Abstract
Objective
To compare, in a primary human papillomavirus screening setting, two different validated human papillomavirus tests, considering their analytical and clinical screening performances.
Methods
In Tuscany, a human papillomavirus screening program was implemented in 2013. Hybrid capture 2 (Qiagen) was used for testing until May 2016, when it was replaced by the cobas® 4800 human papillomavirus test (Cobas; Roche). We evaluated the performance of Hybrid capture 2 and Cobas on: the same screening population in two different periods (before and after changing to Cobas); the same Hybrid capture 2-positive consecutive samples. Discordant samples (Hybrid capture 2-positive/Cobas negative) were typed on the L1 gene (reverse line blot, AB Analitica) and E6/E7 genes (BD Onclarity assay).
Results
In the considered time period (n = 37,775), human papillomavirus positivity was 9.8% and 7.4%, respectively, for Hybrid capture 2 and Cobas (p < 0.0001). At immediate colposcopy, the cervical intraepithelial neoplasia, grade 2 positive predictive value was, respectively, 23.8% and 34% (p < 0.001). At one-year recall, human papillomavirus persistence was, respectively, 40.6% and 62.2% (p < 0.0001). Of Hybrid capture 2-positive re-tested samples (n = 620), 32.4% were Cobas negative. Of discordant samples typed on L1, 7% were positive for the 12 high-risk human papillomavirus. Of the samples found to be negative for the 12 high-risk human papillomavirus types on L1, 14.5% were positive on E6/E7 typing. Among the discordant samples, the only two cervical intraepithelial neoplasia (CIN) grade 3 lesions were non-high-risk human papillomavirus positive on both L1 and E6/E7 typing.
Conclusion
At baseline, Hybrid capture 2 showed greater human papillomavirus positivity and a lower CIN2+ positive predictive value than Cobas, which was more specific than Hybrid capture 2 in detection of high-risk human papillomavirus: 80% of discordant samples were confirmed as high-risk human papillomavirus negative. This higher analytical specificity determined the non-identification of two CIN3 lesions.
Keywords
Introduction
Cervical cancer screening programmes have been running for many years in several developed countries, and cervical cancer incidence and mortality are in constant decline in these countries. Initially, cervical cancer screening was completely based on cytology (Pap test). The development of molecular technologies fed research in this field, leading to the discovery that DNA of human papillomavirus (HPV), a very common sexually transmitted virus, is present in 99.7% of cervical cancers 1 and that persistent infection with specific types of HPV, called oncogenic or high-risk (HR) types, is the necessary, but non-sufficient cause of cervical cancer. 2
During the last 15–20 years, several randomised controlled trials (RCTs) have been conducted throughout Europe to compare cytology-based screening to molecular HPV DNA test-based screening,3–6 with the aim of establishing the effectiveness and feasibility of screening based on an HR-HPV test. From these RCTs, it emerged that the HR-HPV test, which until then was used mainly for the triage of borderline cytological cases (ASC-US: atypical squamous cells of undetermined significance, or L-SIL: low-grade squamous intraepithelial lesion), was more sensitive, but less specific, than cytology in detecting precancerous lesions (CIN: cervical intraepithelial neoplasia) in women over the age of 30–35. Furthermore, it was demonstrated that HPV-based screening provides 60–70% greater protection against invasive cervical carcinomas compared to cytology-based screening. 7 Thus, according to the current European Guidelines, 8 the recommended algorithm of cervical cancer screening includes an HR-HPV test as primary screening test for women over the age of 30–35 and, if positive, a triage test to limit unnecessary colposcopies (in Italy, it is Pap test). Moreover, the optimal time interval for offering HPV primary screening is at least five years. For younger women (age <30–35), the Pap test remains the primary screening test and the HPV test is performed for the triage of borderline cytology and in follow-up protocols.
Nowadays, several HPV DNA tests are commercially available, based on different molecular technologies, HPV target genes, and detected HPV types. In order to be used in the screening context, a HPV test must be clinically validated, 9 to detect only infections potentially associated with precancerous lesions or cancer. In cervical cancer screening, HPV tests detect the DNA of the 12 HR-HPV types (16/18/31/33/35/39/45/51/52/56/58/59) and, depending on the test, HPV with probable (68) or possible (66) risk, according to the International Agency for Research on Cancer (IARC) classification. 10
In Tuscany, a region of Italy, implementation of the HPV primary screening programme has been ongoing gradually since 2013, by age and Local Health Unit. The Cancer Prevention Regional Laboratory of ISPRO (Istituto per lo Studio, la Prevenzione e la Rete Oncologica) in Florence was chosen for centralization of HPV and cytology triage tests coming from the whole of Tuscany. Until May 2016, Hybrid Capture 2 (HC2; Qiagen) was used as the screening test; after a regional public tender, it was replaced by the cobas® 4800 HPV test (Cobas; Roche). These are two HPV tests validated for screening purposes, and characterized by similar but not identical clinical and analytical performances.
