Abstract
At present, there are still significant barriers that impede the clinical use of hESCs and iPS cells, including ethics, immunorejection, tumorigenesis from hESCs, and teratoma formation from iPS cells. It is therefore necessary to search for alternative sources of stem cells. WJ-MSCs originate from embryonic epiblasts and possess properties intermediate between hESCs and adult stem cells. However, the stemness properties of molecules in WJ-MSCs remain unclear compared to those of hESCs. In the present study, we isolated WJ-MSCs by a nonenzymatic method. Further, using microarray analysis by Affymetrix GeneChip and functional network analyses, we determined the degree of expression of stemness genes exhibited by the Human Stem Cell Pluripotency array. We also defined a wide range of stem cell gene expression in the WJ-MSCs in comparison with hESCs. At the same time, the definitive markers of early cardiac precursor cells and more committed progenitors were further characterized in WJ-MSCs. Our results demonstrated for the first time that WJ-MSCs had significant expression of undifferentiated human embryonic stem cell core markers, such as SOX2, NANOG, LIN28, SSEA1, SSEA3, SSEA4, KLF4, c-MYC, CRIPTO, and REX1, with a relatively lower level of expression than in hESCs. We also found WJ-MSCs have high expression of early cardiac transcription factors, such as Flk-1, Isl-1, and Nkx2.5. Functional analysis revealed signature genes of WJ-MSCs with specific roles involved in immune, cytoskeletal, and chemokine regulation, cell adhesion, and cell signaling. Our study indicated that there is a significant overlap between the stemness genes expressed by hESCs and WJ-MSCs. WJ-MSCs harbor a true stem cell population and are promising cells for stem cell-based therapies.
Keywords
Introduction
Ischemic heart failure after myocardial infarction is a leading cause of death worldwide. The progressive and irreversible loss of cardiac myocytes due to myocardial infarction is a feature of ischemic cardiomyopathy despite current therapies such as pharmacological treatment, catheter-based therapy, and surgical interventions (44, 55). Recent experimental and clinical studies, however, have opened a new exciting potential therapeutic strategy involving the transplantation of stem cells into the injured heart (39, 41). Many potential cell sources can be utilized, including skeletal myoblasts (31, 43), mesenchymal stem cells (MSCs) (33, 42, 52), and endothelial progenitor cells (4, 21). It has been demonstrated that MSCs have the potential to differentiate into cardiomyocytes both in vitro and in vivo (28, 40, 49, 57, 64). A number of clinical trials have been performed using autologous bone marrow-derived MSCs. However, the results of these trials have been unsatisfactory because of the poor efficiency of differentiation into mature myocardium and a low number of MSCs in older patients with coronary heart disease (2, 3, 17, 27, 31, 32, 47, 59). The ultimate goals of cell therapy are myocardial regeneration and neovascularization, leading to clinical improvement without severe adverse effects. To this end, it is necessary to search for alternative sources of stem cells that have greater proliferation and differentiation potential.
Pluripotent stem cells derived from the inner cell mass of early stage embryos have provided a prototype for multilineage repair. Currently, human embryonic stem cells (hESCs) have been successfully converted in vitro into functional cardiomyocytes, and transfer of such cells into ischemic animal models has shown successful engraftment (22, 24, 62, 63). However, hESC-derived tissue application in the human is fraught with the problems of ethics, immunorejection, and tumorigenesis. These considerations, along with practical limitations, have precluded adoption of embryonic stem cell platforms (38). On the other hand, the recent landmark discovery that mouse and human somatic cells can be reprogrammed to a ground state of pluripotency by ectopic expression of only a few defined transcription factors offers a novel fascinating route to patient-specific pluripotent cells, without the technical and ethical limitations of somatic cell nuclear transfer (29, 30). However, the therapeutic value of reprogramming remains largely unknown because of the problem of teratoma formation and long-term stability of induced pluripotent stem cell (iPSC)-derived tissues. Therefore, iPSCs still have significant problems that impede their adaptation into the clinic (26).
The human umbilical cord (UC) is an extraembryonic formation that constitutes the essential link between the mother and her fetus during pregnancy. UC is composed of two arteries and a vein, all of which are surrounded by a unique connective tissue stroma, namely, Wharton's jelly (WJ). WJ originates from extraembryonic and/or embryonic mesoderm at day 13 of embryonic development (45). WJ surrounds embryonic blood island cells during their migration to the aorta-gonad mesonephros region from the yolk sac region prior to day E 10.5 (8, 45, 53). Several reports have shown that stem cells with pluripotent differentiation potential, extraembryonic mesoderm stem cells, can be isolated from WJ in humans. These cells fulfill most criteria for MSCs, such as adherence to plastic, expression of characteristic surface markers, and differentiation into cells of mesenchymal origin, including bone, cartilage, adipose tissue, and cardiomyocytes (6, 12, 19, 46, 48, 53, 56). In contrast to bone marrow MSCs, Wharton's jelly mesenchymal stem cells (WJ-MSCs) have greater expansion capability, faster growth in vitro, and may synthesize more and different cytokines (11, 18, 60). It has been found that matrix stem cells derived from porcine umbilical cord or human Wharton jelly express markers characterized in embryonic stem cells, such as the transcription factors Nanog, octamer binding transcription factor-4 (Oct-4), sex determining region Y box 2 (Sox-2), and guanine-adenine-thymine-adenine binding protein 4 (GATA-4), GATA-5, and GATA-6 (10, 37). A more recent study showed that WJ-MSCs, functioning as human extraembryonic mesoderm stem cells, express the key early cardiac transcription factors GATA-4, GATA-5, and GATA-6 (25). Especially, it was also demonstrated that WJ-MSCs express connexin-43 of intercellular gap junctions (56). Moreover, WJ-MSCs express genes with immune regulatory properties, which are immunoprivileged (25, 58). Therefore, WJ-MSCs may represent a rich source of primitive cells derived from the extraembryonic tissue and may be a promising cellular source for cardiac stem cell-based therapy (3, 27, 38, 61).
