Abstract
Background
Vinegar-quenched Testudinis Carapax et Plastrum (TCP) and vinegar-quenched Trionycis Carapax (TC) are made from the shells of Chinemys reevesii and Trionyx sinensis. However, some cheap substitutes from non-medicinal species such as Trachemys scripta, Mauremys sinensis, Chelydra serpentina, and Apalone ferox are often seen on the market, which exposes public health to high risk. Conventional methods are not applicable for the identification of these two products since their intrinsic constituents are largely destroyed after being highly processed.
Objectives
A new quantitative polymerase chain reaction (qPCR) method was developed to detect DNA components of Vinegar-quenched TCP and vinegar-quenched TC based on simple sequence repeat differential genes of animal-derived mitochondria.
Materials and Methods
Six species-specific primer reactions were designed and further validated for specificity, sensitivity, and repeatability. Finally, the developed method was used to assess the authenticity of their commercial products.
Results
The assay exhibited good specificity, and the limit of detection was above 1 × 102 copies/µL. There was good linearity in the concentration, ranging from 1 × 102 to 1 × 107 copies/µL. In addition, the method was validated through repeatability assessment (coefficient variation <1%). While qPCR was applied for the analysis of 36 batches of commercial products of these two tonics, it could effectively identify the authenticity of the products, and consequently, seven batches were fake or adulterated.
Conclusion
This newly proposed method is promising for the quality evaluation of highly processed animal-derived Chinese medicines, which will assist in ensuring safety and efficacy in clinical practice and protect fair trade in the market.
Keywords
Introduction
Trionycis Carapax (TC) is the carapace of Trionyx sinensis Wiegmann and Testudinis Carapax et Plastrum (TCP) is the carapace and plastron of Chinemys reevesii (Gray) (Chinese Pharmacopoeia Committee, 2020), both of which were consumed as important animal-derived medicinal materials in Traditional Chinese Medicine (TCM). They are usually subjected to a vinegar-quenched process, and these highly processed products are more widely administered in clinical practice to nourish Yin and subdue Yang. Vinegar-quenched TCP is often used to improve osteoporosis (Chen et al., 2016), alleviate gastric ulcers (Zhao et al., 2020), and enhance immunity, and vinegar-quenched TC is mainly used to treat liver cancer (Li et al., 2018), liver fibrosis (Sun et al., 2018), strengthen immunity (Chen et al., 2021), as well as inhibit angiotensin I converting enzyme (Liao et al., 2018). They play an important role in Chinese Patent Medicines, like some well-known and well-developed herbal medicines (Lu et al., 2023). However, due to the limitation of Chelonia resources, some fake products made from non-medicinal species, including Trachemys scripta, Mauremys sinensis, Chelydra serpentina, and Apalone ferox, were often seen on the market, which exposed public health to a high risk and caused unfair economic competition. Vinegar-quenched TC and vinegar-quenched TCP are obtained by sand-scalding their clean products until the surface is light yellow, followed by immediate quenching in vinegar (Figure 1). After this deep processing, the complete shape of the carapace is broken, and the external characteristics of the prepared pieces are mostly lost, which increases the difficulty of identification.
