Abstract
Non-small cell lung cancer (NSCLC) is the most common type of lung cancer. Increasing evidence suggests that long non-coding RNAs (lncRNAs) function in the tumorigenesis of NSCLC. LINC00958, a newly identified lncRNA, has been reported to be closely linked to tumorigenesis in several cancers. However, its specific role in NSCLC remains unclear. In this study, we determined the expression of LINC00958 in NSCLC by RT-qPCR analysis and evaluated cell proliferation and migration by CCK-8 and transwell assays, respectively. We established a xenograft tumor model to examine the effect of LINC00958 on tumor growth in vivo. Luciferase reporter assays were performed to determine the interaction between LINC00958 and miR-204-3p and the interaction between miR-204-3p and KIF2A. We found that LINC00958 was up-regulated in NSCLC tissues and cell lines. Down-regulation of LINC00958 inhibited cell proliferation and migration in vitro and suppressed tumor growth in vivo. Besides, miR-204-3p was identified as a target of LINC00958 and miR-204-3p inhibitor could reverse the inhibitory effect of LINC00958 knockdown on proliferation and migration of NSCLC cells. We also validated that KIF2A, a direct target of miR-204-3p, was responsible for the biological role of LINC00958. KIF2A antagonized the effect of miR-204-3p on NSCLC cell proliferation and migration and was regulated by LINC00958/miR-204-3p. Taken together, these data indicate that the LINC00958/miR-204-3p/KIF2A axis is critical for NSCLC progression, which might provide a potential therapeutic target of NSCLC.
Introduction
Lung cancer is one of the most common cancers and a leading cause of cancer-related death 1 . Every year, almost 1.4 million lung cancer cases are diagnosed in the world 2 . Among all the cases, non-small cell lung cancer (NSCLC) accounts for over 85% 3 . In the past decades, a remarkable progress has been made in therapeutic approaches such as surgery, chemotherapy, radiotherapy and immune therapy, but the 5-year survival rate is still dissatisfied 4 –7 . The poor outcome and high mortality are mainly due to asymptomatic early stage and late diagnosis of the disease 8,9 . Thus, it is desperate to explore novel biomarkers for an early diagnosis and treatment of NSCLC.
The ENCODE program has revealed that only a small percentage of the genome encodes proteins and a majority of the genome encodes a large number of non-coding RNAs 10 . Long non-coding RNAs (lncRNAs) are a novel class of transcripts with a length of more than 200 nucleotides 11 . Accumulating data have identified that lncRNAs play a crucial role in diverse biological processes such as cell proliferation, apoptosis, angiogenesis, and tumorigenesis 12 –14 . In recent studies, several lncRNAs have been confirmed to be implicated in NSCLC progression. For example, lncRNA TUSC7 correlates with poor prognosis of NSCLC patients and inhibits cell proliferation 15 . LncRNA NEAT1 was up-regulated in NSCLC cell lines and promotes NSCLC cells proliferation, migration, and invasion 16 . These findings suggested the potential role of lncRNA as a new target for NSCLC 17 . LINC00958 is a newly identified lncRNA and several studies have reported its abnormal expression in multiple types of cancers 18,19 . However, the specific effects of LINC00958 on NSCLC remain unclear.
In this study, we demonstrated for the first time that LINC00958 was significantly up-regulated in NSCLC tissues and cell lines. We also observed the inhibitory effect of LINC00958 knockdown on NSCLC progression through in vitro and in vivo experiments. Mechanically, we revealed that LINC00958 down-regulation inhibited NSCLC cell proliferation and migration partly via regulation of the miR-204-3p/KIF2A axis. Taken together, our study provided new insights into the role of LINC00958 as a promising prognostic biomarker and a therapeutic target for NSCLC.
Materials and Methods
Patients and Tissue Samples
A total of 42 patients from the Second People’s Hospital of Jiaozuo City, the First Affiliated Hospital of Henan Polytechnic University (Jiaozuo, China) participated in the study and provided written informed consent. None of the patients received any adjuvant therapies before the surgery. NSCLC tissues and adjacent non-cancerous tissues were collected from these patients and immediately frozen in liquid nitrogen and stored at −80°C. This study was approved by the Ethics Committee of Henan Polytechnic University.
Cell Lines and Cell Culture
Human NSCLC cell lines (A549, H1299 and PC9) and the normal bronchial epithelial cell line BEAS-2B were obtained from the Chinese Academy of Sciences (Shanghai, China). All cell lines were cultured in RPMI-1640 medium (Gibco, Rockville, MD, USA) supplemented with 10% fetal bovine serum (FBS; Gibco) and 1% penicillin/streptomycin, followed by incubation in a humidified atmosphere with 5% CO2 at 37°C.
Quantitative Real-Time Polymerase Chain Reaction (RT-qPCR)
Total RNA was extracted from tissues or cells using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). Reverse transcription from RNA to cDNA was performed using PrimeScript RT reagent (Takara, Japan). RT-qPCR was carried out using SYBR Green Mixture (Takara) on an ABI 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA, USA). The following primers were used: LINC00958, 5′-CCATTGAAGATACCACGCTGC-3′ (forward) and 5′-GGTTGTTGCCCAGGGTAGTG-3′ (reverse); miR-204-3p, 5′-AGCTGTACAAGTAAGCCTGATCATGTACCCATAGG-3′ (forward) and 5′-GGGAGAGGGGCTTAGCTTATGGGACAGTTATGGGC-3′ (reverse); KIF2A, 5′-GCCTTTGATGACTCAGCTCC-3′ (forward) and 5′-TTCCTGAAAAGTCACCACCC-3′ (reverse); GAPDH, 5′-TGCACCACCAACTGCTTAGC-3′ (forward) and 5′-GGCATGGACTGTGGTCATGAG-3′ (reverse); U6, 5′-CTCGCTTCGGCAGCACA-3′ (forward) and 5′-AACGCTTCACGAATTTGCGT-3′ (reverse). The expression levels of target genes were normalized to internal control and calculated using the 2−ΔΔCt method.
