Abstract
Human embryonic stem cells (hESCs) are advantaged sources for large-scale and homogeneous mesenchymal stem/stromal cells (MSCs) generation. However, due to the limitations in high-efficiency procedures for hESC-MSCs induction, the systematic and detailed information of mesengenesis and early MSC development are largely obscure. In this study, we took advantage of the well-established twist-related protein 1 (TWIST1)-overexpressing hESCs and two small molecular cocktails (CHIR99021, decitabine) for high-efficient MSC induction. To assess the multidimensional biological and transcriptomic characteristics, we turned to cellular and molecular methods, such as flow cytometry (FCM), quantitative reverse transcription–polymerase chain reaction (qRT-PCR), in vitro tri-lineage differentiation, cytokine secretion analysis, in vivo transplantation for acute liver injury (ALI) management, and bioinformatics analyses (eg, gene ontology–biological processes [GO-BP], Kyoto Encyclopedia of Genes and Genomes [KEGG], HeatMap, and principal component analysis [PCA]). By combining TWIST1 overexpression (denoted as T) and the indicated small molecular cocktails (denoted as S), hESCs high-efficiently differentiated into MSCs (denoted as TS-MSCs, induced by T and S combination) within 2 weeks. TS-MSCs satisfied the criteria for MSC definition and revealed comparable tri-lineage differentiation potential and ameliorative efficacy upon ALI mice. According to RNA-sequencing (SEQ) analysis, we originally illuminated the gradual variations in gene expression pattern and the concomitant biofunctions of the programmed hESC-MSCs. Overall, our data indicated the feasibility of high-efficient generation of hESC-MSCs by TWIST1 and cocktail-based programming. The generated hESC-MSCs revealed multifaceted in vivo and in vitro biofunctions as adult BM-MSCs, which collectively suggested promising prospects in ALI management in future.
Introduction
Since the 1970s, mesenchymal stem/stromal cells (MSCs) have been identified from considerable sources (eg, bone marrow, adipose tissue, umbilical cord, and placenta) with unique bidirectional immunomodulation- and hematopoietic-supporting properties, together with tri-lineage differentiation potential toward adipocytes, osteoblasts, and chondrocytes1–3. Of them, bone marrow–derived MSCs (BM-MSCs) had the most range of clinical applications, whereas perinatal tissue-derived MSCs exhibited preferable proliferation and immunomodulatory capacity over adult tissue–derived counterparts4,5. State-of-the-art updates have highlighted the feasibility of large-scale MSC generation from human pluripotent stem cells (hPSCs) including human embryonic stem cells (hESCs) and human-induced pluripotent stem cells (hiPSCs) for both preclinical and clinical purposes2,6. Distinguishing from the aforementioned tissue-derived MSCs, hPSCs-derived MSCs (hPSC-MSCs) revealed robust proliferation capacity and considerable homogeneity exempt from risk of ethics, pathogenic microorganism, and invasive sampling, which thus possess broad prospects in rehabilitation medicine and regenerative medicine 6 .
Human PSCs (HPSCs), characterized by their self-renewal and multi-lineage differentiation potential, have been recognized as promising alternatives for drug discovery, early development, disease remodeling, cellular therapies, and tissue engineering 7 . For instance, in recent years, a variety of functional cell types have been generated from both hESCs and hiPSCs, including megakaryocytes8,9, myocardial cells 10 , muscular cells 11 , and natural killer (NK) cells12,13. Since the year 2005, some investigators have devoted to decoding the biological functions and transcriptomic characteristics of hPSC-MSCs 14 . For a long period of time, studies toward mesengenesis and the underlying molecular mechanisms were largely hindered by the deficiency of procedures for high-efficient hPSC-MSCs generation, which could only be partially achieved by strategies such as continuous passage, scraps, magnetic-activated cell sorting (MACS), and fluorescence-activating cell sorting (FACS)7,14,15. Of note, Wang et al. 16 and Zhang et al. 2 reported a reasonable quantity of MSC induction from hPSCs by utilizing the BMP4- and MSX2-based cell programming, respectively. Subsequently, Wei et al. 6 and Zhang et al. 1 reported the high-efficient generation of hESC-MSCs and hPSC-MSCs by small molecular cocktails, respectively. Interestingly, we recently demonstrated the pivotal role of MSX2-twist-related protein 1 (TWIST1) axis during hPSC-MSCs generation, yet the detailed information of the biofunction and transcriptomic features of TWIST1-programmed hESC-MSCs are still largely obscure 2 .
Therefore, in this study, a well-established procedure was used for evaluating the early mesengenesis and biological characteristics of hESC-MSCs. Notably, a certain proportion of hESC-MSCs were generated by TWIST1 initiation, which was further enhanced by combining with small molecular cocktails (CHIR99021, decitabine [DAC]). The hESC-MSCs revealed typical MSC phenotypes and comparable biofunctions with adult BM-MSCs. With the aid of RNA-sequencing (SEQ), we further investigated the gradually transcriptomic characteristics of MSC generation from the parental hESCs. Finally, acute liver injury (ALI) mice with hESC-MSCs transplantation revealed comparable efficacy and good manifestations as those with BM-MSCs administration.
Materials and Methods
Cell Culture
H1 hESCs (WiCell Research Institute, MADISON, WI, USA) were cultured in E8 medium (Gibco, USA) on matrigel (Corning, USA)-coated 6-well plate (Corning, USA) as we reported before, with several modifications 6 . For hESC-MSCs induction, 2-5 × 104 H1 hESCs were seeded in E8 medium (Gibco, USA) on growth factor reduced (GFR) Matrigel (Corning, USA)-coated 6-well plate (Corning, USA) for 2 days, and then the medium was changed into Dulbecco’s Modified Eagle’s Medium (DMEM)-F12 basal medium (Hyclone, USA) with 1% GlutaMAX (Gibco, USA), 1% nonessential amino acids (NEAA; Sigma, USA), 2% fetal bovine serum (FBS; Gibco, USA), 100 U/ml penicillin/streptomycin (ThermoFisher, USA), 10 ng/ml epidermal growth factor (EGF; Peprotech, USA), and 4 ng/ml basic fibroblast growth factor (bFGF; Peprotech, USA) addition. For MSC induction, H1 hESCs with TWIST1 overexpression were cultured in E8 medium on GFR Material–coated plate for 2 days, and then the medium was replaced with MSC culture medium with 2% FBS and/or DAC (20 nM) addition for 14 days. The H1 hESCs and the derived cells at the indicated time points were observed and collected for the indicated analyses (day 0, day 4, day 7, and day 14).
