Abstract
We aimed to investigate the potential pro-oncogenic properties of GREM1 by Pan-cancer multi-omics analysis. Accumulating evidence has highlighted that GREM1 (Gremlin 1), serves as an inhibitor of BMP (Bone Morphogenetic Protein) family, involve in bone related diseases, carcinogenesis, cell stemness, and cell differentiation. However, the effect and underlying mechanism of GREM1 on the cancer biology remain largely elusive. The mRNA expression of GREM1 were extracted from GTEx (Genotype-Tissue Expression) and TCGA (The Cancer Genome Atlas) database. Analysis of OS (Overall Survival), PFI (Progression Free Interval), DSS (Disease-Specific Survival), and ROC (Receiver Operating Characteristic) were performed to predicted prognostic value of GREM1 in various cancers. The TIMER (Tumor Immune Estimation Resource) online tool was used to investigate the relationship between GREM1 transcriptional level and infiltration of immune cells. KEGG (Kyoto Encyclopedia of Genes and Genomes) analysis and GO (Gene Ontology) analysis were used to investigate the GREM1 related molecular events, and then constructed a PPI (Protein-Protein Interaction) network via the STRING (Search Tool for the Retrieval of Interaction Genes/Proteins) online tool. Western blot was performed to investigate the indicated protein expression. In the present study, our results showed that GREM1 tended to be upregulated in various cancers, which would correlate with the poor prognosis. Mechanistically, our results showed that GREM1 involve in regulating the ECM-receptor interaction pathway, upregulation of MMP activity, angiogenesis, and immune cell infiltration. In vitro studies, our results further showed that BMP agonist significantly decreased the protein level of GREM1 in GES-1 cells and BGC cells, which accompanied by inhibiting migration and proliferation in GES-1 cells and BGC cells. BMP inhibitor significantly promoted GREM1 expression and migration in BGC cells, but not GES-1 cells. GREM1 might serve as a potential and promising prognostic biomarker for drug development and cancer treatment.
Introduction
Global new cases and new deaths of cancer were about 19.3 million and 10 million in 2020, respectively. 1 Among the most common cancers, the mortality rates of breast cancer, prostate cancer and uterine cancer are gradually increasing. Thanks to many advanced progressions in the diagnosis and treatment of cancer, including the innovations of drugs, radiotherapy and chemotherapy techniques, and surgical methods, the 5-year OS rate of cancer has been increased from 49% in the mid-1970s to 68% in 2012–2018. 2 However, the prospects of pancreatic cancer (12%), liver cancer, and esophageal cancer (21%) remain indeterminacy, due to most of these patients usually suffer recurrence and correlate with higher mortality rate. 2 Gastric cancer is a highly malignant digestive system tumor, as the incidence rate of gastric cancer in China accounts for 44% of the world, ranking top1 in the world. 3 Meanwhile, nursing and treatment of cancer patients have result in a heavy global economic burden. Therefore, there is an urgent need to find new methods for diagnosing and treating cancer.
Recently, emerging research have suggested that the GREM1 involved in cancer initiation and progression. GREM1 exhibits a debating role as oncogene and anti-oncogene, which might depend on the specific tumor microenvironment. 4 Several previous studies have suggested that GREM1 involve in EMT (Epithelial-Mesenchymal Transition), fibroblast activation in fibrosis, and tumor angiogenesis.5–7 GREM1 is involved in antagonizing BMP-2, 4, and 7, 4 preventing the interactions between these ligands and their receptors, which in turn to inactivate TGF-β (Transforming Growth Factors-β) signal. 8 Besides, GREM1 was activated by VEGFR2 (Vascular Endothelial Growth Factor Receptor-2) to promote cancer-related invasion and angiogenesis.7,9 In pancreatic cancer, GREM1 inhibited the expression of EMT transcription factors by suppressing BMP activity in adjacent epithelial cells, thereby modulating heterogeneity in pancreatic cancer. 5 Breast cancer-associated fibroblast secreted GREM1 to promote stemness and invasion in breast cancer. 10 GREM1, as a downstream effector, would regulate the growth and metastasis of colorectal cancer by targeting the BMP signaling pathway. 8 In lung tumor microenvironment, fibroblasts-derived GREM1 promoted proliferation of malignant lung adenocarcinoma cells.11,12 However, the underlying molecular events of GREM1 on the cancer biology remains largely unknown.
