Abstract
Adipose-derived mesenchymal stem cells (ADSCs) are ideal sources for the treatment of diabetes, and the differentiation of ADSCs into insulin-producing cells (IPCs) through transfection of exogenous regulatory genes in vitro has been studied in depth. The differentiation of ADSCs is strictly regulated by a variety of transcription factors such as Pdx1, Ngn3, Pax4, Nkx2.2, and Sox9. However, whether these genes can coordinately regulate the differentiation of ADSCs into IPCs is still unknown. In this study, five multigene coexpressing adenovirus vectors (pAdTrack-Pdx1-Ngn3-AdEasy, pAdTrack-Pdx1-Ngn3-Sox9-AdEasy, pAdTrack-Pdx1-Ngn3-Pax4-Sox9-AdEasy, pAdTrack-Pdx1-Ngn3-Nkx2.2-Sox9-AdEasy, and pAdTrack-Pdx1-Ngn3-Nkx2.2-Pax4-AdEasy) were constructed, and then the stocks of the packaged adenoviruses were used to infect the canine ADSCs (cADSCs). Based on results of morphological observation, dithizone staining, sugar-stimulated insulin secretion test, cellular insulin immunofluorescence assays, and the detection of pancreatic β-cell development–related genes in the induced cells, the best induction combination (pAdTrack-Pdx1-Ngn3-Nkx2.2-Pax4-AdEasy) was identified after comparative screening. This study provides a theoretical reference and an experimental basis for further research on stem cell replacement therapy for diabetes.
Keywords
Introduction
Diabetes is one of the most common metabolic diseases in the world. In addition to disturbing people’s daily life, diabetes occurs more often in dogs and cats. According to statistics, the incidence of diabetes in dogs and cats reaches about 0.4% to 1.2%, and the incidence of diabetes in dogs has reached 6% of the total number of clinical cases1,2. Traditional drug therapy and insulin injection therapy cannot treat diabetes dynamically and fundamentally3–5. Islet transplantation can effectively control blood glucose changes and avoid complications, but its clinical application is limited due to donor deficiency and immune rejection 6 . As reported, pluripotent stem cells can be induced to differentiate into cells with insulin-secreting function through targeted induction, which can provide a new method for the treatment of diabetes7,8. Adipose-derived mesenchymal stem cells (ADSCs) are ideal sources for the treatment of diabetes because they are derived from adipose tissue, easy to isolate and culture, and involve fewer medical ethical issues 9 .
Currently, the differentiation of ADSCs into insulin-producing cells (IPCs) in vitro has been studied in depth, and optimistic progress has been made 10 . In current research, transferring one to three exogenous regulatory genes cannot completely activate the cascade regulatory system of mesenchymal stem cells (MSCs) and is not sufficient to promote the differentiation of MSCs into pancreatic β-cells11–13. Moreover, this method needs to be greatly improved in terms of the induction efficiency, IPC maturity, and insulin secretion level. Therefore, screening and optimizing a high-efficiency regulatory gene combination, which can reprogram ADSC differentiation into cells that are sensitive to changes in sugar concentration and whose insulin content and release are comparable to normal pancreatic β-cells, are of great significance for the study of cell transplantation treatment in diabetes.
The differentiation of ADSCs is strictly regulated by a variety of transcription factors. For example, Pdx1 can activate the regulation of early embryonic pancreatic development 14 , Ngn3 can initiate the differentiation of pancreatic precursor cells into pancreatic endocrine cells 15 , Pax4 plays an important regulatory role in the production of islet progenitor cells and the differentiation of islet β-cells 16 , and Nkx2.2 is necessary for the phenotypic determination of mature β-cells 17 . A large number of studies have confirmed that transfection of Pdx1, Ngn3, Pax4, or Nkx2.2 into stem cells can induce their differentiation into IPCs18–20. A Pdx1-Ngn3 combined transfection can mediate the reprogramming of mesenchymal stromal cell differentiation into pancreatic endocrine cells in vitro 12 . In a previous study, we also confirmed that Sox9 can promote the differentiation of canine ADSCs (cADSCs) into islet-like cells. However, whether Pax4, NKX2.2, and Sox9 can coordinate with Pdx1-Ngn3 to regulate the reprogramming process of ADSC differentiation into islet cells is still unknown.
Therefore, in this study, five multigene coexpressing adenovirus vectors based on Pdx1-Ngn3, Pax4, Nkx2.2, and Sox9 were constructed, and the stocks of the packaged adenoviruses were used to infect the cADSCs, respectively. By detecting the differentiation effects of the five multigene adenoviruses on ADSCs, the best inducing gene combination was determined to provide a theoretical and experimental basis for further research on stem cell replacement therapy for diabetes.