Western Blot Analysis
Total protein was extracted by lysis buffer (Sigma, St. Louis, MO, USA) supplemented with PMSF (Biotool, Houston, TX, USA). Then proteins were separated by 12% SDS-PAGE and transferred onto PVDF membranes (Millipore, Billerica, MA, USA). After incubation in skim milk at 37°C, the membranes were incubated overnight at 4°C with primary antibodies against KIF2A and GAPDH, followed by incubation with corresponding secondary antibodies. Protein bands were detected by an ECL kit (Thermo Fisher Scientific, Waltham, MA, USA) and analyzed by the ImageJ software.
Cell Transfection
The shRNA targeting LINC00958, miR-204-3p mimics/inhibitors and corresponding negative controls were purchased from GeneCopoeia (Rockville, MD, USA). The pcDNA3.1-KIF2A and empty vectors were from RiboBio (Guangzhou, China). Transfection of shRNAs, miRNAs or plasmids was conducted using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instructions.
Cell Proliferation Assay
Cell proliferation was assessed using the CCK-8 assay. In brief, 2×103 cells were plated into a 96-well plate. After incubation for different time, CCK-8 solution (Dojindo, Tokyo, Japan) was added to each well and cells were further incubated for 4 h. The absorbance at 450 nm was measured using a microplate reader.
Cell Migration Assay
Cell migration was evaluated using transwell chambers (Corning, New York, USA). 5×104 cells in serum-free medium were added to the upper chamber and 10% FBS was added to the lower chamber. Subsequently, cells were incubated for 24 h and those migrating to the lower surface of the insert were fixed and stained with 0.1% crystal violet. The number of migrating cells from five random fields was counted under a microscope.
In Vivo Xenograft Tumor Assay
Female BALB/c mice aged 5 weeks old were obtained from Shanghai Laboratory Animal Center (Shanghai, China) and kept under specific pathogen-free conditions. All animals were handled with approval of the Animal Care and Use Committee of Henan Polytechnic University. A549 cells transfected with sh-LINC00958 or sh-NC were administered into the left flank of the nude mice via subcutaneous injection. Tumor volume was measured every week and calculated using the following formula: volume (mm3) = length × width 2 /2. Five weeks later, mice were sacrificed, tumors were weighed, and tumor tissues were dissected for further experimental analysis.
Immunohistochemistry (IHC)
Tissue sections were deparaffinized and hydrated using xylene and graded ethanol, respectively, followed by endogenous peroxidase-blocking and antigen retrieval. Then the sections were incubated with polyclonal rabbit anti-KIF2A overnight at 4°C. Subsequently, the sections were treated with appropriate secondary antibody. Finally, the sections were visualized with diaminobenzidine (DAB) and counterstained in hematoxylin.
Luciferase Reporter Assay
Luciferase reporter vectors were constructed using pGL3 vectors (Promega, Madison, WI, USA). 1×104 cells were seeded onto a 96-well plate, followed by co-transfection with pGL3-LINC00958 (WT), pGL3-LINC00958 (MUT), pGL3-KIF2A 3′-UTR (WT), pGL3-KIF2A 3′-UTR (MUT) and miR-204-3p mimics or miR-NC using Lipofectamine 2000 (Invitrogen). After 48 h, luciferase activity was detected by the Dual-Luciferase Reporter Assay System (Promega).
Statistical Analysis
Data were presented as means ± standard deviation (SD). Student’s t-test or one-way ANOVA was used to compare the significant difference of different groups. Statistical analysis was performed using GraphPad Prism and SPSS 20.0 software and P < .05 was considered statistically significant.
Results
LINC00958 Is Overexpressed in NSCLC Tissues and Cell Lines
We examined LINC00958 expression in 42 pairs of NSCLC tissues and corresponding adjacent normal tissues using RT-qPCR analysis. The results showed that LINC00958 had a higher expression level in NSCLC tissues than in the adjacent normal tissues (Fig. 1A). Further investigation was performed in NSCLC cell lines (A549, H1299 and PC9). As shown in Fig. 1B, LINC00958 was significantly overexpressed in A549, H1299 and PC9 cell lines in comparison with the normal bronchial epithelial cell line BEAS-2B. Because A549 cell line exhibited the highest expression of LINC00958, thus, this cell line was selected in the following experiments.
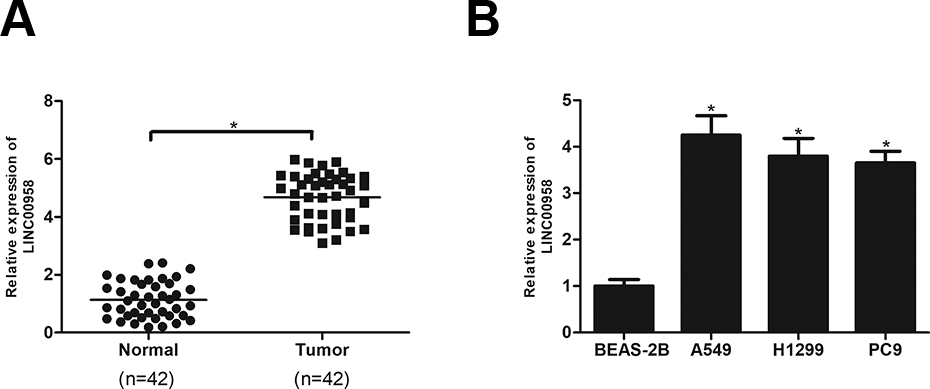
LINC00958 is overexpressed in NSCLC tissues and cell lines. (A) RT-qPCR analysis of LINC00958 expression in 42 pairs of NSCLC tissues and adjacent normal tissues. (B) RT-qPCR analysis of LINC00958 expression in NSCLC cell lines (A549, H1299 and PC9) and the normal bronchial epithelial cell line BEAS-2B. *P < .05
Knockdown of LINC00958 Inhibits NSCLC Cell Proliferation and Migration
To evaluate the role of LINC00958 in NSCLC, we silenced LINC00958 in A549 cells by transfection with sh-LINC00958, which remarkably reduced the expression level of LINC00958 in A549 cells (Fig. 2A). The CCK-8 assay was performed to assess cell proliferation. As shown in Fig. 2B, knockdown of LINC00958 obviously decreased the proliferative rate of A549 cells in comparison with corresponding control cells. Then the transwell assay was performed to detect cell migration. The results showed that the migratory ability of A549 cells was significantly blocked after the expression of LINC00958 was decreased (Fig. 2C).