The main objective of this study was to evaluate, in a primary screening setting with HPV test, the impact of the transition from one analytical method (HC2) to another (Cobas) on laboratory activities and on screening organization, taking into account both clinical and analytical screening performance of the two tests.
Methods
Study population and design
The HPV screening protocol in Tuscany complies with Italian and European guidelines. Women who test HPV positive and have abnormal/inadequate cytology triage are sent directly for colposcopic examination. HPV-positive women with normal cytology triage are invited for a repeat HPV test after one year: women who have a persistent HR-HPV infection undergo colposcopic examination and women cleared of HPV infection are sent to the next screening round.
The study involved two different analyses, as follows.
Evaluation of HPV screening clinical performance in two different periods (first study level)
We considered 37,775 samples from all women (34–64 years old) participating in the HPV primary screening programme of the metropolitan city of Florence from June 2015 to March 2017. We decided to focus on this period to avoid any factor that could influence the characteristics of the target population, taking into account the following variables:
the Tuscany region decided to gradually implement HPV primary screening program by age, starting to invite, in 2013, older women (age range 55–64); in 2014, women aged 45–64; in 2015, all women, aged 34–64; in 2018, reminder letters were sent to women who did not participate in the first screening round, and the second round of cervical cancer screening with HPV test began.
Furthermore, since June 2015 in the Florentine area, all cervical screening samples were collected in ThinPrep medium (Hologic).
All laboratory tests (HPV test and cytology triage) were performed in the Cancer Prevention Regional Laboratory of ISPRO. From June 2015 until May 2016 (first period), samples were analysed on HC2; from June 2016 to March 2017 (second period), samples were analysed on Cobas. Colposcopies were also all performed at ISPRO.
Before and after changing from HC2 to Cobas, we compared at baseline and at one-year recall (when all samples were analysed with Cobas): HPV positivity, cytology and histological results. All comparisons were evaluated using the Chi-square test and 95% confidence interval (CI). A p value < 0.05 was considered statistically significant.
Evaluation of HC2 and Cobas analytical performance (second study level)
A set of 620 consecutive anonymised routine screening HC2-positive samples were randomly chosen to be re-tested on Cobas using residual ThinPrep material. In order to identify the cause of discordant results between HC2 and Cobas, since HC2 has, as molecular target, the full HPV genome length, samples with negative result at the Cobas re-test (discordant samples) were HPV genotyped on specific target genes:
L1 gene by a reverse line blot (RLB) method (Ampliquality HPV-Type Express v3.0 assay; AB Analitica), a HPV test not validated for cervical cancer screening and very useful for identifying possible cross-hybridization as it is characterized by a higher analytical sensitivity than Cobas. E6/E7 genes by a clinically validated screening test (BD Onclarity™ HPV Assay), which provides more “extended” partial genotyping results than Cobas.
The genotyping results were linked to cytological and histological data of baseline and of subsequent tests executed during the follow up.
HR-HPV testing
Hc2 test
The HC2 HR-HPV test is a molecular hybridization and sandwich capture assay that detects the 12 HR-HPV types and type 68, classified as probably carcinogenic to humans (Group 2A) by IARC. 10 This test has full length of HPV genome as target and is a clinically validated HPV screening test. It was performed according to the manufacturer’s protocol.
HC2 utilizes chemiluminescent detection of the target, and its results are expressed as the ratio of the specimen’s light emission in relative light units (RLUs) to the average of three concurrently tested positive controls containing 1 pg/mL HPV DNA, which is the recommended positivity threshold (cut off = 1 RLU/CO).
All samples (collected in ThinPrep) were first processed using QIAsymphony DSP HPV Media kit (Qiagen) on an automated platform (QIASymphony SP, Qiagen), in order to obtain a microplate with the extracted DNA. Subsequently, the microplate was analysed on the Rapid Capture System (RCS, Qiagen) automated system.
