Abstract
Background
The objective of this research is to explore the phytochemical constituents and the potential anti-inflammatory properties of the herb Oxytropis aciphylla.
Materials and Methods
From the ethyl acetate and n-butanol fractions derived from the 70% ethanol extract of O. aciphylla, a total of 17 distinct chemical entities were extracted and subsequently identified. Utilizing network pharmacology, the study predicted the possible anti-inflammatory targets and mechanisms of action of O. aciphylla. The study assessed the anti-inflammatory impact of O. aciphylla extract on RAW 264.7 (a murine macrophage cell line) cells induced with lipopolysaccharide (LPS) by quantifying the concentrations of nitric oxide (NO), tumor necrosis factor-alpha (TNF-α), and interleukin-6 (IL-6).
Results
The isolated compounds included 12 flavonoids, two triterpenoids, one fatty acid, and two sterols. These specifically comprise apigenin, luteolin, kaempferol, isorhamnetin, naringenin, eriodictyol, dihydrokaempferol, butein, 7,4’-homoisoflavanone, apigenin-7-O-β-D-glucopyranoside, isorhamnetin-3-O-β-D-gentiobioside, rutin, azukisapogenol, 3-O-[β-D-glucopyranosyl(1↓2)-β-D-glucuronopyranosyl] azukisapogenol methyl ester, tetracosanoic acid, daucosterol, and β-sitosterol. Network pharmacology analysis suggested that O. aciphylla may exert its anti-inflammatory effects through multiple components, targets, and pathways, such as kaempferol, apigenin, luteolin, AKT1, SRC, TNF, PTGS2, EGFR, MAPK, PI3K-Akt, VEGF, HIF-1, and TNF signaling pathways. The extract of O. aciphylla was found to markedly reduce the levels of NO, TNF-α, and IL-6 in RAW 264.7 cells that were stimulated with LPS, demonstrating a correlation with the dosage administered.
Conclusion
The investigation uncovered, for the first time, the phytochemical constituents and the mechanisms underlying the anti-inflammatory effects of O. aciphylla. These findings lay the groundwork for the scientific development and application of this botanical species as a natural remedy for inflammation.
Introduction
Oxytropis aciphylla Ledeb., a small perennial shrub of the Fabaceae family, is commonly known as Guijianchou and Huzhua. It is a distinctive species found in the desert and grassland ecosystems of the Mongolian Plateau (Committee, 1998). O. aciphylla holds significance in traditional Mongolian medicine, as historical records indicate that its stems and leaves are commonly crushed and boiled to extract a solution for external application in the treatment of abscesses (Committee, 1998; Zhu, 1989).
Recent studies have revealed that the genus Oxytropis, to which O. aciphylla belongs, predominantly contains flavonoids, triterpenes, and alkaloid compounds (She et al., 2012). These compounds are known to demonstrate an extensive array of pharmacological properties, notably their anti-inflammatory (Zhang et al., 2020), analgesic (Zhang et al., 2020), antimicrobial (Narangerel et al., 2021), antitumor (Sheng et al., 2015), and antioxidant (Wang et al., 2010) properties. However, there have been no reported investigations regarding the chemical composition and pharmacological effects of O. aciphylla. Consequently, the objective of this research is to examine the phytochemical constituents of the entire plant of O. aciphylla, investigate its anti-inflammatory mechanisms through network pharmacology, and conduct preliminary pharmacological validation through in vitro experiments. The goal is to discover lead compounds derived from this plant that possess anti-inflammatory activity.
Materials and Methods
Materials and Reagents
The herb of O. aciphylla was collected in May 2018 within Tokto County, Hohhot City, Inner Mongolia Autonomous Region. It was identified as the dried herb of O. aciphylla Ledeb. by Professor Qu from the Department of Pharmacognosy, Inner Mongolia Medical University.
Sephadex LH-20 (Pharmaecia Company), column-layer chromatographic silica gel, high silica gel GF pre-coated plate for TLC, MCI adsorption resin, HP-20 adsorption resin.
Fetal bovine serum (ExCell Bio), mouse TNF-α, IL-6 ELISA kits (Solarbio), CCK-8 cell viability assay kit (TransGen Biotech), RPMI-1640 culture medium, penicillin/streptomycin (Gibco), NO assay kit (Nanjing Jiancheng).
Extraction and Isolation of O. aciphylla
First, 5 kg of dried herb of O. aciphylla was taken and reflux extraction using 70% ethanol–water solution was performed for three cycles, each lasting 2 h. The extracted liquids were combined and the resulting mixture was filtered. The filtrate was concentrated under reduced pressure to obtain a total extract weighing 536.0 g. Water was added to the total extract, and it was repeatedly extracted with ethyl acetate and n-butanol saturated with water, three times for each solvent. Then, the ethyl acetate and n-butanol fractions were separately concentrated, obtaining 135.0 g of ethyl acetate fraction and 206.0 g of n-butanol fraction. They were stored at 4℃ for future use.
The extract from ethyl acetate was separated by silica gel column chromatography and washed with a mixture of dichloromethane–methanol that varied from 100:0 to 50:50. A total of 82 fractions, F1–F82, were obtained by combining the same fractions by TLC. Compound 17 (13 mg) was obtained from the repeated purification of F4 using Sephadex LH-20 gel column chromatography with elution using dichloromethane–methanol (4:6), dichloromethane–methanol (2:8), and pure methanol. Compound 15 (9 mg) was obtained from F5 using Sephadex LH-20 with elution using dichloromethane–methanol (4:6). F16 was purified multiple times using Sephadex LH-20 with elution using dichloromethane–methanol (4:6) and dichloromethane–methanol (2:8) to obtain compound 13 (11 mg). F17 was purified using Sephadex LH-20 with elution using dichloromethane–methanol (4:6) and dichloromethane–methanol (2:8), yielding compound 5 (6 mg) and compound 9 (25 mg). F20 was purified using Sephadex LH-20 with elution using dichloromethane–methanol (4:6) and dichloromethane–methanol (2:8) to obtain compound 1 (45 mg) and compound 7 (22 mg). F22 was purified using Sephadex LH-20 with elution using dichloromethane–methanol (4:6) and dichloromethane–methanol (2:8), yielding compound 3 (30 mg) and compound 6 (8 mg). F24 was purified using Sephadex LH-20 with elution using dichloromethane–methanol (4:6) to obtain compound 8 (12 mg). F27 and F28 were combined and purified using Sephadex LH-20 with elution using dichloromethane–methanol (4:6) to obtain compound 2 (26 mg). F33 was filtered by pumping to give compound 16 (14 mg). F58 was purified using Sephadex LH-20 with elution using dichloromethane–methanol (2:8) to obtain compound 10 (9 mg).
