Abstract
Background
Accurate identification of Chinese herbal medicines is the basis for their research and utilization. Molecular identification can effectively differentiate original plants from counterfeit plants. The quality of genomic DNA is an important factor affecting molecular identification. However, the processing can lead to DNA degradation of the herbal medicines, which can make it difficult for their molecular identification.
Objectives
To establish a genomic DNA degradation model of Gentiana crassicaulis Radix to evaluate the effects of processing methods and storage times on genomic DNA integrity.
Materials and Methods
A genomic DNA degradation model of G. crassicaulis—the original plant source of the Chinese herbal medicine G. crassicaulis Radix—was established using a steam heating method. Genomic DNA integrity of G. crassicaulis Radix was evaluated using capillary electrochromatography (CEC) fingerprinting and polymerase chain reaction (PCR) of DNA barcoding markers following different processing and drying methods, including slicing (sliced roots), no slicing (whole roots), stoving, air drying, and sweating.
Results
CEC fingerprinting and DNA barcoding PCR effectively evaluated genomic DNA integrity. Compared to whole roots, sliced roots better helped maintain genomic DNA integrity. As the storage time increased, the integrity of the genomic DNA reduced; the integrity of the genomic DNA of sliced roots was greater than that of whole roots. Furthermore, the interactions between slicing and drying methods possibly reduced the genomic DNA integrity.
Conclusion
A genomic DNA degradation model and an evaluation system for herbal medicines were established. Our findings can help optimize the method for processing G. crassicaulis Radix and establish the traceability of genuine herbal medicines.
Key Message
The quality of genomic DNA is an important factor affecting molecular identification, which is essential to determine original plants. The results of our study can help develop a method for processing G. crassicaulis Radix and enable the traceability of genuine herbal medicines.
Keywords
Introduction
Ensuring the quality and efficiency of raw herbal medicines remains a major challenge in the use of Chinese medicines. Nevertheless, Chinese medicine practitioners have identified high-quality and highly effective herbal medicines produced under specific natural conditions and ecological environments in long-term practice (Wang et al., 2018). Accurate identification forms the basis for the effective use of herbal medicines. DNA barcoding has received considerable attention in pharmacognosy owing to its high accuracy and efficiency (Kumar et al., 2020; Li et al., 2015; Yu et al., 2018). The Pharmacopoeia of the People’s Republic of China has formulated a series of guiding principles for molecular identification (State Pharmacopoeia Commission of the People’s Republic of China, 2020a). However, out of 2711 traditional Chinese medicines and prescriptions in the Pharmacopoeia of the People’s Republic of China, only four—Fritillariae cirrhosae Bulbus, Dendrobii caulis, Zaocys, and Bungarus parvus—have been identified using molecular identification methods (State Pharmacopoeia Commission of the People’s Republic of China, 2020b).
In practice, the integrity of the genomic DNA template of herbal materials is an important factor that affects amplification efficiency and identification (Tian et al., 2020; Xin et al., 2021). Processing and storage have a significant impact on the integrity of the genomic DNA of herbal materials. During the processing of herbal medicines, processing methods and storage time can damage genomic DNA integrity, affecting the success rate of molecular marker amplification. This is one of the reasons molecular identification is not widely applied (Lo et al., 2015). Thus, an effective system for evaluating the integrity of the genomic DNA of herbal medicines is warranted. The effects of processing methods, such as boiling, steaming, and sweating, on the chemical composition and pharmacological activity of genuine herbal medicines have been extensively studied (Chen et al., 2021; Duan et al., 2022; Jin et al., 2018; Xing et al., 2021; Zhao et al., 2010b). Several drying and processing methods exist for raw Chinese medicines, but their effects on genomic DNA integrity remain unclear. Further studies are needed to ensure the efficiency of molecular identification methods and the safety of the resulting medication.