Pan-cancer analysis, depended on the TCGA database, would help to discover DEG (Differentially Expressed Genes) in various cancers.13,14 The connection between genomic alterations and gene expression profiles in cancer can serve as a foundation for Pan-cancer analysis, which in turn help to uncover commonalities and differences among cancers, provide a crucial scientific basis for discovering new biomarkers, therapeutic targets, and develop personalized and combined therapies. 15
In the present study, we carried out Pan-cancer analysis of GREM1, including gene expression analysis, survival rate analysis, immune infiltration analysis, and enrichment analysis, to explore its roles and effects on regulating tumorigenesis, immune microenvironment, prognosis of various cancers.
Materials and methods
Multiomics analysis of GREM1 expression
GREM1 mRNA expressions were extracted from GTEx (https://www.gtexportal.org/) and TCGA (https://portal.gdc.cancer.gov/) in paired cancer tissues and para-carcinoma tissues, respectively. Then, we entered the GEPIA2 website (http://gepia2.cancer-pku.cn/) to perform the expression analysis and visualize the GREM1 expression results by using log2 (TPM + 1) logarithmic scale transformation of the expression data. By searching for GREM1 and selecting indicated cancers, it would output the expression of GREM1 in various cancer tissues. The protein level of GREM1 was acquired from HPA (Human Protein Atlas, www.proteinatlas.org/) images, these IHC (Immunohistochemistry) images were quantified by IHC toolbox of ImageJ (version 1.53t).
Prognostic analysis
The online Kaplan–Meier curve analysis tool (https://kmplot.com/analysis/) for OS, PFI, and DSS were carried out to evaluate the relationships between GREM1 expression and clinical common outcome in various cancers. 16 Besides, the ROC curves of GREM1 were also plotted in various cancers.
Immune infiltration analysis
The online data derived from TCGA, the TIMER (version 2.0; http://timer.cistrome.org/ online tool were applied to investigate the relationship between GREM1 transcriptional level and infiltration of immune cells via various algorithms, at least including TIMER, EPIC, TIDE, CIBERSORT, CIBERSORT abs, QUANTISEQ, XCELL, and MCPCOUNTER.
Functional enrichment analysis and protein-protein interaction network analysis
The top 100 genes involved in GREM1 regulation with the most similar expression patterns were extracted from the GEPIA2 database. We then performed the KEGG analysis and GO analysis, including BP (Biological Pathways), CC (Cellular Components), and MF (Molecular Functions) were performed to exhibit GREM1 related molecular events. Otherwise, the above 100 genes were used to establish a PPI network via the STRING (version 12.0; https://cn.string-db.org/) online tool, the minimum interaction threshold required was set to 0.4. We also extracted and combined the expression datasets of BRCA, LUAD, HNSC, LUSC, BLCA, OV, PAAD, TGCT, ALL, CHOL, STAD from GEPIA database. Then we performed correlation analysis of GREM1 and CALD1, ACTN1, SYNPO2, CNN1, MYH11, VCL, respectively.
Western blotting
Total protein lysis of GES-1 or BGC cells were prepared by RIPA buffer, the lysis buffer was pre-mixed with protease inhibitors, phosphatase inhibitors, and PMSF (Phenylmethanesulfonyl Fluoride). 10 µg total proteins lysis per well was loaded and separated on 10% sodium dodecyl sulfate-polyacrylamide gels under the constant voltage mode, the separated proteins in gel were then transferred to 0.45 µm PVDF membranes (Millipore) under the constant electric current mode. All membranes were blocked by 5% skimmed milk/TBST (w/v) for 1 h at room temperature, and then following the primary antibodies incubation at 4°C overnight: anti-Grem1 (1:1000, Abnova, H00026585), anti-β-Actin (1:3000, Beyotime, AF0003). After TBST washing three time per 5 min, following secondary antibodies incubation 1 h at room temperature: HRP-linked anti-rabbit (IgG; 1:5000, Beyotime, A0208) or HRP-linked anti-mouse IgG (1:1000, Beyotime, A0216). After TBST washing three time per 5 min, all membranes were incubated with ECL solution (Beyotime, P0018S) and then immediately detected by using the GelDoc XR Bio-Rad.
