Abstract
The incidence and mortality rate of nonsmall cell lung cancer (NSCLC) are continuously increasing. Recently, the important roles of long noncoding ribonucleic acid (lncRNA) zinc finger antisense1 (ZFAS1) in the development of many disease have been proved. However, the roles of ZFAS1 in NSCLC are still not completely understood. Thus, this study aimed to explore the potential roles and underlying mechanisms of lncRNA ZFAS1 in the progression of NSCLC. Our results demonstrated that lncRNA ZFAS1 expression was significantly upregulated in NSCLC tissues and cell lines. Loss-of-function experiments revealed that lncRNA ZFAS1 inhibition could remarkably suppress NSCLC cells proliferation in vitro. Bioinformatic analysis and luciferase reporter assay revealed that lncRNA ZFAS1 directly interacted with miR-590-3p. Rescue experiments showed that miR-590-3p inhibitor reversed the cell proliferation function of lncRNA ZFAS1 knockdown in vitro. Furthermore, we confirmed that lncRNA ZFAS1 inhibited cell division cycle 42 (Cdc42) expression by regulating of miR-590-3p in NSCLC cells. Therefore, our study indicates that lncRNA ZFAS1/miR-590-3p axis is involved in NSCLC cell proliferation. It also suggests that lncRNA ZFAS1 is a putative tumor oncogene in NSCLC.
Introduction
Nonsmall cell lung cancer (NSCLC) accounts for more than 80% of all lung cancer types and is the leading cause of cancer death in the world 1,2 . Despite the enormous improvements made due to the advances in surgical techniques and treatment strategies, the 5-year survival rate of NSCLC is only slightly greater than 15% 3,4 . The outlook for patients with NSCLC is dismal. Therefore, the identification of new specific biomarkers is important for improving the treatment of NSCLC.
Long noncoding ribonucleic acids (lncRNAs) are evolutionarily conserved noncoding RNAs with length greater than 200nt and have no protein-coding capacity 5 . Accumulated evidence showed that lncRNAs play critical roles in various types of cancers. For instance, the lncRNA urothelial cancer associated 1 (UCA1) has been verified to promote proliferation, migration, and immune escape and inhibit apoptosis in gastric cancer by sponging antitumor microRNAs (miRNAs) 6 . LncRNA solute carrier family 16 member 1 (SLC16A1-S1) has been shown to be a novel prognostic biomarker in NSCLC 7 . Lu et al. found that circulating lncRNA ABHD11 antisense RNA 1 (Tail To Tail) (ABHD11-AS1) served as a biomarker for early pancreatic cancer diagnosis 8 . Thus, a lot of evidences are providing the emerging significant roles of lncRNA in tumor progression. However, the functional roles of lncRNAs in NSCLC remain to be clarified.
miRNAs are small noncoding RNAs with 19–22nt in length. They have been reported to play critical roles in regulating diverse cellular processes by interacting with the 3′-untranslated region (3′-UTR) of target messenger RNAs (mRNAs) to suppress expression 9 . Numerous evidences demonstrated that lncRNA containing miRNA binding sites could act as competitive endogenous RNA and some studies showed that lncRNA and miRNA interact 10 –13 . In colon cancer, lncRNA colon cancer associated transcript 2 (CCAT2) has been reported to regulate miR-145 expression by suppressing its maturation process 14 . In breast cancer, lncRNA long intergenic non-protein coding RNA 968 (LINC00968) has been shown to reduce cell proliferation, migration, and angiogenesis through upregulation of prospero homeobox 1 (PROX1) by reducing microRNA (miR)-423-5p 15 . In bladder cancer, zinc finger antisense1 (ZFAS1) promotes cell proliferation and invasion and inhibits cell apoptosis by sponging miR-329 16 . However, the potential molecular mechanism of lncRNAs still remains largely unknown in NSCLC.
In this study, we aimed to explore the regulatory function of ZFAS1 in the progression of NSCLC via regulating miR-590-3p, suggesting the therapeutic potential of ZFAS1/miR-590-3p for future clinical application.
Materials and Methods
Tissues and Cell Culture
Twenty-two pairs of NSCLC and matched tumor-adjacent tissues were collected at the Affiliated Hospital of Inner Mongolia Medical University in China. All the specimens were reviewed and confirmed by pathologists and immediately frozen in liquid nitrogen. All patients provided informed consent forms and agreed to participate in this study.
Two NSCLC cell lines (A549 and HCC827) and human bronchial epithelial cells (16HBE) were purchased from the China Center for Type Culture Collection (CCTCC, Wuhan, China). Cells were maintained in Dulbecco’s modified Eagle’s medium (Gibco, Shanghai, China) enriched with 10% fetal bovine serum (Sigma-Aldrich, Shanghai, China), 100 U/ml penicillin, and 100 mg/ml streptomycin (Sigma-Aldrich, Shanghai, China). Cells were cultured at 37°C with 5% CO2. All cells were subcultured at 90% confluence.
Quantitative Real-Time Polymerase Chain Reaction (PCR) (qRT-PCR)
The total RNA from tissues and cells was extracted using the mirVana miRNA isolation kit (Ambion, Austin, TX, USA) according to the manufacturer’s protocol. A high-capacity RNA-to-cDNA kit (Thermo Fisher Scientific, Shanghai, China) was used to transcribe the total RNA to cDNA. For ZFAS1 expression analysis, the quantification of complementary DNA (cDNA) was carried out using SYBR Green Master Mix (Applied Biosystems, Shanghai, China) with glyceraldehyde 3-phosphate dehydrogenase (GAPDH) as a housekeeping gene for normalization (ZFAS1 forward 5′-AACCAGGCTTTGATTGAACC-3′, reverse 5′-ATTCCATCGCCAGTTTCT-3′; GAPDH forward 5′-GGTCTCCTCTGACTTCAACA-3′, reverse 5′-GTGAGGGTCTCTCTCTTCCT-3′).