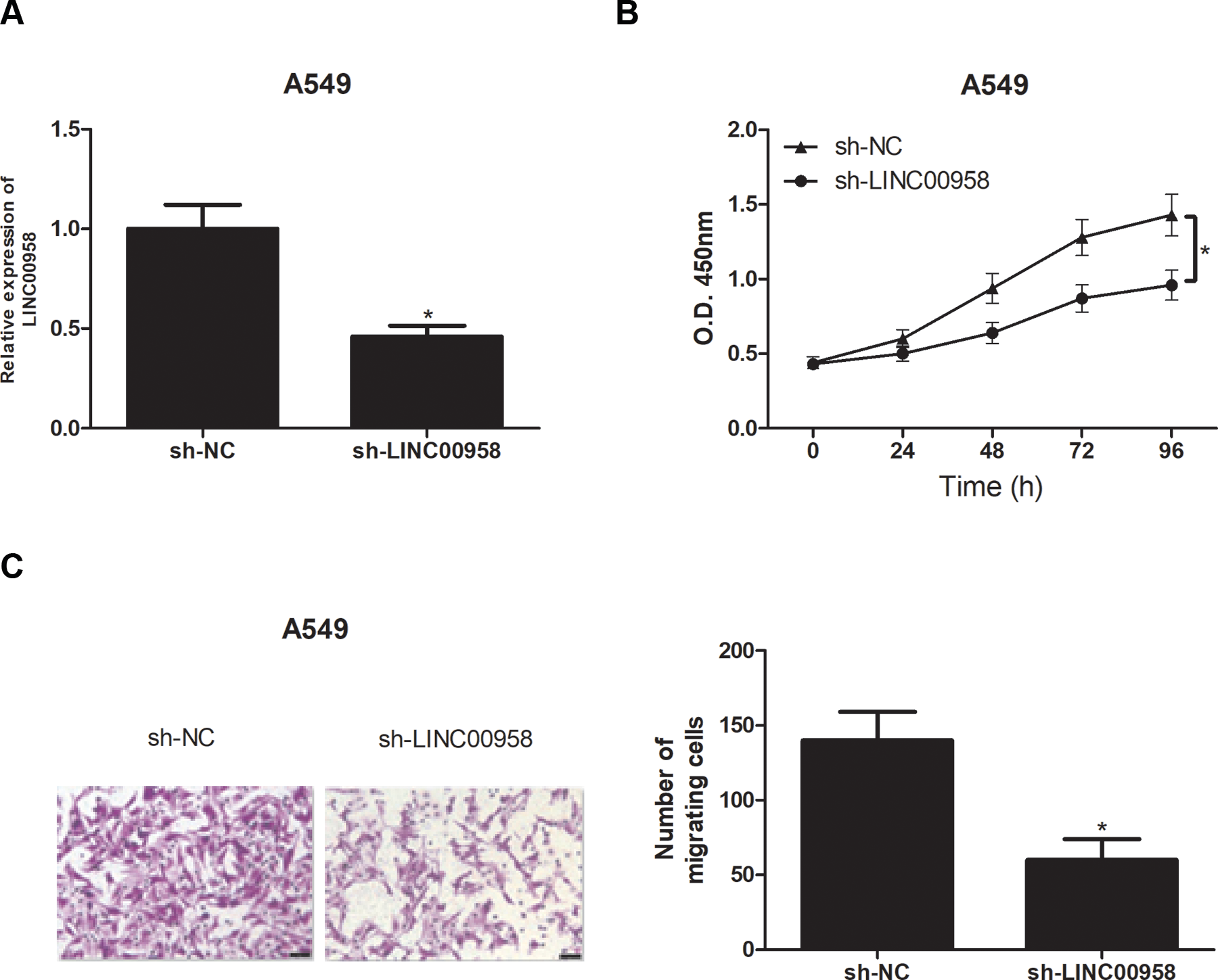
Knockdown of LINC00958 inhibits NSCLC cell proliferation and migration. (A) Relative LINC00958 expression was assessed in A549 cells after transfection with sh-LINC00958 or sh-NC. The proliferation and migration of A549 cells were measured using CCK-8 (B) and transwell (C) assays, respectively. *P < .05
LINC00958 Directly Interacts with MiR-204-3p in NSCLC Cells
Using bioinformatics tools, we found that LINC00958 is a putative target of miR-204-3p (Fig. 3A). To confirm our prediction, we conducted luciferase reporter assays by transfecting LINC00958-WT or LINC00958-MUT constructs into A549 cells with miR-204-3p mimics. As shown in Fig. 3B, C, miR-204-3p mimics up-regulated miR-204-3p expression in A549 cells in comparison with the control group and miR-204-3p overexpression strikingly attenuated the luciferase activity of LINC00958-WT constructs without affecting the luciferase activity of LINC00958-MUT constructs. Furthermore, we found that silencing of LINC00958 increased miR-204-3p expression in A549 cells (Fig. 3D).