Statistical analysis
The Wilcoxon rank sum test 17 was performed to analyze the difference between the two groups, the Spearman rank sum test 18 was performed to analyze the correlation between the two groups. Univariate and multivariate Cox proportional hazards regression were performed to screen factors that affect prognosis. 19 Kaplan Meier analysis 16 and log rank test 20 were performed to carry out survival analysis. P value under 0.05 was defined as statistically significant differences.
Results
The expression of GREM1 in various cancers
We firstly comprehensively investigated the expression patterns of GREM1 in various cancers via TCGA_GTEx data analysis. Our results showed that GREM1 exhibited significant differential expression in most tumors, it was significantly upregulated in 16 types of cancer, including STAD (Stomach Adenocarcinoma), BLCA (Bladder Urothelial Carcinoma), LUAD (Lung Adenocarcinoma), and ESCA (Esophageal Carcinoma). while it was significantly downregulated in 12 types of cancer, including GBM (Glioblastoma Multiforme), ACC (Adrenocortical Carcinoma), KIRP (Kidney Renal Papillary Cell Carcinoma), and CESC (Cervical Squamous Cell Carcinoma and Endocervical Adenocarcinoma; Figure 1a). To further validate this result, we investigated the expression of GREM1 from TCGA data, GREM1 was also upregulated in BLCA, LUAD, LUSC (Lung Squamous Cell Carcinoma), HNSC (Head and Neck Squamous Cell Carcinoma), BRCA, STAD. Otherwise, GREM1 was significantly upregulated in COAD (Colon Adenocarcinoma) while carrying out TCGA data analysis, but there was no significant difference of GREM1 while carrying out TCGA_GTEx data analysis (Figure 1b–h). Our results also showed that the protein level of GREM1 were elevated in STAD and LUAD (Figure 1i).

Relationship between GREM1 expression and cancer prognosis
To further explore the prognostic value of GREM1 in various cancers, the expression of GREM1 and the total OS for 33 cancer types were performed. Our results found that the abnormal GREM1 expression significantly correlated with poor prognosis in ACC, LUSC, KIRP, and KIRC (Kidney Renal Clear Cell Carcinoma), but not LUAD, HNSC, COAD, or BRCA (Figure 2a–j). We further investigated the correlation between GREM1 expression and DSS. Our results showed that GREM1 expression was significantly correlated with the DSS in BLCA, KIRC, KIRP, LGG, PAAD, and UVM (Figure 2k). Besides, the GREM1 expression also significantly correlated with PFI in KIRC, KIRP, LGG, PAAD, OV, and UVM, but not in BLCA, COAD, READ, SARC, and THYM (Figure 2l).

Relationship between GREM1 expression and the prognosis. (a) Analyzing the relationship between GREM1 expression and OS using a forest plot in the indicated cancers. (b–j) Analyzing the relationship between GREM1 expression and OS using survival plots in the indicated cancers. (k) Analyzing the relationship between GREM1 expression and DSS using a forest plot in the indicated cancers. (l) Analyzing the relationship between GREM1 expression and PFI using a forest plot in the indicated cancers.