Materials and Methods
Amplification of Pdx1, Ngn3, Sox9, Pax4, and Nkx2.2 Genes
The coding sequences of Pdx1 (GeneID: 493994), Ngn3 (GeneID: 489022), Sox9 (GeneID: 403464), Pax4 (GeneID: 482268), and Nkx2.2 (GeneID: 485744) were synthesized by Wuhan GeneCreate Biological Engineering Company. In addition, the end of each gene was connected to different 2A sequences (Pdx1+E2A; Ngn3+P2A; Sox9+P2A; Pax4+E2A; Nkx2.2+T2A) and cloned into pUC57. Primers with homologous fragments were designed according to the target gene sequences of the different gene combinations (A: Pdx1+Ngn3; B: Pdx1+Ngn3+Sox9; C: Pdx1+Ngn3+Pax4+Sox9; D: Pdx1+Ngn3+Nkx2.2+Sox9; E: Pdx1+Ngn3+Nkx2.2+Pax4; Supplementary Fig. 1B) and the pAdTrack-CMV vector (Hunan Fenghui Biotechnology Company, Hunan, China) sequences (Table 1) (which contains GFP gene sequences, and the plasmid map can be found in Supplementary Fig. 1A). There was an overlap region of approximately 20 bp between the forward primer of the first gene of each combination and the Bgl II sites of pAdTrack-CMV, and there was an overlap region of approximately 15 bp between the forward and reverse primers of the connected genes. The reverse primer of the last gene had an overlap region of 20 bp with the Hind III sites of pAdTrack-CMV. The target gene fragment was amplified by polymerase chain reaction (PCR) and confirmed by DNA sequencing.
Primer Sequences of Genes With Different Homologous Fragments.

The small font portion of the primer is the overlapping area. F and R represent the forward primers and reverse primers, respectively. U represents universal for all combinations.
A: combination A (Pdx1+Ngn3); B: combination B (Pdx1+Ngn3+Sox9); C: combination C (Pdx1+Ngn3+Pax4+Sox9); D: combination D (Pdx1+Ngn3+Nkx2.2+Sox9); E: combination E (Pdx1+Ngn3+Nkx2.2+Pax4).
Construction of Multigene Coexpression Adenovirus Vectors
The pAdTrack-CMV was digested with Bgl II and Hind III (TaKaRa, Dalian, China) to obtain a linearized vector. The Pdx1, Ngn3, Sox9, Pax4, and Nkx2.2 gene fragments were connected to the linearized vector in the order of each combination according to the instructions of the EasyGeno Rapid Recombination Cloning Kit (Beijing Tian Gen Biotechnology Company, Beijing, China) (Supplementary Fig. 1). The different constructed adenovirus shuttle vectors were verified by segmented PCR (see Table 2 for primer information) and double enzyme digestion experiments.
Primer Sequences of the Recombinant Genes.

The correctly identified shuttle vectors were linearized by Pme I (New England Biolabs, MA, USA) enzyme digestion and then transferred to BJ5183-AD-1 competent cells (containing pAdeasy-1) for recombination. These obtained adenovirus backbone vectors were identified by Pac I (New England Biolabs) restriction enzyme digestion. Using the Advanced Transfection Reagent kit (Zeta Life, San Francisco, CA, USA), the constructed vectors were transfected into 293A cells for adenovirus packaging to obtain infectious multigene coexpression adenovirus stocks.
Canine ADSC Culture
The cADSCs were isolated and identified by the Shaanxi Stem Cell Engineering Technology Research Center 21 . The cryopreserved cADSCs were resuscitated with α-MEM medium and cultured in a 37°C, 5% CO2 incubator. When the cell confluency reached approximately 60%, the successfully constructed multigene adenovirus stock was added to the cells at an MOI (multiplicity of infection) of 100 and then cultured further.
Reverse transcription quantitative PCR
The total RNA was extracted with the TaKaRa MiniBEST Universal RNA Extraction Kit (Code No. 9767). The total RNA was then reverse transcribed into complementary DNA (cDNA) according to the Takara PrimeScript™RT Master Mix (Perfect Real Time) (Code No. RR036Q) instructions. cDNA was used as a template to detect the expression of the target gene by quantitative PCR (qPCR), based on the primers listed in Table 3. The GAPDH gene was used as an internal control. The specific measured methods were carried out as previously described 22 . The verification was repeated three times for each gene of each sample, and the gene expression level was quantified using the 2−ΔΔCt method 23 .
Primer Sequences Used for Real-Time PCR.

Western Blot
A total of 50 μg protein was separated by sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis (PAGE) as previously described 24 . Primary antibodies against the targeted proteins included an anti-beta-actin antibody (ab8227, 1:2,000; Abcam, Cambridge, UK), anti-PDX1 antibody (ab227586, 1:1,000; Abcam), anti-NGN3 antibody (ab176124, 1:1,000; Abcam), anti-NKX2.2 antibody (ab86024, 1:1,000; Abcam), anti-PAX4 antibody (ab101721, 1:1,000; Abcam), and anti-SOX9 antibody (ab26414, 1:1,000; Abcam).
Dithizone Staining
Dithizone staining was performed according to a previously published method 25 . The cells were stained for 15 min with a freshly prepared dithizone (Sigma-Aldrich, MO, USA) working solution at 37°C, and observed and photographed under an optical microscope.