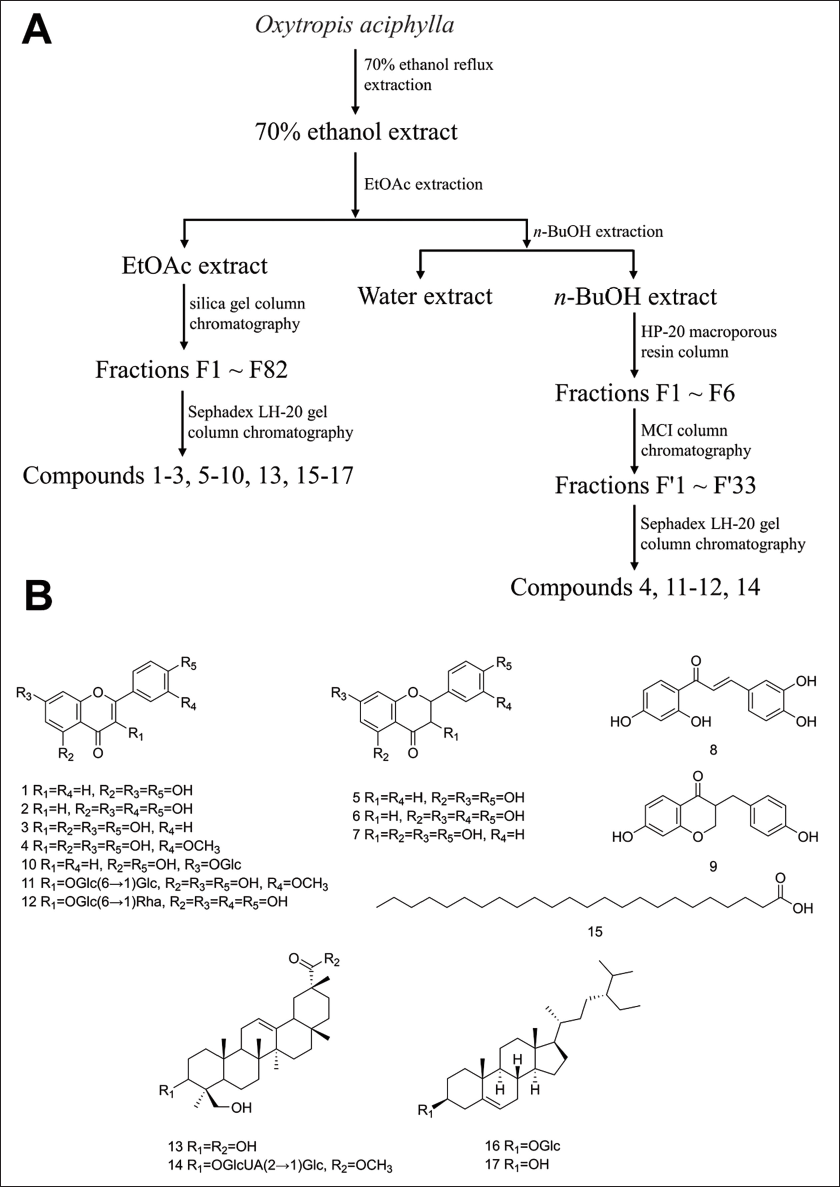
The n-butanol extract was initially separated using an HP-20 macroporous resin column, eluting with a gradient of 0–95% ethanol. Six fractions were collected and combined, designated as F1–F6. F4 (35 g) was further purified using MCI column chromatography with elution using a methanol–water (0:100 to 100:0) gradient. Through TLC analysis, 33 fractions, designated as F’1–F’33, were identified and combined. F’3 was purified using Sephadex LH-20 with elution using methanol–water (4:6). F’4 was purified using Sephadex LH-20 with elution using methanol–water (4:6) and pure methanol to obtain compound 4 (16 mg) and compound 11 (21 mg). F’12 was purified multiple times using Sephadex LH-20 with elution using methanol–water (4:6) and pure methanol, yielding compound 12 (22 mg). F’19–F’22 and F’28–F’29 were purified using Sephadex LH-20 with elution using pure methanol, followed by filtration to obtain compound 14 (6 mg). The extraction and separation process of O. aciphylla is shown in Figure 1(A).
Extraction and Isolation of 17 Compounds from O. aciphylla. (A) Extraction and Separation process of O. aciphylla. (B) 17 Compounds Isolated from O aciphylla.
Screening of Potential Anti-inflammatory Targets of O. aciphylla
The structures of 17 substances were obtained from the online resource PubChem (pubchem.ncbi.nlm.nih.gov). The structures of the substances were entered into the Swiss Target Prediction tool (SwissTargetPrediction.ch) to estimate the potential proteins of the O. aciphylla compounds. Then, “anti-inflammatory” and “inflammation”-related targets were searched in GeneCards (
Building a Network of Interacting Proteins
The information of potential anti-inflammatory targets was submitted to STRING 11 (string-db.org), specifying the organism as “Homo sapiens.” Based on a confidence score of 0.4, protein–protein interaction relationships were retrieved to construct a network. The data file was saved as a TSV file and loaded onto the Cytoscape 3.7.2 program. The cytoHubba plugin was utilized to identify the central targets in the network. The top 20 targets were selected based on Maximal Clique Centrality as the core targets for the anti-inflammatory activity of O. aciphylla.