The effect of GREM1 on regulating tumor immune infiltration
We further investigated the correlation between GREM1 expression and tumor immune infiltrations. Our results showed that there were significant and positive correlation between GREM1 expression and B cells infiltrations in BLCA, COAD, GBM, KIRC, KIRP, LGG, LIHC, LUAD, LUSC, PAAD, SKCM, STAD, THCA, and THYM, while there were significant and negative correlation between GREM1 expression and B cells infiltrations in BRCA, CESC, HNSC, READ, TGCT, and UVM (Figure 3a). There were significant and positive correlation between GREM1 expression and T cell CD4+ infiltrations in HNSC, KIRC, LIHC, PAAD, and SKCM, while there were significant and negative correlation between GREM1 expression and T cell CD4+ infiltrations in LGG, THCA, and THYM (Figure 3b). There were significant and positive correlation between GREM1 expression and T cell CD8+ infiltrations in BLCA, COAD, KICH, KIRC, KIRP, LGG, LIHC, LUAD, LUSC, PAAD, PCPG, PRAD, STAD, and UVM, while there were significant and negative correlation between GREM1 expression and T cell CD8+ infiltrations in BRCA, CESC, HNSC, SKCM, TGCT, and THYM (Figure 3c). There were significant and positive correlation between GREM1 expression and macrophages infiltrations in ACC, BLCA, BRCA, CESC, COAD, ESCA, HNSC, KICH, KIRC, KIRP, LIHC, LUAD, LUSC, MESO, OV, PAAD, PCPG, PRAD, READ, SKCM, STAD, TGCT, THYM, UCEC, UCS, and UVM, while there were significant and negative correlation between GREM1 expression and macrophages infiltrations in LGG and SARC (Figure 3d). There were significant and positive