As described above, although it has been demonstrated that WJ-MSCs express many genes like ESCs that are markers of stemness, the difference in molecular properties between WJ-MSCs and hESC has yet to be established. Even more, the similarities and differences in the global gene profiles between WJ-MSCs and hESCs, as well as which common molecules are shared by them, are still not reported. So in this study, first we examined and compared the global gene expression profiles, and asked whether WJ-MSCs share the expression of a set of core stem marker genes with hESCs by using the Human Stem Cell Pluripotency array. Second, we subcategorized them as signature gene, early cardiac transcription factor, immune property gene, and growth factor and receptor genes, in order to identify major genomic differences and unique biological markers specific to the target cell population compared with human embryonic stem cells by using the Rhesus Affymetrix Gene Chip.
Materials and Methods
Tissue Samples and Cellular Isolation Protocol
The work was conducted following the approval of the Navy General Hospital Ethical Review Board. With the consent of the parents, one fresh human female umbilical cord was obtained from a full-term birth with no complications throughout the pregnancy by caesarian section, aseptically stored in sterile saline, and processed within 6 h from partum to obtain the umbilical cord. After removal of blood vessels, the abundant extracellular matrix of Wharton's jelly, which is a mucous tissue continuum from the subamnion to the perivascular region, was scraped off with a scalpel, finely cut, rinsed in sterile phosphate-buffered saline [PBS, Hyclone, Logan, UT, USA; composition (in mM): 140 NaCl; 2 KCl; 1.5 KH2PO4; 15 Na2HPO4], and transported to the laboratory at room temperature. The Wharton's jelly was then serially cut in a cross-sectional manner, and four to five explants of the matrix, ranging from 1 to 2 cm, were placed in 100-mm tissue culture dishes (Corning, Corning, NY, USA) with 2–3 ml of culture medium [Dulbecco's modified Eagle's medium-F12 (DMEM-F12) low-glucose (Sigma-Aldrich, St. Louis, MO, USA), supplemented with 10% fetal bovine serum (FBS; PAA, Linz, Austria), 1× nonessential amino acids (Sigma), 1× antibiotics–antimycotics (Gibco, Gaithersburg, MD, USA), and 2 mM l-glutamine (Sigma)]. The isolation method made no use of proteases to detach cells from the embedding matrix. Therefore, based on the “mesenchymal” migratory capability of cells, cord Wharton's jelly fragments were left in the culture medium for 14 days, with medium change every second day. Cellular exit from the cord Wharton's jelly and attachment to the plastic surface of the tissue culture slide was monitored by phase-contrast microscopy. Finally, after 15 days of culture, the remnants of the cord Wharton's jelly fragments were removed from the wells, and cells that attached to wells were cultured until they reached confluence. A single human female ESC line was obtained from the Shanghai Institutes for Biological Sciences Chinese Academy of Sciences (Shanghai, China). This cell lineage is not commercially available. In brief, the female ESCs were maintained on a mitotically inactivated (mitomycin C) mouse embryonic fibroblast (Shanghai Institutes for Biological Sciences) feeder layer in DMEM-F12 supplemented with 20% knockout serum replacer, 0.1 mmol/L nonessential amino acids, 1 mmol/L l-glutamine (all from Invitrogen, Carlsbad, CA, USA), 0.1 mmol/L β-mercaptoethanol (Sigma), and 4 ng/ml human recombinant basic fibroblast growth factor (Invitrogen).
Cell Culturing and Passaging
After reaching confluence, primary cells were subcultured routinely in culture medium. Cellular detachment from tissue culture dishes was performed using accutase (Invitrogen) instead of standard trypsin solution. Primary populations of WJ-MSCs were cultivated for up to three to six passages. For long-term storage, WJ-MSCs were also cryopreserved in a medium consisting of 93% FBS and 7% dimethyl sulfoxide (Sigma-Aldrich Inc.). For experiments, cells at passage 3 were used in order to ensure the maintenance of marker expression during cell doubling.
Immunophenotype Analysis
Immunophenotyping of the expanded WJ-MSCs was examined using flow cytometry. The adherent cells were washed with PBS and detached by incubating with trypsin/ ethylenediaminetetraacetic acid (EDTA) (Hyclone) for 3 min at 37°C. The harvested cells were washed using staining buffer containing 4% FBS and 0.1% azide (Sigma) in PBS, and approximately 0.05–0.1 × 106 cells per tube was used for cell surface antigen expression studies. The cells were stained with phycoerythrin/fluorescein (FITC)-labeled monoclonal antibodies against human cluster of differentiation 44 (CD44), CD90, CD105, CD73, CD31, CD45, CD34, CD3, CD4, CD8, CD80, CD86, human leukocyte antigen (HLA)-ABC, and HLA-DR (Beijing Boiynthesis Biotechnology Co, Beijing, China) for 45 min at room temperature. The stained cells were run on a flow cytometer (FACSCalibur, Becton-Dickinson, Franklin Lakes, NJ, USA) equipped with a 488-nm argon laser to confirm WJ-MSC purity. Approximately 10,000 events were acquired and analyzed using the CellQuest software (Becton Dickinson). For viability determination, the cells were stained with 7-amino actinomycin D (7-AAD; BD Pharmingen, USA) and acquired on a flow cytometer. The results were analyzed using WIN MDI v 2.8 software (Becton Dickinson).