For hESC-MSCs passage, the cells were detached by 0.05% Trypsin/EDTA (Gibco) and seeded on GFR Material–coated 6-well plate in MSC culture medium containing DMEM-F12 basal medium (Hyclone, USA), 1% GlutaMAX (Gibco, USA), 1% NEAA (Sigma, USA), 2% FBS (Gibco, USA), 100 U/ml penicillin/streptomycin (ThermoFisher, USA), 10 ng/ml EGF (Peprotech), and 4 ng/ml bFGF (Peprotech). The medium was changed every 2 days. All cells including H1 hESCs and the derivations were maintained at 37°C, 5% CO2.
Construction of H1 hESCs Stable Line With Ectopic TWIST1 Overexpression
For TWIST1 overexpression, the coding sequence of TWIST1 was integrated into the pCDH-GFP-TWIST1 plasmid, and then transduced into H1 hESCs cell line by utilizing the lentivirus-medicated strategy. After 48 h, H1 hESCs with pCDH-GFP-TWIST1 expression were enriched by GFP-based FACS for further analyses.
Flow Cytometry (FCM) Assay
For the quantitative analysis of the proportion of hESC-MSCs, FCM assays were conducted as reported before1,6. In details, cells were detached by 0.05% Trypsin/EDTA (Gibco) at the indicated time points and washed with 1 × PBS (Solarbio, China) twice. After that, the cells were labeled with fluorescence-conjugated monoclonal antibodies (CD44, CD73, CD90, CD105, CD34, and CD45) in dark for 30 min. Then, the cells were washed with 1 × PBS (Solarbio, China) twice, and turned to FACS Canto II (BD Biosci, USA) for detection. CD44, CD73, CD90 and CD105 were used as the positive markers and CD34 and CD45 were used as the negative markers for hESC-MSCs. The proportion of hESC-MSCs was analyzed by FlowJo 10.0 software (Tree Star, USA). The list of antibodies is available in Supplemental Table S1.
qRT-PCR Analysis
A QRT-PCR analysis was conducted as in previous studies6,17. Briefly, hESCs and the derivations at the indicated time points were washed with 1 × PBS (Solarbio, China) and lysed by the TRIZol (Invitrogen Life Technologies, Carlsbad, CA, USA) reagent for total mRNA preparation according to the manufacturer’s constructions. Then, the TransScript Fly cDNA Synthesis SuperMix kit (TransGen, China) was used for cDNA synthetization. After that, SYBR Green PCR Master Mix kit (Qiagen, Germany) and the ABI PRISM 7900 (Applied Biosystems, USA) were used for qRT-PCR analysis. The list of primer sequences is available in Supplemental Table S2.
Tri-Lineage Differentiation of hESC-MSCs and BM-MSCs
Tri-lineage differentiation capacity of the indicated MSCs toward adipocytes, osteoblasts, and chondrocytes was verified, as we recently described, with several modifications1,18. In detail, 5 × 104 hESC-MSCs and BM-MSCs were seeded in 12-well plate wells in the aforementioned MSC culture medium for 2 to 3 days. Then, the medium was replaced with the commercial tri-lineage differentiation medium (adipogenic-, osteogenic-, and chondrogenic-differentiation; Stem Cell Technologies, USA), respectively. After a 2-week induction, the generated adipocytes, osteoblasts, and chondrocytes were dyed with Oil Red O staining (Solarbio, China), Alizarin Red S staining (Solarbio, China), and Alcian Blue staining (Solarbio, China), respectively.
Construction of ALI Mouse Model
An ALI mouse model was conducted as in a previous study with several modifications 19 . Briefly, mice were randomly divided into four groups as follows: the negative control group (NC), the ALI model group (ALI), and ALI mice with TS-MSCs treatment (TS-MSCs) or BM-MSCs treatment (BM-MSCs). In detail, 6-week nonobese diabetic–severe combined immunodeficiency (NOD-SCID) mice received 4% chloral hydrate (0.05 ml per 10 g body weight) through intraperitoneal injection as previously reported20,21. Then, 40 μl sterilized 20% CCl4 solution was instilled into the caudal vein of the aforementioned mice (twice a week for 8 weeks), with the aid of orotracheal tube for ALI induction (the experimental group), while the mice in the control group were treated with equal amount of normal saline. After 8 weeks, the mice in the experimental groups were treated with systemic infusion of 4 × 106 TS-MSCs or BM-MSCs through tail vein injection, while mice in the control group were treated with an equal amount of saline solution. After 2 weeks, all the mice were euthanatized and liver tissues and blood samples were collected to assess the immunoregulatory and therapeutic capacity of TS-MSCs or BM-MSCs. The study was approved by the ethics committee of Gansu Provincial Hospital according to the guideline of Helsinki (2022-088, 2023-120).
RNA-SEQ and Bioinformatic Analysis
RNA-SEQ and bioinformatic analysis was conducted in a recent study1,18. In detail, total mRNAs were extracted from H1 hESCs and derivations at the indicated time points by utilizing TRIzol (Invitrogen, USA) reagent and qualified by NanoDrop (ThermoFisher, USA). The RNA-SEQ analysis was accomplished by Novogene (Tianjin, China). Multifaceted bioinformatic analyses were accomplished by using the online databases and platforms such as gene ontology (GO) assay (http://geneontology.org/) and Kyoto Encyclopedia of Genes and Genomes (KEGG) assay (https://www.kegg.jp/). The detailed information of the differentially expressed genes (DEGs) is available in Supplemental Tables S3–S5.