Cobas test
The cobas 4800 HPV test is a real-time PCR-based assay. It detects the 12 HR-HPV types, and HPV68 and HPV66, classified as probably and possibly a biological agent carcinogenic to humans (respectively Group 2A and 2B). 10 It is a clinically validated HPV screening test 11 , 12 and its targets are the HPV L1 gene and the human β-globin gene as internal control. It performs a partial genotyping, with the identification of HPV16 and HPV18, using four different detection channels (HPV16, HPV18, 12 other HPV types, and human β-globin). A positive result occurs when the CT (cycle threshold) target value is <40.0 for HPV18 and 12 other HPV, and <40.5 for HPV16. The test was performed by an automated system consisting of three instruments: p480 (resuspension and decapping), ×480 (DNA extraction and real-time PCR plate setup), and z480 (amplification, detection, and HPV partial genotyping by real-time PCR). All steps were performed according to the manufacturer’s protocols.
HPV genotyping
Genotyping of HPV L1 gene by RLB method
The Ampliquality HPV-Type Express v3.0 assay is a RLB method that detects the presence of 40 HR- and non-HR-HPV types (6/11/16/18/26/31/33/35/39/40/42/43/44/45/51/52/53/54/55/56/58/59/61/62/64/66/67/68 (subtypes 68a and 68 b)/69/70/71/72/73/81/82/83/84/87/89/90). It is based on the reverse hybridization principle and uses PCR with biotinylated primers for the amplification of a HPV L1 region of 139–145 bp. Another pair of primers is used to amplify the human TST (thiosulfate sulfurtransferase) gene, an internal control for cell adequacy, extraction, and amplification. All steps were performed according to the manufacturer’s protocols.
PCR was performed in a final volume of 25 µL, with 5 µL of extracted DNA obtained by Cobas extractions.
Biotinylated PCR products were denatured with 20 µL of denaturation solution for 5 min and all following steps (hybridization with specific oligonucleotide probes immobilized on membrane strips, stringent washing, streptavidin-conjugated alkaline phosphatase addition and bonding to any previously formed biotinylated hybrid, non-radioactive reaction using a chromogenic substrate) were performed using automatic instrumentation (Autoblot). Strips were analysed with specific software and checked further visually, using an interpretation grid supplied with the kit. The samples were considered positive for a HPV type if the precipitate was viewed at the specific position on the strip.
Genotyping of HPV E6/E7 genes by BD onclarity HPV assay
The BD Onclarity HPV assay is a HPV DNA real-time PCR, targeting the viral genes E6 and E7 of 14 HPV types (12 HR-HPV, HPV66, and 68).13 It provides an extended genotyping, allowing individual detection of six types (HPV16/18/31/45/51/52) and the detection of eight types in three distinct groups: HPV33/58, HPV56/59/66, and HPV35/39/68. The assay also detects human beta-globin gene as internal control for sample adequacy and assay performance.
This HPV test was performed with the BD Viper LT system in the laboratory of the IEO (European Institute of Oncology, Milan) on discordant samples that were negative for the 12 HR-HPV types on L1 gene RLB genotyping, using DNA from Cobas extractions: 30 µL of Cobas extracts were transferred in the BD extraction tubes, diluted in a total volume of 400 µL (200 µL of elution buffer plus 200 µL of neutralization buffer), and 150 µL of this dilution were used for the PCR reactions.
Cytology triage
Pap tests of samples with positive result from the HPV screening test were prepared using an automatic platform (ThinPrep 5000 processor with Autoloader, Hologic) and interpreted using the classification “The Bethesda System (TBS) for reporting cervical cytology” of 2014.
Results
Evaluation of HPV screening clinical performance in two different periods (first study level)
In the Florentine area, 37,775 HPV screening samples were collected in ThinPrep from June 2015 to March 2017: 17,137 (45.4%) were tested on HC2 and 20,638 (54.6%) on Cobas. The average ages of women in the two groups were similar (46.5 vs. 46.1 years old). The values of screening indicators and other parameters compared between the two groups of samples (HC2 vs. Cobas) at baseline are showed in Figure 1 and detailed in Table 1.