Differentiation Studies
Osteogenic Potential
For osteogenic differentiation, the third-passage WJ-MSCs were plated in a six-well chamber slide (Corning) with a count of 3,000–10,000 cells per well. After plating the cells in DMEM-F12 supplemented with 10% FBS or 24 h, osteogenesis was induced by replacing with differentiation medium [DMEM-F12 supplemented with 10% FBS, 0.1 μM dexamethasone (Sigma-Aldrich), 10 mM β-glycerophosphate (Sigma-Aldrich), and 50 μM ascorbic acid (Sigma-Aldrich)]. The differentiation medium was changed every 2 to 3 days. After 21 days of differentiation, the cells were checked for calcium deposition with the Von Kossa technique (Sigma).
Chondrogenic Potential
For chondrogenic differentiation, third-passage WJ-MSCs were harvested and differentiated into chondrocytes in pellet cultures at a density of 0.25–0.5 × 106 in a polypropylene tube (Corning) containing chondrogenic differentiation medium, which was DMEM-F12 supplemented with 0.1 μM dexamethasone (Sigma-Aldrich), 10 mM β-glycerophosphate (Sigma-Aldrich), and 50 μM ascorbic acid (Sigma-Aldrich). The tubes were incubated at 37°C in a 5% CO2 incubator. The cells grew as pellets, which were fed with fresh chondrogenic medium every 2 to 3 days. Chondrogenic pellets were harvested after 14 to 28 days in culture. The harvested chondrogenic pellets were fixed with 10% formalin (Sigma-Aldrich) and paraffin-embedded for histological processing. Thin sections, approximately 4–10 μm, were also stained with Safranin O (Sigma-Aldrich).
Adipogenic Potential
For adipogenic differentiation, third-passage WJ-MSCs were plated in a six-well chamber slide with a count of 3,000–10,000 cells per well. Cells were incubated in DMEM-F12 supplemented with 10% FBS, 1 μM dexamethasone, 0.5 mM methylisobutylxanthine, 10 μg/ml insulin, and 100 μM indomethacin for 3 weeks and assessed by Oil Red O staining (Sigma-Aldrich). All inducing agents were procured from Sigma-Aldrich.
Cardiomyogenic Potential
The differentiation of cardiomyogenic (CMG) cells from WJ-MSCs was performed. The third-passage WJ-MSCs were seeded into 35-mm dishes (Corning) at a density of 20,000 cells/cm2 in DMEM-F12 containing 10% FBS and 1% antibiotics (100 U/ml penicillin and 250 ng/ml streptomycin). On the second day, the medium was changed, and the cells were exposed to the medium with 20% FBS or treated for 24 h with the medium consisting of 20% FBS and 5-azacytidine (5-AZ; 3 μmol/L; Sigma-Aldrich). The medium was changed twice a week for 4 weeks or more until the experiment was terminated. Control cells were cultured in DMEM-F12 supplemented with 5% FBS and antibiotics. CMG cells showed spontaneous beating after 5-AZ treatment for 1–2 weeks. 4′,6-Diamidino-2-phenylindole (DAPI; Sigma)-positive cardiomyogenic cells were stained with mouse monoclonal antibodies against sarcomeric α-actin, troponin T (TnT), connexin-43, and GATA-4 at 1:200 dilution (Beijing Biosynthesis Biotechnology Co., Ltd.) and FITC-conjugated secondary antibody (Beijing ZGB, Biotechnology Co., Ltd., Beijing, China; 1:100).
Microarray Experiment
RNA Extraction From WJ-MSCs and ESCs
Total RNA was extracted from three different samples each of third-passage WJ-MSCs and hESCs using Trizol reagent (Sigma-Aldrich). RNA was further purified using Qiagen RNeasy Mini Kit according to the manufacturer's instructions (Qiagen, Germantown, MD, USA). RNA quality was assessed by formaldehyde agarose gel (Sigma-Aldrich) electrophoresis (Bio-Rad, Hercules, CA, USA) and was quantitated spectrophotometrically (Thermo Scientific, Waltham, MA, USA).
Microarray Analysis
An aliquot of 2 μg of total RNA was used to synthesize double-stranded cDNA, then produce biotin-tagged cRNA using the MessageAmpTM II aRNA Amplification Kit (Takara, Shiga, Japan). The resulting bio-tagged cRNA were fragmented into strands of 35–200 bases in length according to the protocols from Affymetrix (Santa Clara, CA, USA). The fragmented cRNA was hybridized to a Human Genome U133 Plus 2.0 array (Affymetrix), containing 47,000 transcripts. Hybridization was performed at 45°C with rotation for 16 h (Affymetrix GeneChip Hybridization Oven 640). The GeneChip arrays were washed and then stained (streptavidin–phycoerythrin) on an Affymetrix Fluidics Station 450, followed by scanning on a GeneChip Scanner 3000 (Affymetrix). All samples were prepared in three biological repeats.
Normalization and Data Analysis
The hybridization data were analyzed using GeneChip Operating software (GCOS 1.4; Affymetrix). The scanned images were first assessed by visual inspection, then analyzed to generate raw data files saved as CEL files using the default setting of GCOS 1.4. An invariant set normalization procedure was performed to normalize the different arrays using a DNA-chip analyzer (dChip; Affymetrix). In a comparison analysis, 13,392 genes were determined to be significantly differentially expressed with a selection threshold of ratio >2.0 or ratio <0.5 in the output result.