Statistical Analysis
Statistical analysis was performed by utilizing the Prism 6.0 (GraphPad Software, USA). Briefly, all the data follow the normal distribution pattern. As we described before1,4, the unpaired t test and one-way analysis of variance (ANOVA) test were used for the analysis of two different unpaired groups and multiple unpaired groups, respectively. Statistically significant differences were considered when P < 0.05. Data were shown as mean ± standard error of the mean (SEM; N = 3 independent experiments), NS, not significant, and *P < 0.05; **P < 0.01; ***P < 0.001.
Results
TWIST1 Overexpression Facilitates hESC-MSCs Generation and Early Differentiation
To better understand the biological effect during MSC generation and early germ layer specification, the H1 hESCs stable line with ectopic TWIST1 overexpression was used for further investigations (Fig. 1A, B). In MSC culture medium with 2% FBS, we intuitively observed the typical morphological changes from compact clones into spindle shape by TWIST1 overexpression (Fig. 1C). With the aid of FCM assay, a certain proportion of MSCs was generated from hESCs with TWIST1 overexpression (denoted as T-MSCs), which was distinguished from those in the control group (Fig. 1D).
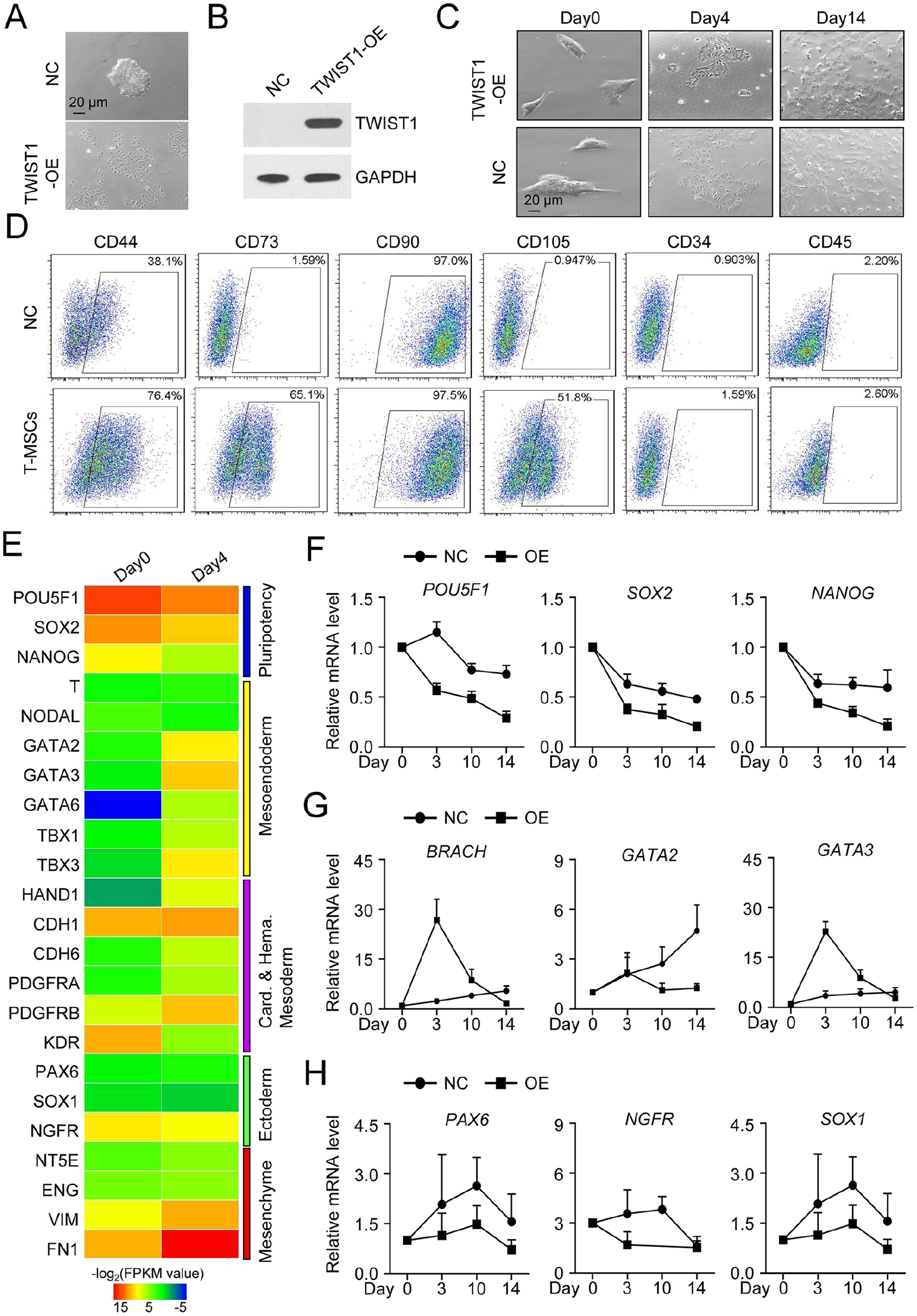
Induction of T-MSCs by TWIST1-overexpression with MSC-like phenotype. (A) Representative phase contract images of H1 hESCs without (upper) and with (bottom) TWIST1-overexpression, respectively. Scale bar = 20 μm. (B) Western-blotting bands of TWIST1 in H1 hESCs without (left) and with (right) TWIST1-overexpression, respectively. (C) Representative phase contract images of H1 hESCs without (upper) and with (bottom) TWIST1-overexpression under MSC induction medium during hESC-MSCs differentiation. (D) Representative FCM diagrams of H1 hESC-derived cells (upper, without TWIST1-overexpression) and T-MSCs (bottom, with TWIST1-overexpression). (E) Representative HeatMap diagrams of pluripotency-, mesoendoderm-, and MSC-associated biomarker expression in H1 hESCs (day 0) and the derivations (day 4). (F-H) qRT-PCR analysis of pluripotency- (POU5F1, SOX2, and NANOG) (F), mesoendoderm- (BRACH, GATA2, and GATA3) (G), and ectoderm- (PAX6, NGFR, and SOX1) (H) associated biomarker expressions during hESC-MSCs induction (mean ± SEM, n = 3) without (NC) and with TWIST1-overexpression. T-MSCs: T-mesenchymal stem/stromal cells; hESCs: human embryonic stem cells; FCM: flow cytometry; qRT-PCR: quantitative reverse transcription–polymerase chain reaction; SEM: standard error of the mean; NC: negative control group; TWIST1: twist-related protein 1.