Glucose-Stimulated Insulin Secretion
A low-glycemic serum-free Dulbecco’s Modified Eagle Medium (DMEM) (containing 5.6 mmol/l glucose), a high-glycemic serum-free DMEM (containing 25 mmol/l glucose), or a KCl stimulation solution (containing 30 mmol/; KCl) was added to the cells for incubation for 30 min. After that, the canine Insulin Quantikine ELISA Kit (DINS00) was used to detect insulin secretion in the cell supernatant or cell culture fluid, and then normalized by cell count. The insulin stimulation release index, SI, is the ratio of insulin secretion after high-glucose stimulation to that after low-glucose stimulation.
Immunofluorescence Assay
The immunofluorescence staining was performed according to a previously published method25,26. The cells were fixed in 4% paraformaldehyde for 20 to 30 min, triple-washed with phosphate-buffered saline (PBS), stained with rabbit antibodies against dog insulin (ab63820, 1:100; Abcam), and then stained with DyLight594-labeled goat anti-rabbit IgG secondary fluorescent antibodies (35560, 1:100; Invitrogen, Carlsbad, CA, USA). In addition, Hoechst 33342 (Beijing Solarbio Science & Technology Company, Beijing, China) was used to counterstain the cell nuclei, after which the cells were photographed with an inverted fluorescence microscope.
Statistical Analysis
Statistical analysis was performed using SPSS 20.0. A one-way analysis of variance or t test was used to assess the difference between the two groups or between multiple groups. A value of P < 0.05 was considered statistically significant.
Results
Identification and Packaging of Multigene Coexpression Adenovirus Vectors
First, we cloned specific gene fragments that contain different homology arms (A: Pdx1-Ngn3; B: Pdx1-Ngn3-Sox9; C: Pdx1-Ngn3-Pax4-Sox9; D: Pdx1-Ngn3-Nkx2.2-Sox9, and E: Pdx1-Ngn3-Nkx2.2-Pax4) from pUC57 plasmids (Supplementary Fig. 2) and then reconnected these fragments with the vector pAdTrack-CMV that was digested by Bgl II and Hind III to obtain different adenovirus shuttle vectors. The obtained vectors were verified by segmented PCR and double enzyme digestion (Bgl II and Hind III), and the electrophoresis results showed that all vectors were consistent with the expected product size (Supplementary Figs. 3 and 4A), which proved that the five groups of adenovirus shuttle vectors were successfully constructed.
Afterward, the five groups of adenovirus shuttle vectors, treated with Pme I enzyme, were transferred into BJ5183-AD-1 competent cells to obtain different adenovirus backbone vectors. These obtained adenoviral backbone vectors were processed by the restriction endonuclease Pac I for reverse verification, and the electrophoresis results showed that two bands of approximately 30 kb and 5 kb were consistent with the expected product sizes (Supplementary Fig. 4B).
Then the adenovirus backbone vectors treated with Pac I enzyme were purified, recovered, and then transfected into 293A cells, respectively. A small amount of green fluorescence was observed in the cells after 24 h of culture, and green fluorescence was seen in the cells in the full field view after 48 h (Supplementary Fig. 4C). When the cells were continuously cultured until they were observed to be in the shape of grape clusters, we collected the cell suspension and performed repeated freezing and thawing to collect the successfully packaged A, B, C, D, and E combination adenovirus stocks.
Expression of Exogenous Genes in cADSCs After Infection With Multigene Adenovirus Stocks
We used the successfully constructed adenovirus stocks of the five combinations of A, B, C, D, and E to infect the cADSCs, respectively, and then assess the expression levels of the exogenous genes in the cADSCs after 4 days of culture. The results from the reverse transcription quantitative (RT-qPCR) and western blotting showed that the expression of the corresponding exogenous transfected gene in the cells after infection was significantly higher than that of the control group, and the trends of the messenger RNA (mRNA) level and the protein level of the target gene were roughly the same (Fig. 1). The results indicated that these five combinations of adenoviruses can successfully introduce different exogenous genes into cADSCs.