GO and KEGG Enrichment Analysis
The core anti-inflammatory targets of O. aciphylla were imported into the DAVID database (david.ncifcrf.gov). The identifier was chosen to be the “official gene symbol,” and the list type was picked as “genelist.” The species were specified as Homo sapiens for conducting GO and KEGG enrichment analyses, and the top 20 results based on the count value were selected to identify the main signaling pathways and functions involved in the anti-inflammatory properties of O. aciphylla.
Cell Culture
The RPMI-1640 complete medium with 10% fetal bovine serum was used to culture the RAW264.7 cells. The cells were kept at 37°C with 5% CO2.
Cell Viability Assay Using CCK-8
Logarithmic phase RAW 264.7 cells were adjusted to a concentration of 1 × 104 cells/mL and seeded in a 96-well plate. After 24 hours of incubation, the cells were treated with different concentrations of the alcohol extract solution of O. aciphylla. The control group was treated with an equal volume of medium. After 24 hours of drug treatment, the cell viability was determined using the CCK-8 assay.
Assessment of NO, TNF-α, and IL-6 Production in Cells
Logarithmic phase RAW 264.7 cells were adjusted to a concentration of 1 × 105 cells/mL and seeded in a 24-well plate. The cells underwent pre-treatment with various doses of O. aciphylla alcohol extract and indomethacin (100 µM) for 2 hours, following 24 hours of incubation. After adding 1 µg/mL of LPS, the cells continued to incubate for another 24 hours. The model group was treated only with LPS, while the control group was treated only with the medium. The supernatant was collected for measurement. NO and ELISA assay kits were used to measure the NO, TNF-α, and IL-6 concentrations in the supernatant, following the kit’s guidelines. The indomethacin group (INM) served as the positive control, while the model group (MX) served as the negative control.
Statistical Methods
SPSS 22.0 software performed the statistical analysis of the experimental data. The data visualization was done using the GraphPad Prism 9.0 software. The mean ± standard deviation (mean ± SD) showed the results. A one-way ANOVA compared the groups. For homogeneous variances, the LSD test was applied, while for heterogeneous variances, Dunnett’s T3 test was applied. The level of significance was p < 0.05 for significant differences, p < 0.01 for highly significant differences, and p < 0.001 for extremely significant differences.
Results and Discussion
Structural Identification
Compounds 1–2, 4–11, and 13–15 were determined for their relative molecular mass through mass spectrometry (MS), and their structures were identified through nuclear magnetic resonance (NMR). Compounds 3, 12, 16, and 17 were identified by comparing them with standard substances using TLC and HPLC. Figure 1(B) displays the structures of 17 compounds extracted from O. aciphylla.
Compound 1: Yellow powder (purity > 98%), ESI-MS m/z: 271[M+H]+, molecular formula: C15H10O5. 1H-NMR (500 MHz, DMSO-d6) δ: 7.93 (2H, d, J = 8.5 Hz, H-2’, 6’), 6.92 (2H, d, J = 8.5 Hz, H-3’, 5’), 6.77 (1H, s, H-3), 6.48 (1H, d, J = 2.0 Hz, H-8), 6.20 (1H, d, J = 1.5 Hz, H-6); 13C-NMR (125 MHz, DMSO-d6) δ: 181.7 (C-4), 164.1 (C-2), 163.7 (C-7), 161.4 (C-5), 161.1 (C-4’), 157.3 (C-9), 128.4 (C-2’, 6’), 121.1 (C-1’), 115.9 (C-3’, 5’), 103.7 (C-10), 102.8 (C-3), 98.8 (C-6), 93.9 (C-8). The above data were in general agreement with literature reports (Zhang et al., 2017), so compound 1 was determined to be apigenin.
Compound 2: Yellow powder (purity > 98%), ESI-MS m/z: 287[M+H]+, molecular formula: C15H10O6. 1H-NMR (500 MHz, MeOD) δ: 7.40 (2H, m, H-2’, 6’), 6.92 (1H, d, J = 8.5 Hz, H-5’), 6.65 (1H, s, H-3), 6.46 (1H, d, J = 2.0 Hz, H-8), 6.22 (1H, d, J = 1.5 Hz, H-6); 13C-NMR (125 MHz, MeOD) δ: 183.9 (C-4), 166.4 (C-2), 166.1 (C-7), 163.3 (C-5), 159.5 (C-9), 151.0 (C-4’), 147.1 (C-3’), 123.8 (C-1’), 120.3 (C-6’), 116.8 (C-5’), 114.2 (C-2’), 105.4 (C-10), 104.0 (C-3), 100.2 (C-6), 95.1 (C-8). The above data were in general agreement with literature reports (Zhang et al., 2017), so compound 2 was determined to be luteolin.
Compound 3: Yellow powder (purity > 98%), soluble in methanol, dark spot under UV254, non-fluorescent under UV365, 10% ethanol sulfate solution shows yellow color, HCI-Mg color reaction is red. TLC identification with kaempferol standard (petroleum ether-ethyl acetate, trichloromethane–methanol, two kinds of unfolding agent system), in the same color development process, the Rf value is consistent; at the same time, mixed with the kaempferol standard for spotting, TLC identification, showing a single spot; using the HPLC method for sample determination, this sample showed consistent retention time with the kaempferol standard under the same chromatographic conditions, and the physico-chemical properties of kaempferol reported in the literature were basically consistent (Cai et al., 2007), so compound 3 was determined to be kaempferol.
Compound 4: Light yellow powder (purity > 98%), ESI-MS m/z: 315[M-H]-, molecular formula: C16H12O7. 1H-NMR (500 MHz, DMSO-d6) δ: 7.76 (1H, d, J = 2.0 Hz, H-2’), 7.68 (1H, dd, J = 8.5, 2.0 Hz, H-6’), 6.93 (1H, d, J = 8.5 Hz, H-5’), 6.46 (1H, d, J = 2.0 Hz, H-8), 6.18 (1H, d, J = 2.0 Hz, H-6), 3.81 (3H, s, 3’-OCH3); 13C-NMR (125 MHz, DMSO-d6) δ: 176.3 (C-4), 164.7 (C-7), 161.1 (C-9), 156.7 (C-5), 149.3 (C-4’), 147.8 (C-2), 147.0 (C-3’), 136.3 (C-3), 122.5 (C-1’), 122.2 (C-6’), 116.0 (C-2’), 112.3 (C-5’), 103.4 (C-10), 98.8 (C-6), 94.1 (C-8), 56.3 (-OCH3). The above data were in general agreement with literature reports (Yan et al., 2017), so compound 4 was determined to be isorhamnetin.