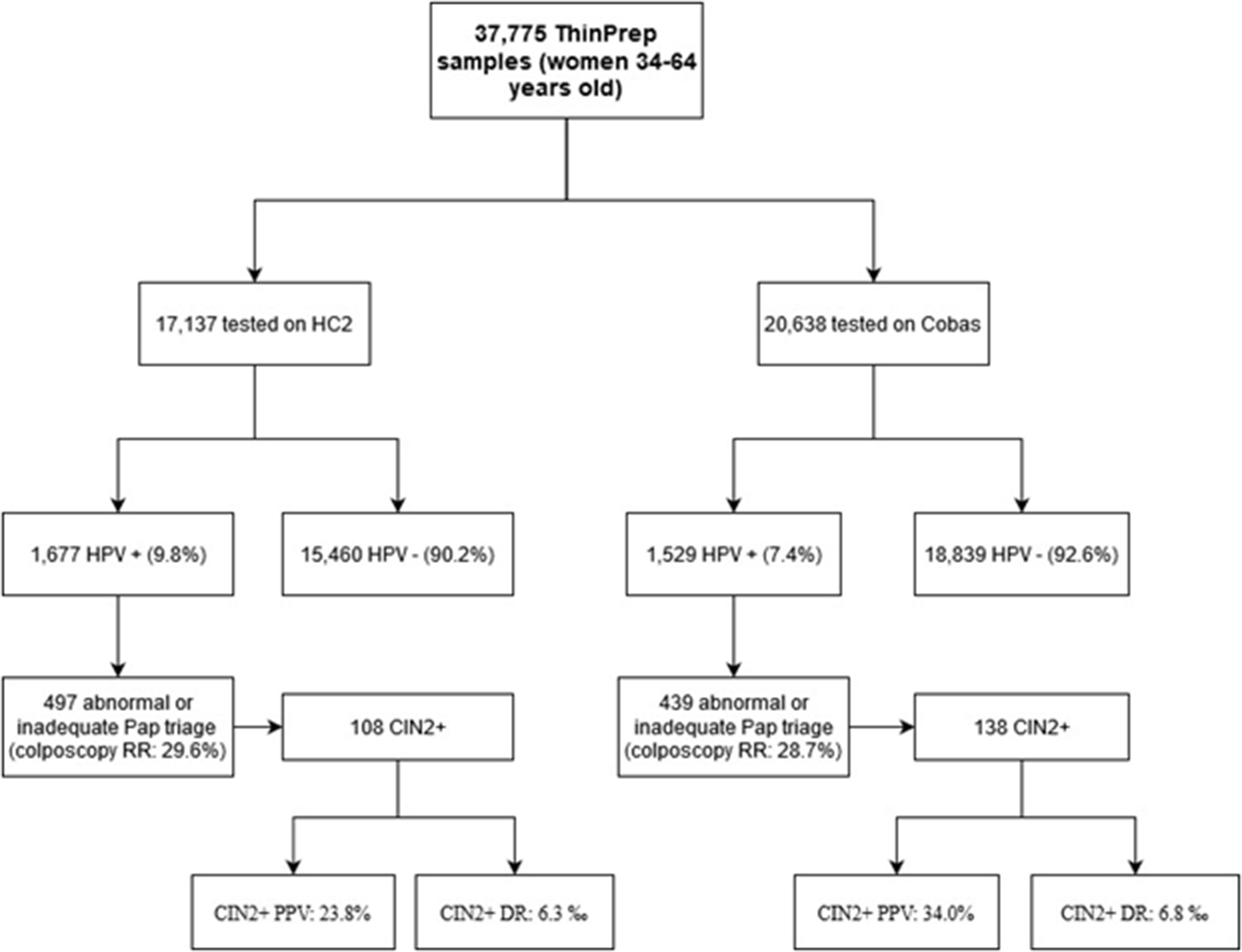
Flowchart of the evaluation of HC2 versus Cobas clinical performance in the same screening population at baseline.
Screening indicators and other parameters compared between the two groups of samples (Hybrid Capture 2 (HC2) vs. Cobas) at baseline.
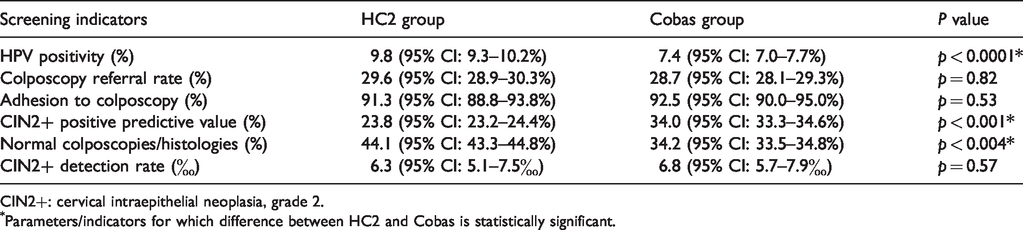
CIN2+: cervical intraepithelial neoplasia, grade 2.
