Abstract
Background
Trillium govanianum Wall. ex D. Don occurring at high altitudes in the Himalayas and known by the common name ‘Nag Chhatri,’ has high medicinal value. Its rhizome samples have substantial therapeutic potential and possess anti-cancer and anti-aging properties. Due to similar common names and morphology, there is an issue with adulteration and misidentification of actual rhizome samples with rhizomes of other species. So, there is a need to develop valid scientific methods to identify raw herbal samples of authentic ‘Nagchatri’ (T. govanianum).
Objectives
The present study focused on developing botanical, molecular, and chemical standards for identifying raw herbal samples of T. govanianum.
Materials and Methods
Samples were collected from four locations in the Northwestern Himalayas. Botanical characters were studied by stereomicroscope and compound microscope. Molecular identification was done by DNA barcoding and chemical identification by chemical fingerprinting (TLC and HPTLC), LC-MS profiling, and NMR studies.
Results
The botanical characters involved detailed morpho-anatomical characterization of herbal samples. The DNA barcoding exhibited a 100% identification match of generated barcode sequences with NCBI database sequences for all the selected markers. Chemical profiling revealed diosgenin, makisterone A, and 20-hydroxy ecdysone as major constituents in the rhizome sample, confirming the authenticity of the species.
Conclusion
The integrated authentication approach employing botanical standardization, DNA barcoding, and chemical profiling will be a promising tool for accurately identifying the genuine raw herbal material of T. govanianum.
Introduction
The need for raw herbal drugs in the herbal sector is rising globally. 1 Most crude plant drugs used for different herbal preparations are mainly procured from wild populations. 2 The trade of raw herbal drugs is generally in dried form, unregulated, with unknown background information, and often encountered with intentional or unintentional adulteration. 3 The adulteration problem is generally more prevalent with species of high market demand in the national and international herbal market. Among several herbal drug samples traded from the Indian Himalayan region, Trillium govanianum Wall. ex D. Don has recently gained popularity in the Indian herbal market due to its enormous medicinal potential. This genus is a member of the Melanthiaceae family, including 11 species from Asia and 38 from North America. 4 From India, T. govanianum and Trillium tschonoskii Maxim. have been reported from the Himalayas so far, of which the former is endemic to the Himalayas. 5 T. govanianum is a small (10–30 cm height) perennial medicinal herb growing in coniferous forests and moist, shady places. The plant is characterized by an aerial, erect, solitary, and unbranched stem from an underground rhizome; a stem bearing three apical leaves; and a single terminal stalked flower on a peduncle from the center leaves. T. govanianum, a native of the Himalayan region, grows at an altitude range of 2,700–4,000 m above sea level, with a geographical distribution from Sikkim, Darjeeling, and Himachal Pradesh to Kashmir. The species also grows in the adjacent countries, i.e., Nepal, Bhutan, China, Afghanistan, Pakistan, and Tibet. This species is known as Nag Chhatri, Himalayan Trillium, Chotasatwa, Sheethkar, Triflower, Matarzela, Birthroot, and Satwa. 6 T. govanianum has rich medicinal potential and is used in traditional Indian, Chinese, and Pakistani systems of medicine. 7
The rhizome of T. govanianum is rich in essential phytochemicals such as steroids, saponins, and several other compounds such as diosgenin and trillarin. 8 The compound from “rhizome” trillarin is believed to be used in preparing sex hormones, cortisone, and allied preparations in menstrual flow regulation and curing stomach problems. Another compound, ’diosgenin,’ has been reported with anti-cancer and anti-aging properties. Additionally, this substance is used as a substrate for synthesizing a number of steroidal drugs.9, 10 Compared to other medicinal plants, T. govanianum accumulates almost triple the amount of diosgenin. 8 Rhizomes also possess anti-bacterial, anti-inflammatory, radical scavenging, anti-cancer, analgesic, and β-glucuronidase inhibitory activities.11–13 Rhizomes are reported to cure dysentery, backache, menstrual and reproductive disorders, wound healing, inflammation, sepsis, skin boils, infections, kidney problems, skin allergy, cold, cough, and headache; they are also used against worms, cancer, and as antiseptic.6, 11, 14 Fresh rhizome samples have been reported to be edible in some regions of the Kashmir Himalayas of India. 8
T. govanianum is highly sought after for medicinal use in domestic and foreign markets because of its medicinal qualities. 15 From 2010 onwards, the species has seen an increase in illegal commercial trade in Himachal Pradesh and Uttarakhand. 16 The unsustainable overharvesting, illegal trade, and smuggling of rhizome samples have resulted in decreasing populations of T. govanianum from the Indian Himalayas (categorized as a threatened species), and the market prices have also risen to USD50–315 per kg.17, 18 Due to the high demand and low availability, there is much risk of adulteration. Other plants, such as Paris polyphylla Sm. of the same family Melanthiaceae, are also exploited in the name of Nagchhatri. 19 Being members of the same family, rhizome samples of P. polyphylla and T. govanianum showed morphological similarities. Another ayurvedic drug, “Swet Vacha,” was reported as adulterant drugs (rhizome samples of Cryptocoryne spiralis (Retz.) Fisch. Ex Wydler) with confusion to genuine drugs (rhizome samples of T. govanianum and P. polyphylla) when sold in the market. 20 To regulate illegal trade and adulteration of the genuine sample, there is a need to develop the proper identification standards for herbal drug samples. Additionally, accurate identification and standardization of raw herbal drugs are necessary for the efficacy and safety of plant-based medicines in the herbal drug sector. 21 Standardization of herbal drugs generally includes botanical studies, chemical characterization, and DNA fingerprinting. Reference standards aid the identification and authentication of the quality and purity of herbal medications based on botanical and chromatographic investigations as well as barcode sequences. 22 The qNMR method has also been known to identify chemical compounds in various processed herbal preparations. 23 Another novel method is DNA barcoding, which identifies species quickly, precisely, and automatically by employing short gene regions and species-specific markers. 24 Botanical identification (macroscopic and microscopic characters) provides an easy, fast, reliable, efficient, cost-effective, and authentic method of identifying raw herbal drug samples. 25
The present study is focused on the botanical characterization, development of chemical fingerprints, and DNA barcode sequences of T. govanianum to develop an authentic reference standard for the easy and quick identification of herbal samples. The botanical study involved the description of morpho-anatomical traits of herbal samples. Besides, DNA barcoding with primers matK, ITS, trnH-psbA, and rbcL was used for species identification. Chemical identification involved the development of thin layer chromatography (TLC), high performance thin layer chromatography (HPTLC), and liquid chromatography-mass spectroscopy (LC-MS) profiles, along with NMR quantification. Detailed macroscopic and microscopic characterization of raw drug samples supported by chromatographic fingerprints and barcode sequence data is more authentic for identifying rhizome samples of T. govanianum in fresh, dried, fragmented, or powdered form. The reference standards developed in the current study will be useful in the easy, quick, and cost-effective identification of crude herb samples traded in the herbal market.