Gentiana crassicaulis Duthie ex Burk is the basic plant source of G. crassicaulis Radix, which is one of the sources of Gentiana macrophyllae Radix (Qinjiao) and is used to treat pain caused by rheumatoid arthritis, stroke hemiplegia, muscle cramps, joint pain, and damp-heat jaundice (State Pharmacopoeia Commission of the People’s Republic of China, 2020b). In recent years, given the huge demand for these resources and the lack of availability, this species has been listed on the List of National Key-Protected Wild Medicinal Species (Zhou et al., 2007). There is a long history of cultivation and production of G. crassicaulis in Lijiang, Yunnan Province, which is one of the main production areas for G. crassicaulis Radix (Ho, 1988). We previously studied the genetic background and quality of G. crassicaulis Radix obtained from Lijiang, and the quality was found to be exceptional (Kang et al., 2021; Lu et al., 2019; Ni et al., 2016; Ni et al., 2017; Ni & Zhao, 2018; Wang et al., 2009; Wang et al., 2017; Zhang et al., 2014; Zhao et al., 2010a; Zong et al., 2020).
In practice, G. crassicaulis Radix may be mixed with other species, particularly with their roots with characteristics such as “fleshy, usually several contiguous and twisted into a cylindric stout structure” (Ho, 1988). The Chinese Pharmacopoeia specifies a general processing method for G. macrophyllae Radix (Qinjiao), but there is no uniform standard (State Pharmacopoeia Commission of the People’s Republic of China, 2020c). Moreover, there are obvious differences in the processing methods employed in local and pharmaceutical factories.
The quality of genomic DNA is an important factor for the molecular identification of G. crassicaulis Radix under different processing methods and storage times. In the present study, we aimed to establish a genomic DNA degradation model of G. crassicaulis Radix to evaluate the effects of processing methods and storage times on genomic DNA integrity. We consider that the study provides a scientific basis for the production and evaluation of G. crassicaulis Radix. In addition, traditional Chinese medicine can be divided into different types according to medicinal parts. G. crassicaulis Radix is a radix herb and the method established in this study may provide a reference for DNA integrity evaluation of other radix materia medica.
Materials and Methods
Sampling Plot
The sampling plot was located in Lijiang, Yunnan Province, China, which is the conventional planting area of G. crassicaulis, with a longitude of 99° 29’ 12.8" E, a latitude of 27° 9’ 24" N, and an altitude of 2843 m.
Materials
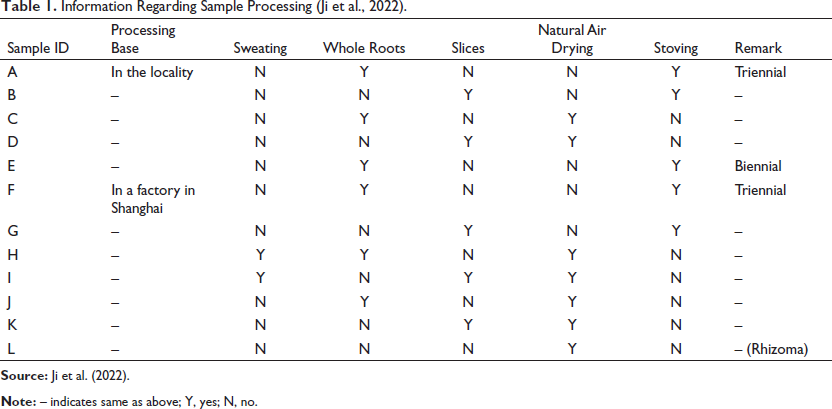
Three-year-old and two-year-old plants, which met the standard of harvesting, were selected from the sampling plot. The original plant was identified as G. crassicaulis Duthie ex Burk by taxonomic identification, and voucher specimens were deposited in the Herbarium of Medicinal Plants at the Shanghai University of Traditional Chinese Medicine. Original medicinal materials containing roots and rhizomes were collected from the sampling plot on 3 and 4 December 2020 and then processed locally (Diandong Group, Yulong County, Lijiang, Yunnan) and in a pharmaceutical factory (Shanghai Kangqiao Pharmacy, Shanghai), respectively. When the materials were subjected to local processing, the drying method consisted of stoving (45°C) and air drying (20°C). Each drying group was further divided into sliced and whole root groups, based on whether the sample was sliced. When the materials were processed in a pharmaceutical factory, the drying method consisted of stoving, air drying, and sweating, with the rhizomes removed. Sweating is a special treatment used for the drying of traditional Chinese medicinal herbs. Usually, herbs are slightly sun-dried, baked, or steamed and then stacked and wrapped to increase their temperature and make the internal water overflow (Chen et al., 2021). Each drying group was further divided into sliced and whole root groups, and the one with rhizomes removed was designated as a separate group. The samples were divided into 12 groups, labeled A–L (Figure 1, Table 1).