Real-Time Quantitative Polymerase Chain Reaction Analysis
Selected genes, including embryonic stem cell marker genes and early cardiovascular-specific transcription factor genes from microarray experiments, were further confirmed by qRT-PCR (Table 1). Total RNA was extracted from three different samples each of WJ-MSCs and hESCs. Total RNA was reversed-transcribed using a QuantiTect Reverse Transcription kit (Applied Biosystems, Foster City, CA, USA). The qRT-PCR reactions were performed with Platinum SYBR Green qPCR superMix UDG (Invitrogen) using gene-specific primers (listed in Table 1; Sangon, Shanghai, China) following the manufacturer's instructions. PCR was performed in the Bio-Rad DNA Engine Opticon-2 (Bio-Rad, CA, USA) under the following conditions: 50°C for 2 min, 95°C for 10 min, 40 cycles (15 s at 95°C, 1 min at 60°C). Target genes were assayed in triplicate on each plate. Glyceraldehyde phosphate dehydrogenase (GAPDH) was used as a housekeeping gene to normalize, and then it was quantified in WJ-MSCs relative to the expression level in ESCs, after running dilution standards for every assay to ensure the ratio of the differences in cycle thresholds (2–ΔΔCt) was equivalent to the fold change calculated from the standard curve.
Primers Used in Real-Time RT-PCR for Stemness Molecular Markers and Early Cardiac Transcription Factors Used in the Analysis of WJ-MSCs and hESCs

WJ-MSCs, Wharton's jelly mesenchymal stem cells; hESCs, human embryonic stem cells; OCT4, octamer binding transcription factor-4; SOX2, sex determining region Y box 2; PODXL, podocalyxin-like; SMAD3, mothers against decapentaplegic homolog 3; KDR, kinase insert domain receptor; NKX2.5, NK2 homeobox 5; ISL1, islet-1; KIT, v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog; APJ, apelin receptor; APLN, apelin; THY1, thymocyte cell surface antigen; GAPDH, glyceraldehyde 3-phosphate dehydrogenase.
Statistical Analysis
Data were presented as means ± standard error of the mean. Statistical comparisons were performed using either the Student's two-tailed t test (unpaired/paired) or the Wilcoxon matched pairs test as appropriate according to data distribution. Values of p < 0.05 were considered significant. Data analysis and graphical representations were performed by using GraphPad Prism 5 software (GraphPad, San Diego, CA, USA). The differentially expressed genes were mapped to Gene Ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways based on the human accession ID, using MAS (molecule annotation system, http://bioinfo.capitalbio.com/mas) platform. GO terms and KEGG pathways with false discovery rate (FDR)-corrected values of p < 0.05 were considered statistically significant.
Results
Morphology and Immunophenotypic Characterization
WJ-MSCs isolated from human Wharton's jelly with phase-contrast microscopy displays a uniform spindleshaped morphology similar to fibroblastoid cells (Fig. 1A). In vitro differentiation analysis confirmed that WJ-MSCs exhibited the capacity to differentiate into various cell types, such as osteoblasts, chondrocytes, and adipocytes (Fig. 1B–D).

Morphology and multilineage differentiation capacity of WJ-MSCs. (A) Wharton jelly-derived mesenchymal stem cells (WJ-MSCs) showed a homogeneous spindle-shaped morphology. (B) Osteogenesis was analyzed by von Kossa staining for mineral nodule deposition. (C) Chondrogenesis was assessed by Safranin O staining for proteoglycan deposition. (D) Adipogenesis was observed by the presence of lipid vesicles and confirmed by oil red O staining.
The population doubling time of WJ-MSCs at passages 5–6 was 30–36 h in this study. For further characterization of WJ-MSCs, a panel of surface markers was tested using flow cytometric analysis. WJ-MSCs were negative for CD31 (endothelial cell marker), CD34, CD45 (both as hematopoietic markers), HLA-DR (human leukocyte antigen class II), and CD3, CD4, CD8, CD80, and CD86, whereas they were positive for CD90, CD44 (both as adhesion markers), CD105, CD73 (both as mesenchymal markers), and HLA-A, -B, and -C (class I) (Fig. 2).

Immunophenotype of WJ-MSCs by flow cytometric analysis. Representative histograms are demonstrated. The WJ-MSCs were positive for cluster of differentiation 44 (CD44), CD73, CD90, CD105, and human leukocyte antigen (HLA)-A, -B, and -C and negative for CD31, CD34, CD45, CD3, CD4, CD8, CD80, CD86, and HLA-DR.
To confirm the in vitro capacity of WJ-MSCs to differentiate into cardiomyocytes, 2 weeks after 5-AZ treatment, we examined the expression of several cardiac myocyte markers. DAPI staining was used to identify WJ-MSCs (blue nucleus with multiple nucleoi). These cells stained positive for the cardiac specific proteins sarcomeric α-actin, TnT, the cardiac specific transcription factor GATA-4, and the intercellular connection and electrical coupling factor connexin-43 (Fig. 3).

Confocal microscopy of fluorescent immunohistochemical staining of DAPI-labeled WJ-MSCs induced by 5-AZ after 2 weeks. Expression of (A) α-actin, (B) troponin T, (C) GATA-4, and (D) connexin-43. Nuclei identified by DAPI (a), all proteins— red fluorescence (b), merged image (c). 5-AZ, 5-azacytidine.
Similarities and Differentiation of Gene Expression Profiles Between WJ-MSCs and hESCs
Using high-density oligonucleotide microarrays, we analyzed the global gene expression of WJ-MSCs and hESCs. Further comparison of gene expression in WJ-MSCs and hESCs indicated that 7,417 genes showed a twofold difference in expression. Of these, 2,951 were twofold or higher in WJ-MSCs than hESCs. The other 4,466 genes were downregulated in WJ-MSCs compared with ESCs. To generate the list of similar and differentially expressed genes, a fold-change ranking method was used to perform significance analysis of microarrays (54). After filtering with p < 0.05, we ranked genes by no fold changes and chose the top 50 genes that were expressed similarly between WJ-MSCs and hESCs as shown in Figure 4. Molecular function, included major similar genes involved in ribosomal structural constituents (34 genes), protein binding (31 genes), RNA binding (19 genes), and oxidoreductase activity (7 genes). The top biological process associated with genes in this network, included those involvement in translational elongation, ribosomal RNA (rRNA) processing, ribosomal small subunit synthesis, mitochondrial electron transport, nicotinamide adenine dinucleotide (NADH)–ubiquinone interaction, ribosome biogenesis, assembly, and translational initiation. (Please contact the authors if you wish to view the functional network plots of this from the MetaCore program.)