Materials and Methods
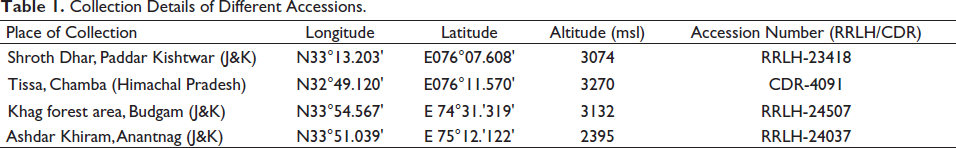
Plant samples were obtained by field visits in the Western Himalayas, India: from three different locations in Jammu and Kashmir and one in Himachal Pradesh. Herbarium sheets were prepared by regular herbarium procedure. 26 The herbarium specimens were submitted to the Regional Research Laboratory (RRLH) at CSIR-IIIM, Jammu, India. The various details of plant specimens, including collection details, accession numbers, etc., of individual specimens, are given in Table 1. Furthermore, the rhizome samples were oven-dried and submitted to the Crude Drug Repository (CDR) at CSIR-IIIM, Jammu. To carry out botanical identification studies, samples were placed in a fixative of formalin (5 ml) + acetic acid (5 ml) + 70% ethanol (90 ml). For DNA barcoding, leaf and rhizome samples were collected in airtight polybags containing silica gel. For chemical profiling, air-dried rhizome samples were used.
Collection Details of Different Accessions.
Botanical Characterization
The botanical study involved the study of the macroscopic, anatomical, and powder characteristics of rhizome, adventitious root, stem, and leaf samples. Morphological characters of rhizome samples, such as shape, size, color, surface, texture, node and internode pattern, adventitious roots, fracture, and transversely cut surface of the rhizome and root, were observed under a stereomicroscope (Model: Leica S9i). Anatomical studies of the samples were performed as per Kumar et al.’s 27 procedure, with some modifications. Freehand transverse sections (three sections of each plant part) of different accessions were obtained by the razor blade. The sections were sequentially dehydrated in ethanol of different grades, i.e., 30%, 50%, and 70% (10–15 min each), and then stained in safranin for 5–10 min. This was followed by their decolorization in 70% alcohol (5-10 min), staining in fast green (2 min), dehydration in 90% alcohol for 4–5 min, and finally in absolute alcohol (4–5 min). Canada balsam was finally used to mount the sections, which were photographed and analyzed using a compound microscope (LEICA, Germany: Model DM 750) with an attached camera (LEICA ICC50 E). In the powder study, organoleptic and microscopic characters (characteristic cell types and cell contents) of rhizome samples were studied. All micrometric measurements were done using LEICA LAS V 4.9.0 software.
Molecular Identification
Generation of DNA Barcode Sequences
The DNA purification kit was used to extract genomic DNA from leaf and rhizome samples by following the instructions from the manufacturer (Wizards Genomic DNA Purification Kit, Promega). The PCR thermocycler (Applied Biosystems, Veriti Thermal Cycler) was used to amplify selected barcode markers (ITS, matK, rbcL, and psbA-trnH) using specific primers. A PCR reaction with a total volume of 20 µL comprised of 1 µL template gDNA (60–80 ng/µL), primers (0.5 µL forwards and reverse primers, each 10 pmols/µL), 10 µL Master Mix (Promega GoTaq Colourless, Master Mix, 2X), and 8 µL nuclease-free water. The details of genomic DNA isolation and PCR amplification conditions are given in Table S1. The list of primers with sequence details is shown in Table S2. The PCR products were amplified and then examined on a 1.2% agarose gel along with a 100-bp ladder as size markers in tris acetate EDTA buffer in an electrophoretic unit. The PCR products were then purified by “Wizard SV Gel and PCR Clean-Up System” and used for bidirectional sequencing by Sanger sequencing.
Sequence Alignment and Phylogenetic Analysis
After Sanger sequencing, the chromatograms of all marker sequences were analyzed using Geospiza’s Finch TV version 1.4.0 and were assembled with required manual editing to get the final sequences. The final barcode sequences were used for phylogenetic relationship analysis and species identification of various accessions. The unknown query sequences were matched with the reference sequence database in the NCBI-BLAST (Basic Local Alignment Tool) tool from GenBank. 28 The top hit in the BLAST results corresponds to the species to be identified.
Some barcode sequences of species under study and adulterant species available in the NCBI database were retrieved in FASTA format for phylogenetic analysis. The barcode sequences generated from the current study and the sequences from the NCBI database were subjected to maximum likelihood (ML) phylogenetic tree analysis to study the relationship among different accessions. For phylogenetic analysis, the barcode sequences were aligned using MUSCLE, which is installed in MEGA7. The ML method was employed for constructing a phylogenetic tree. A 1000 bootstrap replicates were calculated for generating the trees. Genetic distance was also calculated among different accessions of T. govanianum.