Information Regarding Sample Processing (Ji et al., 2022).
Construction of the DNA Degradation Model of G. crassicaulis Radix
The whole root of G. crassicaulis Radix air-dried in a pharmaceutical factory was selected as the sample for the degradation model. The degradation of genomic DNA was accelerated by steam heating and was stimulated during storage by controlling the heating time. Eight different heating times were used: 1, 2, 3, 4, 5, 6, 10, and 15 min.
Water vapor was used as the heating medium. The samples were placed into the steam drawer when the water in the steaming device was boiling, the lid was immediately placed to ensure that the steam temperature (100°C) was stable, and the time was recorded. After each heating time, the samples were removed from the steaming device and immediately placed in a pre-prepared ice container for 5 min for rapid cooling. The samples were then placed in a dryer at room temperature (20°C) for 4 h.
Extraction of Total Genomic DNA
The epidermis of the heated samples was removed to prevent exogenous DNA contamination of the surface. In each group, the same mass materials were cut according to three diameter specifications (above 8, 3–8, and below 3 mm), and 0.2 g was precisely taken after mixing. In the detection of different processing groups, 5 replicates were selected for each group. Quartz sand was added to account for one-tenth of the weight of the material and uniformly ground to a powder in the presence of liquid nitrogen. The appropriate grinding time was determined by evaluating the damage to the genomic DNA at 15, 30, 45, 60, 90, and 120 s. In total, 0.1 g of powder was precisely weighed, and genomic DNA was extracted using a modified cetyltrimethylthemmonium bromide (CTAB) method (Doyle & Doyle, 1987). The concentration of genomic DNA was determined using a microvolume UV-visible spectrophotometer by measuring the absorbance of the sample at the wavelengths 260 and 280 nm (Q5000, Quawell Technology Inc., San Jose, CA, USA).
Capillary Electrochromatography Fingerprinting
Capillary electrochromatography (CEC) fingerprinting was performed to determine the integrity of genomic DNA in each sample. The Qseq100 bioanalyzer (Taiwan BiOptic Inc., New Taipei City, China) was used for CEC fingerprinting, which was performed at Hangzhou Houze Biotechnology Co., Ltd. (Hangzhou, China).
The integrity of genomic DNA was analyzed by calculating the ratio of DNA fragments of different lengths. M1 and M2, markers of 20 and 1000 bp, respectively, were used as reference standards. Genomic DNA fragments were classified as short (20–1000 bp), medium (1000–20,000 bp), and long (over 20,000 bp) fragments. The integrity and degradation of the genomic DNA were assessed by comparing the proportion of fragments in the three aforementioned length ranges.