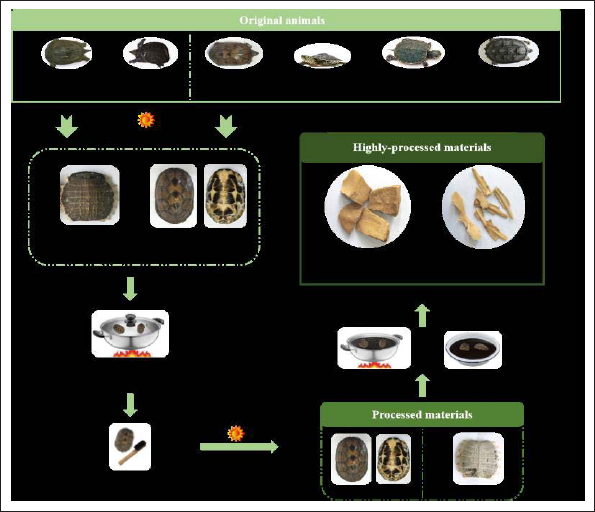
Conventional identification based on morphology is not suitable for the authenticity of highly processed products and their powder. Rather than proteins and small molecules, specific DNA fragments are present in most biological samples even after being processed and can be readily extracted from even a very small portion of tested products. Therefore, they are an ideal and powerful tool for the identification of original material existing in final products (Han et al., 2018; Kim & Kang, 2023; Liu et al., 2022; Lo & Shaw, 2018). In recent decades, polymerase chain reaction (PCR)-based methods to verify the origin of species have been considered the most preferred technology due to the favorable specificity and stability of DNA fragments (Geng, 2015). Previous studies (Yang et al., 2018) showed that only the authenticity of medicinal materials and raw products of TCP or TC could be identified by simple PCR; however, none of the species was detected in 10 out of 12 batches of commercial Vinegar-quenched TCP products. Authentication of the highly processed animal-derived Chinese medicines cannot be accomplished yet due to improper primers as well as insufficient limitation of detection as desired, which often leads to false negative results. Accordingly, the development of applicable and convenient approaches is crucial for the identification of highly processed Chelonia animal carapaces.
SYBR Green-based qPCR is widely used because of its convenience in designing the assay and its relatively low setup and running costs. SYBR Green dye intercalates into double-stranded DNA to monitor the amplification of the target gene specifically initiated by specific primers (Kang & Tanaka, 2018; Rio-Lavín et al., 2021). For instance, specific primer pairs and probes were designed based on the mitochondrial cytochrome b gene, followed by the establishment of a qPCR detection method to detect fresh donkey meat and different processed donkey meat products (Kim et al., 2020). In addition, a convenient qPCR method was well developed to identify 10 shrimp varieties using five gene-specific primers and distinguish mislabeled seafood or seafood adulterated with other varieties (Mondal & Mandal, 2020).
In this study, an SYBR Green-based qPCR assay was proposed for the identification of Vinegar-quenched TC and Vinegar-quenched TCP, which targeted the two ends of the simple sequence repeats (SSR) of six Chelonia species, and the specificity, sensitivity, and repeatability were determined. At last, the commercial products available have been quantitatively detected using the newly established method.
Materials and Methods
Materials
In this study, a total of 20 batches of six animal origins from Chelonia, including C. reevesii, M. sinensis, T. scripta, C. serpentina, T. sinensis, and A. ferox, were collected in recent years (Table S1), and all the species were verified by COI barcoding before any subsequent experiments. The protocols are described in the Chinese Pharmacopoeia (Ch.P., 2020 edition) (Figure 1). Ethical approval (No. 2017-03-03) was obtained from the Animal Ethics Committee of Jiangsu University prior to the experiments. The medicinal materials were purchased from aquaculture markets and breeding bases in different regions, but not wild animals from the environment.
In addition, 36 batches of commercial vinegar-quenched TC and vinegar-quenched TCP were purchased from TCM hospitals (Table S2).
Reagents
Proteinase K from Tritirachium album (Lot#020M8627) was purchased from Sigma-Aldrich (USA). DNA extraction phenol reagent (Lot#1027Z011) was purchased from Solarbio (China). BeyoFastTM SYBR Green Qpcr Mix (2×, High ROX) (Lot#030521220105) was purchased from Beyotime (China). PEASY®-Basic Seamless Cloning and Assembly Kit (Lot#P61026) was purchased from TransGen Biotech (China). Other reagents were all of analytical grade and supplied by Sinopharm Chemical Reagent Co. Ltd.