Heat map of top 50 of common genes expressed by human WJ-MSCs and ESCs. Log2 expression levels are shown in a green-black-red gradient. ESCs, embryonic stem cells.
After filtering with p < 0.05, we ranked genes by twofold changes and chose the top 25 genes that were either upregulated or downregulated, respectively, in WJ-MSCs and hESCs as shown in Tables 2 and 3. Upregulated genes were involved in molecular functions, including protein binding, sequence-specific DNA binding, transcription factor activity, collagen binding, extracellular matrix structural constituent formation, platelet-derived growth factor binding, and ligand regulated transcription factor activity. The biological processes associated with genes included integrin-mediated signaling pathways, cell-matrix adhesion, anterior/posterior pattern formation, peptide cross-linking, collagen fibril organization, cell adhesion, development, and T-cell secretory granule organization. (Please contact the authors if you wish to view the functional network plots of this from the MetaCore program.)
Top 25 Genes With Greater Than Twofold Upregulation in hWJ-MSCs Than hESCs

The signal intensity on the microarray is reported. Significantly differentially expressed with a selection threshold of ratio >2.0 or ratio <0.5 in the output result.
Top 25 Genes With Lower Than 2.5-Fold Downregulation in WJ-MSCs Than ESCs
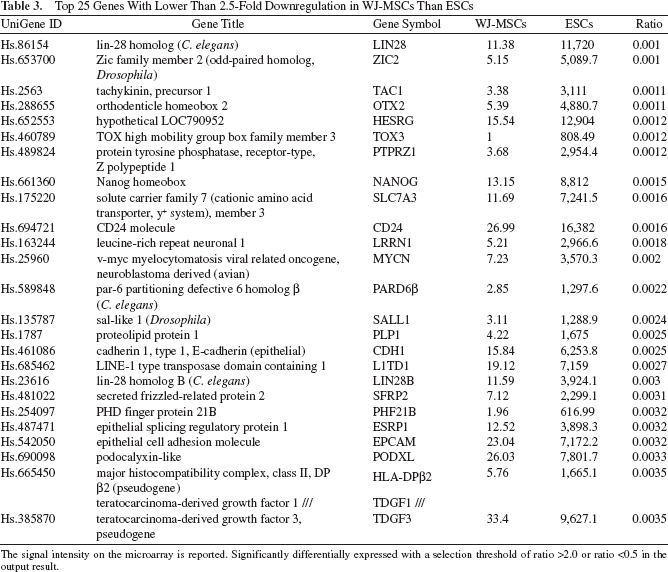
The signal intensity on the microarray is reported. Significantly differentially expressed with a selection threshold of ratio >2.0 or ratio <0.5 in the output result.
Downregulated genes were involved in molecular functions, including zinc ion binding, transcription factor activity, metal ion binding, L-lysine transporter activity, and cationic amino acid transporter activity. The biological process associated with genes in the network included regulation of transcription, pre-B-cell differentiation, positive regulation of B-cell apoptosis, positive regulation of T-cell homeostatic proliferation, negative regulation of T-cell homeostatic proliferation, and B-cell receptor transport into membrane rafts. (Please contact the authors if you wish to view the functional network plots of this from the MetaCore program.)
Validation of Microarray Results with Real-Time Quantitative PCR
To verify and add to the array results, pivotal genes from microarray analysis that have been implicated in the stemness and early cardiovascular development were selected and subjected to quantitative RT-PCR. The genes selected were octamer binding transcription factor-4 (OCT4), sex determining region Y box 2 (SOX2), NANOG, podocalyxin-like (PODXL), mothers against decapentaplegic homolog 3 (SMAD3), Brachyury, kinase insert domain receptor (KDR), NK2 homeobox 5 (NKX2.5), islet-1 (ISL1), v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog (KIT), apelin receptor (APJ), apelin (APLN), and thymocyte cell surface antigen (THY1). The mean (±SEM) relative expression (n = 4 sample each) of each of these genes, as compared with the expression by quantitative PCR, is shown in Figure 5, which confirmed the good correlation between Affymetrix gene expression results and real-time quantitative PCR.
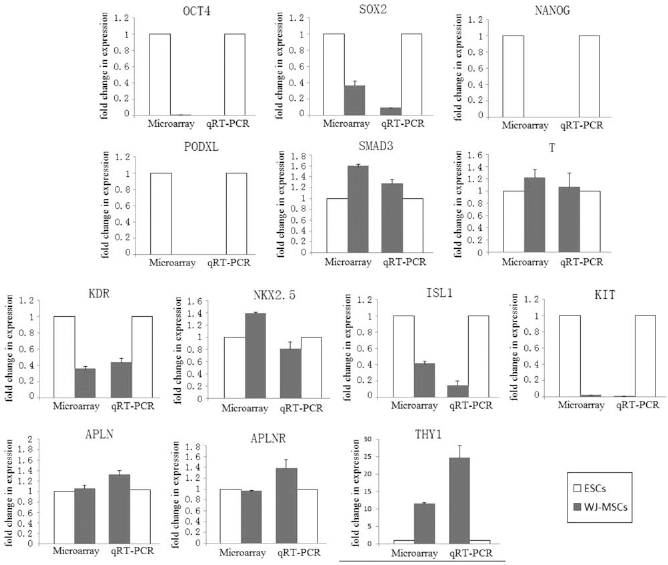
Comparison of gene expression by microarray and qRT-PCR. Comparisons of octamer binding transcription factor-4 (OCT4), sex determining region Y box 2 (SOX2), NANOG, podocalyxin-like (PODXL), mothers against decapentaplegic homolog 3 (SMAD3), Brachyury, kinase insert domain receptor (KDR), NK2 homeobox 5 (NKX2.5), islet-1 (ISL1), v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog (KIT), apelin receptor (APJ), apelin (APLN), and thymocyte cell surface antigen (THY1). Gene expression patterns as determined by microarray and qRT-PCR analysis. The x-axis indicates the WJ-MSCs and control hESCs, and the y-axis represents fold changes determined by microarray and qRT-PCR analysis.