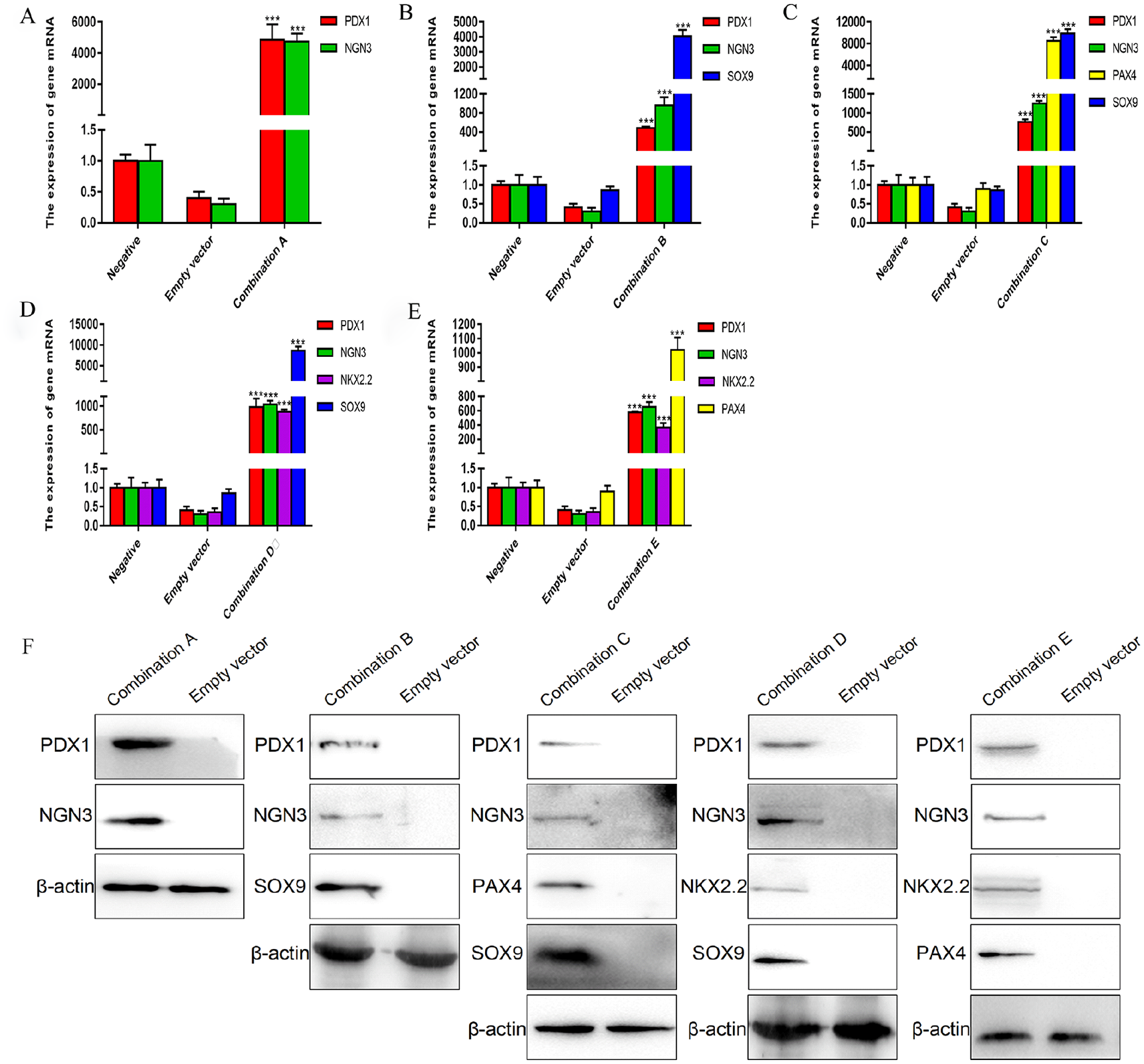
The expression levels of exogenous genes in cADSCs after infection with multigene adenovirus stocks. (A–E) The mRNA expression levels of exogenous genes in cADSCs after infection with the five multigene adenovirus stocks; (F) the protein expression levels of exogenous genes in cADSCs after infection with the five multigene adenovirus stocks. cADSCs: canine adipose-derived mesenchymal stem cells; mRNA: messenger RNA. ***P < 0.001 versus the negative group.
Expression of GFP in cADSCs After Infection With Multigene Adenovirus Stocks
To understand the effect of the multigene coexpressing adenoviruses on cADSCs, we observed the cell morphology and green fluorescent protein (GFP) expression in cADSCs during the infection process by microscopic evaluation (Fig. 2). A large amount of relatively weak GFP expression was detected in the cADSCs infected with the five combination adenoviruses after 2 days. The fluorescence intensity of GFP in the cells infected by each combination of adenovirus increased significantly after 4 days of infection. At the end of the 30-day induction, the adenovirus-infected cells in each group appeared clustered and showed a spherical shape of pancreatic islet-like cells. However, the empty vector-infected cells did not clump by the 30th day, and their GFP expression decreased significantly.