Subsequently, a qRT-PCR analysis was performed and the results showed that TWIST1 overexpression further accelerated the downregulation of pluripotency-associated biomarkers (POU5F1, SOX2, and NANOG) and the upregulation of MSC-associated biomarkers (NT5E, ENG, VIM, and FN1; Fig. 1E, F). Instead, mesendoderm-associated markers (BRACH, GATA2, and GATA3) were transiently upregulated by TWIST1 overexpression, whereas only moderate variations were found in the expression of neuroectoderm-related genes (PAX6, NGFR, and SOX1) between the control group and the TWIST1-overexpression group (Fig. 1G, H). Taken together, our data indicated the promoting effect of TWIST1 in facilitating MSCs generation and early differentiation toward mesoendoderm.
High-Efficient Generation of hESC-MSCs by TWIST1 Overexpression and the Indicated Small Molecule Cocktail Co-Stimulation
Considering the promoting effect of TWIST1 during early differentiation, we turned to verify the feasibility of high-efficient hESC-MSC generation by TWIST1 overexpression and small molecular cocktails. For this purpose, DAC and CHIR9901 were combined with TWIST1 overexpression for hESC-MSC induction (denoted as TS-MSCs) and it was found that TS-MSCs reveal more typical spindle morphology over that of T-MSCs (Fig. 2A). The FCM data further confirmed the high-efficient generation of MSCs from hESCs within the 14-day induction (Fig. 2B, C). Furthermore, qRT-PCR analysis indicated the further upregulation of MSC-related genes (NT5E, ENG, VIM, and FN1) in TS-MSCs over T-MSCs (Fig. 2D).
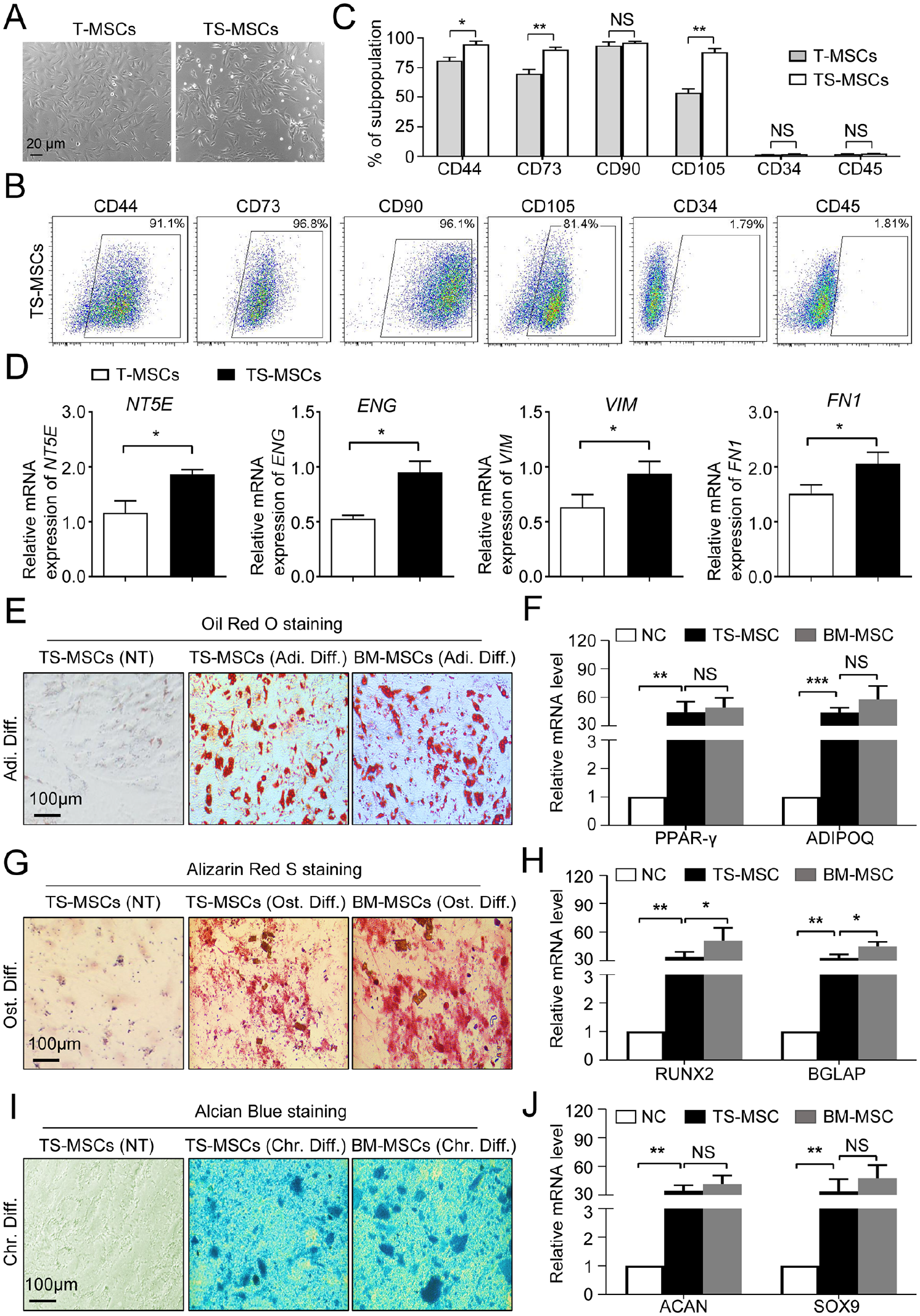
High-efficient generation of TS-MSCs by TWIST1-overexpression and small molecular cocktail combination. (A) Representative phase contract images of T-MSCs and TS-MSCs, respectively. Scale bar = 20 μm. B-C Representative FCM diagrams (B) and statistical analysis (C) of MSC-associated (CD44, CD73, CD90, CD105) and hematopoietic-associated (CD34, CD45) biomarkers in T-MSCs and TS-MSCs, respectively. (D) qRT-PCR analysis of MSC-associated biomarkers (NT5E, ENG, VIM, and FN1) in T-MSCs and TS-MSCs. (E-F) Adipogenic differentiation assessment of TS-MSCs and BM-MSCs identified by Oil Red O staining (E) and qRT-PCR analysis of the indicated genes (ADIPOQ, PPAR-γ) (F). (G-H) Osteogenic differentiation assessment of TS-MSCs and BM-MSCs identified by Alizarin Red S staining (G) and qRT-PCR analysis of the indicated genes (RUNX2, BGLAP) (H). (I-J) Chondrogenic differentiation assessment of TS-MSCs and BM-MSCs identified by Alician Blue staining (I) and qRT-PCR analysis of the indicated genes (SOX9, ACAN) (J). All data were shown as mean ± SEM (n = 3). T-MSCs: TWIST1-overexpression induced-mesenchymal stem/stromal cells from H1 hESCs; TS-MSCs: H1 hESC differentiation towards MSCs by TWIST1-overexpression and small molecular cocktail combination; FCM: flow cytometry; qRT-PCR: quantitative reverse transcription–polymerase chain reaction; BM-MSCs: bone marrow-derived mesenchymal stem/stromal cells; SEM: standard error of the mean; NS: not significant; TWIST1: twist-related protein 1. *P < 0.05; **P < 0.01.