Analysis of DNA Barcoding Amplification Efficiency
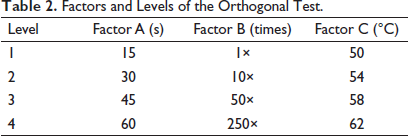
Candidate markers included rbcL, psbA-trnH, ycf15, trnL-F, and ITS. The optimal parameters were selected using an orthogonal test with three factors and four levels. The polymerase chain reaction (PCR) products were detected using agarose gel electrophoresis, and the band brightness in each group was quantified using Image Lab 3.0 (Bio-Rad Laboratories, Inc., Hercules, CA, USA). Orthogonal test parameters included the grinding times during genomic DNA extraction (factor A, s), DNA template concentration (1000 ng/µL), dilution times (factor B, times), and PCR annealing temperatures (factor C, °C) at four levels (Table 2). PCR conditions were established by evaluating the amplification results of the five markers (rbcL, psbA-trnH, ycf15, trnL-F, and ITS) and were amplified using the following primers: rbcL, 5′-ATGTCACCACAAACAGAAAC-3′, and 5′-TCGCATGTACCTGCAGTAGC-3′; psbA-trnH, 5′-GTTATGCATGAACGTAATGCTC-3′, and 5′-CGCGCATGGTGGATTCACAATCC-3′; ycf15, 5′-TACTTCCCAAACAG-ATCCTCCTAC-3′, and 5′-TTCGGCACTTAACC-ATTCAG-3′; trnL-F, 5′-GGTTCAAGTCCCTCTATCCCC-3′, and 5′-ATTTGAACTGGTGACACGAG-3′; and ITS, 5′-GGAAGTAGAAGTCGTAACAAGG-3′, and 5′-TCCTCCGCTTATTGATATGC-3′. The markers with the highest score in combination with the orthogonal test parameters were selected.
Factors and Levels of the Orthogonal Test.
The genomic DNA of the samples under different processing conditions was used as the template, and PCR was performed under the optimized conditions. Amplification results were assessed using agarose gel electrophoresis and quantified using Image Lab 3.0.
Assessment of Genomic DNA Degradation under Different Processing Methods
The genomic DNA of G. crassicaulis under different processing methods (Figure 1, Table 1) was extracted, and CEC fingerprinting and PCR amplification of DNA barcoding markers were performed. To identify the effect of specific variables associated with the processing methods (sliced or not) and drying methods (stoving, air drying, and sweating) on genomic DNA integrity, analysis of variance (ANOVA) was performed using SPSS 14.0 (SPSS Inc., Chicago, IL, USA).
Principal Component Analysis and Orthogonal Partial Least Squares Discriminant Analysis
Data regarding the proportion of short fragments in genomic DNA were used as variables for Principal Component Analysis (PCA) modeling and were divided into the sliced root and whole root groups for Orthogonal Partial Least Squares Discriminant Analysis (OPLS-DA). In OPLS-DA, a regression model was constructed between the multivariate data and a response variable (class information). R2X and R2Y represented the accuracy of the model. X and Y represented the interpretation rate of the matrices of the built model; Q2 represented the predictive ability of the model. The analyses were performed using SIMCA 14.1 (Sartorius A.G, Goettingen, Germany).
Assessment of Genomic DNA Integrity during Storage
To assess the effect of storage time on genomic DNA integrity, CEC fingerprinting and PCR amplification of DNA barcoding markers were performed on 12 groups of G. crassicaulis Radix samples with different storage times [0 (when processing was completed), 3, 8, and 12 months] in a pharmaceutical factory.
Results
Degradation of Genomic DNA under Different Heating Times
Assessment of the integrity of genomic DNA extracted at six different grinding times revealed that the genome gradually degraded as the grinding time increased. Considering the extraction efficiency and genomic DNA integrity, the grinding time for genomic DNA extraction was set to 45 s (Figure S1).
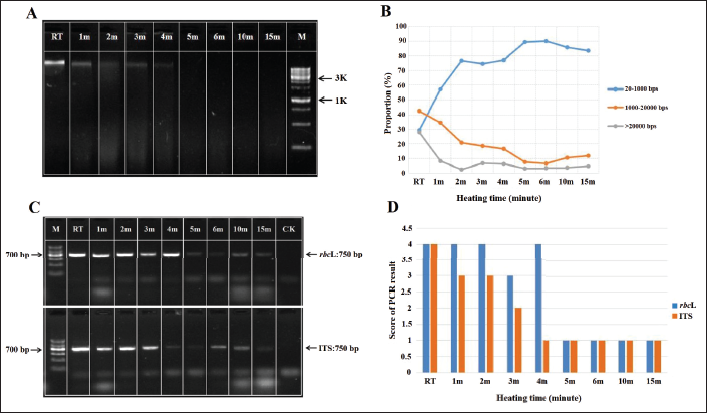
Assessment at eight different heating times revealed that the genomic DNA bands of the sample genome gradually blurred as heating time increased, indicating that the genome gradually degraded. After steaming for 4 min, the bands almost disappeared; therefore, this was considered the most sensitive period for genome degradation (Figure 2A).