Compound 5: White needle-like crystals (purity > 98%), ESI-MS m/z: 273[M+H]+, molecular formula: C15H12O5. 1H-NMR (500 MHz, MeOD) δ: 7.32 (2H, d, J = 8.0 Hz, H-2’, 6’), 6.83 (2H, d, J = 8.0 Hz, H-3’, 5’), 5.88 (2H, dd, J = 7.5, 2.0 Hz, H-6, 8), 5.34 (1H, dd, J = 13.0, 3.0 Hz, H-2), 3.13 (1H, dd, J = 17.0, 13.0 Hz, H-3a), 2.71 (1H, dd, J = 17.0, 3.0 Hz, H-3e); 13C-NMR (125 MHz, MeOD) δ: 197.5 (C-4), 168.4 (C-7), 165.5 (C-5), 164.9 (C-9), 159.1 (C-4’), 131.4 (C-1’), 129.1 (C-2’, 6’), 116.3 (C-3’, 5’), 103.4 (C-10), 97.1 (C-6), 96.2 (C-8), 80.5 (C-2), 44.1 (C-3). The above data were in general agreement with literature reports (Zhang et al., 2017), so compound 5 was determined to be naringenin.
Compound 6: Yellow powder (purity > 98%), ESI-MS m/z: 287[M-H]-, molecular formula: C15H12O6. 1H-NMR (500 MHz, MeOD) δ: 6.91 (1H, d, J = 1.5 Hz, H-2’), 6.79 (1H, d, J = 1.5 Hz, H-6’), 6.77 (1H, d, J = 7. 5 Hz, H-5’), 5.90 (1H, d, J = 2.0 Hz, H-8), 5.88 (1H, d, J = 2.0 Hz, H-6), 5.28 (1H, dd, J =13.0, 3.5 Hz, H-2); 13C-NMR (125MHz, MeOD) δ: 197.8 (C-4), 168.4 (C-7), 165.5 (C-5), 164.9 (C-9), 146.9 (C-4’), 146.5 (C-3’), 131.8 (C-1’), 119.3 (C-6’), 116.4 (C-5’), 114.8 (C-2’), 103.4 (C-10), 97.1 (C-6), 96.2 (C-8), 80.6 (C-2), 44.2(C-3). The above data were in general agreement with literature reports (Wang et al., 2018), so compound 6 was determined to be eriodictyol.
Compound 7: Yellow powder (purity > 98%), ESI-MS m/z: 287[M-H]-, molecular formula: C15H12O6. 1H-NMR (500 MHz, MeOD) δ: 7.36 (2H, d, J = 2.0 Hz, H-2’, 6’), 6.83 (2H, d, J = 2.0 Hz, H-3’, 5’), 5.92 (1H, d, J = 2.0 Hz, H-8), 5.88 (1H, d, J = 2.0 Hz, H-6), 4.98 (1H, d, J = 11.5 Hz, H-2), 4.54 (1H, dd, J = 11.5, 2.0 Hz, H-3); 13C-NMR (125MHz, MeOD) δ:198.6 (C-4), 168.8 (C-7), 165.4 (C-5), 164.6 (C-9), 159.3 (C-4’), 130.4 (C-2’, 6’), 129. 4 (C-1’), 116.2 (C-3’, 5’), 101.9 (C-10), 97.4 (C-6), 96.4 (C-8), 85.0 (C-2), 73.7 (C-3). The above data were in general agreement with literature reports (Wang et al., 2010), so compound 7 was determined to be dihydrokaempferol.
Compound 8: Yellow needle-like crystals (purity > 95%), ESI-MS m/z: 271[M-H]-, molecular formula: C15H12O5. 1H-NMR (500 MHz, MeOD) δ: 7.95 (1H, d, J = 8.5 Hz, H-6’), 7.74 (1H, d, J = 15.5 Hz, H-β), 7.55 (1H, d, J = 15.0 Hz, H-α), 7.20 (1H, d, J = 2.0 Hz, H-2), 7.12 (1H, dd, J = 8.0, 2.0, H-6), 6.83 (1H, d, J = 8.0 Hz, H-5), 6.43 (1H, dd, J = 8.5, 2.0, H-5’), 6.29 (1H, d, J = 2.0 Hz, H-3’); 13C-NMR (125MHz, MeOD) δ: 167.6 (C-4’), 166.4 (C-2’), 150.0 (C-4), 146.9 (C-3), 146.1 (C-β), 133.3 (C-6’), 128.5 (C-1), 123.6 (C-6), 118.4 (C-α), 116.7 (C-5), 115.9 (C-2), 114.8 (C-1’), 109.2 (C-5’), 103.9(C-3’). The above data were in general agreement with the literature reports (Junior et al., 2008); therefore, compound 8 was determined to be butein.
Compound 9: Light-yellow powder (purity > 95%), ESI-MS m/z: 271[M+H]+, molecular formula: C16H14O4. 1H-NMR (500 MHz, MeOD) δ: 7.73 (1H, d, J = 9.0 Hz, H-5), 7.05 (2H, d, J =8.5 Hz, H-2’, 6’), 6.85 (2H, d, J = 8.5 Hz, H-3’, 5’), 6.50 (1H, dd, J = 9.0, 2.0 Hz, H-6), 6.30 (1H, d, J = 2.5 Hz, H-8), 4.14 (1 H, dd, J = 11.5, 7.5 Hz, H-2b), 3.06 (1H, dd, J = 13.5, 4.5 Hz, H-9a), 2.78 (1H, m, H-3), 2.63 (1H, dd, J = 13.5, 10.5 Hz, H-9b); 13C-NMR (125MHz, MeOD) δ: 195.1 (C-4), 166.6 (C-7), 165.4 (C-8α), 157.2 (C-4’), 131.2 (C-2’, 6’), 130.5 (C-1’), 130.3 (C-5), 116.4 (C-6), 114.7 (C-3’, 5’), 103.6 (C-4α, 8), 70.8 (C-2), 48.6 (C-3), 33.1 (C-9). The above data were in general agreement with the literature reports (Morales-Serna et al., 2010); therefore, compound 9 was determined to be 7,4’-homoisoflavanone.