Chemical Identification
The chemicals used in the LC-MS study, including water, acetic acid, formic acid, MS-grade acetonitrile, and other required chemicals/solvents used for extraction, were of analytical quality. All chemicals were purchased from Merck in Germany.
Extraction Procedure
The attached soil and debris from the collected plant material were removed by gentle brushing and then grated into small pieces. This grated plant material was evenly distributed on a plastic sheet in a clean, adequately ventilated room at 25°C, and the relative humidity was maintained at 65(±5)%. The grated and completely dried plant material is then fed into the mechanical grinder to convert it into a 1 mm particle size. This pulverization process leads to deeper solvent penetration as the surface area increases, resulting in effective extraction. To extract a range of secondary metabolites from the plant matrix, the polar organic solvent methanol was used for extraction.
The dry powder (10.0 g) was placed in methanol, subjected to sonication for 2 h, and then stored overnight. The extraction process was carried out three times (24 h each) using the same solvent to produce a colorless and transparent solvent. Soxhlet extraction was not employed because the long-term boiling of plant materials in the organic solvent may lead to the formation of unnecessary pseudometabolites known as artifacts, particularly in organic solvents, and the loss of bioactivity of potent secondary metabolites. The combined extract was then dried at 40°C in a vacuum evaporator after being filtered using Whatman filter paper (No. 2). The dried extract was then stored in an airtight jar at −20°C until further use in various analytical techniques such as TLC, HPTLC, and LC-MS analysis.
HPTLC Analysis
The methanol extract of T. govanianum (100 mg) was dissolved in 1 mL of methanol, centrifuged (10 min), and filtered through a 0.2-µm PTFE filter. An aliquot of 10 µL of the test solution was loaded using a CAMAG LINOMAT 5 instrument and Hamilton syringe.
The chamber was saturated with mobile phase for 15 min before chromatography. After that, the loaded plate was placed in a mobile, phase-filled development chamber and raised to 7 cm. The solvent from the plate was evaporated by air drying, and the plate was then placed in the “CAMAG REPROSTAR 3” photo-documentation chamber to take pictures at two different UV wavelengths (254 nm and 366 nm). After derivatization, the plate was once again fixed in scanner stage “CAMAG TLC SCANNER 3,” and then scanning was carried out under 500 nm of white light.
LC-MS Analysis
The samples were prepared in a volumetric flask with methanol:water (50:50, v/v). A disposable membrane filter (millipore) with a 0.25 m pore size was used to filter the crude extract, and then methanol was used to make appropriate dilutions. Both the working and stock solutions were kept at 4°C. A liquid chromatographic system “Agilent 1260, USA,” with a column heater, auto-sampler, and quaternary solvent delivery system, was used. A Merck Chromolith fast gradient RP18e column (100 × 4.6 mm) was used for chromatographic separation. The mobile phase consisted of [A] formic acid (0.1% aq.): ACN, (1.0%) v/v/v, and [B] Acetonitrile. Mobile phase was initiated with B-0%, B-20% (4.0 min), B-50% (15 min), B-50% (20 min), B-70% (25 min), B-70% (35 min), B-85% (38 min), B-85% (42 min), B-0% (45 min), and B-0% (47 min) for a gradient elution. A flow rate was noted at 0.5 mL/min, with a 1 µL volume of injection, and the temperature of the column at 30°C. A triple quad LC-MS system from Agilent, model number 6410B, was used to identify a hybrid triple quadrupole mass spectrometer equipped with Turbo V sources. The analyses used positive and negative modes (ESI) and electrospray ionization sources. Following operating parameters were used: a step size (0.1 amu), a scan range (110–1300 amu), the capillary voltage of 4,000V, an ion source temp of 300°C, nebulizer (50 psi), and gas flow of 13 L/min. Throughout every case, nitrogen was used. The Agilent Mass Hunter software (version B.04.00) was used to acquire and process the data.
Isolation and Characterization of Compounds 1–3
To isolate major metabolites, 1 kg of methanol-soaked, dried, powdered sample was kept overnight for 24 h and filtered the following day. The same extraction process was carried out three times. The combined extract was then filtered and dried at 40°C in a vacuum evaporator, yielding a crude extract (110 g). The 100-g methanol extract was mixed in water (300 mL) and extracted with petroleum ether (300 mL), and petroleum extract from the upper phase was removed (10.1 g). Further partitioning of the residue was done using chloroform (27.1 g) and n-butanol (52.5 g). A chloroform fraction (25.0 g) purified by the use of column chromatography (of silica gel, 100–200 mesh), eluted by a gradient of petroleum ether: ethyl acetate (100:0 to 0:100, 250 mL of each fraction), and concentrated, giving 18 combined fractions (Fr.- to Fr.18), based on the TLC profile. Fraction 9 (4 g) was chromatographed further with a gradient of petroleum ether-ethyl acetate (90:10 to 0:100, 50 mL of each fraction) to afford five subfractions (Fr. 9A-9E) over silica gel.