correlation between GREM1 expression and monocytes infiltrations in BLCA, BRCA, CHOL, COAD, ESCA, HNSC, LIHC, OV, PAAD, PCPG, PRAD, READ, SKCM, STAD, TGCT, THCA, THYM, UCS, and UVM, while there were significant and negative correlation between GREM1 expression and monocytes infiltrations in ACC, LGG, LUAD, LUSC, MESO, and SARC (Figure 3e).
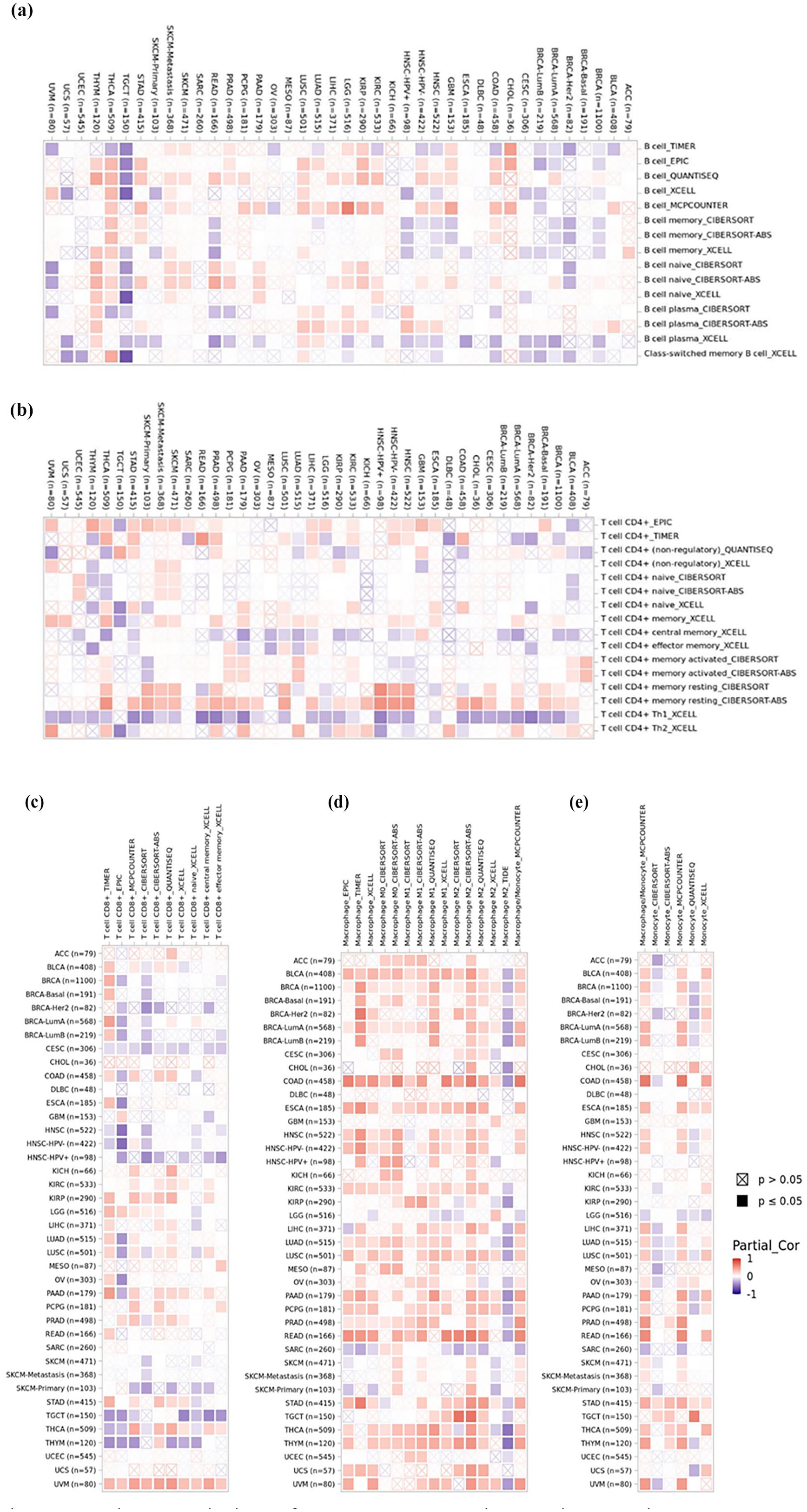
The correlation of GREM1 expression and tumor immune infiltration. (a–e) Heatmaps of correlations between GREM1 expression and B cells, T cell CD4+, T cell CD8+, macrophage, and monocyte in TIMER2 database, respectively.
Functional enrichment analysis and protein interaction analysis of GREM1 Target genes
To investigate the biological function of GREM1, we performed GO/KEGG analysis. Our results showed that GREM1 targeted genes be involved in regulating cell substrate adhesion, focal adhesion, actin binding, and vascular smooth muscle contraction (Figure 4a and b). Besides, we constructed a PPI network of these genes to showed positive interactions between CALD1, ACTN1, SYNPO2, CNN1, MYH11, and VCL proteins (Figure 4c). As we acquired gene expression data of the GREM1 positive cancers (including BRCA, LUAD, HNSC, LUSC, BLCA, OV, PAAD, TGCT, ALL, CHOL, STAD) from the GEPIA database to confirm the above correlations of GREM1 and CALD1, ACTN1, SYNPO2, CNN1, MYH11, VCL, respectively. The R values were 0.63, 0.39, 0.49, 0.51, 0.53, and 0.46, respectively, and each P value was less than 0.05 (Figure 4d–i).
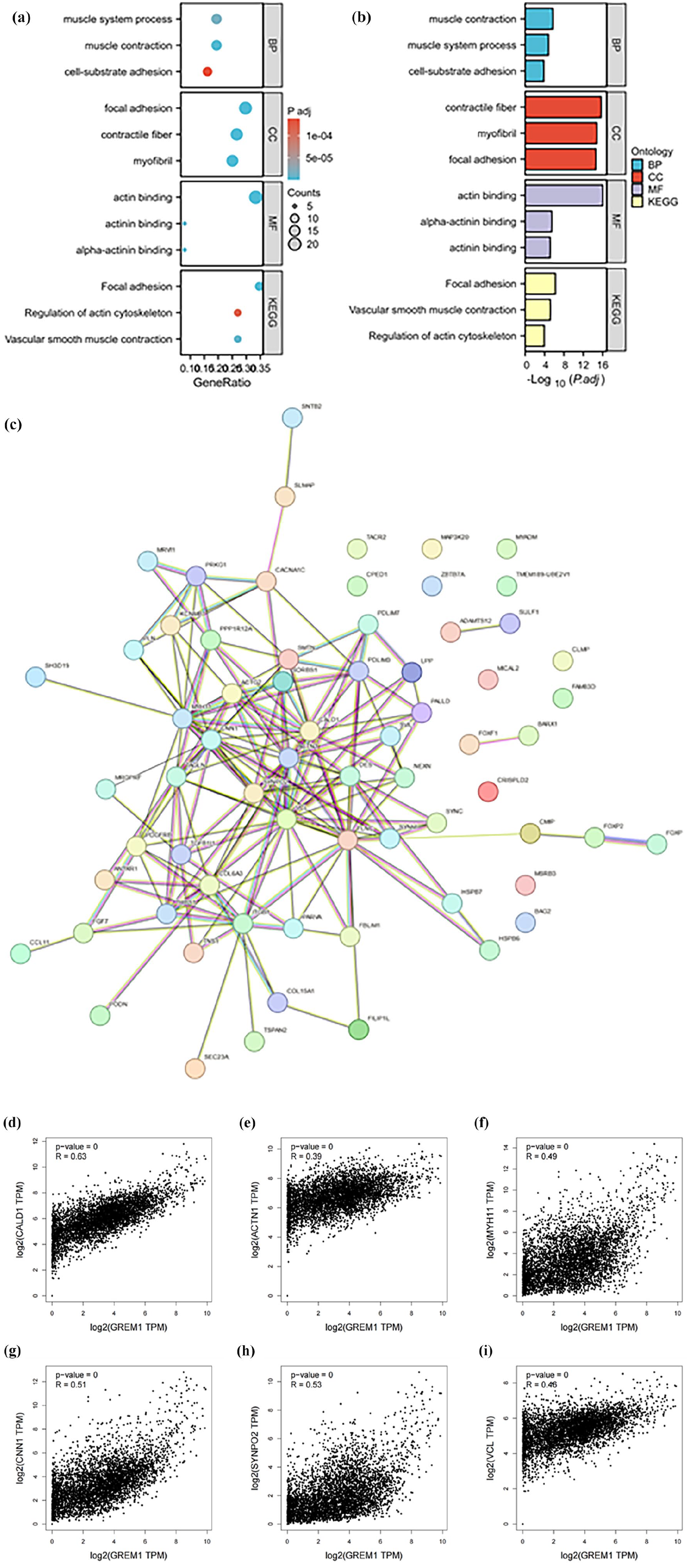