Compound 10: Yellow powder (purity > 98%), ESI-MS m/z: 433[M+H]+, molecular formula: C21H20O10. 1H-NMR (500 MHz, MeOD) δ: 12.96 (1H, s, 5-OH), 10.40 (1H, s, 4’-OH), 7.97 (2H, J = 8.5 Hz, H-2’, 6’), 6.95 (2H, d, J = 8.5 Hz, H-3’, 5’), 6.87 (1H, s, H-3), 6.84 (1H, d, J = 2.5 Hz, H-8), 6.45 (1H, d, J = 2.0 Hz, H-6), 5.07 (1H, d, J = 8.0 Hz, H-1”); 13C-NMR (125 MHz, MeOD) δ: 182.5 (C-4), 164.7 (C-2), 163.4 (C-7), 161.8 (C-5), 161.6 (C-4’), 157.4 (C-9), 129.1 (C-2’, 6’), 121.5 (C-1’), 116.5 (C-3’, 5’), 105.8 (C-10), 103.6 (C-3), 100.4 (C-6), 100.0 (C-1”), 95.3 (C-8), 77.7 (C-5”), 76.9 (C-3”), 73.6 (C-2”), 70.0 (C-4”), 61.1 (C-6”). The above data were in general agreement with literature reports (Zhang et al., 2017); therefore, compound 10 was determined to be apigenin-7-O-β-D-glucopyranoside.
Compound 11: Yellow needle-like crystals (purity > 98%), ESI-MS m/z: 663[M+Na]+, molecular formula: C28H32O17. 1H-NMR (500 MHz, MeOD) δ: 7.94 (1H, d, J = 2.0 Hz, H-2’), 7.63 (1H, dd, J = 8.5, 2.0 Hz, H-6’), 6.91 (1H, d, J = 8.5 Hz, H-5’), 6.40 (1H, d, J = 2.0 Hz, H-8), 6.20 (1H, d, J = 2.0 Hz, H-6), 5.54 (1H, d, J = 7.0 Hz, H-1’’’), 4.84 (1H, d, J =8.0 Hz, H-1’’); 13C-NMR (125 MHz, MeOD) δ: 179.7 (C-4), 166.0 (C-7), 163.2 (C-5), 158.8 (C-9), 158.5 (C-2), 151.0 (C-3’), 148.6 (C-4’), 134.9 (C-3), 124.1 (C-6’), 123.2 (C-1’), 116.2 (C-5’), 114.5 (C-2’), 105.9 (C-10), 104.3 (C-1’’’), 101.4 (C-1’’), 99.9 (C-6), 94.8 (C-8), 79.9 (C-3’’), 78.4 (C-3’’’), 78.2 (C-5’’’), 77.9 (C-4’’’), 77.2 (C-5’’), 75.3 (C-2’’’), 74.8 (C-2’’), 71.4 (C-4’’), 70.1 (C-6’’’), 62.4 (C-6’’). The above data were in general agreement with literature reports (Wang et al., 2017); therefore, compound 11 was determined to be isorhamnetin-3-O-β-D-gentiobioside.
Compound 12: Yellow powder (purity > 98%), soluble in methanol, dark spot under UV254, non-fluorescent under UV365, 10% ethanol sulfate solution shows yellow color, HCI-Mg color reaction is red. TLC identification with rutin standard (trichloromethane–methanol, methanol–water, two kinds of unfolding agent system), in the same color development process, the Rf value is consistent; at the same time, mixed with rutin standard for spotting, TLC identification, showing a single spot; using the HPLC method for sample determination, this sample showed consistent retention time with rutin standard under the same chromatographic conditions, and the physico-chemical properties of rutin reported in the literature were basically consistent (Cheng et al., 2000); therefore, compound 12 was determined to be rutin.
Compound 13: White powder (purity > 95%), ESI-MS m/z: 473[M+H]+, molecular formula: C30H48O4. 1H-NMR (500 MHz, DMSO-d6) δ: 5.19 (1H, brs, H-12), 4.96 (1H, d, J = 12.0 Hz, H-24a), 3.81 (1H, dd, J = 11.5, 4.5 Hz, H-3), 3.83 (1H, d, J = 12.0 Hz, H-24b), 2.51 (1H, dd, J = 14.0, 7.0 Hz, H-18), 1.54 (3H, s, H-23), 1.49 (3H, s, H-30), 1.20 (3H, s, H-27), 0.95 (9H, s, H-25, 26, 28); 13C-NMR (125 MHz, DMSO-d6) δ: 180.0 (C-29), 144.3 (C-13), 122.6 (C-12), 79.1 (C-3), 63.5 (C-24), 55.8 (C-5), 47.6 (C-9), 45.9 (C-19), 42.6 (C-4, 20), 42.0 (C-14), 41.6 (C-18), 40.1 (C-8), 38.6 (C-1), 36.7 (C-10), 35.9 (C-21), 33.0 (C-7), 32.5 (C-17), 29.2 (C-22), 28.3 (C-2, 28), 27.6 (C-16), 26.8 (C-27), 26.0 (C-15), 23.7 (C-23), 23.3 (C-11), 19.7 (C-30), 19.1 (C-6), 16.8 (C-26), 16.0 (C-25). The above data were in general agreement with literature reports (Yao et al., 2007); therefore, compound 13 was determined to be azukisapogenol.