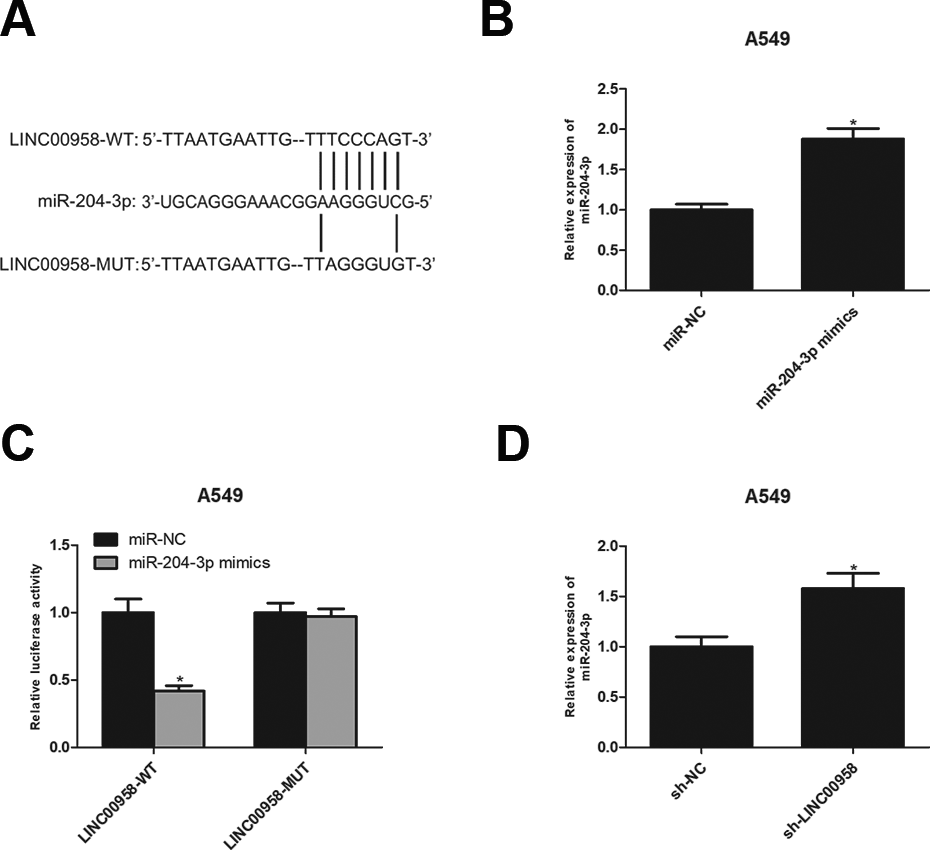
LINC00958 directly interacts with miR-204-3p in NSCLC cells. (A) The potential binding sites between LINC00958 and miR-204-3p. (B) The expression of miR-204-3p was detected in A549 cells transfected with miR-204-3p mimics or miR-NC. (C) Luciferase reporter assay of A549 cells co-transfected with LINC00958-WT or LINC00958-MUT and miR-204-3p mimics or miR-NC. (D) The expression of miR-204-3p was detected in A549 cells after LINC00958 knockdown. *P < .05.
MiR-204-3p Is Down-Regulated In NSCLC and Negatively Correlated with LINC00958
We measured miR-204-3p expression in NSCLC tissues and cell lines using the RT-qPCR analysis. The results showed that miR-204-3p was significantly decreased in NSCLC tissues in comparison with the adjacent normal tissues (Fig. 4A). Consistently, miR-204-3p expression in A549, H1299 and PC9 cell lines was much lower than in the normal bronchial epithelial cell line BEAS-2B (Fig. 4B). In addition, miR-204-3p expression was inversely correlated with LINC00958 level in NSCLC tissues (Fig. 4C).
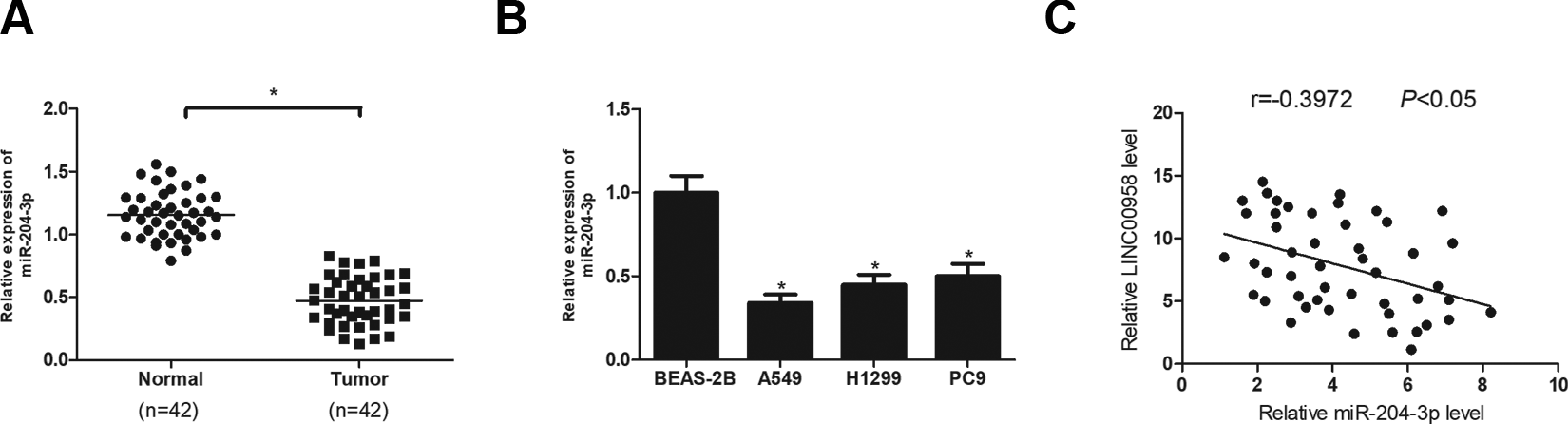
MiR-204-3p is down-regulated in NSCLC and negatively correlated with LINC00958. (A) RT-qPCR analysis of miR-204-3p expression in NSCLC tissues and adjacent normal tissues. (B) RT-qPCR analysis of miR-204-3p expression in A549, H1299, PC9 and BEAS-2B cell lines. (C) The correlation between miR-204-3p and LINC00958 level in NSCLC tissues. *P < .05.