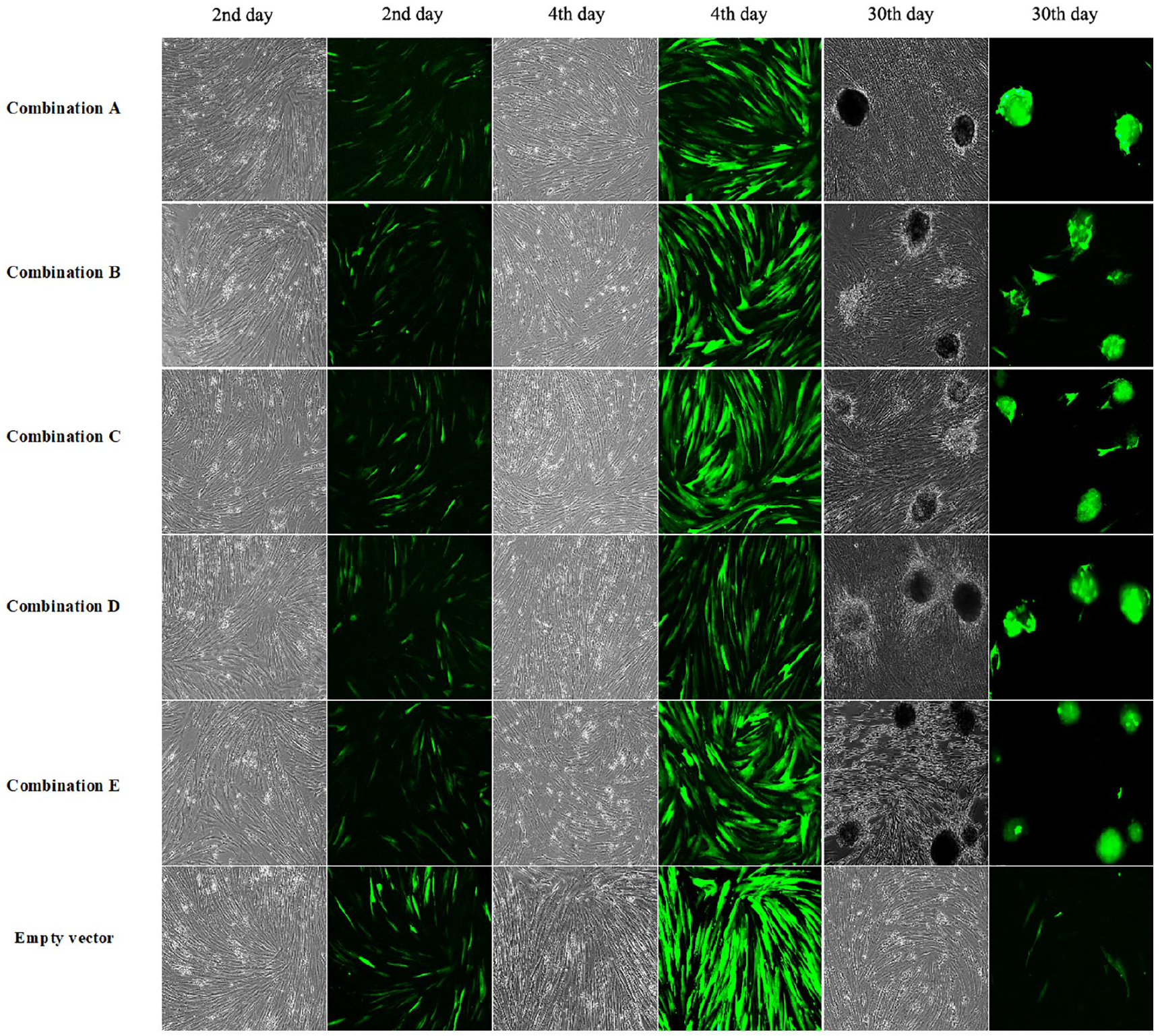
The cell morphology change (bright field) and green fluorescent protein expression (fluorescent images) in canine adipose-derived mesenchymal stem cells after infection with multigene adenovirus stocks (100× magnification) on the 2nd, 4th, and 30th day.
After induction for 30 days, the number of cell agglomerates in the combinations of A, B, C, D, and E were significantly higher than those of the empty control group (Fig. 3). For example, there were approximately 103 cell clusters in 106 cADSC cells, the diameter of the cell cluster was approximately 98 μm, and each cell cluster contained approximately 105 cells in group E. This phenomenon of cell clustering indicates that the cells had a preference to differentiate into pancreatic islet-like cell clusters.

Statistical analysis of the cell agglomerates after infection with multigene adenovirus stocks for 30 days. ***P < 0.001 versus the empty vector control group.
Identification of Islet-Like Cell Clusters After Infection of cADSCs With Multigene Adenovirus Stocks
We used dithizone staining to identify whether the cADSC cell group induced by multigene coexpression adenovirus infection had the characteristics of islet-like cells. The cell clusters after induction for 30 days with A, B, C, D, or E combinations all showed scarlet red staining after dithizone staining, while the cells in the empty vector control group did not show scarlet staining (Fig. 4). This result indicated that the cell clusters induced by multigene coexpression adenovirus infection exhibited the characteristics of islet cells.

Dithizone staining of canine adipose-derived mesenchymal stem cells after infection with multigene adenovirus stocks (100× magnification).
Insulin Secretion of cADSCs After Infection With Multigene Adenovirus Stocks
To further determine whether the cell clusters that were induced by the infection of the different multigene coexpression adenoviruses had the function to produce insulin, we performed glucose stimulation tests on canine pancreatic islet cells and the cells that were infected with adenoviruses with different combinations of multigene (Fig. 5). The results of insulin secretion in the cell supernatant and cell fluid showed that the insulin contents of the cells of five groups were significantly higher than those of the empty vector group but lower than those of the islet cells (Fig. 5A, B). The insulin secretion measured in the cell fluid was slightly higher than that of the supernatant, and the insulin secretion levels after KCl stimulation and high-glucose stimulation were similar. The insulin secretion of the E group cells was the largest among the five induction groups, and the insulin stimulation release index SI of the E group cells (SI in the supernatant was 2.00; SI in the cell fluid was 2.10) was also higher than that of the cells of other four groups but lower than that of the islet cells (SI in the supernatant was 2.65; SI in the cell fluid was 2.51) (Fig. 5C).