Further purification of fraction 9D (1 g) was done by column chromatography (230–400 mesh silica gel) and elution using 5% ethyl acetate in petroleum ether, giving 15 fractions of 50 mL volume each. A precipitate (white colored) was formed in the last three fractions, which on purification by washing with hexane (HPLC grade) yielded 180 mg of compound
Results
Botanical Characterization
Morphological Characters
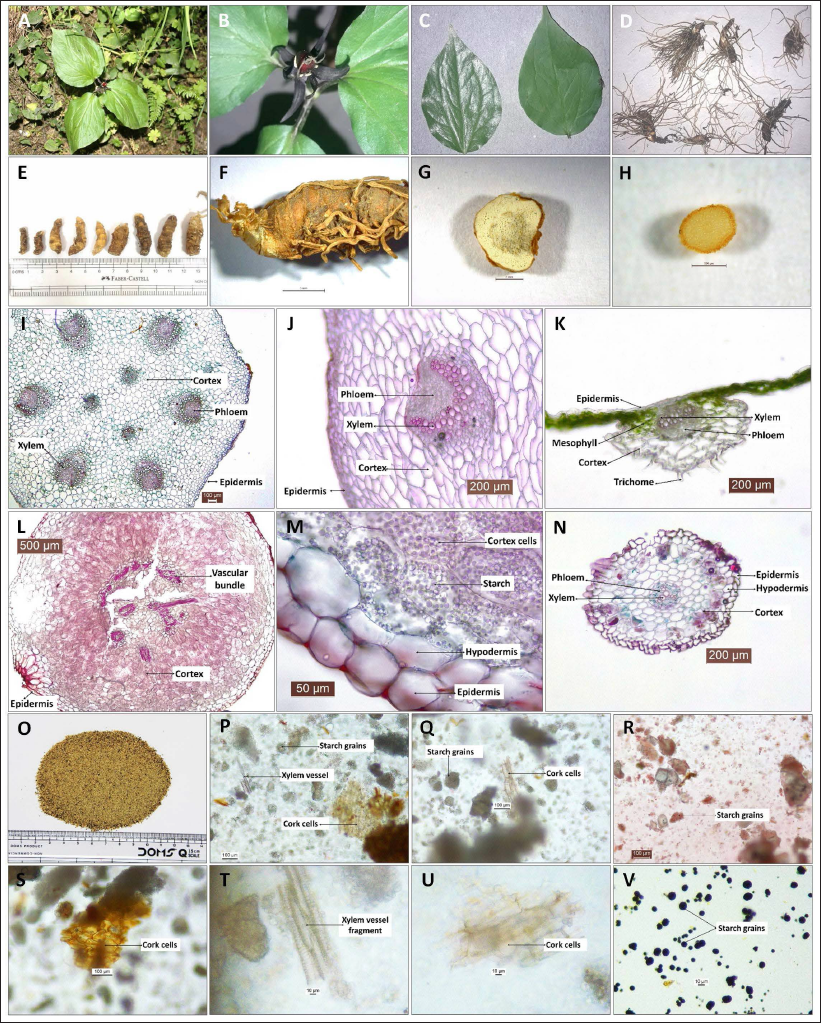
The plant is a small herb with three cordate leaves (Figure 1A). Flowers are purple-colored and present singly over the leaves on a small peduncle (Figure 1B). Leaves are green, petiolated, simple, ovate to cordate-shaped, margin entire, tip pointed, and lamina broad with three main veins arising from the leaf base and forming reticulate venation (a few small veins may also appear at the base with leaf maturity) (Figure 1C). Fresh rhizomes possess several threads like unbranched adventitious roots (Figure 1D), while the dried rhizome surface is observed with scars from fallen roots (Figure 1E). Dried rhizomes are light brown in color, stout, length (2.5–4 cm), thickness (0.5–1.3 cm), slightly curved or twisted, with nodes and internodes (nodes form ring-like markings on the surface) (Figure 1E and F). The transversally cut rhizome surface was circular to irregular in outline, the internal matrix was creamish colored, and there were light brown, ring-like, broken circular patches in the center (Figure 1G). The transversely cut surface of a fresh root appeared uniformly light green in color with less distinction between internal tissues (Figure 1H).
Macroscopic and Microscopic Details of Rhizome of Trillium govanianum. (A) Plant Habit, (B) Close View of the Flower, (C) Leaf Samples (Dorsal and Ventral View), (D, E) Raw Rhizome Sample, (F) Rhizome Surface View, (G) Rhizome Cut Surface View, (H) Adventitious Root Cut Surface View, (I) TS of the Stem, (J) TS of the Stem (vascular bundle enlarged view), (K) TS of the Leaf, (L) TS of the Rhizome, (M) TS of Rhizome (enlarged epidermal region), (N) TS of the Adventitious Root, (O) Rhizome Powder Sample, and (P–V) Common Microscopic Structures in Rhizome Powder Study.
Anatomical Characters
Transverse Section (TS) of Stem
The TS of the stem was circular, with a single-layered epidermis as the outermost layer having square-shaped cells, inner single-layered hypodermis, and a central mass of cortex comprised of the oval to deformed parenchyma cells with intercellular spaces. The cortex region comprised the outer ring of six oval-shaped vascular bundles and the central ring of three vascular bundles smaller than the outer ones (Figure 1I). Vascular bundles consisted of crescent-shaped xylem enclosing phloem in the cavity (Figure 1J).
TS of Leaf
TS of leaf showed basic dicot anatomical arrangement with midrib and lamina regions. Lamina is comprised of the single-layered upper epidermis and the inner, less distinct layers. Midrib consisted of the single-layered upper and lower epidermis with trichomes on the lower epidermis and followed by the inner cortex zone. The cortex zone was 2–3 layers thick on the adaxial side with small and compact cells, while the abaxial side had 3-4 layers of cortex cells with comparatively larger cell sizes. The cortex region surrounded inner oval-shaped vascular bundles, which consisted of xylem (adaxial side) and phloem (abaxial side) (Figure 1K).
TS of Rhizome
TS of rhizome was circular in outline (Figure 1L), with an outermost single-layered epidermis with rectangular to square-shaped cells, followed by 1–2 layer of large hypodermal cells and central multilayered cortex zone comprised of starch-containing parenchymatous cells. Epidermal cells were comparatively smaller than hypodermal cells (Figure 1M). Cortex cells are comprised of hexagonal to oval-shaped parenchymatous cells with intercellular spaces. The central region of the cortex was observed with a broken, circular ring of vascular bundles with less distinct vascular tissues (Figure 1L).
TS of Adventitious Root
The TS of the adventitious root was circular. The outermost epidermal layer was a thin, broken, irregular with deformed cells, followed by single-layered hypodermis, central broad cortex region (4–5 cells wide), and a central circular vascular region. The cortex is composed of polygonal-shaped, compactly packed parenchyma cells. The contents of a few outer cortex cells were found to be brownish. The vascular area includes tri-radiate xylem with larger lumen diameters and the xylem grooves occupied by phloem (Figure 1N).