Functional enrichment analysis and protein interaction analysis of GREM1 target genes. (a) GO/KEGG analysis of GREM1 targeted genes. (b) Significance of GO/KEGG analysis of GREM1 targeted genes. (c) Protein-protein interaction networks of GREM1 targeted genes. (d–i) The correlation analysis of GREM1 and CALD1, ACTN1, SYNPO2, CNN1, MYH11, VCL, respectively.
Expression of GREM1 in gastric cancer
Our results further showed that BMP agonist (SJ000291942) significantly reduced GREM1 protein level in GES-1 cells, while BMP inhibitor (LDN-193189) had no effect on GREM1 expression (Figure 5a). In BGC cancer cells, BMP agonist significantly reduced GREM1 protein level, while lower dose BMP inhibitor significantly increased GREM1 expression, but higher dose BMP inhibitor had no effect on GREM1 expression (Figure 5b). Higher dose BMP agonist significantly reduced wound healing rate of GES-1 cells, but lower dose BMP agonist or BMP inhibitor had no effect on wound healing of GES-1 cells (Figure 5c and e). BMP agonist also significantly reduced wound healing rate of BGC cells, while lower dose BMP inhibitor significantly promoted wound healing of BGC cells, but higher dose BMP inhibitor had no effect on wound healing of BGC cells (Figure 5d and f). The results further showed that BMP agonist dose-dependently and significantly reduce proliferation of GES-1 cells and BGC cells (Figure 5g and h). BMP inhibitor slightly, dose-dependently, and significantly decrease proliferation of GES-1 cells, BMP inhibitor also showed an inhibition effect on proliferation of BGC cells, but lower dose BMP inhibitor seemed to be more effective. (Figure 5g and h).