DNA Extraction
All samples were cleaned with 70% ethanol (EtOH) and pulverized after the removal of EtOH by air drying, then ground into a fine powder. Sodium dodecyl sulfate (SDS)-based protocols, as described by Yang et al. (2018), were used to extract DNA from each sample. In detail, 50 mg of the homogenized sample was mixed with 995 µL of extraction buffer (100 mM NaCl, 10 mM Tris–HCl, 25 mM ethylenediaminetetraacetic acid [EDTA], 0.5% SDS) and 5 µL proteinase K (20 mg/mL), and the mixture was incubated at 56°C for 3 h. After centrifugation at 12,000 rpm for 10 min, 900 µL of the supernatant was transferred to a new clear tube. An equal volume of phenol chloroform isoamyl alcohol solution and chloroform isoamyl alcohol solution were sequentially mixed with the supernatant for further purification. Then, the supernatant was precipitated by 850 µL of isopropanol and 85 µL of 5 M potassium acetate. The supernatant was removed after centrifugation, and the resulting DNA pellet was washed with 500 µL of 70% EtOH and finally reconstituted in 25 µL of Tris–EDTA (TE) buffer for subsequent experiments. SDS acts as an anionic surfactant to lyse cells at high temperatures (50°C–60°C) and denature proteins. Meanwhile, protease K was added to further degrade proteins and release nucleic acids from the sample matrix. The concentration and purity of the resulting DNA were determined by a NanoDrop1000 analyzer (Thermo Fisher Scientific, USA). The purified DNA was stored at −20°C for subsequent experiments.
Primers Design
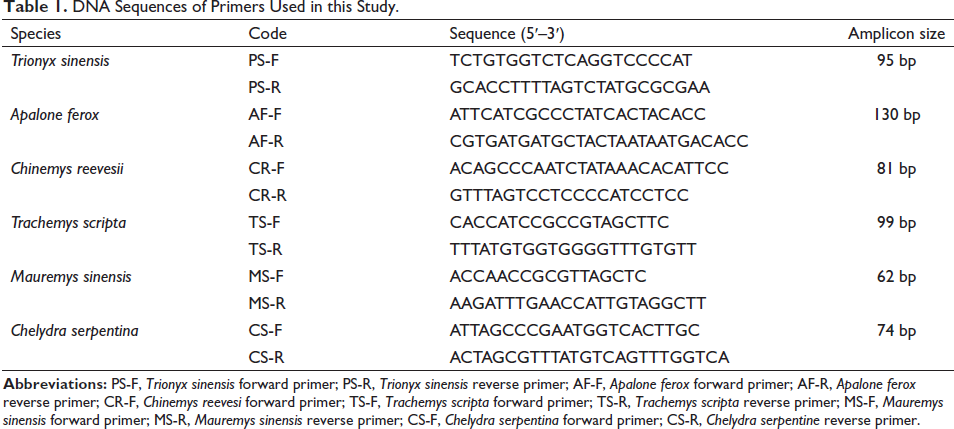
Complete mitochondrial genome sequences of six species, including C. reevesii, T. scripta, M. sinensis, C. serpentina, T. sinensis, and A. ferox (Accession No.: NC_006082.1, NC_011573.1, NC_016685.1, NC_011198.1, NC_006132.1, NC_014054.1) were individually imported into SSRHunter1.3 software to retrieve their respective SSR sequences, and primers were designed by Oligo 7 software using the SSR sequences (ca. 300 bp) as templates for each species. Then, primer sets were further evaluated by Oligo and online NCBI Primer-BLAST (Figure S1), and the resulting primer candidates (Table 1) were synthesized by Sangon Biotech (Shanghai) Co. Ltd.
Top 20 Most Active Countries Publishing Articles on CGRM, Based on Citations.