Insulin secretion and insulin stimulation release index of canine adipose-derived mesenchymal stem cells after infection with multigene adenovirus stocks. (A) Insulin secretion in the supernatants; (B) insulin secretion into the cell culture fluid; (C) insulin stimulation release index. ***P < 0.001 versus the empty vector control group.
In addition, the insulin immunofluorescence results of the cells after infection with the five groups of multigene adenovirus stocks all showed the red fluorescence (Fig. 6). However, only blue-stained nuclei were observed in the cells infected with the empty vector. The result confirmed that infection of the cADSCs with the five multigene coexpressing adenoviruses could induce the production of IPCs.
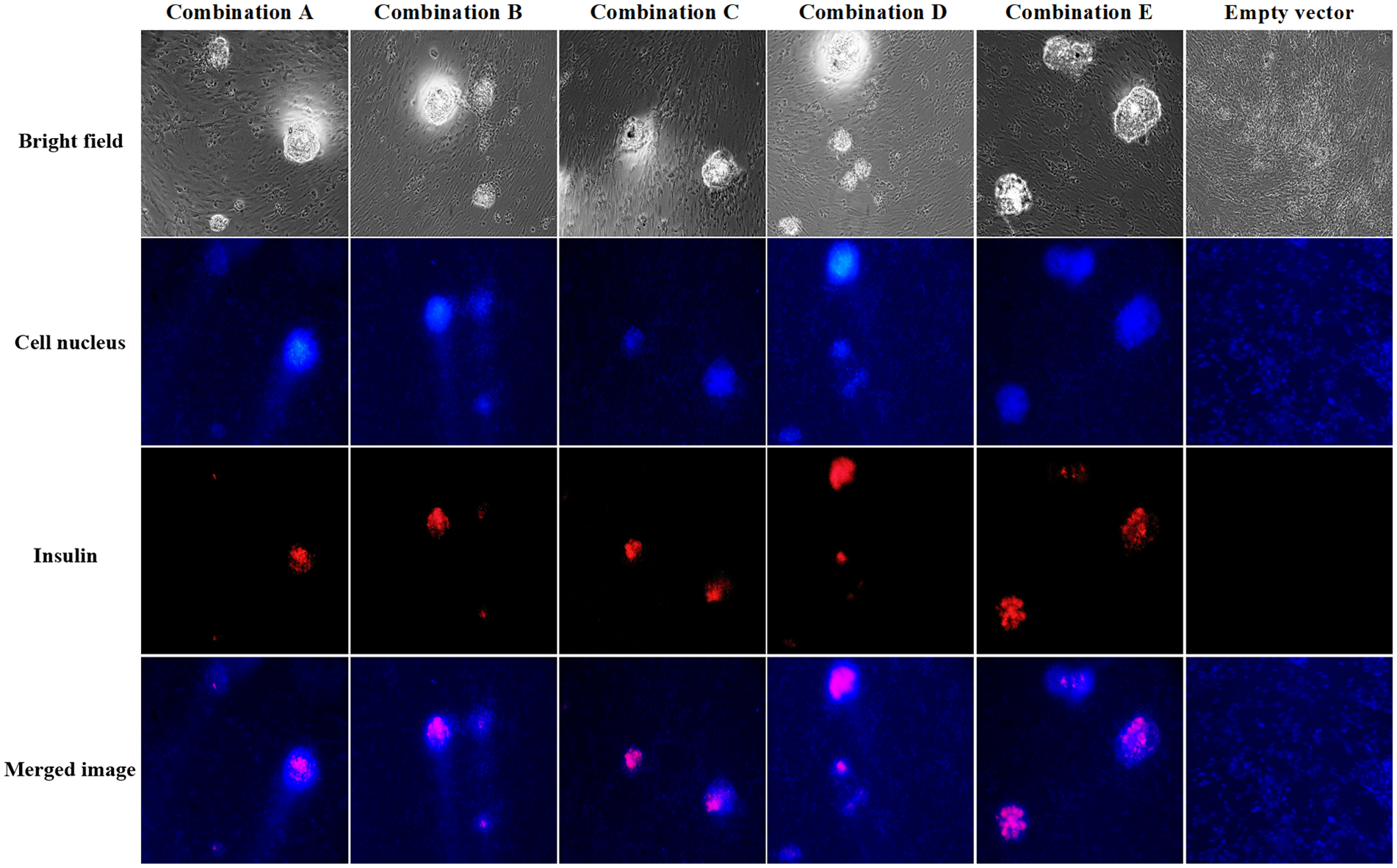
Insulin immunofluorescence staining of canine adipose-derived mesenchymal stem cells after infection with multigene adenovirus stocks (100× magnification).