Expression of GREM1 in gastric cancer. (a) The effect of BMP agonist or inhibitor on regulating GREM1 protein level in GES-1 cells, β-ACTIN was using as loading control. (b) The effect of BMP agonist or inhibitor on regulating GREM1 protein level in BGC cells, β-ACTIN was using as loading control. (c) The effect of BMP agonist or inhibitor on wound healing in GES-1 cells. (d) The effect of BMP agonist or inhibitor on wound healing in BGC cells. (e) Quantification of wound healing percentage from (c). (f) Quantification of wound healing percentage from (d). (g) Cell-Counting-Kit-8 assay to analyze the effect of BMP agonist or inhibitor on proliferation of GES-1 cells. (h) Cell-Counting-Kit-8 assay to analyze the effect of BMP agonist or inhibitor on proliferation of BGC cells.
Discussion
Accumulating studies have made great efforts to screen the potential and promising cancer hub genes that fundamentally improved drug designs and cancer treatments. In the present study, we first used multiple online databases such as TGCA, TIMER2, and GEPIA to explore the expression and related bio-functions of GREM1 in various cancers. Our results and previous studies showed GREM1 was elevated in most tumor tissues, which tended to exhibit a oncogene characteristic during cancer progression.4,21 High expression of GREM1 is associated with poor prognosis of breast cancer.10,22 In line with previous studies, our results show a positive correlation between GREM1 expression and poor prognosis in nine types of cancer. Bao et al. 23 found that GREM1 was increased in NSCLC and served as an oncogene in NSCLC occurrence and development. GREM1 promoted immune cell infiltration and correlated with poor prognosis in NSCLC 23 ; Jiang et al. 24 showed that GREM1 expression was positively correlated with poor prognosis in bladder cancer, GREM1 down-regulation significantly inhibited proliferation, apoptosis, migration, invasion, and EMT in bladder cancer cells. These research findings collectively indicate that GREM1 tends to serve as a promising oncogenic candidate for pan-cancer screening.
The immune microenvironment mainly contributed to the occurrence and development of tumors. Recently, emerging studies have shown GREM1 responded for tumor initiation, occurrence and development. Yet its specific mechanism of its action in the immune microenvironment is still not fully understood. Our results suggested that GREM1 expression significantly and positively correlated with B cells, CD4+T cells, CD8+T cells, macrophages, and monocytes in various cancers, such as PAAD, STAD, and THCA, which provide a potential application of CAR-T (Chimeric Antigen Receptor T-Cell Immunotherapy) or CAR-M (Chimeric Antigen Receptor Macrophage Immunotherapy) in GREM1 positive expression cancers.
Trend of gastric cancer burden in China becomes worse, as the incidence rate of gastric cancer in China accounts for 44% of the world, ranking top1 in the world. 3 After Pan-cancer analysis in the present study, our results showed that GREM1 was upregulated in gastric cancer. Taken these together, they reasoned us to focus and investigate the molecular events of GREM1 in gastric cancer. Previous study showed that GREM1 acted as the inhibitor of BMPs. 4 Because of specific agonist or inhibitor of GREM1 is still not available, we then intervened with BMP pathway inhibition/activation to mock GREM1 activation or inactivation in the normal gastric mucosal epithelial cells (GES-1 cells) and gastric cancer cells (BGC cells). The results showed that BMP agonist had a negative regulation on the expression of GREM1, thus our finding together with those of previous study suggested that there is a reciprocal inhibition between GREM1 and BMP signaling. However, there was a difference of BMP inhibitor on regulating GREM1 expression in the normal gastric mucosal epithelial cells and gastric cancer cells, our results showed that BMP inhibitor had no effect on GREM1 expression in the normal gastric mucosal epithelial cells, which it would tend to increase GREM1 expression in the gastric cancer cells, thus these results reason us to raise our concerns when apply any agency would inhibit BMP signaling in gastric cancer treatment.25–32
Previous study showed that GREM1 expression in gastric cancer is associated with EMT, angiogenesis, and lymph-angiogenesis, and affects the progression and prognosis of gastric cancer. 33 In the present study, our results showed that BMP agonist significantly decreased migration and proliferation in the normal gastric mucosal epithelial cells and gastric cancer cells, based on the reciprocal inhibition between GREM1 and BMP signaling, GREM1 inhibition might contribute to decrease migration and proliferation in these cell lines. Interestingly, BMP inhibitor tended to promote migration in the gastric cancer cells, but not the normal gastric mucosal epithelial cells, thus these results reason us to develop specific GREM1 inhibitor for gastric cancer treatment in the future.