Powder Study of Rhizome
The powder sample was creamish white with a few black fragments (Figure 1O), granular texture, characteristic aromatic odor, and tasted slightly sweet with a bitter tinge. Microscopic examination of the powder sample revealed cork cell fragments, parenchyma cell fragments, a few xylem vessel fragments, golden yellow fragments, a few raphid acicular crystals, and starch grains (Figure 1P–V).
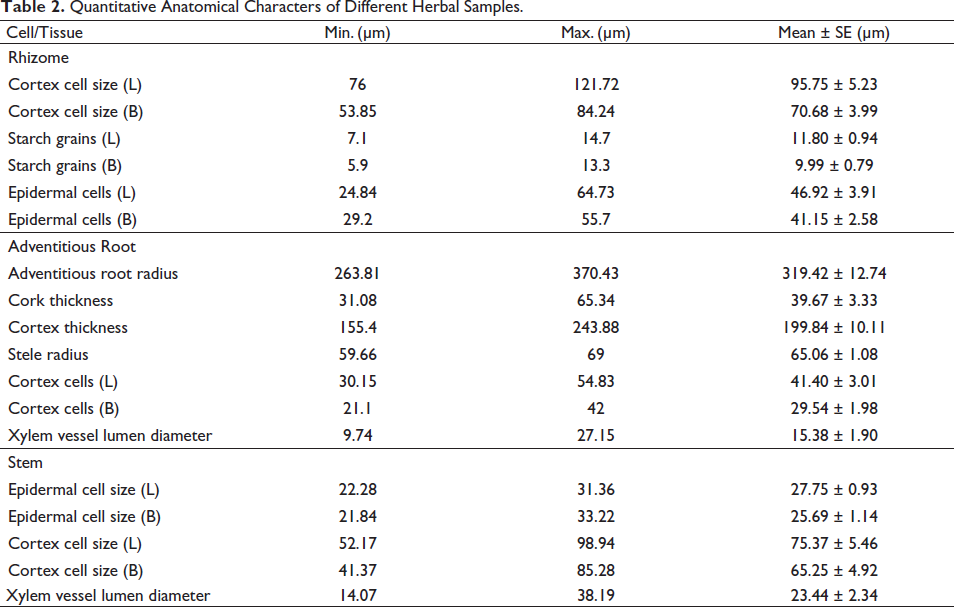
Quantitative anatomical characteristics of different herbal samples (rhizome and stem) are shown in Table 2.
Quantitative Anatomical Characters of Different Herbal Samples.
Molecular Identification (by DNA Barcoding)
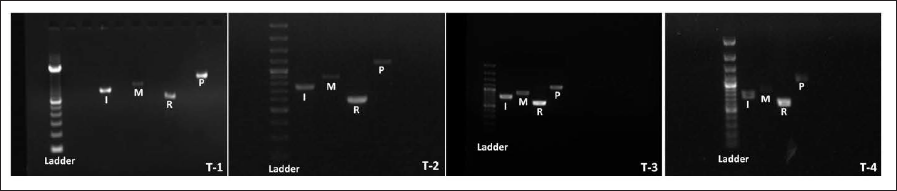
In the present study, four accessions of T. govanianum from Western Himalaya (Table 1) were analyzed for their authentication using four barcode markers, namely, ITS, matK, rbcL, and psbA-trnH. The electrophoretic gel photographs of different accessions are provided in Figure 2. In this analysis, the annealing temperature of 54°C for ITS and matK; and 52°C for rbcL was found to be optimum for developing a high-intensity molecular size band. The sequence length of ITS, matK, rbcL, and psbA-trnH varied from 649–720, 761–785, 525–573, and 976–1,060 bases, respectively, and were submitted in the database of NCBI (Table 3). The nucleotide frequencies in the ITS region were A=21.20%, T=21.20%, G=28.80%, and C=28.80%. For matK region, nucleotide frequencies were A=33.83%, T=33.83%, G=16.17%, and C=16.17%. For rbcL region, the nucleotide frequencies were found to be A=28.08%, T=28.08%, G=21.92%, and C=21.92%, and for psbA-trnH region, the nucleotide frequencies were found to be A=32.78%, T=32.78%, C=17.22%, and G=17.22%.
Accessions Numbers of Sequences Submitted in the NCBI Database from the Present Study.
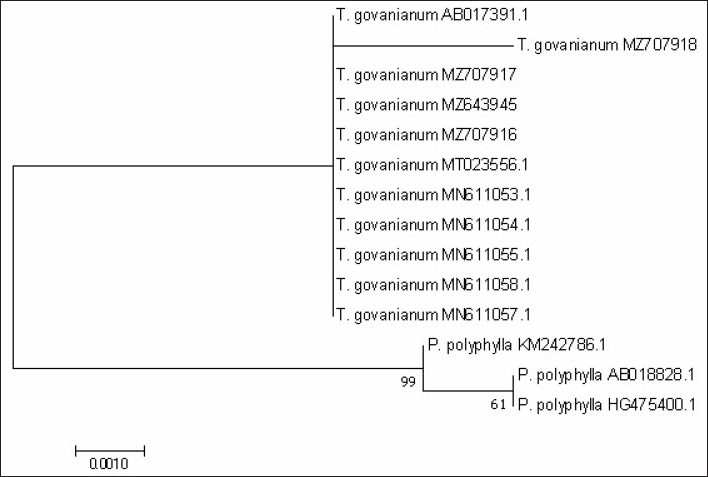
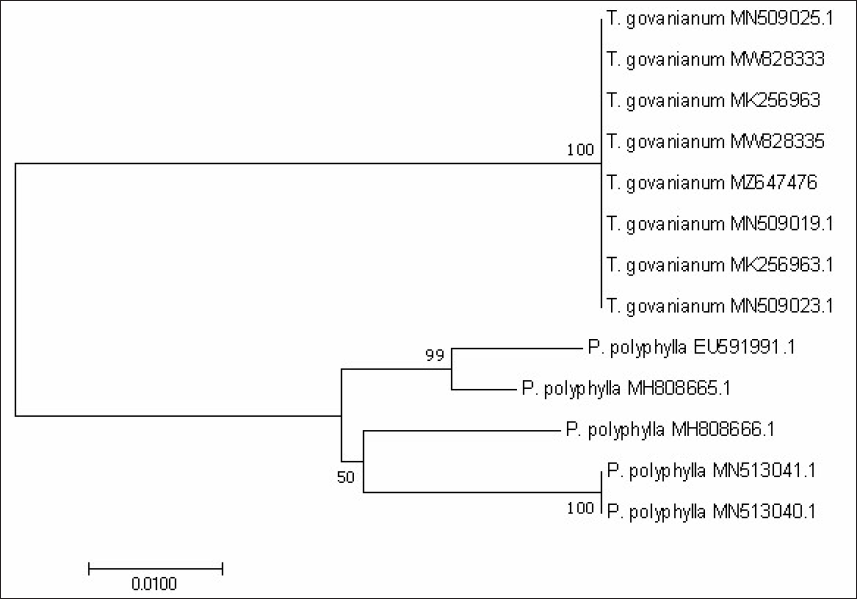
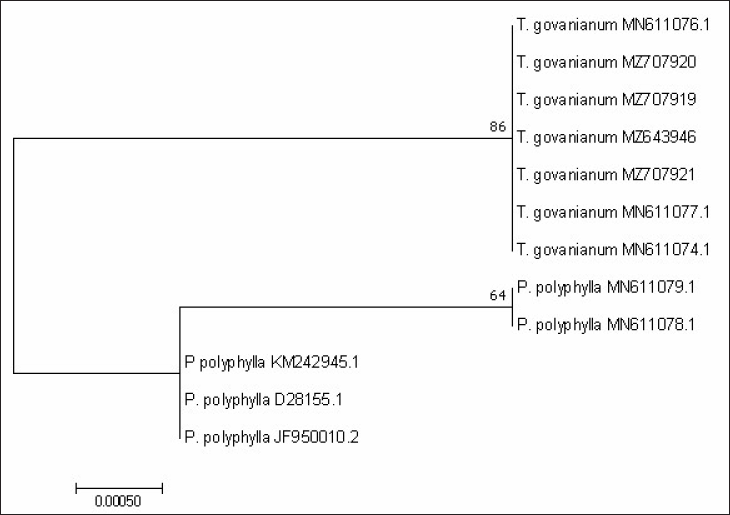
From BLAST analysis of the ITS, matK, and rbcL markers sequences, species were identified as T. govanianum. The psbA-trnH barcode sequences were unavailable in the NCBI database and were submitted for the first time. For the studied barcode regions, the maximum intraspecific genetic distance among the T. govanianum accessions is less than the smallest interspecific distance (Tables S3–S6), indicating the barcode gap because there are no overlaps in intra- and interspecific distances, thus discrimination of the two species from each other. It could also be found from the phylogenetic tree that the sequences