Next, the tri-lineage differentiation potential–related assays were performed to verify the maturation of TS-MSCs. As shown by the Oil Red O staining, TS-MSCs revealed comparable adipogenic differentiation with BM-MSCs, which was confirmed by adipogenic-associated gene expression (PPAR-γ, ADIPOQ; Fig. 2E, F). However, we found that the osteogenic differentiation potential of TS-MSCs was moderately weaker than BM-MSCs according to Alizarin Red S staining and qRT-PCR analysis of the corresponding genes (RUNX2, BGLAP) (Fig. 2G, H). As to chondrogenic differentiation potential, it was found that there were minimal differences between the indicated two MSCs as shown by the Alcian Blue staining and the corresponding gene expression (ACAN, SOX9; Fig. 2I, J). Collectively, TWIST1 overexpression and cocktail stimulation were adequate for high-efficient hESC-MSCs generation with considerable trilineage differentiation potential.
Multifaceted Characteristics of the Variations in Gene Expression Profiling During hESC-MSCs Generation
Having clarified the biological features of TS-MSCs, we are curious about the variations at the transcriptomic level during hESC-MSCs generation. For this purpose, we enriched mRNAs from the initial hESCs (day 0) and the derivations (day 4, day 7, and day 14) during hESC-MSCs induction. Intuitively, we observed the similarity in total gene expression distribution based on fragments per kilobase of transcript per million (FPKM) values (Fig. 3A). According to correlation analysis, we noticed the most distant similarity between hESCs (day 0) and hESC-MSCs (day 14), which was further confirmed by the principal component analysis (PCA) diagram (Fig. 3B, C). Meanwhile, with the aid of Circos diagrams, we noticed a limited number of genes with random variations during hESC-MSCs generation (Fig. 3D).
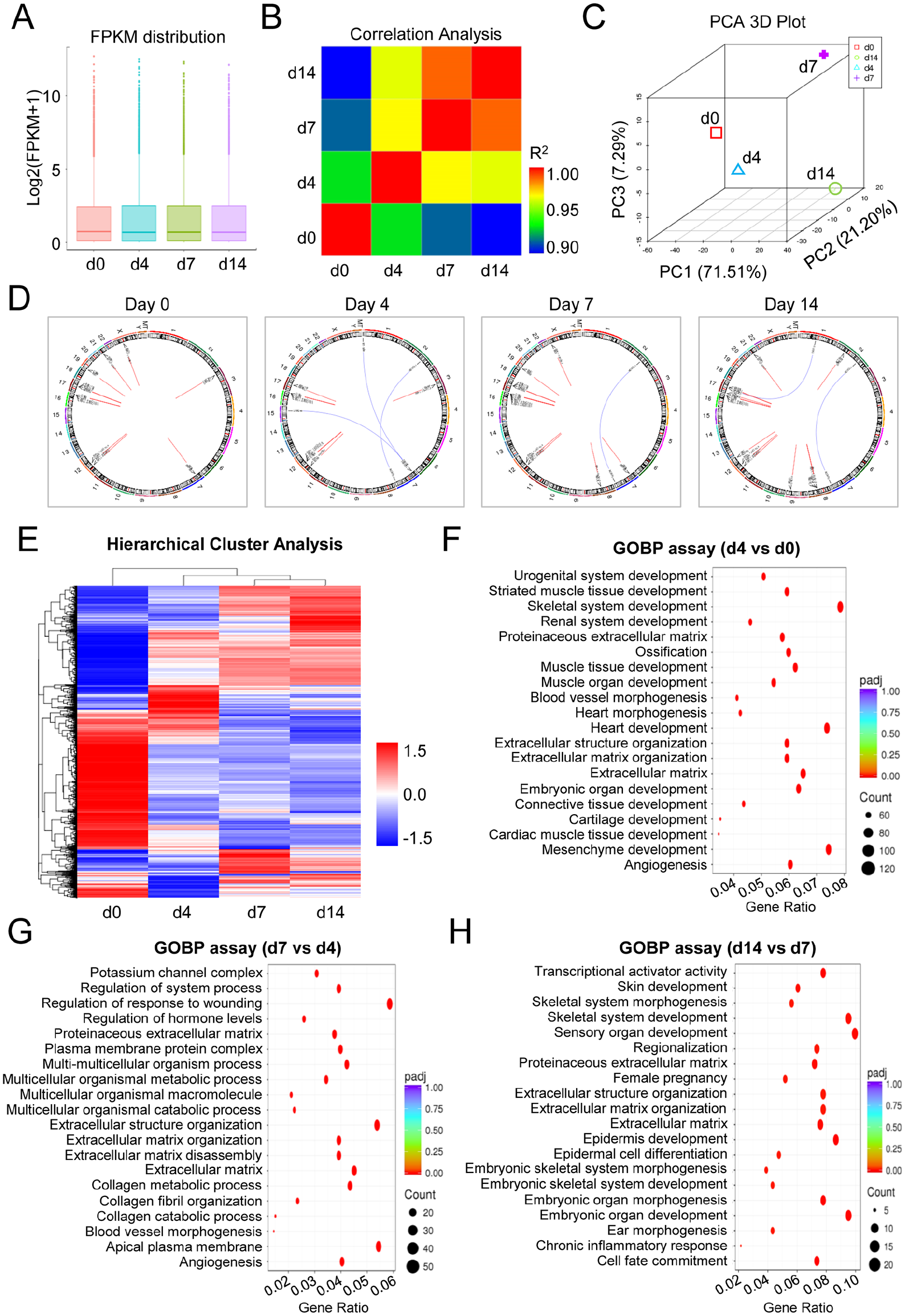
Gene expression profiling of hESC-MSCs during in vitro induction. (A) The distributions of gene expression in H1 hESCs (day 0) and the derivations (day 4, day 7, and day 14) based on log2 (FPKM + 1). (B) Correlation analysis of H1 hESCs and the derivations by HeatMap diagram. (C) PCA of H1 hESCs and the derivations. (D) The variations of the indicated loci regional distributions of gene fusion events and SNP variations were shown by Circos diagrams during TS-MSCs induction. (E) HeatMap diagrams revealed the hierarchical cluster analysis of H1 hESCs and the derivations. (F-H) GO-BP analysis of DEGs between the indicated groups during TS-MSCs induction, including day 4 versus day 0 (F), day 7 versus day 4 (G), and day 14 versus day 7 (H). FPKM: fragments per kilobase of transcript per million; hESCs: human embryonic stem cells; MSCs: mesenchymal stem/stromal cells; TS-MSCs: H1 hESC differentiation towards MSCs by TWIST1-overexpression and small molecular cocktail combination; PCA: principal component analysis; GO-BP: gene ontology–biological processes; DEGs: differentially expressed genes; SNP: single nucleotide polymorphism.