Core Gene Expression of Stemness in WJ-MSCs, Compared with hESC Controls
The International Stem Cell initiative (ISCI) characterized 59 human embryonic stem cell lines from 17 laboratories worldwide. Accordingly, we compared the expression difference of the stemness genes between WJ-MSC and hESC samples (Table 4). WJ-MSCs expressed most of the stemness marker genes, but most of these were expressed at lower levels than in hESCs. However, some key core genes such as thiosulfate sulfurtransferase [TST or stage-specific embryonic antigen 3 (SSEA3)], transducing-like enhancer of split 1 [TLE1 or enhancer of split groucho-like protein 1 (ESG1)], HLA-E, HLA-B, HLA-C, HLA-G, and HLA-A were higher than in hESCs (Table 4).
Core Gene Expression of Stemness in WJ-MSCs, Compared With hESCs

The signal intensity on the microarray is reported. Significantly differentially expressed with a selection threshold of ratio >2.0 or ratio <0.5 in the output result.
Expression of Early Cardiac Transcription Factor Genes in WJ-MSCs, Compared with hESC Controls
As shown in Table 5, most of the early cardiovascular progenitor genes were expressed in WJ-MSCs. Moreover, some key cardiac transcription factors, such as T box 3 (TBX3), TBX2, TBX20, chemokine (C-X-C motif) receptor 5 (CXCR5), NKX2-5, MEF2C (myocyte enhancer factor 2C), TBX1, KITLG (stem cell factor; SCF), CXCR3, TBX4, CXCR6, TBX5, TBX6, TBX19, TBX10, and PECAM1 (platelet/endothelial cell adhesion molecule 1; CD31), were expressed at higher levels than in hESCs.
Expression of Early Cardiac Transcription Factor Genes in WJ-MSCs, Compared With hESCs

The signal intensity on the microarray is reported. Significantly differentially expressed with a selection threshold of ratio >2.0 or ratio <0.5 in the output result.
Functional Analysis of Signature Gene Expression in WJ-MSCs, Compared with hESC Controls
To generate the list of differentially expressed signature genes, a fold-change ranking method was done as described above. Similarities in pathways affected ribosomes, adherens junctions, focal adhesion, and tight junctions in WJ-MSCs and hESCs. However, upregulation of pathways was involved in extracellular matrix (ECM)–receptor interaction, nicotinate and nicotinamide metabolism, focal adhesion, TGF-β signaling pathway, and pyrimidine metabolism in WJ-MSCs compared with hESCs. Downregulation in the pathway was involved in endometrial cancer, hedgehog signaling pathway, adherens junction, cell adhesion molecules (CAMs), tight junction, and Wingless-type mouse mammary tumor virus (MMTV) integration site family (Wnt) signaling pathway in WJ-MSCs compared with hESCs. (Please contact the authors if you wish to view the functional network plots of this from the MetaCore program.)
Functional Analysis of Immune Property Gene Expression in WJ-MSCs, Compared with hESC Controls
In this study, we demonstrated greater expression of genes that regulate immune properties in WJ-MSCs than in hESCs, as shown in Table 6, such as: 5′-nucleotidase, ecto (CD73), endoglin, Thy-1 cell surface antigen (Thy-1), CD44 molecule, membrane metallo-endopeptidase (MME), CD14 molecule, major histocompatibility complex class I, G (HLA-G), selectin P, and integrin (granule membrane protein antigen CD62). These data indicate that a proper immune characterization of WJ-MSC lines should be based on a more extended panel of marker genes, spanning HLA typing to costimulatory molecule expression.
Expression of Immune Property Genes in WJ-MSCs, Compared With hESCs

The signal intensity on the microarray is reported. Significantly differentially expressed with a selection threshold of ratio >2.0 or ratio <0.5 in the output result.
Functional Analysis of Growth Factors and Receptor Gene Expression in WJ-MSCs, Compared with hESC Controls
WJ-MSCs have been shown to express a broad spectrum of cytokines and growth factor genes as shown in Figure 6. After ranking genes by twofold or more changes, we found that ACTA2 (actin, a2, smooth muscle, aorta), TGFI, IL-6 (interleukin 6), VEGFC (vascular endothelial growth factor C), IL-1B, IL-7, TGF-β2, HRAS (v-Ha-ras Harvey rat sarcoma viral oncogene homolog), NGF (nerve growth factor), HGF (hepatocyte growth factor), BMP6 (bone morphogenetic protein 6), MMP1 (matrix metallopeptidase 1), IL-8, PDGFC (platelet-derived growth factor C), HIF1A (hypoxia inducible factor 1, a subunit), BMP4, IFNb1 (interferon, β1), FGF7 (fibroblast growth factor 7), (keratinocyte growth factor) KGF (IL)15, and THOP1 (thimet oligopeptidase 1) exhibited much higher expression in WJ-MSCs than in hESCs.

Heat map of the expression of growth factors and receptor gene in WJ-MSCs, compared with hESCs. Log2 expression levels are shown in a green–black–red gradient.