The Expression of Genes Related to the Development of Islet β-Cells After Infection of cADSCs With Multigene Adenovirus Stocks
To understand the regulation process of cADSC differentiation into islet-like cells induced by different multigene adenoviruses, we tested the expression levels of pancreatic β-cell development–related genes (Pdx1, Pax4, Gata4, and Nkx2.2), insulin secretion–related genes (Nkx6.1, MafA, SLC30A8, ABCC8, KCNJ8, and G6PC2), and insulin formation–related genes (PCSK1, PCSK2, and Insulin) in the differentiated cells. As shown in Fig. 7, the expression levels of islet β-cell development–related genes (Pax4 and Nkx2.2), insulin secretion–related genes (Nkx6.1, ABCC8, and KCNJ8), and insulin formation–related genes (PCSK1, PCSK2, and Insulin) in the cells infected with the five multigene coexpressing adenoviruses were significantly higher than in the cells of the empty vector group. Among them, the gene expression levels of Insulin, PCSK2, and SLC30A8 of groups B and E were significantly higher than those of groups A, C, and D, and the gene expression of group E was higher than that of group B. These results indicate that the E group infection had the highest efficiency in inducing cADSCs to differentiate into pancreatic islet cells. When compared with the pancreatic islet cell group, the expression levels of Pdx1, MafA, Nkx2.2, Pax4, Gata4, Insulin, and ABCC8 were still lower in group E.

The mRNA expression levels of islet β-cell development–related genes of canine adipose-derived mesenchymal stem cells after infection with multigene adenovirus stocks. *P < 0.05, **P < 0.01, ***P < 0.001 versus the empty vector control group. mRNA, messenger RNA.
Discussion
ADSCs contain biologically active substances that regulate blood sugar balance, such as leptin, adipocytokines, and visfatin, and are better sources of seed cells for diabetes cell transplantation therapy compared with other MSCs27,28. Many studies have reported that transferring key genes that regulate the development of islet β-cells into ADSCs can induce them to differentiate into IPCs18,29,30. However, only part of the cascade-regulated gene expression can be found in the induced IPCs, and the ability to secrete insulin from IPCs is limited. The main reason may be that the screening of exogenous regulatory genes has not been sufficiently comprehensive. Currently, many researchers often simultaneously overexpress multiple foreign genes in a target cell to study their mutual synergy or inhibition. Previous studies have reported that the combined induction of multiple different exogenous genes can promote the differentiation of MSCs into IPCs with insulin secretion function, and the induction effect of multiple genes was significantly better than that of induction with a single gene 12 .
In this study, we used the 2A sequence to carry out the recombination connection between multiple foreign genes 31 . Adenovirus vector–mediated gene transfer has the advantages of instantaneous high-level transgene expression and nonintegration into chromosomes 32 . Therefore, we had successfully constructed five multigene coexpression adenovirus vectors using this ideal transgenic tool. We tested the expression of exogenous genes after infection of cADSCs with these adenovirus stocks. It was found that the expression levels of the exogenous genes in the cells after infection with the five multigene adenoviruses were significantly higher than those of the control group, and the expression levels of exogenous genes in the different multigene groups were also significantly different. As previous studies have reported, there are many factors, such as size, location, number, and type of the target gene, that can affect the final gene expression in the infected cells in 2A peptide–mediated coexpression of multiple genes 33 . We used different 2A peptides to connect multiple genes, and as the cleavage efficiencies of the different 2A peptides are different, this might cause more uncertainty in the cleavage efficiency 34 . Thus, this result may be due to the connection sequence of the different genes and the different cutting efficiency of the 2A peptide when we constructed the vectors. Interestingly, the results of RT-qPCR and western blotting revealed that the expression levels of the last gene in multiple combinations were higher than those of the other genes, and the specific reasons for this phenomenon still need to be further explored.
Cell differentiation is affected by both the combination of gene control programs and the microenvironment where the cell is located. Among these, genetic programming is the decisive factor that controls the differentiation of stem cells, and the external environment can only play a role by influencing the selective expression of genetic programming within the cell. To study the genetic control program for the differentiation of ADSCs into pancreatic β-cells, we infected cADSCs with the adenovirus stocks of A, B, C, D and E combinations and then tested the induction effect of each combination. At first, we found that cells in all five groups showed pancreatic islet-like cell clusters after induction with the five polygenic adenoviruses, and the cell immunofluorescence results were positive for dithizone and insulin staining. However, by comparing the amount of insulin secretion, SI, and the mRNA expression of islet development–related genes, we found that the E combination (pAdTrack-Pdx1-Ngn3-Nkx2.2-Pax4-AdEasy) had the best induction effect, and the induction effect of the B combination (pAdTrack-Pdx1-Ngn3-Sox9-AdEasy) was better than that of the A (pAdTrack-Pdx1-Ngn3-AdEasy), C (pAdTrack-Pdx1-Ngn3-Pax4-Sox9-AdEasy) and D (pAdTrack-Pdx1-Ngn3-Nkx2.2-Sox9-AdEasy) combinations. These findings indicate that the coordinated expression of Pdx1, Ngn3, Nkx2.2, and Pax4 plays an important role in the gene control program that induces the differentiation of cADSCs into IPCs. At the same time, it also showed that Sox9 had a synergistic effect with Pdx1 and Ngn3, but when Nkx2.2 or Pax4 was added to the three-gene combination (Pdx1-Ngn3-Sox9), antagonism may have occurred and led to poor induction effects. The specific reason needs to be further experimentally studied. In addition, although the exogenous genes Pdx1, Ngn3, Pax4, Nkx2.2, and Sox9 were successfully introduced into cADSCs in the form of different combinations and induced cADSCs to differentiate into IPCs, the insulin secretion level of cells that were infected with the adenovirus carrying the best gene combination of Pdx1-Ngn3-Nkx2.2-Pax4 was still extremely significantly lower than that of the islet cells, which is far from the goal of a clinical diabetes treatment.