of four markers of all accessions of T. govanianum could be grouped in the same clade, while the P. polyphylla formed a different clade (Figures 3–6), thereby discriminating the two species from each other. The accession numbers of sequences submitted in the database of NCBI from our study are given in Table 3. The accession numbers of sequences for phylogenetic analysis retrieved from the NCBI database are provided in Table S7.


Chemical Analysis
The TLC fingerprints were developed using methanol extracts of rhizome samples of T. govanianum. A literature review was done to develop the TLC profile of rhizome crude extract of T. govanianum, and for chromatographic separation of chemical compounds of crude extracts, varied solvent combinations were tried. Among the combinations tried, the most suitable solvent combination of Ethyl acetate/n-Hexane with solvent ratios ranging from (0.5:9.5 v/v to 2.0:8.0 v/v) was observed with comparatively clear bands. Of the varied solvent combinations, the solvent of Ethyl acetate:n-Hexane (1.5:8.5, v/v) was observed with the most desirable results. The TLC of crude extract of rhizome in selected solvent (total run of 7.0 cm) was observed with 6 bands 0.9, 2.8, 3.5, 4.0, 5.3, and 6.0 with respective retention factor (R
f
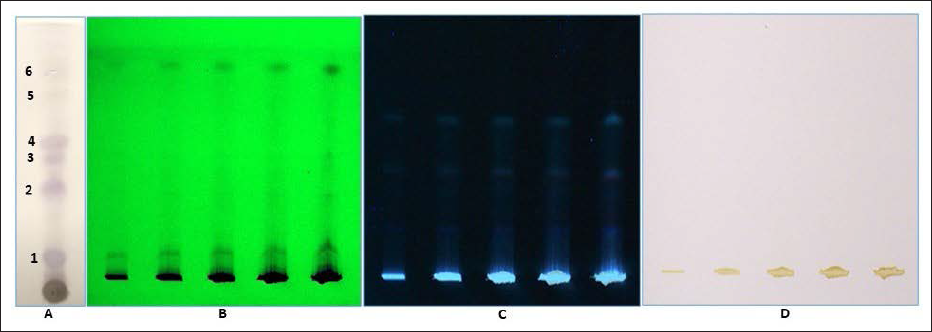
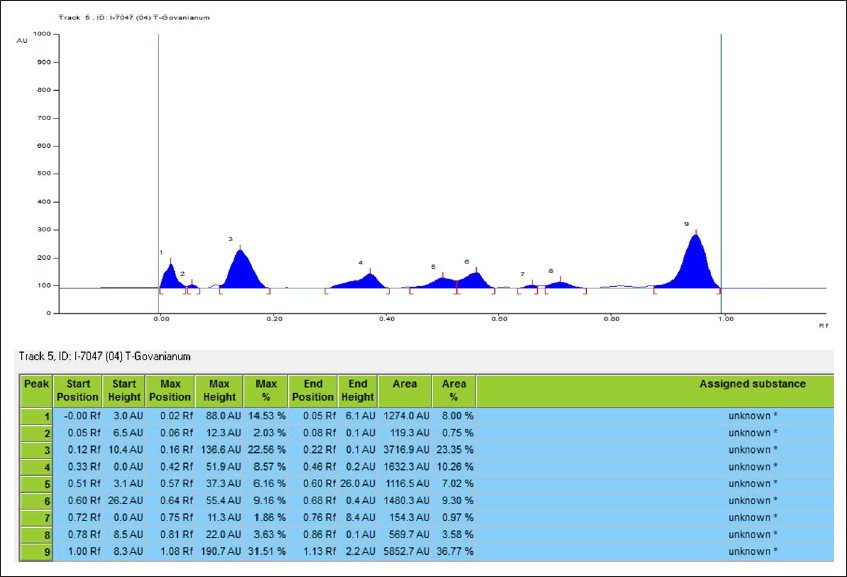
) values of 0.12, 0.40, 0.50, 0.57, 0.75, and 0.85, respectively. Furthermore, HPTLC was also used to identify raw rhizome samples of T. govanianum. The photo documentation for HPTLC fingerprinting of the studied species at different wavelengths is shown in Figure 7. Crude extract of rhizome samples examined in HPTLC under UV (254 and 366 nm), the chromatogram of sample solution exhibited three bands each while no bands under white light. Crude extract examined in HPTLC showed different bands of phytoconstituents at different retention factors (Rf), i.e., (0.00) to (1.00), respectively, which is shown in Figure 8. In the present study, LC-MS profiles were developed for rhizome samples to identify the characteristic compounds of species. As it was challenging to identify all the peaks from the LC-MS chromatogram, three compounds at retention time (Rt) 26.3, 17.25, and 13.1 were determined from their mass data and UV pattern as Diosgenin (
Photo Documentation of Chromatographic Fingerprinting Profile of Methanolic Crude Extract of Rhizome of T. govanianum: (A) TLC Fingerprint; (B-D) HPTLC Fingerprint of the Extracts of T. govanianum at 254nm (B), 366 nm (C), and White Light (D).