Discussion
WJ-MSCs originate from embryonic epiblasts and possess properties between hESCs and adult stem cells, and are thought to be more primitive than MSCs from any other adult tissue sources (8, 45, 53, 56). They represent an important and rich source of primitive cells and are now considered as an alternative source of stem cells (8, 45, 53, 56). However, the properties of stemness molecules in WJ-MSCs, compared with that in hESCs, have not previously been clarified. Furthermore, both similarities and differences in global gene profiles between WJ-MSCs to hESCs were not previously established. In the present study, we isolated WJ-MSCs by a nonenzymatic method. WJ-MSCs express a set of core mesenchymal stem cell markers and are able to differentiate into osteoblasts, chondrocytes, and adipocytes. Further, using microarray analysis by Affymetrix GeneChip and functional network analyses, we determined the degree of expression of stemness genes exhibited by the Human Stem Cell Pluripotency array, containing markers of characterized undifferentiated stem cells. We also defined a wide range of stem cell gene expression in the WJ-MSCs and compared the degree of expression with hESCs. At same time, the definitive markers of early cardiac precursor cells and more committed progenitors were further characterized in WJ-MSCs.
Our results demonstrated for the first time that WJ-MSCs had significant expression of the undifferentiated human embryonic stem cell core markers, such as SOX2, NANOG, LIN28, SSEA1, SSEA3, SSEA4, KLF4, c-MYC, TDGF1, and ZFP42, with a relatively lower level of expression than in hESCs. It is also a surprise that we found that WJ-MSCs have high expression of early cardiac transcription factors, such as KDR, islet 1 (Isl-1), and Nkx2.5, some of which exhibit higher levels of expression in WJ-MSCs than in hESCs. Functional analysis revealed the signature gene of WJ-MSCs with specific roles involved in immune regulation, cytoskeletal regulation, chemokine regulation, cell adhesion, and cell signaling.
The key characteristics of stem cells are long-term self-renewal and a capacity to differentiate into diverse mature tissues, which favor their use in regenerative medicine. Human MSCs have been isolated from bone marrow, adipose tissues, cord blood, amniotic fluid, amniotic membrane, placenta, umbilical cord tissues, and Wharton's Jelly of the umbilical cord (18, 28, 40, 49, 54, 57, 64). Tsai et al., using microarray analysis of MSCs derived from four different origins including amniotic fluid, amniotic membrane, cord blood, and bone marrow, found that within each group of MSCs from the same origin, the variability of the gene expression levels was smaller than that between groups of different origins, and functional genomic studies revealed the specific roles of MSCs from different origins (48, 54). However, so far there have been no reports on the global profile of genes in WJ-MSCs and their functional analysis, and no comparison with hESCs. In the present study, we used high-density oligonucleotide microarrays, Human Genome U133 Plus 2.0 array containing 47,000 transcripts, and functional network analyses. The Affymetrix microarray data were analyzed with GeneSifter software. There were 7,417 genes that showed a twofold or greater difference in expression between the WJ-MSCs and hESCs. Of these, 2,951 were twofold or higher in WJ-MSCs than in ESCs. The remaining 4,466 genes were downregulated in WJ-MSCs compared with ESCs. After filtering with a threshold of p < 0.05, we ranked genes by the number of fold changes and chose the top 50 genes with similar expression between WJ-MSCs and hESCs. These genes are involved in molecular functions, such as coding for structural ribosome constituents, protein binding, RNA binding, oxidoreductase activity, and nucleotide binding.
The ISCI was established by the International Stem Cell Forum to carry out a comparative study of a large and diverse set of hES cell lines derived and maintained in different laboratories worldwide. The present ISCI study analyzed 59 independently derived hES cell lines from 17 laboratories in 11 countries. All lines exhibited similar expression patterns for several markers of human embryonic stem cells (51). In our study, WJ-MSCs were isolated from Wharton's jelly in an indistinct region of umbilical cord, which is a continuum from the subamnion to the perivascular region. In contrast to the gene markers established by ISCI, we found that WJ-MSCs expressed the majority of stemness core markers, such as NANOG, SOX2, and LIN28, which notably maintain the undifferentiated ES cell state, although the levels were lower than the signal intensity reported on the microarray. Moreover, the group of genes whose expression correlated closely with that of NANOG and appear to be characteristic of the hES cell, such as MYC, TDGF1, growth differentiation factor 3 (GDF3), γ-aminobutyric acid (GABA) A receptor, β3 (GABRB3), and THY1, are also expressed by WJ-MSCs. To our surprise, we found that SSEA1, SSEA3, and SSEA4, which were another common set of markers generally to monitor the presence of pluripotent stem cells set up by ISCI, were highly expressed in WJ-MSCs, in comparison to the hESCs. The findings agree with recent reports in which it was found that MSCs derived from porcine or human Wharton's jelly of umbilical cord expressed some embryonic stem cells marker genes such as OCT-4, GATA4, KIT, Laminin, γ1 (LAMC1), LIN28, Noggin, nuclear receptor subfamily 6, group A, member 1 (NR6A1), POU5F1, runt-related transcription factor 2 (RUNX2), and SOX17 (10, 15). Therefore, these data have demonstrated that WJ-MSCs have properties of primitive pluripotent stem cells and have undergone an unprecedented characterization process in our studies that further extend the similarities with respect to hESCs. The fact that WJ-MSCs remain behind and coexist with the hESCs suggests that it is possible that, during development, cells in the Wharton's jelly migrate into the fetus along with the primordial germ cells and hematopoietic cells to the aorta-gonad mesonephros (AGM) region (8, 53). Moreover, in our study, WJ-MSCs express CXCR3, CXCR4, and CXCR6, which are the chemokine and surface receptors for function of the migration and homing ability. Thus, the WJ-MSCs express some genes characteristic of primitive stem cells including embryonic stem cells.
