bonds forming per residue (Figure 4G). Highlighting the dynamic interaction between daidzin and HER2 provides compelling evidence for the stable interaction between daidzin and HER2, identifying potential binding hotspots and dynamic hydrogen bonding patterns. BCs that are positive for HER2 demonstrate a particularly aggressive clinical profile, exhibiting limited responsiveness to standard chemotherapy and a heightened propensity for recurrence and metastasis.
Significantly, some research has highlighted the enduring impacts of trastuzumab, whether administered independently or in conjunction with chemotherapy, on patients diagnosed with HER2-positive BC. Across these studies, a consistent finding emerges—trastuzumab demonstrates a substantial influence on disease-free and overall survival (Cameron et al., 2017; Testa et al., 2020; Tolaney et al., 2019). Two novel drugs designed to target HER2 have been incorporated into the clinical treatment landscape for HER2-positive BCs: (a) trastuzumab emtansine, an antibody-drug conjugate pairing trastuzumab with the cytotoxic agent emtansine—a microtubule inhibitor, and (b) pyrotinib, an irreversible pan-ERB RTK inhibitor specifically targeting HER1, HER2, and HER4. Notably, adjuvant trastuzumab emtansine demonstrated a 60% lower risk of recurrence of invasive BC or mortality compared to trastuzumab alone in patients with HER2-positive early BC who exhibited residual invasive disease post neoadjuvant therapy completion (von Minckwitz et al., 2019).
Most of the isoflavones studied were found to meet Lipinski, Pfizer, Ghose, and GoldenTriangle criteria, indicating drug-like qualities, according to ADMET data. Genistein has the highest HIA potential in the Boiled-Egg plot; however, daidzein, glycitein, and genistein have different BBB penetration capacities. Interestingly, all isoflavones are unlikely PGP substrates, indicating negligible medication absorption interference. The PGP pessimistic predictions for daidzin, genistin, and glycitin show they will not inhibit medication absorption. These detailed ADMET profiles suggest that soy-derived isoflavones like daidzin, genistin, and glycitin can supplement the existing BC drugs’ without any risks. Toxicity predictions suggest minimal overall risk, with glycitin and daidzin showing the lowest cytotoxicity. Daidzein might also have the least potential for immune system effects.
Conclusion
Our study explored BC treatment and identified potential therapeutic proteins associated with this prevalent and challenging disease. Through various databases, RNA-Seq data analysis, and PPI networks, essential proteins such as HER2, CRMP2, CA15.3, and ESR1 were identified for BC progression. Furthermore, molecular docking analyses revealed strong binding affinities, with the CRMP2–genistin complex exhibiting the highest binding energy, closely followed by the HER2–daidzin complex. Further, a comprehensive 50 ns MDS revealed the dynamic behavior, solvent exposure, structural integrity, and stability of the HER2 protein in a complex with daidzin. This study presents promising initial results for daidzin as a potential BC therapeutic candidate. However, further research is necessary to validate its efficacy, elucidate its mechanisms of action, and assess its safety and specificity in vivo.
Footnotes
Abbreviations
ADMET: Absorption, distribution, metabolism, excretion, and toxicity; CA15.3: Carcinoma-associated mucin; CRMP2: Collapsin response mediator protein 2; DPYSL2: Dihydropyrimidinase like 2; EGF: Epidermal growth factors; ESR1: Estrogen receptor 1; ESR2: Estrogen receptor 2; FPKM: Fragments per kilobase of exon per million; HER2: Human epidermal growth factor receptor-2; HIA: Human intestinal absorption; HPA: Human Protein Atlas; LOS: Linux operating system; MAPK: Mitogen-activated protein kinase; MDS: Molecular dynamics simulation; MUC1: Glycoprotein mucin 1; PIEK: Phosphatidylinositol-3 kinase; PPI: Protein–protein interaction; Rg: Radius of gyration; RMSD: Root mean square deviation; RMSF: Root mean square fluctuation; RTKs: Receptor tyrosine kinases; TCGA: The Cancer Genome Atlas.
Acknowledgments
This research has been funded by Scientific Research Deanship at University of Ha’il - Saudi Arabia through project number RG-21 121.
Authors’ Contributions
Conceptualization, SAA (Syed Amir Ashraf) and AEOE (Abd Elmoneim O Elkhalifa); methodology, MA (Mohd Adnan), FA (Fauzia Ashfaq), and MSB (Mirza Sarwar Baig); validation, AMA (Amir Mahgoub Awadelkareem), MS (Manojkumar Sachidanandan), and MIK (Mohammad Idreesh Khan); formal analysis, MSB and SAA; investigation, SAA, MSB, FA, and MA; data curation, MK (Mohammed Kuddus) FSA (Fahad Saad Alhodieb), MIK, AMA, and MS; writing—original draft preparation, AEOE, AMA, MSB, FA, MA, and MK; writing—review and editing, SAA, FA, MA, MS, FSA, and AMA; software, MIK, MK, FSA, MSB, and AEOE; visualization, MS, MK, FSA, FA, MA, MIK, and AMA; supervision, AEOE and SAA; project administration, AEOE and SAA. All authors have revised the manuscript and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval and Informed Consent
No humans or animals were used in the present research.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research has been funded by the Scientific Research Deanship at the University of Ha’il, Ha’il, Saudi Arabia, through project number RG-21121.
Supplementary Material
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