To further verify the potential differences in gene expression profiling, we conducted a hierarchical cluster analysis and found gradual variations in affinity during hESC-MSCs generation (Fig. 3E). Furthermore, with the aid of gene ontology–biological processes (GO-BP) analysis, we observed detailed variations in differentially expressed genes (DEGs) between hESCs and the derivations. As shown by the GO-BP analysis, the DEGs between day 0 and day 4 were mainly involved in mesenchyme development, muscle development, heart morphogenesis, and extracellular matrix (ECM; Fig. 3F). According to the DEGs between day 7 and day 4, we further observed the enrichment of ECM- and collagen-associated biological processes such as ECM disassembly, ECM organization, extracellular structure organization, collagen metabolic process, and collagen catabolic process (Fig. 3G). As to those enriched between day 7 and day 14, the DEGs were principally involved in skeletal development and ECM organization (Fig. 3H). Taken together, these findings revealed the multidimensional similarities and differences in transcriptomic features during hESC-MSCs induction.
TS-MSCs Revealed Comparable Efficacy Upon Mice With ALI
To evaluate the in vitro and in vivo biofunction of TS-MSCs, we measured the secretion of the indicated cytokines in the culture supernatant and the efficacy upon the ALI model, respectively. According to enzyme-linked immunosorbent assay (ELISA) data, we found that TS-MSCs showed minimal differences in IL-6, TGF-β1 and TNF-α secretion compared with BM-MSCs, whereas there was less IL-8 secretion in the supernatant (Fig. 4A). Subsequently, we took advantage of the ALI mouse model to assess the similarities and differences in disease remission. As shown by the survival rate curve, mice with ALI revealed a sharp decline in survival rate, which could be substantially improved with systemic infusion of TS-MSCs or BM-MSCs (Fig. 4B). According to the general morphological score, we further observed the significant remission of ALI mice after TS-MSCs or BM-MSCs administration compared with the ALI group (Fig. 4C). Based on pathological sections, we intuitively observed the infiltration of inflammatory cells in liver tissue of ALI mice, and the pathological manifestations could be dramatically alleviated with TS-MSCs or BM-MSCs infusion (Fig. 4D). Furthermore, with the aid of qRT-PCR analysis, we noticed that the abnormally elevated expression of proinflammatory factors (Il-1β, Tgf-β1) in ALI mice was suppressed after TS-MSCs or BM-MSCs injection, and no significant differences were observed between the two indicated groups (Fig. 4E). Conversely, the declined levels of liver-related factors (Alb, Hgf) in ALI mice were effectively improved after TS-MSCs or BM-MSCs infusion (Fig. 4F). Collectively, the data suggested the feasibility of TS-MSCs for the remission of ALI mice, which thus held promising prospects in disease remodeling.