Parameters/indicators for which difference between HC2 and Cobas is statistically significant.
The HPV positivity was 9.8% (1677/17,137) for HC2 and 7.4% (1529/20,368) for Cobas (p < 0.0001); 480 HC2-positive women (28.6%) had abnormal cytology triage and 17 (1%) inadequate; 418 Cobas-positive women (27.3%) had abnormal cytology triage and 21 (1.4%) inadequate. Hence, the colposcopy referral rates (RRs) were similar: 29.6% for HC2 and 28.7% for Cobas. The distribution of cytology triage abnormalities for HC2-positive women was: 78.3% (376/480) L-SIL, 9.8% (47/480) ASC-H (atypical squamous cell – cannot exclude high grade), 10.8% (52/480) H-SIL (high-grade squamous intraepithelial lesion) and 1.1% (5/480) AGC (atypical glandular cells); for Cobas positive women, it was: 81.6% (341/418) L-SIL, 5.3% (22/418) ASC-H, 12.4% (52/418) H-SIL and 0.7% (3/418) AGC. The only statistically significant difference between the two groups of women was for ASC-H frequency (p < 0.02).
Adhesion to colposcopy was high: 91.3% (454/497) in HC2 group and 92.5% (406/439) in Cobas group.
At immediate colposcopy, 108 cervical intraepithelial neoplasia, grade 2 (CIN2+) lesions were found in women tested with HC2 and 138 in women tested with Cobas (respectively PPV 23.8% and 34.0%, p < 0.001), 145 versus 126 CIN1/condiloma, 200 versus 139 normal colposcopies/histologies (p < 0.004) and 1 versus 2 inadequate histologies.
The detection rate (DR) of CIN2+ at baseline was similar between the two groups (without any statistically significant difference): 6.3‰ (108/17,137) for HC2 group and 6.8‰ (138/20,368) for Cobas group.
According to the screening protocol, HPV-positive women with normal cytology triage were recalled after one year to repeat HPV test, apart from a small group of women who were not included because they were part of a research study. Consequently, 914 of 1180 women of HC2 group and 1051 of 1090 women of Cobas group with normal cytology triage at baseline were invited to the one-year recall. The compliance to recall was 82.5% (754/914) for HC2 group and 82.2% (864/1051) for Cobas group.
The HPV positivity at one-year recall (when all samples were analysed with Cobas) was statistically different between the two groups: 40.6% (306/754; 95% CI: 37.1–44.1%) for HC2 group (women analysed at baseline with HC2) and 62.2% (537/864; 95% CI: 59.0–65.4%) for Cobas group (women analysed at baseline with Cobas); p < 0.0001. The overall compliance to colposcopy at one-year recall was 91.5% (771/843) and 82 CIN2+ was found (32 in HC2 group and 50 in Cobas group; PPV: 10.6%).
As shown in Table 2, overall (baseline and one-year recall) CIN2+ PPV and DR were similar between the two groups: respectively for HC2 and Cobas, 19.0% (140/738) versus 21.1% (188/893) and 8.2‰ (140/17,137) versus 9.2‰ (188/20,368).
Overall (baseline and one-year recall) screening indicators compared between the two groups (Hybrid Capture 2 (HC2) vs. Cobas).

CIN2+: cervical intraepithelial neoplasia, grade 2.
Evaluation of HC2 and Cobas analytical performance (second study level)
620 HC2-positive routine screening samples were re-tested with Cobas: 419 were confirmed HPV positive, while 201 gave a negative HPV result (discordant samples). The positive agreement between HC2 and Cobas was 67.6%.
36/201 (17.9%) discordant samples had abnormal cytology and the results of colposcopic and histological examination were: 2 (5.6%) CIN2+, 17 (47.2%) CIN1 and 17 (47.2%) normal colposcopies. In Table 3, cytology triage and CIN2+ cases at histological examination of discordant samples are reported.
Cytology triage results of discordant samples (Hybrid Capture 2 (HC2) positive but Cobas negative) and cervical intraepithelial neoplasia, grade 2 (CIN2+) cases at histological examination (both CIN3).
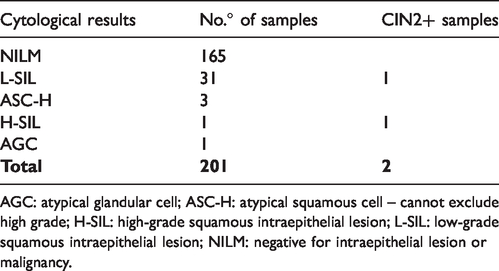
AGC: atypical glandular cell; ASC-H: atypical squamous cell – cannot exclude high grade; H-SIL: high-grade squamous intraepithelial lesion; L-SIL: low-grade squamous intraepithelial lesion; NILM: negative for intraepithelial lesion or malignancy.