MiR-204-3p Inhibitor Reverses the Effect of LINC00958 Knockdown on Proliferation and Migration of NSCLC Cells
To provide further insight into the link between LINC00958 and miR-204-3p in NSCLC progression, A549 cells were co-transfected with sh-LINC00958 and miR-204-3p inhibitor. Cell proliferation and migration were determined for these cells. The CCK-8 and transwell assays showed that down-regulation of miR-204-3p by transfection with miR-204-3p inhibitor significantly abrogated the inhibitory effect of LINC00958 knockdown on proliferation (Fig. 5A) and migration (Fig. 5B) of A549 cells.
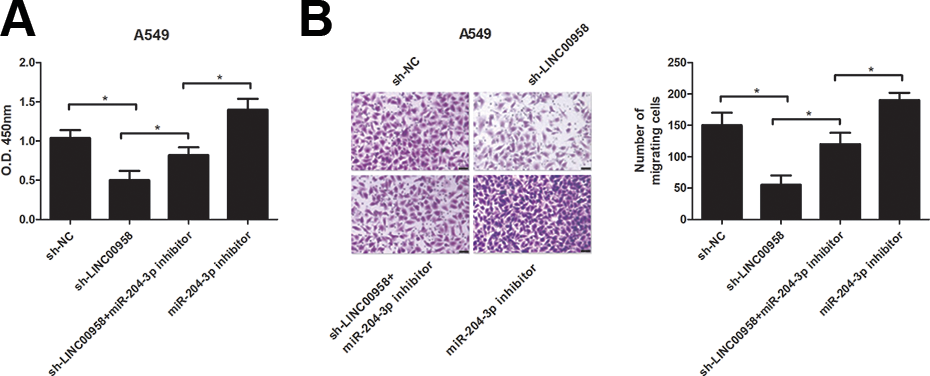
MiR-204-3p inhibitor reverses the effect of LINC00958 knockdown on proliferation and migration of NSCLC cells. The CCK-8 and transwell assays were conducted to determine proliferation (A) and migration (B) respectively in A549 cells co-transfected with sh-LINC00958 and miR-204-3p inhibitor. *P < .05.
KIF2A Is a Direct Target of MiR-204-3p
Bioinformatics analysis presented that KIF2A was a potential target of miR-204-3p (Fig. 6A). The prediction was confirmed by luciferase reporter assays. The results showed that miR-204-3p up-regulation obviously reduced the luciferase activity of KIF2A-3′-UTR-WT but exerted little effect on the luciferase activity of KIF2A-3′-UTR- MUT (Fig. 6B). We further investigated the correlation between the expression levels of KIF2A, miR-204-3p and LINC00958 in NSCLC tissues. We found that KIF2A was negatively correlated with miR-204-3p expression (Fig. 6C), while it was positively correlated with LINC00958 expression (Fig. 6D). Moreover, we determined whether miR-204-3p regulated KIF2A expression in NSCLC cells. As expect, KIF2A expression was markedly decreased by transfection with miR-204-3p mimics in A549 cells (Fig. 6E). Subsequently, KIF2A mRNA expression was detected in NSCLC tissues and cell lines. The results revealed that KIF2A had a higher expression level in NSCLC tissues (Fig. 6F) and cell lines (Fig. 6G) than in the corresponding control group.
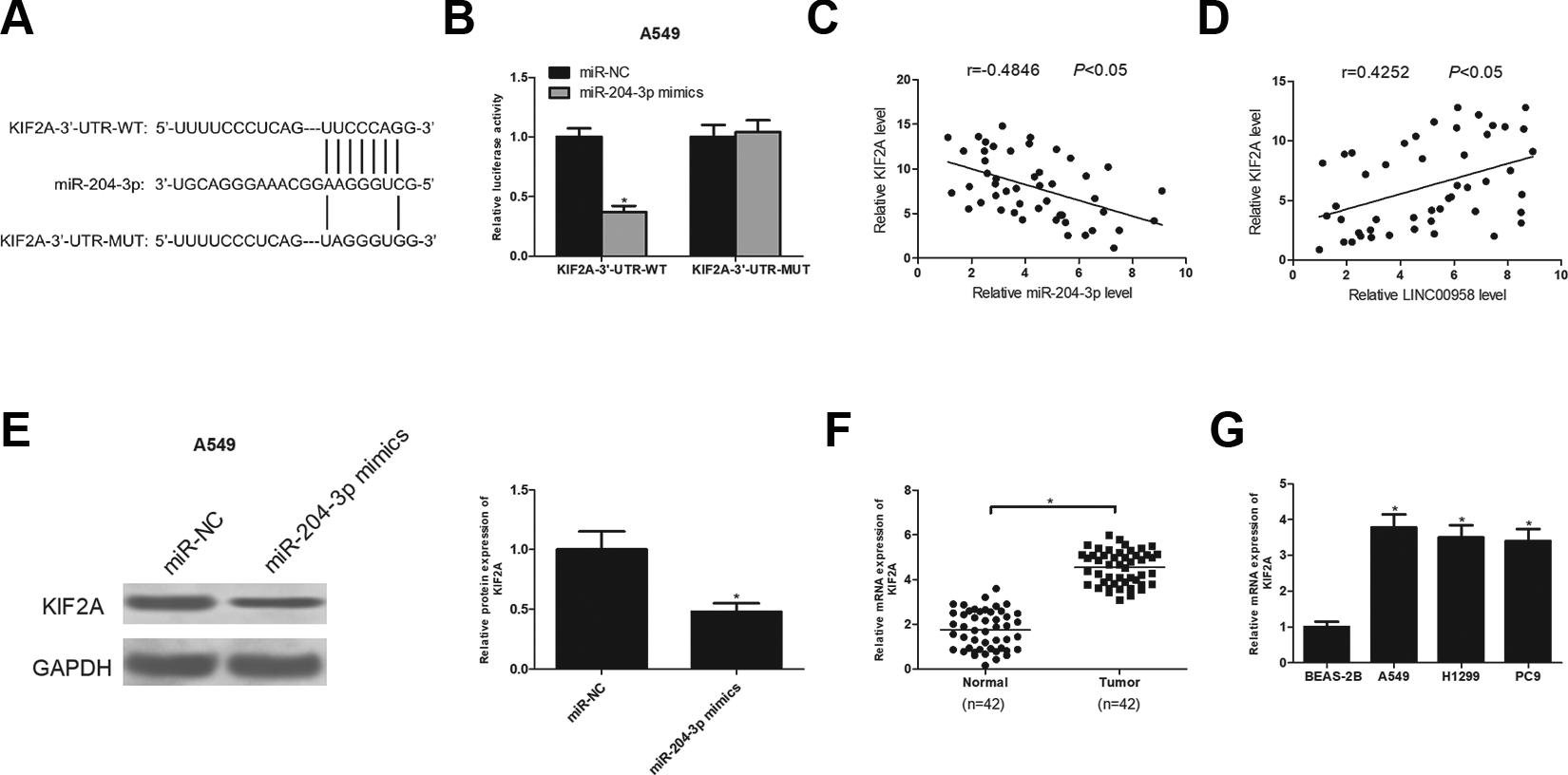