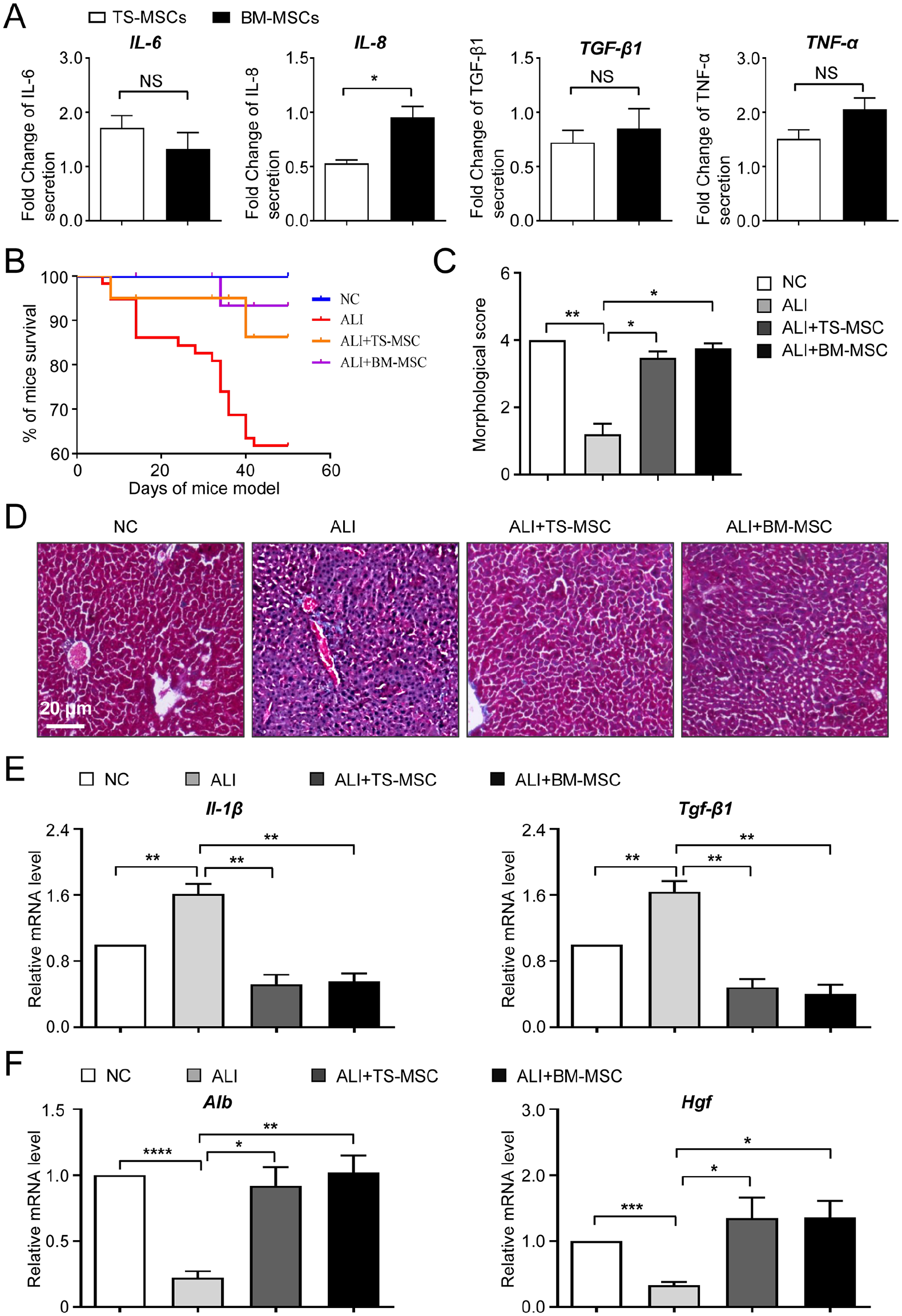
The variation of cytokine secretion and the efficacy upon ALI between TS-MSCs and BM-MSCs. (A) ELISA of representative cytokines and inflammatory factors (IL-6, IL-8, TGF-β1, and TNF-α) in TS-MSCs and BM-MSCs. (B) The overall survival of mice in the indicated groups. (C) The statistical analysis of the morphological scores in the indicated groups. (D) The pathological morphology of liver tissues in the indicated groups by H&E staining. (E-F) The relative mRNA expression levels of the indicated inflammatory factors (E) and liver-associated factors (F) in the liver tissues of mice in the indicated groups. All data were shown as mean ± SEM (n = 4). ALI: acute liver injury; ELISA: enzyme-linked immunosorbent assay; H&E: hematoxylin and eosin; MSCs: mesenchymal stem/stromal cells; BM-MSCs: bone marrow-derived mesenchymal stem/stromal cells; SEM: standard error of the mean; TS-MSCs: H1 hESC differentiation towards MSCs by TWIST1-overexpression and small molecular cocktail combination; NS: not significant. *P < 0.05; **P < 0.01; ****; P < 0.0001.
Discussion
HPSCs are exceptional alternative sources of MSCs for regenerative medicine and disease remodeling. However, the deficiency of procedures for high-efficient hPSC-MSC generation largely hinders the early mesenchymal development and translational research1,2,6. Herein, TWIST1-overexpressing hESCs and small molecular cocktails (DAC and CHIR99021) were used for high-efficient MSCs generation within 14 days. The hESC-MSCs displayed typical cytomorphology and immunophenotypes of MSCs, and comparable tri-lineage differentiation potential with BM-MSCs. With the aid of RNA-SEQ and bioinformatics analyses, the variations in transcriptomic features during hPSC-MSC induction were intuitively observed. In addition, the manifestations of mice with ALI were efficaciously rescued by hPSC-MSC administration. Collectively, the results indicated the feasibility of high-efficient hPSC-MSC generation by TWIST1-based programming with ameliorative effect upon ALI.
TWIST1, short for twist basic helix-loop-helix (bHLH) transcription factor 1, is a well-established regulator during embryogenesis and tumor initiation through orchestrating the epithelial-mesenchymal transition (EMT) process. For instance, current updates have indicated the unappreciated roles of the TWIST1-E-protein heterodimers in guiding lineage differentiation of ESCs and, in particular, early germ layer specialization toward mesoderm and neural crest cells 22 . In consistence, the results revealed the promoting effect of TWIST1-overexpression upon hESCs differentiation toward mesoendoderm and the resultant MSCs according to the indicated lineage biomarker expression. Interestingly, muscle- and myocardium-associated biological process by exogenous TWIST1 expression on day 4 of hESC-MSCs induction was also specifically enriched, which was consistent with the report on smooth muscle cells (SMCs) generation by endothelial TWIST1 expression 23 . Furthermore, we previously identified the programming effect of MSX2-TWIST1 in facilitating hPSC-MSCs generation, and verified the crucial role in mesengenesis by interfering endogenous TWIST1 expression. In detail, the deficiency of hPSC-MSCs programming due to MSX2 knockout was efficiently rescued by TWIST1 overexpression 2 . In this study, the TWIST1-overexpressing hESCs were further used to illuminate the detailed information during MSC induction, including reduced pluripotency, germ layer specification, ECM accumulation, and collagen organization, together with the continuous variations in gene expression profiling. To our knowledge, there is no other literature upon dissecting the programming effect of TWIST1 during hPSC-MSCs. In a word, the findings in this study would benefit the further investigations of early mesengenesis and MSCs development.