All discordant samples were genotyped with two different molecular tests, and the flowchart in Figure 2 summarizes the procedures and results of HPV genotyping.
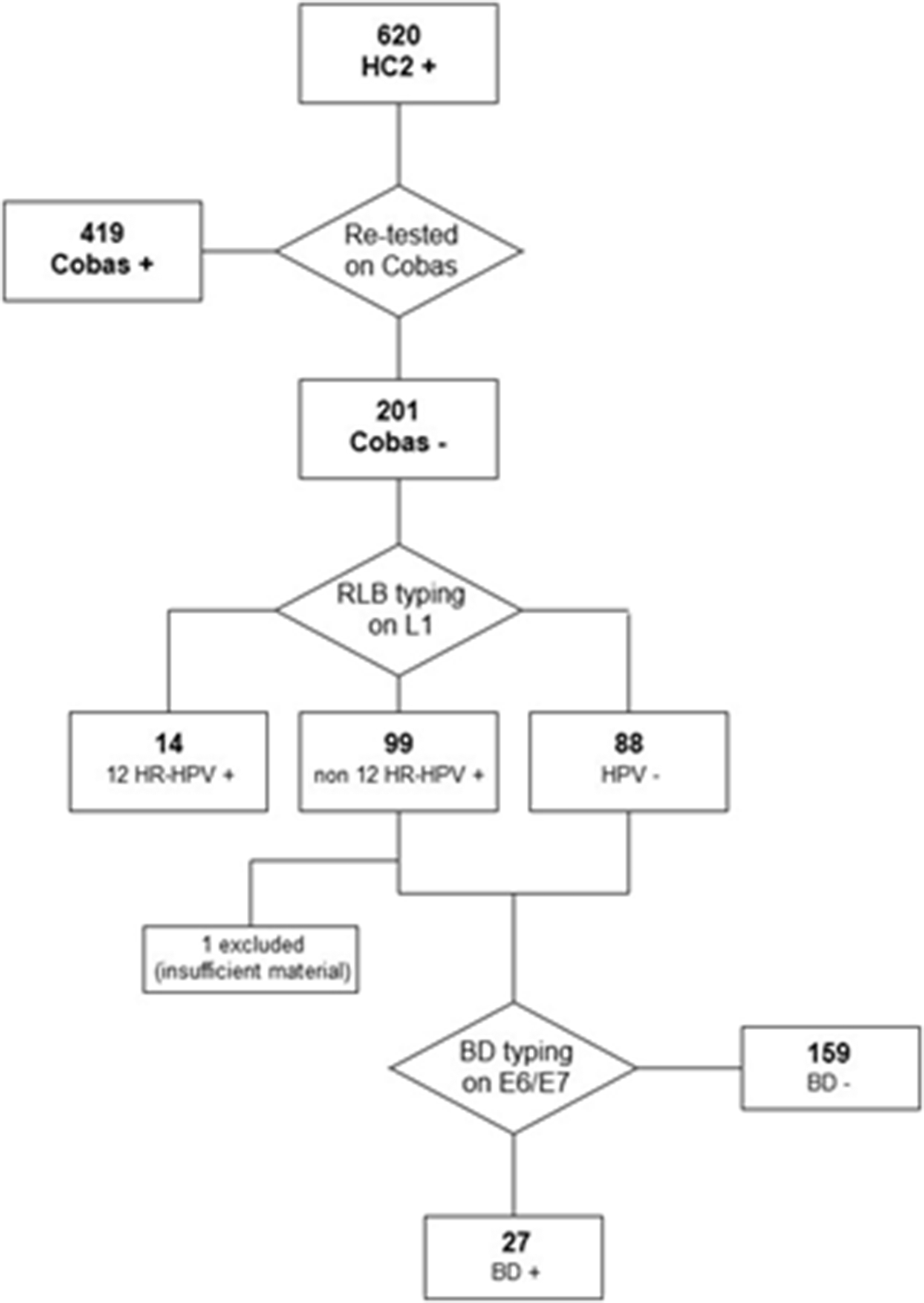
Flowchart of HPV genotyping procedures and results.