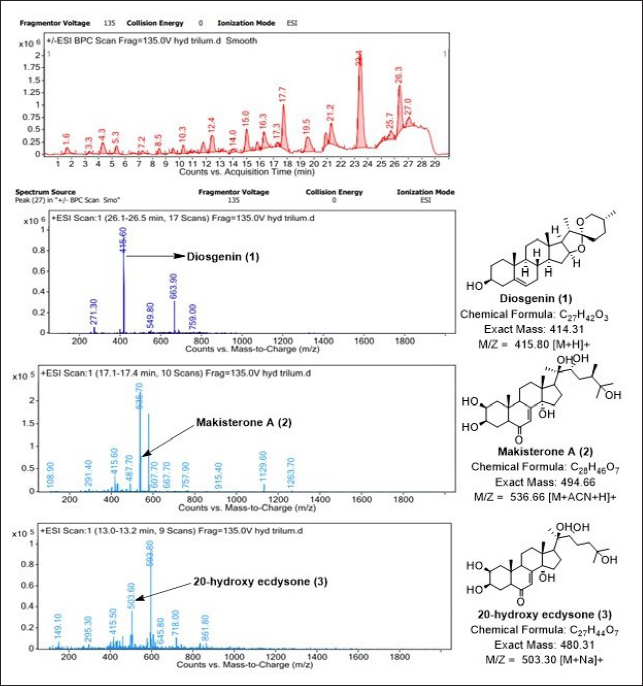
NMR spectroscopic data of marker compounds from the crude rhizome extract of T. govanianum.
White solid:
1H NMR (CD3OD, 400MHz): δ 5.81 (1H, d, J=2.6 Hz, H-7), 3.95 (1H, q, H-3α), 3.84 (1H, ddd, J=12.0, 4.0, 3.2 Hz, H-2α), 3.33 (1H, d, J=2.6 Hz, H-22), 3.15 (1H, ddd, J=11.2, 7.0, 2.6 Hz, H-9), 2.39 (1H, dd, J=9.5, 8.0 Hz, H-17), 2.38 (1H, dd, J=13.0, 4.5 Hz, H-5), 2.13 (1H, dt, J=13.0, 13.0, 4.8 Hz, H-12α), 1.99 (1H, H-15α), 1.95 (1H, H-16α), 1.88 (1H, ddd, J=12.8, 4.6, 2.3 Hz, H-12β), 1.81 (1H, H-11β), 1.79 (2H, H-1α/24a), 1.75 (1H, H-4α), 1.73 (1H, H-16β), 1.70 (1H, H-4β), 1.69 (1H, H-11α), 1.66 (1H, H-24a), 1.60 (1H, H-15β), 1.43 (2H, dd, J=13.3, 12.3 Hz, H-1β/24B), 1.28 (1H, dddd, J=13.0, 11.5, 11.0, 4.6 Hz, H-23b), 1.191 (3H, s, 26-CH3), 1.199 (3H, s, 21-CH3), 1.205 (3H, s, 27-CH3), 0.97 (3H, s, 19-CH3), 0.89 (3H, s, 18-CH3).
Discussion
T. govanianum has recently attracted immense attention as the species is used in the Indian and Chinese medicine systems to treat many health-related ailments. This species has been studied in several areas, including chemical composition, pharmacology, and population genetic diversity.10, 32, 33 Because of its limited distribution in the Himalayan region, a large-scale wild extraction has been observed in the study area due to rising market demand. As a result, adulteration or substitution of this plant with the rhizomes of P. polyphylla has become a problem related to quality control in the herbal drug industry. For regulated and controlled trade at national and international borders, the traded herbal samples are required with quick and authentic identification methods. When pharmaceutical plants are replaced or tainted with adulterant plants, it reduces the value of the original substance and, in some situations, makes it lethal. Therefore, there is a need to develop a quick and reliable technique for identifying raw herbal samples. Botanical standards can be helpful in the identification of such traded crude herbs: The present study included the study of vascular characteristics for rhizome, adventitious root, leaf, and stem. From the microscopic examination of the rhizome, observed characters, including cortex cells, golden yellow fragments, starch grains, a few raphid acicular crystals, and starch grains, were partially in line with the previous report. 7 Various qualitative and quantitative botanical characteristics described in the present study can play an important role in specimen identification when received in crude, fresh, or dry form.