Assessment of Genomic DNA Integrity using CEC
The integrity of the genomic DNA of each sample after eight different heating times was assessed using high-performance CEC. The electrophoretic pattern is shown in Figure S2, with the length distribution of the genomic DNA fragments displayed as peak plots. Genomic DNA was gradually fragmented into shorter fragments after heating.
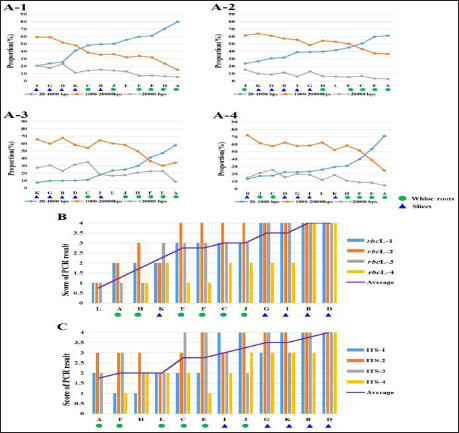
As the heating time increased, the proportion of short fragments also rapidly increased (Figure 2B). Upon heating for 2 min, over 70% of the DNA fragments became short. Upon heating for more than 4 min, the proportion of short fragments was stable at 80%–90%, accounting for most genomic DNA fragments. Correspondingly, the proportions of medium and long fragments gradually decreased as the heating time increased. The proportion of both fragments was less than 20% after 4 min. These results indicate that the genomic DNA integrity gradually decreased as the heating time increased, with most of the genomic DNA being degraded at 4 min. Thus, a genomic DNA degradation model was successfully constructed.
Assessment of Genomic DNA Integrity based on DNA Barcoding Amplification Efficiency
Based on the orthogonal test results (Table S1), rbcL and ITS were selected as markers. The conditions were as follows: the grinding time for DNA extraction was set to 45 s, the extracted genomic DNA (1000 ng/µL) was 10-fold diluted and used as the PCR template, and the annealing temperature for PCR was set to 54°C.
The effect of genomic DNA integrity on DNA barcoding amplification efficiency was analyzed using PCR. G. crassicaulis Radix was heated at eight different durations, and genomic DNA was extracted as the template. The rbcL and ITS fragments were amplified after the genomic DNA was homogenized. The genomic DNA of G. crassicaulis Radix extracted at eight different heating times was used as the template, and rbcL and ITS fragments were amplified after homogenization. The results were analyzed using agarose gel electrophoresis. The brightness of the bands was quantified. As evident by the results of electrophoresis (Figure 2C) and quantitative analysis (Figure 2D), the brightness of the PCR bands for rbcL and ITS gradually decreased as the heating time increased, indicating that the PCR efficiency gradually decreased. Upon heating for >4 min, the brightness of the bands decreased.
Assessment of Genomic DNA Degradation under Different Processing Methods
Sixty samples were collected from 12 groups of G. crassicaulis Radix using different processing methods (five samples per group). Genomic DNA was extracted from the samples under optimized conditions. Agarose gel electrophoresis revealed obvious differences in the brightness of the genomic DNA bands (A–L). The brightness of whole roots (A, H, and J) and rhizomes (L) was weaker than that of the sliced roots (Figure 3A).