Compound 14: White powder (purity > 95%), ESI-MS m/z: 839[M+H]+, molecular formula: C44H70O15. 1H-NMR (500 MHz, MeOD) δ: 5.23 (1H, brs, H-12), 4.69 (1H, d, J = 7.5 Hz, H-1’), 4.52 (1H, d, J = 8.0 Hz, H-1’’), 4.12 (2H, d, J = 12.0 Hz, H-24), 3.42 (1H, t, J = 6.0 Hz, H-3); 13C-NMR (125 MHz, MeOD) δ: 182.8 (C-29), 172.3 (C-6’’), 145.5 (C-13), 124.0 (C-12), 105.0 (C-1’’), 104.4 (C-1’), 92.6 (C-3), 80.9 (C-2’’), 78.2 (C-3’), 78.1 (C-5’), 76.6 (C-5’’), 75.6 (C-3’’), 72.9 (C-2’, 4’’), 70.4 (C-4’), 64.3 (C-24), 62.0 (C-6’), 57.4 (C-5), 48.5 (C-9), 47.5 (C-19), 44.6 (C-20), 42.9 (C-4), 42.0 (C-14, 18), 41.1 (C-8), 39.7 (C-1), 37.5(C-10), 37.1 (C-21), 34.0 (C-7), 33.6 (C-17), 30.4 (C-22), 28.7 (C-28), 27.2 (C-16), 26.5 (C-2, 15, 27), 24.9 (C-23), 22.9 (C-11), 19.8 (C-30), 19.5 (C-6), 17.4 (C-26), 16.2 (C-25). The above data were in general agreement with literature reports (Rongqi et al., 1988); therefore, compound 14 was determined to be 3-O-[β-D-glucopyranosyl(1↓2)-β-D-glucuronopyranosyl] azukisapogenol methyl ester.
Compound 15: White powder (purity > 98%), ESI-MS m/z: 367[M-H]-, molecular formula: C24H48O2. 1H-NMR (500 MHz, CDCl3) δ: 2.35 (2H, t, J = 7.5 Hz, H-2), 1.63 (2H, m, H-3), 1.25 (38H, m, H-4 – 23), 0.88 (3H, t, J = 6.5 Hz, H-24); 13C-NMR (125 MHz, CDCl3) δ: 178.6 (C-1), 22.7–33.8 (C-2 – 23), 14.1 (C-24). The above data were in general agreement with literature reports (Li et al., 2009); therefore, compound 15 was determined to be tetracosanoic acid.
Compound 16: White powder (purity > 98%), insoluble in methanol and trichloromethane, no dark spots under UV254, no fluorescence under UV365, 10% sulfuric acid ethanol solution shows purple color. TLC identification with daucosterol standard (trichloromethane–methanol, methanol–water, two kinds of unfolding agent system), in the same color development process, the Rf value is consistent; at the same time, mixed with daucosterol standard for spotting, TLC identification, showing a single spot, and the physico-chemical properties of daucosterol reported in the literature were basically consistent (Wang et al., 2008); therefore, compound 16 was determined to be daucosterol.
Compound 17: White flaky solid (purity > 98%), easily soluble in methanol and trichloromethane, no dark spots under UV254, no fluorescence under UV365, 10% ethanol sulfate solution shows purple color. TLC identification with β-sitosterol standard (petroleum ether-ethyl acetate, trichloromethane–methanol, two kinds of unfolding agent system), in the same color development process, the Rf value is consistent; at the same time, mixed with β-sitosterol standard for spotting, TLC identification, showing a single spot, and the physico-chemical properties of β-sitosterol reported in the literature were basically consistent (Wang et al., 2008); therefore, compound 17 was determined to be β-sitosterol.
Core Inflammatory Targets of O. aciphylla
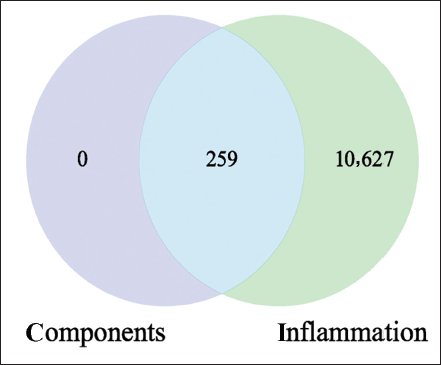
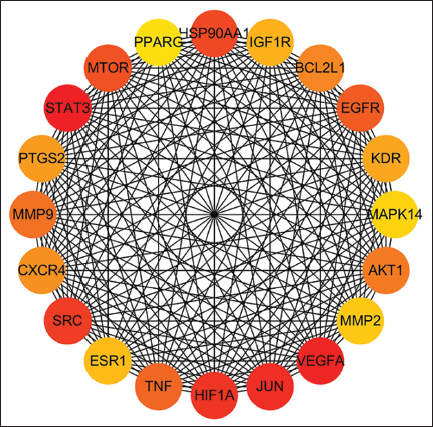
A total of 259 potential targets were predicted for the chemical constituents of O. aciphylla. By querying the database, 10,886 inflammation-related targets were obtained. The results indicate that all these 259 targets are potential anti-inflammatory targets of O. aciphylla (Figure 2). Based on the PPI network analysis of the potential anti-inflammatory targets of O. aciphylla, 20 core targets were selected (Figure 3).
Potential Anti-inflammatory Targets of O. aciphylla.
Core Targets in the PPI Network.
GO and KEGG Analysis
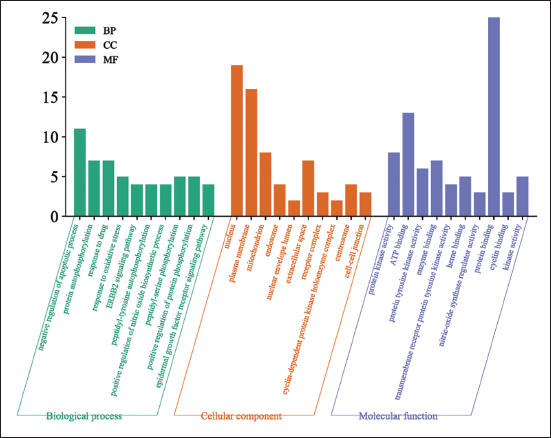
The GO analysis results are presented in Figure 4, indicating that the anti-inflammatory properties of O. aciphylla mainly involve biological processes such as negative regulation of the apoptotic process and protein autophosphorylation. Additionally, they impact cellular components such as the nucleus and mitochondrion, as well as molecular functions such as protein kinase activity and ATP binding.