State-of-the-art literatures have also highlighted the molecular mechanism underlying MSC induction. For example, Deng and colleagues reported the inhibitory effect of IKK/NF-κB signaling and p65 cascades during hESC differentiation into MSCs 24 . Sánchez et al. 25 and Chen et al. 26 reported the small molecule SB431542-based mesengenic induction of hESCs and hiPSCs to generate MSCs through inhibition of TGF-β–smad2/3 cascades, respectively. Instead, Zhang et al. 2 and Wang et al. 27 verified the role of MSX2 and the BMP signaling in accelerating hESC-MSC induction through the neural crest cell and trophoblast interstage, respectively. Of note, Wei et al. 6 and Zhang et al. 1 took advantage of small molecular library-based screening for high-efficient hPSC-MSCs generation, and identified two types of cocktails, including the JAK/STAT antagonist (JNKi) and the DNA methylation inhibitor (DAC), and the JAK/STAT or BRD4 inhibitors (LLY-507, AZD5153). Herein, with the aid of qRT-PCR and RNA-SEQ analysis, we further confirmed the transient upregulation of EMT factors (eg, SNAI1, SNAI2, ID1, ID2, ZEB1, ZEB2, and MSX2) and the involvement of TGF-β signaling, Wnt signaling, and Hedgehog signaling during hESC-MSCs induction, which would benefit the further investigation of hESC-MSC development and the concomitant molecular mechanisms.
ALI caused by drug overdose (eg, acetaminophen, idiosyncratic drug, and herbal drugs) or inflammatory cytokines is an important and uncommon cause of liver failure and microcirculatory dysfunction, which has caused significant clinical problem in the Western world28,29. Considering the limitations in efficacy and the side effects, pioneering investigators have turned to stem cell–based innovative cytotherapies29,30. For instance, Hu and the colleagues indicated the ameliorative effect of adult MSCs and the derivations (eg, exosomes, extracellular vesicles) upon ALI mice31,32. In this study, we took advantage of the CCl4-inducted ALI mouse model, and verified the feasibility of hESC-MSCs in meliorating pathological manifestations and inflammatory response in ALI mice. In addition, MSC-derived exosomes (MSC-exo) have been recognized as splendid nanovesicles with diverse components (eg, microRNAs, mRNAs, cirRNAs, cytokines, anti-inflammatory factors), which thus play a crucial role for mediating the therapeutic effect of MSCs. Therefore, it would be interesting to explore the feasibility of ALI treatment with hESC-MSC-exo infusion in future33,34. Collectively, these data suggested promising prospects of hESC-MSCs in liver disease management, such as for ALI and the resultant acute liver failure (ALF) in future.
Proverbially, MSCs are heterogeneous cell populations with unique bidirectional immunomodulatory- and hematopoietic-supporting capacities, which thus hold promising prospects in disease management and the concomitant regenerative medicine. Of note, Zhang et al. 35 and Zhao et al. 4 respectively reported the variations in the efficacy upon ALF and acute graft-versus-host disease (aGvHD), which indicated the potential impact of heterogeneity of MSCs in preclinical and clinical practice. Interestingly, more and more investigators put forward the subsets of MSCs with potentially unique bioactivity. For instance, Battula et al. 36 verified that the MSCA-1+CD56+ subset showed superiority over the negative counterparts in differentiation toward chondrocytes and pancreatic-like islets, whereas Wei et al. 37 demonstrated the VCAM-1+ (CD106+) subset with preferable immunomodulatory property and proangiogenic potential over the corresponding negative subset. Collectively, these data indicated the potential advantages of homogeneous subpopulation rather than the heterogeneous counterpart in regenerative medicine. In this study, we turned to TWIST1-based programming for the preparation of homogeneous hESC-MSCs, which would supply new approaches for the further development of large-scale and clinical-grade MSCs in future. Nevertheless, considering the current progress of the experimental efforts to generate such cells from hESCs, there is still a long way to go before clinical grade MSCs for homologous applications when compared with generating MSCs from bone marrow or adipose tissue of a patient or even from umbilical cord.
Supplemental Material
sj-docx-1-cll-10.1177_09636897231218383 – Supplemental material for Multifaceted Characterization of Human Embryonic Stem Cell-Derived Mesenchymal Stem/Stromal Cells Revealed Amelioration of Acute Liver Injury in NOD-SCID Mice
Supplemental material, sj-docx-1-cll-10.1177_09636897231218383 for Multifaceted Characterization of Human Embryonic Stem Cell-Derived Mesenchymal Stem/Stromal Cells Revealed Amelioration of Acute Liver Injury in NOD-SCID Mice by Youlai Zhang, Ying He, Rufei Deng, Zhenyu Jiang, Leisheng Zhang, Yuanlin Zeng and Lijin Zou in Cell Transplantation
Footnotes
Ethical Approval
The study was approved by the ethics committee of Gansu Provincial Hospital according to the guideline of Helsinki (2022-088, 2023-120).
Statement of Human and Animal Rights
This article does not contain any studies with human subjects. Meanwhile, the studies upon ALI mouse model were approved by the ethics committee of Gansu Provincial Hospital according to the guideline of Helsinki (2022-088, 2023-120).
Statement of Informed Consent
Not applicable.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Natural Science Foundation of China (No. 81660364, 81760343, and 82260031) and Gansu Provincial Hospital Intra-Hospital Research Fund Project (22GSSYB-6).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