Real-time PCR
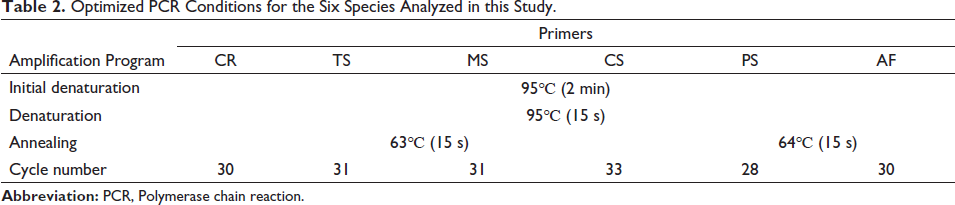
The DNA of vinegar-quenched TC and vinegar-quenched TCP was largely degraded after high processing including sand scalding and vinegar tempering, so the amount of template DNA in the samples has been much reduced. Accordingly, a more sensitive qPCR was adopted for the identification of six Chelonia species. In brief, the reaction was conducted in a total volume of 20 µL, containing 10 µL of SYBR Green Qpcr Mix, 2 µL of forward and reverse primers, 2 µL of DNA template (10 ng/µL), and 6 µL of deionized distilled water. At the cycling stage, initial denaturation was performed at 95°C for 2 min, followed by amplification for 35 cycles (95°C, 15 s; 64°C, 15 s). Then, the reaction system was kept at 95°C for 15 s prior to incubation at 60°C for 1 min. At the end of PCR, the temperature was increased to 95°C at a slope of +0.3°C/s, and maintained for 15 s. The annealing temperature and cycle number were optimized to establish the optimal amplification conditions (Table 2), and the specificity and sensitivity of the primer were investigated for method validation.
Top 20 Most Active Countries Publishing Articles on CGRM, Based on Citations.
Construction of Recombinant Standard Plasmid
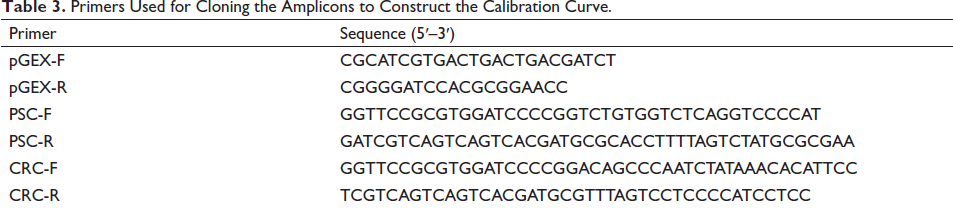
The DNA amplicons of C. reevesii or T. sinensis were subjected to PCR by the primer sets and then inserted into the amplified pGEX-4T-1 (Fenghui Biotechnology, China) plasmid by the pGEX primer set (Table 3) using the pEASY-Basic Seamless Cloning and Assembly Kit. Primer pGEX, PSC, and CRC were used to amplify the pGEX-4T-1 plasmid and the amplicons of C. reevesii and T. sinensis, respectively. The resulting recombinant vector was transformed into Escherichia coli Top10 competent cells (Sangon Biotech, China), and the correct cloning was confirmed by Sanger sequencing (Sangon Biotech, China). DNA was quantified by the NanoDrop1000 analyzer (Thermo Fisher Scientific, USA). Moreover, the standard plasmid copy number (N) was calculated by the following formula:
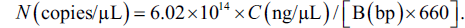
Top 20 Most Active Countries Publishing Articles on CGRM, Based on Citations.

C is the DNA concentration, and B is the sum of base numbers in the plasmid and in the insert.
Calibration Curve
The standard plasmid was diluted 10-fold serially and amplified under optimized qPCR conditions at a concentration ranging from 1 × 102 to 1 × 107 copies/µL. To construct a calibration curve with abscissa as the logarithm of plasmid copy numbers and ordinate as the Ct value.
Specificity Assay
The DNA (10 ng/µL) purified from self-made vinegar-quenched TC or vinegar-quenched TCP and their counterfeits was used as a template, and a qPCR assay was then carried out to verify the specificity under the optimized conditions. The DNA amplicons were sequenced by Sangon Biotech (Shanghai) Co. Ltd. to verify the accuracy of the specific amplification.
Sensitivity Assay
The DNA was extracted from six standard plasmids, and dissolved into TE buffer. Then, 10-fold dilution was performed six times, which resulted in the DNA solution ranging from 10 to 1 × 107 copies/µL. The sensitivity of the method was investigated under optimized reaction conditions.