Results of the GO Enrichment Analysis.
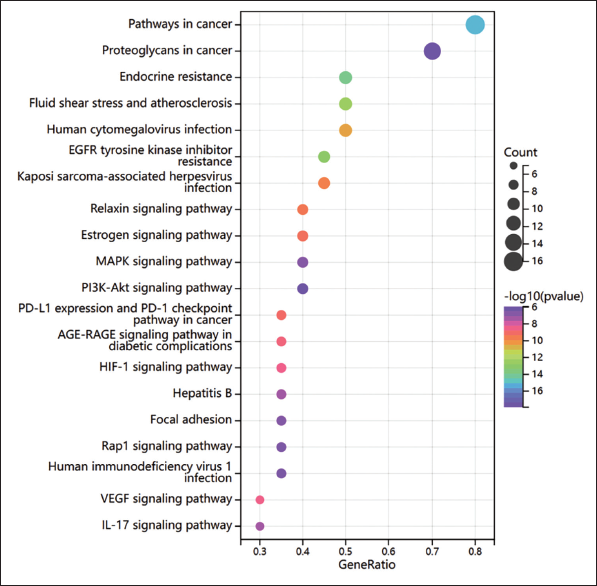
The KEGG analysis results, as shown in Table S1, and the visualization results, shown in Figure 5, reveal that the anti-inflammatory effects of O. aciphylla primarily involve signaling pathways such as EGFR, MAPK, and PI3K-Akt. According to earlier studies, TNF-α triggers the NF-κB and MAPK signaling pathways, which results in the production of adhesion molecules, the attraction and activation of neutrophils in the body, and the generation of inflammatory immune complexes that start the inflammatory process (Dostert et al., 2019).
Results of the KEGG Enrichment Analysis.
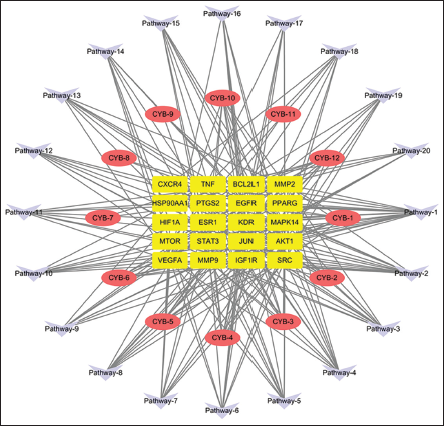
“Components–Targets–Pathways” Network
Cytoscape 3.7.2 software built the “Components–Targets–Pathways” network. The result is depicted in Figure 6. This network comprises a total of 52 nodes, consisting of 12 compounds, 20 targets, and 20 pathways, interconnected by 246 edges. Compounds such as kaempferol (degree value 10), apigenin (degree value 9), luteolin (degree value 9), eriodictyol (degree value 9), and naringenin (degree value 9) with higher degrees are potential core components responsible for the anti-inflammatory effects exhibited by O. aciphylla. Studies reveal that apigenin significantly inhibits miR-33 expression in a concentration- and time-dependent manner, which enhances the production of the ATP-binding cassette transporter A1 and cholesterol efflux. This, in turn, lowers lipid levels and mitigates inflammation (Ren et al., 2018). Additionally, it inhibits hypoxia-inducible factor 2α (HIF-2α)-induced expression of matrix metalloproteinase 3 (MMP3), IL-6, and COX-2 in chondrocytes, suppressing the release of aggrecan, thereby exerting protective effects on the extracellular matrix (Cho et al., 2019).
“Components–Targets–Pathways” Network Diagrams.
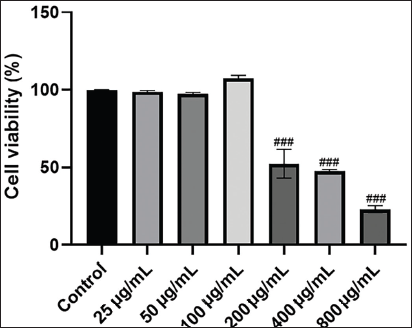
Impact of O. aciphylla on Cell Proliferation
As shown in Figure 7, the O. aciphylla extract did not show any cytotoxic effect on RAW 264.7 cells at doses up to 100 µg/mL. Therefore, these doses (25, 50, and 100 µg/mL) were selected for further experiments.
Impact of O. aciphylla on Cell Proliferation Assessed by CCK-8 Assay. The results are depicted as mean ± SD. n = 6. In comparison to the control group, ###p < 0.001.
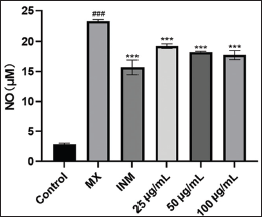
Influence of O. aciphylla on Cellular NO Production Levels
The impact of O. aciphylla on NO production in RAW 264.7 cells is depicted in Figure 8. LPS treatment caused a significant increase (p < 0.05) in cell NO production compared to the blank control group. In contrast to the model group, the positive drug, indomethacin, and O. aciphylla at low, medium, and high doses significantly reduced NO production (p < 0.05), displaying a dose-dependent relationship. NO, an essential inflammatory mediator expressed in large quantities during the inflammatory response, has a strong connection to inflammation (Cheng et al., 2021). Inducible nitric oxide synthase (iNOS) can induce NO production, and an excess of NO can lead to cell damage, tissue necrosis, and the promotion of the occurrence and development of inflammatory diseases (Sharawy & Lehmann, 2020). O. aciphylla can alleviate inflammation by inhibiting the generation of NO.
Effect of O. aciphylla on Cellular TNF-α and IL-6 Levels
The outcomes of the influence of O. aciphylla on TNF-α and IL-6 levels in RAW 264.7 cells are depicted in Figure 9. In comparison to the blank control group, cellular levels of TNF-α and IL-6 increased significantly following LPS treatment (p < 0.05). In contrast to the model group, both the positive drug indomethacin and O. aciphylla extract substantially decreased cellular TNF-α and IL-6 levels (p < 0.05), exhibiting a dose-dependent pattern. Notably, the decline in IL-6 levels was more pronounced than that of TNF-α.