All 201 discordant samples were typed using AB Analitica RLB: 14/201 (7%) samples were found to be positive for the 12 HR-HPV types, of which one with abnormal cytology and without any CIN2+ lesion; 88/201 (43.8%) samples were HPV negative, of which five with abnormal cytology and none with CIN2+ lesion; 99/201 (49.2%) were positive for HPV types different from the 12 HR-HPV (1 HPV66 and 20 HPV68), of which 30 (30.3%) with abnormal cytology and 2 with CIN3 lesions (1 HPV26 positive with H-SIL cytology, 1 HPV54/73/90 positive with L-SIL cytology). These results are detailed in Table 4.
Reverse line blot (RLB) genotyping and cytology results of Hybrid Capture 2 (HC2) versus Cobas discordant samples.
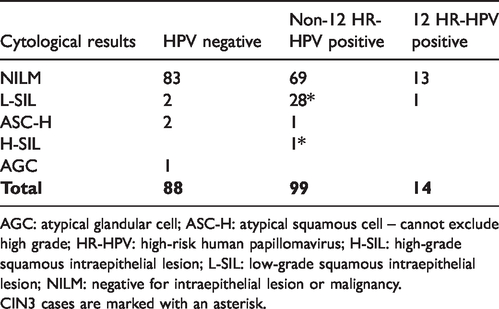
AGC: atypical glandular cell; ASC-H: atypical squamous cell – cannot exclude high grade; HR-HPV: high-risk human papillomavirus; H-SIL: high-grade squamous intraepithelial lesion; L-SIL: low-grade squamous intraepithelial lesion; NILM: negative for intraepithelial lesion or malignancy.
CIN3 cases are marked with an asterisk.
Amongst the 14 discordant samples found to be positive for the 12 HR-HPV types, 5 were positive for HPV52, the type for which Cobas has the lower analytical sensitivity. Moreover, 11/14 (78.6%) of these samples had a low RLU/CO on HC2 (<3).
Amongst the HPV68 positive samples on RLB (20/201, 10%), it was found that all of them were HPV68a positive and none HPV68b positive.
b.
186 of 187 discordant samples that were found to be HPV negative or not positive to the 12 HR-HPV types on RLB (one excluded because of insufficient material) were analysed on BD Onclarity. 27/186 samples (14.5%) gave a positive result (4 to HPV33/58 and 23 to HPV39/68/35), of which 7/27 (25.9%) had abnormal cytology (all L-SIL) but were negative for CIN2+. The range of HC2 ratio (RLU/CO) for these samples was 3.14–3732.96. Amongst the 23 samples positive for HPV39/68/35 on BD Onclarity, 16 (69.6%) were found to be HPV68a positive on RLB. The only two cases of CIN2+ (CIN3) also gave a negative result for HR-HPV on BD Onclarity.
We decided to monitor the follow up (24–36 months) of the discordant cases, searching subsequent exam results (HPV test and/or cytology) in our screening database. The results of subsequent tests were available for 139/201 (69.1%) discordant samples: only 13/139 (9.3%) gave a positive result on Cobas testing and only two of them had abnormal cytology (both L-SIL).
Discussion
HC2 and Cobas 4800 are two clinically validated tests for HPV screening of cervical cancer and they are based on two different technologies. In this study, we considered HPV screening samples from the Florentine area, and we analysed HPV screening parameters in the two periods before and after the transition from one test to the other. In addition, we analysed the analytical performance, re-testing some routine HC2-positive samples on Cobas and deepening the analysis on discordant samples.
Evaluation of HPV screening clinical performance in two different periods (first study level)
The use of HC2 as primary screening test, compared to Cobas, registered at baseline a greater HPV positivity, a lower CIN2+ PPV and a higher frequency of normal colposcopies and histologies (all statistically significant differences), while the DRs of CIN2+ were similar. Indeed, although the colposcopy referral rate (determined by cytology triage) were comparable, HC2 testing sent more women for colposcopy since it determined a higher HPV positivity. This resulted in CIN2+ PPV being lower, losing in specificity but not in sensitivity, since the DR of CIN2+ remained stable.
The higher positivity of the HC2 test could be partially explained by the well-known cross-hybridization of this test with non HR-HPV types,14–17 unlike Cobas, which is characterized by a lower rate of cross-hybridization and a higher analytical specificity. In fact, as shown by our data, non HR-HPV types detected by HC2 increased HPV positivity, but determined a lower PPV for CIN2+ lesions at the immediate colposcopy, since these are women with low-risk HPV infection mostly associated with no pre-neoplastic lesions. Moreover, considering that after the transition from STM collection medium to ThinPrep in our laboratory we observed a higher HC2 positivity rate (9.8% vs. 7.3%, data not shown), we might hypothesize that application of the HC2 test on ThinPrep could cause an increase in the HC2 cross-hybridization rate.