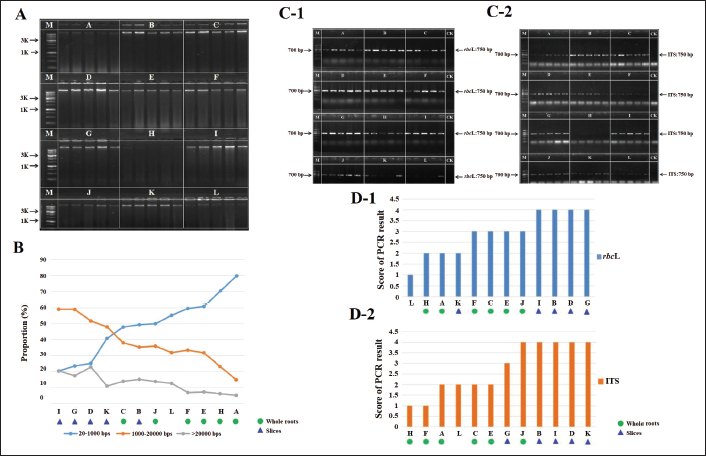
The genomic DNA of the samples was detected using CEC, and the results of each group were averaged and assessed according to the proportion of fragments of different lengths. The results revealed that the genomic DNA integrity of most sliced root groups was higher than that of whole root groups (Figure 3B). Two-factor ANOVA revealed that genomic DNA integrity was significantly affected by whether the herbs were sliced or not (p = 0.000). The effects of drying methods (stoving, air drying, and sweating) on genomic DNA integrity were not significant (p = 0.099). In addition, ANOVA revealed an interaction between slicing and drying methods in reducing genomic DNA integrity (p = 0.022).
rbcL and ITS fragments were amplified from the genomic DNA of the samples. The results revealed differences in the brightness of the bands among the different processed samples (A–L) (Figure 3C and D). The brightness of whole roots (A, H, and J) and rhizomes (L) was lower than that of sliced roots; this finding was consistent with that of the electrophoretic analysis of genomic DNA.
PCA and OPLS-DA5
Two principal components with large contribution rates were extracted, which explained 76.3% of the original data. The samples were analyzed using these two new variables, and a PCA score map was obtained (Figure 4A). The results revealed that sliced roots could be distinguished from whole roots.
(A) Principal Component Analysis Score Plots of the Samples. (B) Orthogonal Projections to Latent Structures Discriminant Analysis Score Plots of the Samples (80 mm × 10.14 mm).
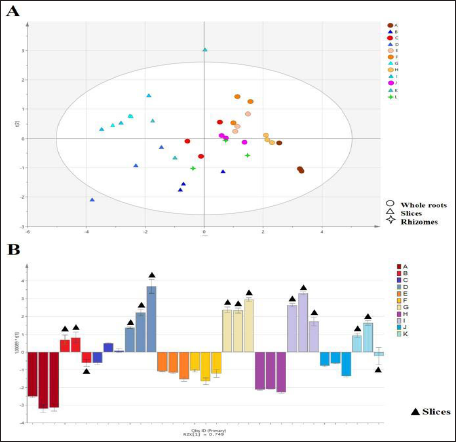
The corresponding values of OPLS-DA models were R2X = 74.9%, R2Y = 67.8%, and Q2 = 66.7%. The OPLS-DA score map (Figure 4B) revealed that sliced root and whole root groups were distributed in two different quadrants, indicating differences in genomic DNA integrity between the groups.
Assessment of Genomic DNA Integrity during Storage
The results revealed that the genomic DNA integrity of most sliced roots in the four samples with different storage times was higher than that of whole roots (Figure 5A, Table S2). Moreover, under different storage times, the brightness of the bands of most sliced roots was greater than that of the bands of whole roots (Figure 5B and C), consistent with the CEC trend. In summary, as the storage time increased, the degree of genome degradation of whole roots became greater than that of sliced roots.