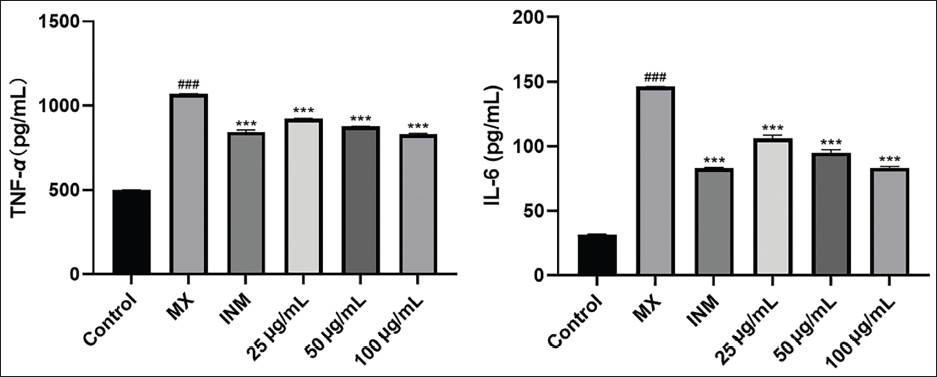
TNF-α, known for its potent pro-inflammatory effects, is one of the most robust inflammatory factors within the body (Sclavons et al., 2013), and it plays a coordinating and regulating role in inducing other cytokines involved in the inflammatory response, such as IL-6. Thus, the inhibition of TNF-α can effectively suppress the occurrence and development of inflammation (Moreira-Tabaka et al., 2012). Various cells, such as macrophages, endothelial cells, and some T lymphocytes, produce IL-6 (Sen et al., 2011), which is a cytokine that indicates the level of inflammation (Savale et al., 2009). IL-6 has multiple effects on the immune system, such as stimulating B lymphocytes to mature and differentiate, increasing the ability of NK cells to kill pathogens, activating bone marrow-derived cells, enhancing the inflammatory activity of neutrophils, and triggering inflammation (Hirano, 2010). Therefore, inhibiting the secretion of inflammatory factors by macrophages is a crucial step in alleviating the inflammatory response.
Conclusion
In this study, a comprehensive chemical analysis of O. aciphylla was conducted. From the 70% ethanol extract of O. aciphylla, a plant species in the Fabaceae family, we obtained and characterized 17 compounds. These compounds encompassed 12 flavonoid components, namely, apigenin, luteolin, kaempferol, isorhamnetin, naringenin, eriodictyol, dihydrokaempferol, butein, 7,4’-homoisoflavanone, apigenin-7-O-β-D-glucopyranoside, isorhamnetin-3-O-β-D-gentiobioside, and rutin. Additionally, two pentacyclic triterpenoid compounds, azukisapogenol and 3-O-[β-D-glucopyranosyl(1↓2)-β-D-glucuronopyranosyl] azukisapogenol methyl ester; one fatty acid compound, tetracosanoic acid; and two steroidal components, daucosterol and β-sitosterol, were identified. Notably, compounds 1 through 17 were isolated for the first time from O. aciphylla. Among these, compounds 5 to 9, 11, and 15 were also isolated for the first time within this genus.
Network pharmacology analysis indicates that O. aciphylla may exert its anti-inflammatory effects through active components such as kaempferol, apigenin, and luteolin, acting on targets such as AKT1, MAPK14, TNF, PTGS2, and EGFR, to regulate signaling pathways including MAPK, PI3K-Akt, VEGF, and HIF-1. Simultaneously, the ethanolic extract of O. aciphylla significantly inhibits the release of inflammatory factors, namely, TNF-α, IL-6, and the inflammatory mediator NO, in LPS-induced RAW264.7 cells, indicative of its anti-inflammatory properties. The network pharmacology results were consistent with the in vitro experiments, as some of the core targets (such as MAPK14, TNF, and PTGS2) and pathways (like the MAPK signaling pathway) were directly related to the inflammatory factors (NO, TNF-α, and IL-6) measured in the cell supernatant. The inflammatory response is worsened by the production of inflammatory mediators and factors, which are induced by the activation of inflammatory pathways. These findings suggest that O. aciphylla has a potential anti-inflammatory effect by modulating multiple targets and pathways.
In summary, this study provides the basis for exploring the anti-inflammatory potential and the chemical constituents of O. aciphylla. The results from network pharmacology provide a theoretical basis for understanding its anti-inflammatory mechanisms. Moreover, this study also provides some preliminary insights into the potential clinical applications of O. aciphylla extract as a novel anti-inflammatory agent. However, there are still many challenges and limitations that need to be addressed before the extract can be used in clinical practice. First, the molecular mechanisms of the anti-inflammatory effect of O. aciphylla and its isolated compounds are still unclear and need further investigation. Second, the safety and toxicity of O. aciphylla and its isolated compounds have not been assessed. Therefore, more studies are needed to investigate the pharmacokinetics, pharmacodynamics, and toxicology of O. aciphylla extract and its isolated compounds in animal models.
Abbreviations
IL-6: Interleukin-6; RAW 264.7: A murine macrophage cell line; LPS: Lipopolysaccharide; TNF-α: Tumor Necrosis Factor-alpha; NO: Nitric Oxide.
Authors’ Contributions
Hao Qian and Jiaqiong Liu contributed equally to this work. They performed the conceptualization, methodology, investigation, data curation, writing (original draft), and visualization. Xianqi Meng performed the software, validation, formal analysis, and writing, review, and editing. Xiaoqin Wang performed the supervision, project administration, funding acquisition, and writing (review and editing).
Footnotes
Acknowledgments
The authors would like to thank the National Natural Science Foundation of China (No. 81760686) for supporting this study.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China under the grant number 81760686.
Statement of Informed Consent and Ethical Approval
Not applicable as the study did not involve any animal or human subjects.
Supplementary Material
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