KIF2A is a direct target of miR-204-3p. (A) The potential binding sites between KIF2A and miR-204-3p. (B) Luciferase reporter assay of A549 cells co-transfected with KIF2A-3′-UTR-WT or KIF2A-3′-UTR-MUT and miR-204-3p or miR-NC. (C) The correlation between KIF2A and miR-204-3p expression in NSCLC tissues. (D) The correlation between KIF2A and LINC00958 expression in NSCLC tissues. (E) The protein expression of KIF2A was detected in A549 cells after transfection with miR-204-3p or miR-NC. (F) RT-qPCR analysis of KIF2A mRNA expression in NSCLC tissues and adjacent normal tissues. (G) RT-qPCR analysis of KIF2A mRNA expression in A549, H1299, PC9 and BEAS-2B cell lines. *P < .05.
KIF2A Antagonizes the Effect of MiR-204-3p on NSCLC Cell Proliferation and Migration and Was Regulated by LINC00958/MiR-204-3p
We explored whether miR-204-3p-regulated NSCLC progression was mediated by KIF2A. A549 cells were transfected with miR-204-3p mimics alone or together with KIF2A expression vectors. The CCK-8 and transwell assays indicated that miR-204-3p mimics dramatically reduced the proliferative and migratory abilities of A549 cells while KIF2A transfection antagonized these inhibitory effects (Fig. 7A, B). Besides, we investigated how LINC00958/miR-204-3p affected KIF2A expression in NSCLC cells. The results showed that the protein expression of KIF2A was strikingly decreased by LINC00958 knockdown in A549 cells and the regulatory effect was reversed in response to miR-204-3p down-regulation in A549 cells (Fig. 7C).
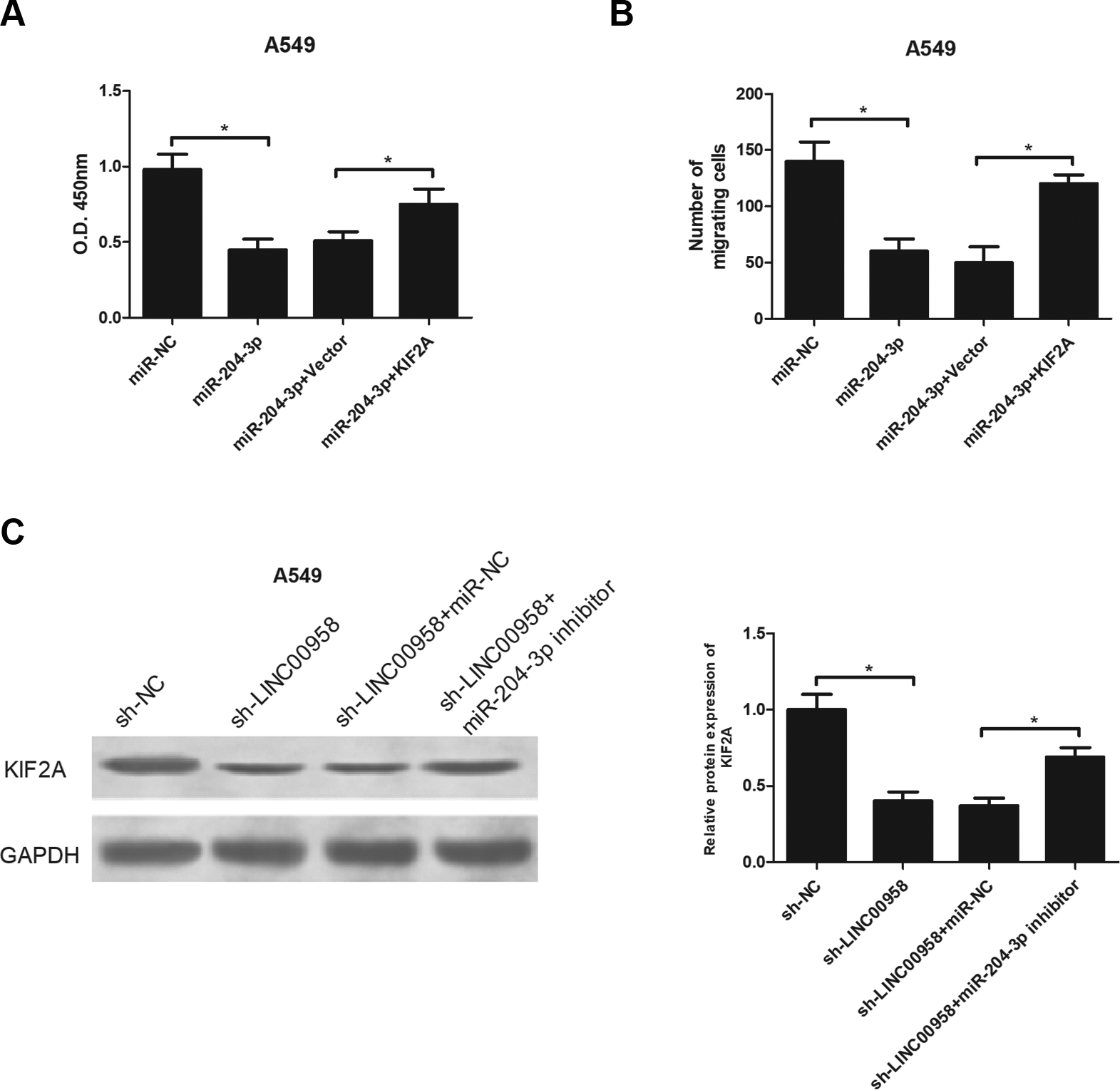
KIF2A antagonizes the effect of miR-204-3p on NSCLC cell proliferation and migration and was regulated by LINC00958/miR-204-3p. A549 cells were transfected with miR-204-3p mimics alone or together with KIF2A expression vectors, followed by detection of cell proliferation (A) and migration (B) by the CCK-8 and transwell assays, respectively. (C) The protein expression level of KIF2A was detected by the western blot analysis in A549 cells after different treatment. *P < .05.