Discussion
Genomic DNA integrity is inevitably reduced during the storage of herbal medicines. Genome fragmentation directly affects the efficiency of molecular identification. For example, a study on the integrity of the ginseng genome in ginseng decoction revealed that as the decoction time increased, the ginseng genome gradually fragmented, and the PCR detection efficiency decreased (Lo et al., 2015). Further assessment of the genomic DNA degradation in herbal medicines and the establishment of a comprehensive evaluation method are required (Pang et al., 2013; Pinya et al., 2022; Senapati et al., 2022; Xiong et al., 2018). In the present study, we used accelerated experimental methods with different heating times to assess the degradation trend in the G. crassicaulis Radix genome during storage. By heating herbal medicines, the degradation trend of genomic DNA under natural conditions could be simulated well. The genome of the herbal medicines showed obvious degradation after heating for approximately 4 min. CEC fingerprinting combined with PCR of DNA barcoding markers can effectively evaluate genomic degradation. The results revealed a degradation trend during storage. There are many Traditional Chinese medicines with root as a medical part, such as Platycodonis Radix (JieGeng), Salviae Miltiorrhizae Radix et Rhizoma (DanShen), Astragali Radix (HuangQi), and so on, which have similar morphological characteristics and texture. The method established in this study may provide a reference for genome integrity evaluation of other radix materia medica and has certain reference significance for their effective molecular identification.
Sweating is a special treatment used in the drying of traditional Chinese medicinal herbs, which has important effects on the quality of medicinal herbs (Yang et al., 2019). As to G. macrophyllae Radix (Qinjiao), there are two kinds of processing methods in the Chinese Pharmacopoeia, namely, sweating and without sweating. According to Chen et al. (2021), the contents of effective components (gentiopicroside and loganin acid) in the G. macrophyllae Radix samples dried after sweating treatment were significantly lower than those without sweating. In this study, the results showed that the DNA integrity was not affected significantly by sweating or not. Therefore, the necessity of sweating treatment in the processing of G. macrophyllae Radix may be insufficient.
On the basis of ensuring the content of pharmacologically active substances, processing methods should be reasonably selected to reduce damage to genomic DNA integrity and to ensure the accurate identification of G. crassicaulis Radix. The results revealed that drying fresh slices could effectively reduce the degradation ratio of the genome during processing. High water content in medicinal herbs will promote biochemical reactions during processing methods and may accelerate DNA degradation (Faller et al., 2019; Yan et al., 2016). Compared with whole roots, the sliced freshly roots could be dried faster after slicing and were in line with the water content requirement in the Chinese Pharmacopoeia. Therefore, the DNA degradation of whole roots is greater than that of sliced roots.
Previous chemical experiments have revealed that the stability of secondary metabolites is increased when the roots are sliced before drying. There were obvious differences between sliced roots and whole roots, although the content of index components both met the Pharmacopoeia requirements (Ji et al., 2022; Xu et al., 2018). The results were consistent with the results of DNA degradation analysis in this study, that is, the sliced roots were superior to the whole roots, and the drying method had little effect. Considering the content of active substances and genomic DNA integrity, it is recommended that herbal medicines be sliced before drying for processing. Drying methods could be selected according to the actual conditions.
Conclusion
In this study, a genomic DNA degradation model and an evaluation system for herbal medicines were preliminarily established to evaluate the genomic DNA integrity during storage and processing. The sampling time needs to be extended to further analyze the fitting of the actual situation with the model. The findings of this study can provide the basis for optimizing the method for processing G. crassicaulis Radix and establishing the traceability of genuine herbal medicines. Furthermore, our results may help facilitate the accurate molecular identification, quality evaluation, and drug safety assessment of Chinese herbal medicines.
Footnotes
Availability of Data and Materials
The datasets supporting the conclusions of this study are included within the article and its additional files. All materials are available from the corresponding author upon request.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Statement of Ethical Approval and Informed Consent
Not applicable.
Funding
This work was supported by the National Natural Science Foundation of China [grant numbers 82073959 and 81173654] and the Traditional Chinese Medicine research project of the Shanghai Municipal Health Commission [grant number 2022QN030].