Repeatability Assay
The DNA template from different species was used for the repeatability test to evaluate the variation within and between groups at the time of measurement. For intragroup repeatability, three replicate samples were tested in the same run. In addition, the DNA was tested in three independent runs for intergroup repeatability. The repeatability and stability of the method were measured by coefficient of variation (CV).
Application of Method to Commercial Products
Seventeen batches of commercial vinegar-quenched TCP and 19 batches of commercial vinegar-quenched TC were subjected to the newly developed qPCR assay for the purpose of species identification. Accordingly, genuine products were differentiated from counterfeits and adulterants.
Results
Calibration Curve
As shown in Figure S2, the calibration curve of PS was drawn as y = −3.090x + 35.08 (r2 = 0.9941, eff% = 110.6), and that of CR was drawn as y = −3.203x + 36.58 (r2 = 0.9956, eff% = 105.2), indicating a good linear relationship in the concentration ranging from 1 × 102 to 1 × 107 copies/µL. Moreover, negative controls should have a Ct > 38. The limits of quantification of the DNA template from C. reevesii and T. sinensis were both 1 × 102 copies/µL.
Specificity Assay
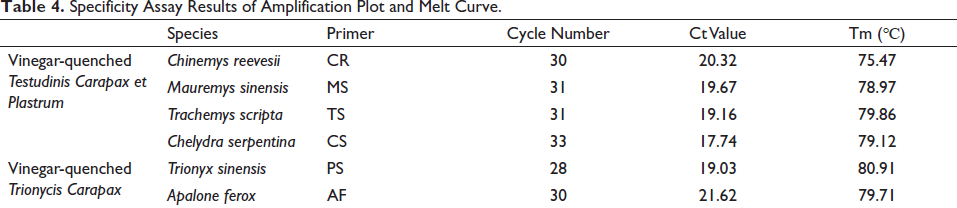
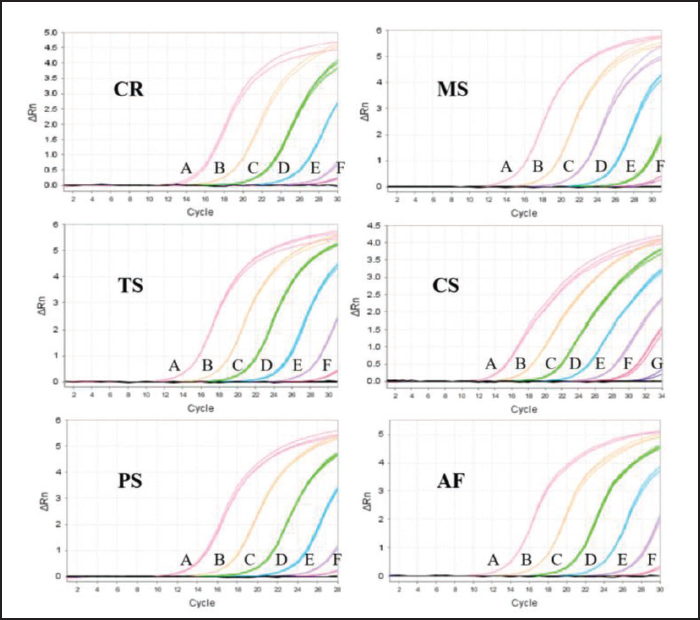
All the sequences of DNA amplicons and their homology rates against the available corresponding gene sequences are presented in Table S3. An investigation of the specificity of these primer sets was performed on a 10 ng/µL DNA template of self-made vinegar-quenched samples. It showed that strong fluorescent signals were achieved after qPCR of the DNA samples from their corresponding species, and no measurable amplification was observed for the negative control and other species DNA. On the other hand, a single peak was shown in all melt curves without a visual primer dimer (Figure 2). The cycle number and Ct value of the amplification plot and Tm of the melt curves obtained in the specificity assay were summarized in Table 4. In addition, the Tm of DNA amplicons was in line with the theoretical parameters, suggesting that the specific primers used were reliable and feasible. These results demonstrated that the method has good specificity and could be employed to identify the species of vinegar-quenched TC or vinegar-quenched TCP regardless of their destroyed morphological characteristics.