The same transcription factor exhibits different effects at different developmental stages in the pancreas, and the in vivo microenvironment can facilitate further induction and maturation of pancreatic β-cells to compensate for the damaged pancreatic β-cells and regulate blood sugar levels35,36. Therefore, in the follow-up research, we will further optimize the induction program by adding exogenous activin 37 , controlling the time point of gene expression 38 , creating a cell microenvironment, and ultimately screening different combinations of transgene induction.
Conclusion
In this study, five groups of multigene coexpressing adenoviruses were used to infect cADSCs to induce them to differentiate into IPCs. Through morphological observation, dithizone staining, sugar-stimulated insulin secretion test, cellular immunofluorescence, and RT-qPCR assessment in the induced cells, the best induction combination (pAdTrack-Pdx1-Ngn3-Nkx2.2- Pax4-AdEasy) was identified after comparative screening. This study provides a theoretical reference and an experimental basis for further exploration of more efficient diabetic cell transplantation programs.
Supplemental Material
sj-jpg-1-cll-10.1177_09636897221081483 – Supplemental material for The Roles of Different Multigene Combinations of Pdx1, Ngn3, Sox9, Pax4, and Nkx2.2 in the Reprogramming of Canine ADSCs Into IPCs
Supplemental material, sj-jpg-1-cll-10.1177_09636897221081483 for The Roles of Different Multigene Combinations of Pdx1, Ngn3, Sox9, Pax4, and Nkx2.2 in the Reprogramming of Canine ADSCs Into IPCs by Dengke Gao, Pengxiu Dai, Zhixin Fan, Jinglu Wang and Yihua Zhang in Cell Transplantation
Supplemental Material
sj-tif-2-cll-10.1177_09636897221081483 – Supplemental material for The Roles of Different Multigene Combinations of Pdx1, Ngn3, Sox9, Pax4, and Nkx2.2 in the Reprogramming of Canine ADSCs Into IPCs
Supplemental material, sj-tif-2-cll-10.1177_09636897221081483 for The Roles of Different Multigene Combinations of Pdx1, Ngn3, Sox9, Pax4, and Nkx2.2 in the Reprogramming of Canine ADSCs Into IPCs by Dengke Gao, Pengxiu Dai, Zhixin Fan, Jinglu Wang and Yihua Zhang in Cell Transplantation
Supplemental Material
sj-tif-3-cll-10.1177_09636897221081483 – Supplemental material for The Roles of Different Multigene Combinations of Pdx1, Ngn3, Sox9, Pax4, and Nkx2.2 in the Reprogramming of Canine ADSCs Into IPCs
Supplemental material, sj-tif-3-cll-10.1177_09636897221081483 for The Roles of Different Multigene Combinations of Pdx1, Ngn3, Sox9, Pax4, and Nkx2.2 in the Reprogramming of Canine ADSCs Into IPCs by Dengke Gao, Pengxiu Dai, Zhixin Fan, Jinglu Wang and Yihua Zhang in Cell Transplantation
Supplemental Material
sj-tif-4-cll-10.1177_09636897221081483 – Supplemental material for The Roles of Different Multigene Combinations of Pdx1, Ngn3, Sox9, Pax4, and Nkx2.2 in the Reprogramming of Canine ADSCs Into IPCs
Supplemental material, sj-tif-4-cll-10.1177_09636897221081483 for The Roles of Different Multigene Combinations of Pdx1, Ngn3, Sox9, Pax4, and Nkx2.2 in the Reprogramming of Canine ADSCs Into IPCs by Dengke Gao, Pengxiu Dai, Zhixin Fan, Jinglu Wang and Yihua Zhang in Cell Transplantation
Footnotes
Acknowledgements
We thank the editors from American Journal Experts for editing the English text of a draft of this manuscript.
Author’s Note
Yihua Zhang is also affiliated to Baioupai Tianjin Biotechnology Ltc., Tianjin, China.
Author Contributions
D.G. carried out the experiments, analyzed the data, and drafted the manuscript. Y.Z. designed the study, reviewed the data, and revised the manuscript. P.D., Z.F. and J.W. conducted parts of the experiments. All authors have read and agreed to the published version of the manuscript.
Ethical Approval
Ethical Approval is not applicable for this article.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (project number: 31872529) and the Key Project of Natural Science Foundation of Shaanxi Province (project number: 2019JZ-16) in the design of study, data analysis, and writing of the manuscript.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