The most important finding in this study was that prominent functional genes that may be involved in cardiovascular formation and regeneration are expressed in WJ-MSCs. We found higher expression of mesoderm and specialized cardiac progenitor cell genes, including Brachyury T, mesoderm posterior 1 (Mesp1), KDR, Nkx2.5, and Isl-1. We also demonstrated that WJ-MSCs express GATA-4 and connexin-43, which are typically expressed in embryonic and myocardial cells. Indeed, our in vitro study showed that after 2 weeks of treatment with the DNA demethylation agent 5-azacytine, the WJ-MSCs differentiated into cardiomyocyte-like cells as shown in Figure 3. Immunostaining showed that differentiated cells were strongly positive for cardiac α-actin, TnT, connxin43, and GATA-4.
At present, some studies have demonstrated that the formation of the mature multichambered heart requires the contribution of diverse cell types with specialized function, including cardiomyocytes, endothelial cells, and vascular smooth muscle cells (20, 50, 61). Precursors for heart-forming cells in the vertebrate mesoderm transition from expressing Brachyury T to expressing mesoderm posterior 1 (Mesp1) when they enter the precardiac mesoderm stage of development (61). During Mesp1 cell migration, cardiac progenitor cells expand rapidly to form the anterior and lateral plate mesoderm, where they generate a crescent-shaped structure called the cardiac crescent (61). In the cardiac crescent stage, the heart precursor cells commit to the cardiac lineage and become cardiac progenitor cells expressing key developmental transcription factors such as KDR, Nkx2.5, and Isl-1 (7, 20, 50, 61). Studies have identified that a diverse set of human fetal ISL-1+ cardiovascular progenitors can give rise to the cardiomyocyte, smooth muscle and endothelial cell lineages (7, 20, 50). KDR is a cardiovascular progenitor that represents one of the earliest stages in mesoderm specification to the cardiovascular lineage (20, 61). Yang et al. found that KDR+ embryonic stem cell-derived population can differentiate into cardiac, endothelial, and vascular smooth muscle. When plated in monolayer cultures, KDR+ cells differentiate to generate populations consisting of greater than 50% contracting cardiomyocytes (62). Nkx2.5 is a key transcription factor in cardiac development. It has been shown that cells in the first heart field, marked by Nkx2.5, give rise to the left ventricle and portions of the right and left atrium (7, 20, 50, 61). Moretti's study found the ISL1+/NKX2.5+/KDR+ mesodermal progenitors can effectively induce cardiovascular cell types (34). Therefore, a subpopulation of WJ-MSCs, as KDR+/ Isl-1+/Nkx2.5+ WJ-MSCs, may hold tremendous promise for cardiovascular regeneration medicine.
Recent studies found that the therapeutic capacity of MSCs to treat a wide spectrum of diseases has been attributed to their potential to differentiate into many different reparative cell types (33, 42, 52); however, the efficiency of transplanted MSCs to differentiate into functional reparative cells in the injured tissues or organs and the therapeutically relevant numbers have never been adequately documented or demonstrated (13, 16). Recent reports have suggested that some of these reparative effects are not mediated by differentiation of MSCs but rather by paracrine factors secreted by MSCs (9, 23). At present, our study found that WJ-MSCs express huge number of cytokines, chemokines, and growth factors at the transcript level by using a high-throughput genome-wide gene expression assay. The predominance of cytokines and chemokines such as TGF-β, VEGF, HGF, BMP, PDGF, FGF, MMP, and interleukins (IL-1–IL-17), which are involved in important signaling pathways in cardiovascular biology, bone development, and hematopoiesis, the majority of which have already been implicated in many aspects of cardiovascular, musculoskeletal, and hematopoiesis biology. For example, TGF-β is critical for correct heart development cardiac remodeling, progression to heart failure, and vascularization (5, 14). Janus kinase 3–signal transducer and activator of transcription 3 (JAK-STAT) signaling is associated with cardioprotection (36). Therefore, our present study also supports the notion that WJ-MSCs mediate paracrine effects for cardiac repair and regeneration.
On the other hand, we also found that WJ-MSCs exhibit properties of gene profiles unique to embryonic stem cells. Thus, these cells expressed a set of molecules, such as HLA-G and class 1 HLA molecules, but not class 2 HLA molecules, which support their ability to potentially induce immune tolerance (58). Especially in WJ-MSCs, the expression of HLA-G, which is one the main molecules responsible for induction of tolerance of natural killer (NK) cells toward self-cells, as well as in the process of tolerance of the mother's immune system toward the semiallogeneic embryo, is a key feature of the allogenesis stem cell transplantation process (35).
Conclusion
WJ-MSCs, as the dominant cells of extraembryonic derived tissue, share several features of gene expression compared to embryonic stem cells. They express in common a set of core stemness gene markers with hESCs, albeit in low levels. They also share the expression of early cardiac transcription factors, at levels even higher than hESCs. Like hESCs, they express CXCR3 and CXCR4 and so are likely to possess migratory and homing ability. Moreover, WJ-MSCs express huge numbers of cyokines, growth factors, and chemokines similar to hESCs. However, WJ-MSCs exhibit a gene profile unique to embryonic stem cells, in that they express HLA-G and class 1 HLA molecules and lack class 2 HLA molecules. It is therefore important to consider the benefits of WJ-MSCs, which are not ethically sensitive, have differentiation potential, and do not have the worrying issue of teratoma formation. Thus, WJ-MSCs constitute a true stem cell population and are promising cells for stem cell-based therapies.
Footnotes
Acknowledgments
This study was supported by a grant from the National Advanced Technology Development Plan of China (863 plan) (2006AA02Z469 and 2011AA020109) and the National Natural Science Foundation of China (81170094). We thank Dr. Joel S. Karliner, Professor of Medicine, University of California San Francisco, and Staff Cardiologist, Department of Veterans Affairs Medical Center, San Francisco, CA, USA, for valuable comments and helpful suggestions in writing and revising this manuscript. The authors declare no conflict of interest.