It is common knowledge that BMPs regulates embryonic development and organ formation under physiological conditions. 34 In pathological conditions, aberrant activation of BMPs related pathways can be observed in the occurrence and progression of pan-cancer. The BMP-4 related signaling pathway has been shown to inhibit the VEGF related genes. 35 BMP-7 target Smads/MAPK14 pathway to promote HCC proliferation and migration. 36 However, our results further showed that the BMP inhibitor also decreased proliferation in these cell lines. Actually, BMP inhibitor could not be equal to GREM1 activation in the normal gastric mucosal epithelial cells or gastric cancer cells. Because our results also found another GREM1 targeted candidates, at least including CALD1, ACTN1, and CNN1. Previous study showed that RECK inhibited CALD1 activation to decrease proliferation, migration, and invasion in gastric cancer cells. 37 ACTN1 upregulation increase proliferation, migration, and invasion in gastric cancer cells. 38 Our previous study also found that CNN1 contributed to angiogenesis in gastric cancer cells. 39 Thus, GREM1 might bypassed BMP signaling to activate CALD1, ACTN1, and CNN1 to promote proliferation, migration, and invasion in gastric cancer cells.
In summary, we have conducted a Pan-cancer analysis of GREM1 to show its multiple roles in prognosis and immunocytes infiltration, we also found GREM1 was important to proliferation and migration in gastric cancer. Thus, the present study in turn provided a new prognostic screen biomarker, potential and promising target for drug research and development and cancer treatment.
Novelty and limitation
In the present study, we uncovered the expression profiles of GREM1 in various cancers, which contributed to cancer cell proliferation, migration, and tumor immune infiltration. Thus, GREM1 tended to serve as an oncogenic effect for cancer prevention, diagnosis, and treatment.
However, the findings of the present study may be limited by existing data analysis, more dataset and bigger samples need to recruit to confirm our results in the future. In vitro study, it needs more loss or gain function assay to further confirm the role of GREM1 on regulating proliferation, migration, chemotherapy resistance in various cancers, including but not limit to gastric cancer cells.
Conclusion
In the present pan-cancer analysis, our results showed that GREM1 tended to be upregulated in various cancers to serve as an oncogene to result in poor prognosis. In the potential molecular events, GREM1 would upregulate its target genes, at least including CALD1 and CNN1 to play important roles in regulating angiogenesis, immune cell infiltration, cancer cell migration and proliferation. Moreover, our results rose a concern that BMP inhibitor elevated GREM1 level to promote cancer cells migration. Thus, it would need to monitor GREM1 expression while apply BMP inhibitor related therapies in future.
Footnotes
Acknowledgements
We acknowledge the help and support from KY Chen and N Li, both from Science and Technology Innovation Center, Guangzhou University of Chinese Medicine, for manuscript proofreading. However, the responsibility for the content and any remaining errors, omissions, and inaccuracies is our own.
Author contributions
Huafeng Pan conceived and designed the study. Menglu Zhu, Hengli Zhou, Yue Zhuo, Changhua Liu, Jiaxin Li, Peiyao He, Naihua Liu, Ziming Zhao performed bioinformatics analysis. Menglu Zhu and Naihua Liu prepared the manuscript, Menglu Zhu submitted the manuscript. All authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by National Science and Technology Major Project of China (2024ZD0521000), Innovation Team and Talents Cultivation Program of National Administration of Traditional Chinese Medicine (ZYYCXTD-C-202208), Guangdong Provincial Key Laboratory of TCM Pathogenesis and Prescriptions of Heart and Spleen Diseases (2022B1212010012), Special Project for Research and Development in Key areas of Guangdong Province (2020B1111100011). All funders had no role in the design of the study and collection, analysis, interpretation of data and in writing the manuscript, the decision to submit the manuscript for publication.
Availability of data and materials
The data used to support the findings of this study are available from the corresponding author upon request.