The chemical characterization of rhizome samples also proved helpful in species identification. From TLC chromatograms, the Rf values provided essential information related to the phytochemical polarity and clues for their separation process. Owing to the automatization and high resolution of HPTLC methods, it has been extensively utilized to analyze and fingerprint plant extracts and standardization. The inclusion of HPTLC by Pharmacopeia indicates the importance of this technique for the authentication and validation of plant extracts. 34 The main feature of HPTLC is that it generates chromatographic fingerprints that can be compared with authentic markers in color and position. The TLC and HPTLC results were insufficient in determining the profile of crude extracts and chemical constituents. This may be because the extract was rich in non-active UV chemical constituents. Thus, to identify phytoconstituents in T. govanianum extract, additional studies of LC-MS profiling were done. LC-MS profiling is widely used to identify chemical constituents in natural product extracts.35, 36 The proton nuclear magnetic resonance ( 1 H NMR) quantification method has been reported as suitable for the characterization and quality control of herbal medicine. 37 The chemical compounds (Diosgenin, Makisterone A, and 20-hydroxy ecdysone) identified in the present study were major metabolites in T. govanianum and were comparable with published data.38, 39 In terms of biological activities, the chemical compounds identified in the present study have some useful biological activities. “Diosgenin” is a typical initial intermediate for the synthesis of steroidal compounds, oral contraceptives, and sex hormones and is also reported to be helpful for the treatment of disorders, including leukemia, inflammation, hypercholesterolemia, and cancer, 40 genotoxic and mutagenic effects, 41 anti-cancerous,42–45 anti-diabetic activities, 46 anti-inflammatory, antioxidant, and antiangiogenic activities. 47 Makisterone A reported attenuating cholestasis via increasing farnesoid X receptor (FXR) activation. 48 “Makisterone A” has bioinsecticidal potential in preventing stored-product insect pests. 49 The compound “20-hydroxy ecdysone” has anti-cancer properties, 50 regulating immune response51, 52 and cholinesterase inhibitory activities. 53
These identified compounds can potentially be used as a chemical marker in species identification in addition to molecular and botanical data. Also, DNA barcoding is known to make identification more practical in samples lacking diagnostic botanical characters or samples with difficulty differentiating based on the phytochemical-based approach.28, 54
The present study developed DNA barcode sequences to identify T. govanianum for selected four markers. The four markers distinguished T. govanianum from P. polyphylla in the present study. Among all the markers, the ITS and rbcL regions have been considered efficient for delimiting species in most plant species. 55 However, the matK region has not been considered ideal in many plant groups due to its low PCR success rate and sequencing. 56 Molecular identification of T. govanianum by developing DNA barcode sequences is available in some studies.57, 58 Osaloo and Kawano 57 observed less resolution in the rbcL marker-based tree in the phylogenetic analysis of Trillium and other allied species. Islam et al. 58 also observed some limitations in distinguishing T. govanianum from adulterant samples by DNA barcode-based method. Although the three DNA barcode markers used in this study were useful in distinguishing the two studied species, more genes are recommended to validate other Trillium species. According to Thakar and Sharma, 59 DNA barcode sequence data for T. govanianum is scanty in nucleotide databases. There is a need to develop DNA barcode sequence data for medicinal plants in trade to aid in easy and fast authentication.
Defining a species needs more data than mere DNA barcode sequences. 60 DNA-based identification is considered a potent herbal identification technique with limited use in identifying samples in a powdered or dried state or processed raw materials or extracts. 61 Molecular identification only by BLAST can have low identification if not incorporated by any other evidence. 62 Quality control and correct identification of herbal samples require an integrative approach using various methods such as botanical, chemical, and DNA-based studies. 22 Presently, there are several appropriate testing methods for identification studies of crude herbal drugs, but the microscopic examination is still considered a fundamental and economical way to identify source materials accurately and efficiently. 63 Anatomical study aids in the distinction of herbal samples, which are challenging to be distinguished by morphological characteristics. 64 The combined approach used in the current study will help in rapid, easy, and practical authentication for identifying rhizome samples of T. govanianum irrespective of physical form.
Conclusion
Considering the medicinal importance and the problem of adulteration and illegal trade of raw rhizome samples of T. govanianum, there is a need to characterize the primary herbal samples traded in the market. The present study involved a detailed characterization of the species for their botanical, molecular, and chemical profiles. The specific traits described in this study can be employed as standards to identify raw herbal samples sold in dried or fresh form. These detailed reference standards will be helpful to taxonomists, pharmaceutical industries, academicians, and researchers for taxonomic studies of raw herbal samples. The reference standards will help to regulate the illegal trade of herbal samples and also help to resolve the problems regarding adulteration and misidentification of crude herbal samples. The characteristic compounds identified from herbal samples have been used for various biological activities and can further be explored for their therapeutic potential, especially in anti-cancerous activities.
Summary
T. govanianum Wall. ex D. Don grows in the Himalayas, and the rhizomes were reported with high medicinal potential. There is the problem of adulteration and misidentification of rhizome samples with other species.
The herbal samples need correct identification by using authentic reference standards. The present study involved the development of botanical, molecular, and chemical standards.
The botanical study provided morpho-anatomical characterization of herbal samples. The DNA barcoding exhibited a 100% identification match of generated barcode sequences with NCBI database sequences using four markers. Chemical profiling revealed Diosgenin, Makisterone A, and 20-hydroxy ecdysone as major constituents in the rhizome sample, confirming the authenticity of the species and potentially being used for the development of pharmaceuticals.
The integrated authentication approach will be promising in accurately identifying the genuine raw herbal material.
Footnotes
Abbreviations
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship and/or publication of this article.
Statement of Informed Consent and Ethical Approval
Necessary ethical clearances and informed consent was received and obtained respectively before initiating the study from all participants.