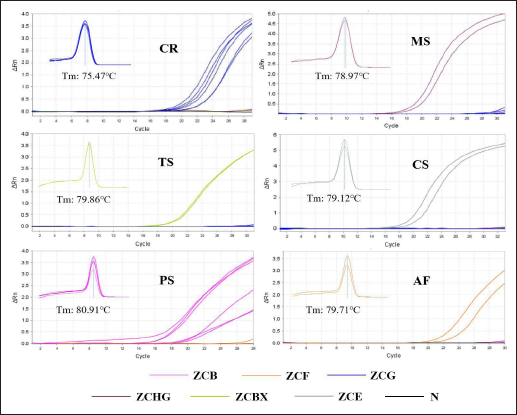
Specificity Assay Results of Amplification Plot and Melt Curve.
Sensitivity Assay
The sensitivity of each primer set against their corresponding species was investigated on the DNA template extracted from the self-made sample, and a series of concentrations ranging from 10 to 1 × 107 copies/µL were subjected to the qPCR amplification. As shown in Figure 3, a clear fluorescence signal was detected after the amplification against CR, TS, MS, AF, and PS when the DNA concentration of these samples was 1 × 102 copies/µL or above. Furthermore, CS was even able to generate a fluorescence signal against 10 copies/µL of the C. serpentina DNA template. The results demonstrated that the qPCR assay was highly sensitive and could be used to detect trace amounts of DNA from six Chelonia species.
Repeatability Assay
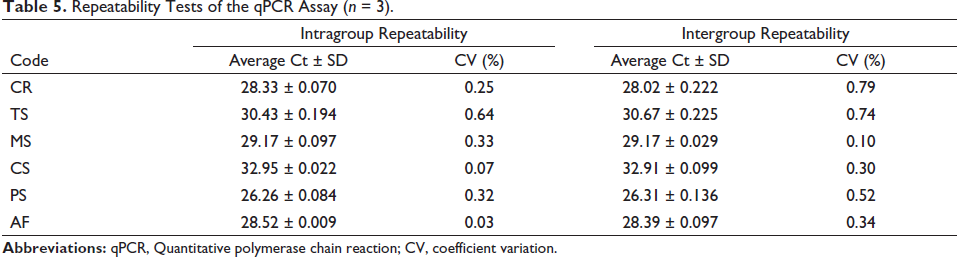
Consequently, the intragroup CV for six species ranged from 0.03% to 0.64%, and the intergroup CV varied from 0.10% to 0.79% (Table 5). Both their CVs were less than 1%, indicating the high accuracy and good repeatability of the qPCR method.
Repeatability Tests of the qPCR Assay (n = 3).
Application of Methods for Species Identification of Commercial Products
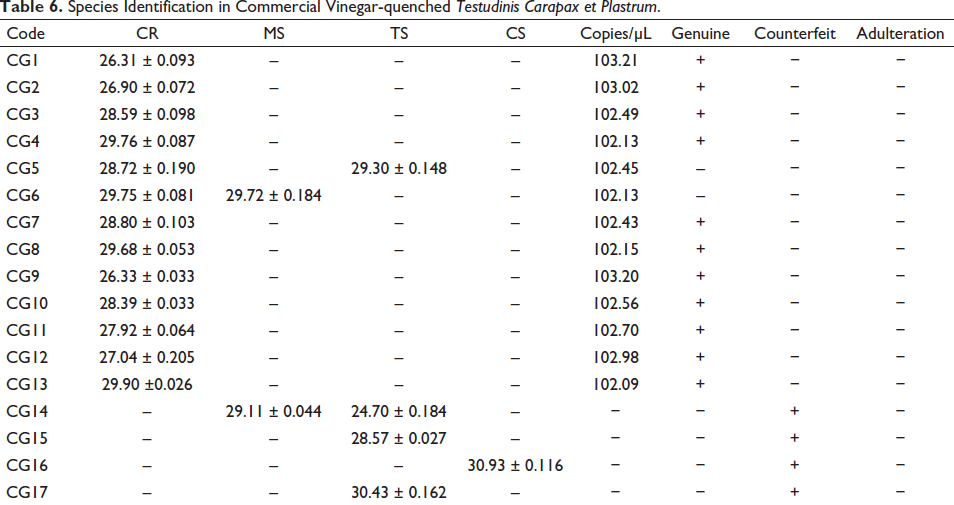
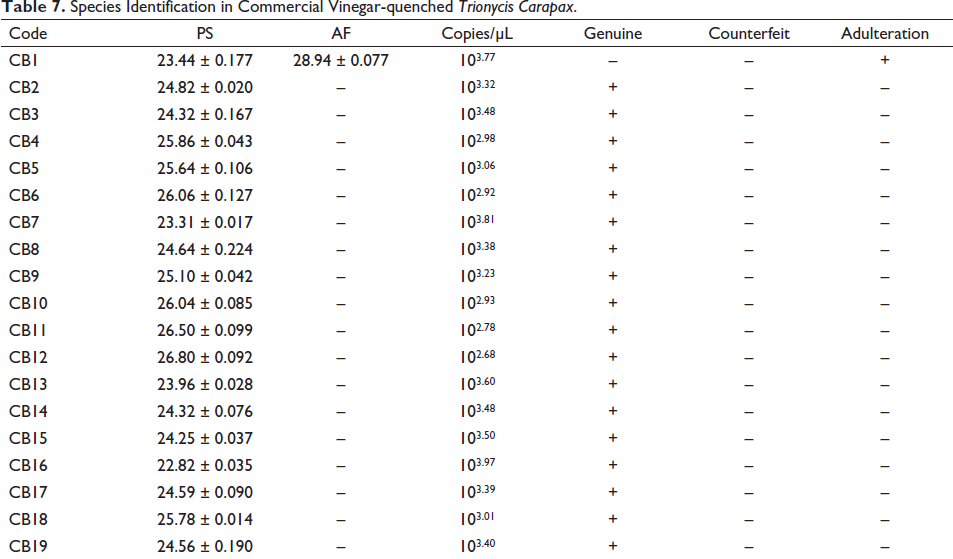
Eleven batches of vinegar-quenched TCP, including CG1–CG4 and CG7–CG13 have been amplified with the primer CR solely. The results suggested the presence of C. reevesii but not the other three fake species in these samples, and accordingly, they were authenticated to be genuine. However, another two batches (CG5 and CG6) have been amplified with the primers CR and TS or MS, indicating the presence of T. scripta in CG5 and M. sinensis in CG6 in addition to C. reevesii, and hence they were both authenticated to be adulterants. Another four batches (CG14–CG17) were not amplified with the primer CR; instead, they were amplified with the primer TS and MS or TS or CS. The results implied the lack of C. reevesii in these four products, and therefore they were authenticated to be counterfeit (Table 6). On the other hand, 18 out of 19 batches of Vinegar-quenched TC have been amplified uniquely with the primer PS. The results showed the presence of T. sinensis but not A. ferox in these samples, so they were authenticated to be genuine. However, one batch (CB1) was amplified with the primers PS and AF. It suggested the presence of A. ferox in addition to T. sinensis in this batch, and accordingly, CB1 was authenticated to be adulterant (Table 7).
Species Identification in Commercial Vinegar-quenched Testudinis Carapax et Plastrum.
Species Identification in Commercial Vinegar-quenched Trionycis Carapax.