Knockdown of LINC00958 Inhibits NSCLC Cell Growth In Vivo
The impact of LINC00958 on NSCLC cell growth was examined with a tumor xenograft assay. Nude mice were injected subcutaneously with A549 cells after transfection with sh-LINC00958 or sh-NC. As shown in Fig. 8A, tumor volume was reduced in the LINC00958 knockdown group in comparison with the sh-NC group. After dissection, tumor weight was measured. As shown in Fig. 8B, tumor weight was lower in sh-LINC00958 group than in the sh-NC group. LINC00958 and miR-204-3p expression levels in the excised tumors were assessed by RT-qPCR analysis. LINC00958 expression was down-regulated in the sh-LINC00958 group in comparison with the sh-NC group while miR-204-3p expression was greatly up-regulated in the sh-LINC00958 group in comparison with the control group (Fig. 8C, D). Moreover, KIF2A mRNA and protein levels were also examined. The results showed that the expression of KIF2A at both mRNA and protein levels was significantly reduced in the sh-LINC00958 group in comparison with the sh-NC group (Fig. 8E, F). In addition, the immunohistochemical analysis confirmed decreased staining of KIF2A derived from xenograft tumors in the sh-LINC00958 group compared with the sh-NC group (Fig. 8G).
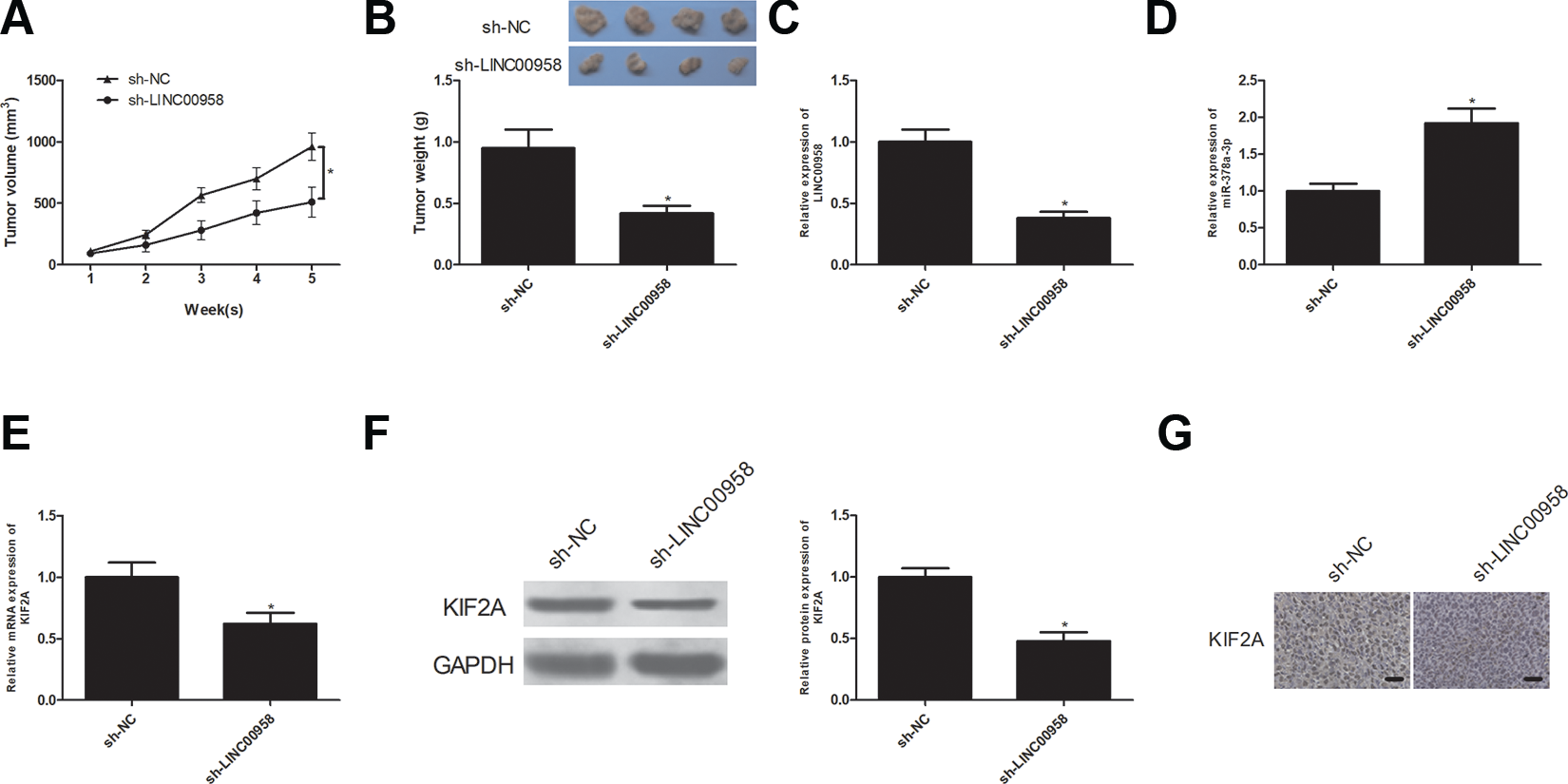
Knockdown of LINC00958 inhibits NSCLC cell growth in vivo. Changes of tumor volume (A) and weight (B) in mice bearing A549 cells with sh-LINC00958 or sh-NC transfection. (C) RT-qPCR analysis of LINC00958 expression levels in the excised tumor tissues. (D) RT-qPCR analysis of miR-204-3p expression levels in the excised tumor tissues. (E, F) RT-qPCR and western blot analysis of KIF2A mRNA and protein expression levels in the excised tumor tissues. (G) The representative images of KIF2A expression in xenograft tumors by IHC. *P < .05.