At one-year recall, the HPV positivity was statistically different (much higher for Cobas group: 62.2% vs. 40.6%). The lower positivity of the HC2 group samples could also be explained by the cross-hybridization that characterizes the HC2 test. The use of the same HPV screening test for both groups (HC2 and Cobas) at one-year recall determined that the results of overall screening indicators were substantially the same between the two groups.
Evaluation of HC2 and Cobas analytical performance (second study level)
Re-testing HC2 positive screening samples with Cobas, the positive agreement was 67.6%, so we identified about one-third of discordant samples (HC2 positive/Cobas negative). This result reflects the decrease in HPV positivity that has been found after the introduction of the Cobas test in Tuscan routine screening, and it is confirmed by another study,
18
in which it was reported that HPV assays disagreed in about one-third of all positive cases. Analysing the discordant samples with other HPV tests that perform a complete or partial genotyping (AB Analitica RLB and BD Onclarity, respectively), we obtained the following results:
on AB Analitica RLB (L1 as target gene), more than 90% gave a result of HPV negative or not positive to the 12 HR-HPV types on BD Onclarity (E6/E7 as target genes), 85% of these samples also gave a negative result.
We decided to type the discordant samples also on E6/E7 genes because the Cobas test and AB Analitica RLB have the same HPV target gene (L1), in contrast to HC2 which targets the whole HPV genome, and this could explain the major concordance registered between Cobas and AB Analitica. Overall, 79.6% of discordant samples (HC2 positive/Cobas negative) resulted as negative to the 12 HR-HPV types and 77.6% as negative to all HPV types that are the target of Cobas (12 HR-HPV + HPV66 + HPV68). The possible reasons why the Cobas test did not detect 22.4% of discordant samples that resulted as positive to its HPV target types on other HPV tests are: (1) Cobas test and BD Onclarity Assay have a different target gene (L1 vs. E6/E7) and (2) RLB typing (AB Analitica) has a higher analytical sensitivity, in fact it is not used as a cervical cancer screening test.
Amongst the discordant cases (HC2 positive/Cobas negative), over 80% had a normal cytology and we found only two CIN3 lesions that were associated with non-HR-HPV types. Moreover, using the oncogenic E6/E7 genes as target, we were not able to increase the clinical sensitivity for CIN2+ and only 9% of discordant cases (HC2 positive/Cobas negative) were found to be HPV positive on Cobas testing in subsequent examinations. Thus, the use of the Cobas test would have determined a lower rate of false positives and, consequently, a lower rate of referral to colposcopy.
HPV68 is a target type of both HC2 and Cobas, but about 10% of discordant samples were HPV68a positive on RLB typing (with no cases positive for HPV68b). So, it is not clear if HPV68a and HPV68b are both detected with the same sensitivity by Cobas.
Conclusions
The parallel comparison of the HC2 and Cobas tests, both on the same population and on the same samples, allowed us to deepen our understanding of the clinical and analytical performance of these two validated screening tests.
The use of HC2 as primary screening test, compared to Cobas, has registered:
at baseline: greater HPV positivity, similar colposcopy RR and CIN2+ DR, lower CIN2+ PPV and higher frequency of normal colposcopies/histologies at one-year recall (all samples re-analysed with Cobas): lower HPV positivity.
Cobas was shown to be more specific than HC2: overall, about 80% of discordant samples (HC2 positive/Cobas negative) were found to be HR-HPV negative by typing on L1 and E6/E7 genes. However, the higher analytical specificity of Cobas led to the missed identification of those few cases of CIN3 lesions, detected by HC2 due to its cross-hybridization which thus increases its clinical sensitivity.
Footnotes
Author contributions
Alessandra Mongia, Giampaolo Pompeo and Francesca Carozzi contributed to the study conception and design. Data collection and analysis, material preparation and molecular analysis were performed by Alessandra Mongia and Giampaolo Pompeo; Leonardo Ventura and Fabio Bottari have performed respectively screening data collection and BD Onclarity molecular analysis. The first draft of the manuscript was written by Alessandra Mongia and Giampaolo Pompeo. All authors contributed with comment and correction on previous versions of the manuscript. All authors read and approved the final manuscript that was written by Alessandra Mongia and Giampaolo Pompeo.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