Discussion
In this study, six species-specific primers were designed, and a new qPCR method was developed for the identification of vinegar-quenched TCP and vinegar-quenched TC. The calibration curves of PS and CR indicated a good linear relationship in the concentration ranging from 1 × 102 to 1 × 107 copies/µL (r2 > 0.99), which could realize the quantitative detection of the original species DNA in these two animal-derived tonics. In the optimized qPCR conditions, the specificity assay showed that the strong fluorescent signals were only achieved after the amplification of the DNA samples from their corresponding species, and high accuracy was verified by sequencing the DNA amplicon obtained from the specific amplification. Moreover, the detection limits of the DNA template from C. reevesii, T. scripta, M. sinensis, A. ferox, and T. sinensis were all 1 × 102 copies/µL, while C. serpentina was at 10 copies/µL, and the CV was all were less than 1% in the repeatability assay. These results demonstrated that the novel qPCR method had good performance in terms of specificity, sensitivity, and repeatability. Consequently, we effectively identified the authenticity of 36 batches of commercial products made from these two tonics by different manufacturers. Among 17 batches of commercial vinegar-quenched TCP, 64.7% (11/17) were considered genuine, 11.8% (2/17) as adulterant, and 23.5% (4/17) as counterfeit. In addition, 94.7% (18/19) were considered genuine and 5.3% (1/19) were adulterants among 19 batches of commercial vinegar-quenched TC. Compared with previous studies, the designed species-specific primers can usefully and accurately detect the species of these two highly processed products, and the qPCR method enables more rapid quantification and performs better for authentication purposes.
In summary, the results above indicated that there are some adulterants and counterfeits in the commercial vinegar-quenched TCP and vinegar-quenched TC, so it is necessary to establish a proper method to detect and quantify their DNA components. In our study, the species-specific primers could rapidly and reliably identify six Chelonia species of highly processed products. Real-time PCR has crucial advantages in the identification of Chinese medicines, but it usually requires highly specialized equipment and a long analysis cycle. Therefore, the combination of nucleic acid detection and other cutting-edge technologies will be further studied. For instance, to enhance the detection limitation and achieve absolute DNA quantitation, droplet digital PCR could be employed, which could be appropriate for the identification of highly processed Chinese medicine or Chinese Patent Medicine. Furthermore, to achieve efficient on-site identification of commercial Chinese medicine products, visual and more rapid detection methods could be developed, such as loop-mediated isothermal amplification (Jawla et al., 2021; Zhao et al., 2022).
Conclusion
In this study, we established a reliable and convenient qPCR method to detect the DNA from C. reevesii, T. sinensis, and their substitutes for the authentication of commercial Vinegar-quenched TC and Vinegar-quenched TCP products. It was well demonstrated for its stronger species specificity, higher sensitivity, and shorter time, in addition to its desired applicability to adulteration tests in commercial products. In particular, the method performed well in the assay of counterfeits of vinegar-quenched TC and vinegar-quenched TCP products. The newly proposed method could be employed as a routine detection means for the authenticity identification of these highly processed medicines, which will help ensure safety and efficacy in clinical practice and protect fair trade.
Footnotes
Abbreviations
AF-F: Apalone ferox forward primer; AF-R: Apalone Ferox reverse primer; BLASTn: basic local alignment search tool nucleotide; CR-F: Chinemys reevesi forward primer; PS-F: Trionyx sinensis forward primer; PS-R: Trionyx sinensis reverse primer; TCP: Testudinis Carapax et Plastrum; TCM: traditional Chinese medicine; TC: Trionycis Carapax.
Declaration of Conflicting Interests
The authors declare no conflict of interest.
Funding
This work was supported by the National Natural Science Foundation of China (82173937), the Science and Technology Planning Project of Zhenjiang City (SH2021062) and Jiangsu Pharmaceutical Association-Aosaikang Project (A202133).
Statement of Informed Consent and Ethical Approval
Necessary ethical clearances and informed consent was received and obtained respectively before initiating the study from all participants.
Summary
A new qPCR method was developed to detect DNA components of vinegar-quenched TCP and vinegar-quenched TC, based on simple sequence repeat differential genes of animal-derived mitochondria. Six species-specific primer reactions were designed and confirmed to have high specificity and repeatability. The limit of detection was above 1 × 102 copies/µL. There was a good linearity in the concentration ranging from 1 × 102 to 1 × 107 copies/µL. In particular, the newly method performed well in the assay of counterfeits of vinegar-quenched TC and vinegar-quenched TCP products.