Discussion
With a high incidence and mortality rate, lung cancer is considered a “leading killer” in the world 20 . As the most common type of lung cancer, NSCLC has attracted an enormous attention 21 . Currently, NSCLC patients at an advanced stage suffer from a poor prognosis and low survival rate due to treatment failure 8,9 . Thus, in-depth understanding of molecular mechanisms underlying NSCLC progression will be of great help for developing effective therapeutic strategies. Increasing evidence demonstrates that lncRNAs play an important role in development and progression of diverse cancers including NSCLC 22 –25 . For instance, TFPI2AS1 was reported to be markedly up-regulated in NSCLC and significantly increase NSCLC cell proliferation and migration 26 . H19 was elevated in NSCLC cell lines and its inhibition suppressed NSCLC progression, suggesting its prognostic and therapeutic value in NSCLC 27 .
LINC00958, a newly identified lncRNA, has been reported to be aberrantly expressed in several types of cancers. For example, Guo et al. revealed that LINC00958 was highly expressed in glioma and its ectopic expression was correlated with poor prognosis of glioma patients 18 . Seitz et al. demonstrated that LINC00958 was overexpressed in bladder cancer and it knockdown reduced both cell viability and migration 19 . Consistently, our study showed that LINC00958 was significantly up-regulated in NSCLC tissues and cell lines and its down-regulation inhibited NSCLC cell proliferation and migration. Furthermore, the suppressive effect of LINC00958 on NSCLC was confirmed by the xenograft tumor assay which indicated that LINC00958 knockdown reduced NSCLC cell growth in vivo. These observations provide new evidence in support of the oncogenic role of LINC00958 in cancer progression.
A growing number of studies have demonstrated that lncRNAs could function as competitive endogenous RNAs (ceRNAs) by binding miRNAs, thus regulating their expression and biological functions 28,29 . Such ceRNA networks formed by the interaction between lncRNAs and miRNAs have been reported in the development of various cancers including NSCLC. For instance, lncRNA PTAR sponged miR-101 to further regulate miR-101 expression and consequently affect NSCLC progression 30 . In this study, we performed bioinformatics analysis and found that LINC00958 was a putative target of miR-204-3p. Many studies have reported involvement of miR-204-3p in tumorigenesis. For example, miR-204-3p was uncovered to be lowly expressed in hepatocellular cancer and inhibit cell proliferation and induce cell apoptosis 31 . MiR-204-3p was also found down-regulated in colon cancer and its overexpression suppressed cell proliferation, migration and invasion 32 . In this study, we conducted luciferase reporter assays and confirmed the interaction between miR-204-3p and LINC00958. Moreover, we found that miR-204-3p expression was decreased in NSCLC and inversely correlated with LINC00958. Additionally, miR-204-3p inhibitor reversed the effect of LINC00958 knockdown on NSCLC cell proliferation and migration. These data indicated that LINC00958 participated in NSCLC development by binding to miR-204-3p and mediating its function.
KIF2A, an M-type nonmotile microtubule depolymerase, is a crucial player in carcinogenesis 33 –35 . The bioinformatics analysis presented that KIF2A was a direct target of miR-204-3p. Our study showed that KIF2A was negatively correlated with miR-204-3p expression while it was positively correlated with LINC00958 expression. Besides, KIF2A antagonized the effect of miR-204-3p on NSCLC progression and was regulated by LINC00958/miR-204-3p. These findings suggested that LINC00958 functioned as a tumor suppressor in NSCLC via regulation of the miR-204-3p/KIF2A axis.
In conclusion, we demonstrated that LINC00958 was up-regulated in NSCLC tissues and cell lines. Down-regulation of LINC00958 inhibited cell proliferation and migration in vitro and suppressed tumor growth in vivo. Mechanically, we revealed that LINC00958 influenced NSCLC progression partly by sponging miR-204-3p and regulating KIF2A expression. Taken together, our study suggested the value of LINC00958 as a potential therapeutic target for NSCLC treatment.
Footnotes
Authors’ Contributions
Qing Wang designed the study and prepared the manuscript. Kai Li collected and analyzed the data. Xiaoliang Li performed experiments. All authors read and approved the final manuscript.
Availability of Data and Materials
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
Ethics Approval
Ethical approval was obtained from the Ethics Committee of Henan Polytechnic University.
Statement of Human and Animals Rights
All experimental procedures with human subjects in this study were conducted in accordance with the Ethics Committee of Henan Polytechnic University. All animal experiments were approved by the Animal Care and Use Committee of Henan Polytechnic University
Statement of Informed Consent
All patients involved in the study provided written informed consent.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