To determine the expression of miR-590-3p, total RNA was reverse-transcribed using the Taqman advanced miRNA cDNA synthesis kit (Applied Biosystems, Shanghai, China) according to the manufacturer’s protocols. U6 small nuclear RNA was used as the endogenous reference for normalization. U6 was reverse-transcribed by Taqman microRNA reverse transcription kit (Applied Biosystems, Shanghai, China) following the manufacturer’s protocol. All the specific primers for miR-590-3p expression are commercially available from Applied Biosystems (Shanghai, China). The expression levels of ZFAS1 and miR-590-3p were defined from the threshold cycle (C t), and relative expression levels were calculated using the 2−ΔΔ C t method 17 .
Cell Transfection
We purchased the ZFAS1-small interfering RNA (si-ZFAS1) and negative control siRNA (si-NC) from GenePharma (Shanghai, China). We constructed the ZFAS1 overexpression vector as previously described 18 . The empty pcDNA3.1 vector (Invitrogen, Carlsbad, CA, USA) was used as the control (Ctl-OE). In order to overexpress or knockdown miR-590-3p, we used the commercially available lentiviral vectors to construct the miR-590-3p-mimic vector (miR-590-3p-OE) and the miR-590-3p inhibitor vector (miR-590-3p inhibitor; GenePharma, Shanghai, China). The empty vector was used as miRNA control (miR-con). The cell transfection was carried out according to previous studies 18 .
Luciferase Reporter Assay
ZFAS1 wild-type luciferase reporter vector (ZFAS1-WT) containing miR-590-3P–targeted sequence and its mutant in the seeded region (ZFAS1-MUT) were constructed by Applied Biosystems (Foster City, CA, USA). To verify the targeted interrelation between ZFAS1 and miR-590-3p, cells were transfected with ZFAS1-WT or ZFAS1-MUT, together with miR-590-3p-mimic or miR-con using Lipofectamine 3000 Transfection Reagent (Invitrogen, Shanghai, China). After transfection for 48 h, cells were lysed and measured by the Dual-Luciferase Reporter Assay Protocol (Promega, Madison, WI, USA).
Cell Viability Assay
The proliferation of cell lines after transfection with plasmids, siRNAs, mimics, or inhibitor was determined using the Cyquant assay (Thermo Fisher Scientific, Shanghai, China). Cells were seeded in the 96-well plate (BD Biosciences, Shanghai, China) at a density of 5,000 cells per well. Plates were frozen at certain indicated times (24, 48, 72, and 96 h) after following incubation. To the wells, 100 µl of freshly prepared Cyquant solution was added and incubated in the dark for 45 min at room temperature. Plates were read at excitation at 497 nm and emission at 520 nm. Data are reported as fold change of 0 h.
RNA-Binding Protein Immunoprecipitation (RIP) Assay
RIP assays were carried out to determine the interaction between ZFAS1 and miR-590-3p as in the previous study 19,20 . We used EZMagna RIP RNA-binding protein immunoprecipitation kit (Millipore, Chengdu, Sichuan, China) according to the manufacturer’s protocol. Cells were subjected to different treatments and then lysed using RNA lysis buffer with 1X protease inhibitor cocktail. The cell lysate was incubated with magnetic beads conjugated with human Ago2 antibody (Millipore, Chengdu, Sichuan, China) or negative control immunoglobulin G (IgG) (Abcam, Beijing, China) to immunoprecipitate the RNA-induced silencing complex. After overnight incubation at 4°C, the coprecipitated RNAs were reverse-transcribed and analyzed by qRT-PCR.
Western Blotting
Cells were lysed in ice-cold RIPA Lysis and Extraction buffer (Thermo Fisher Scientific, Shanghai, China). Western blotting was performed according to a standard method 21 . All the experiments were repeated thrice. All the results are from separate blots to avoid possible problems related to incomplete stripping.
All of the primary antibodies (Cdc42 and GAPDH) and the secondary antibodies (antirabbit IgG conjugated with horseradish peroxidase (HRP)) are commercially available from Abcam (Cambridge, MA, USA).
Statistical Analysis
Statistical analyses were performed using the SPSS 22.0 software (SPSS Inc, Chicago, IL, USA). Student’s paired t-test (for two-group data) or post-analysis of variance (ANOVA) followed by Student–Newman–Keuls (for more than two-group data) was used for the analysis of differences. P < 0.05 was considered as statistically significant. All data were expressed as mean ± standard error (SE).
Results
ZFAS1 Was Significantly Upregulated in NSCLC Tissues
We first determined the expression level of lncRNA ZFAS1 in 18 paired human NSCLC and adjacent normal tissues by qRT-PCR. ZFAS1 expression was found to be significantly upregulated in NSCLC tissues compared with the adjacent normal tissues (Fig. 1A, P < 0.05). The results are consistent with previous studies 19,20 . We also examined the ZFAS1 expression level in A549, HCC827, and 16HBE cells. As shown in Fig. 1B, the expression levels of ZFAS1 in A549 and HCC827 were significantly higher than in 16HBE (P < 0.05).
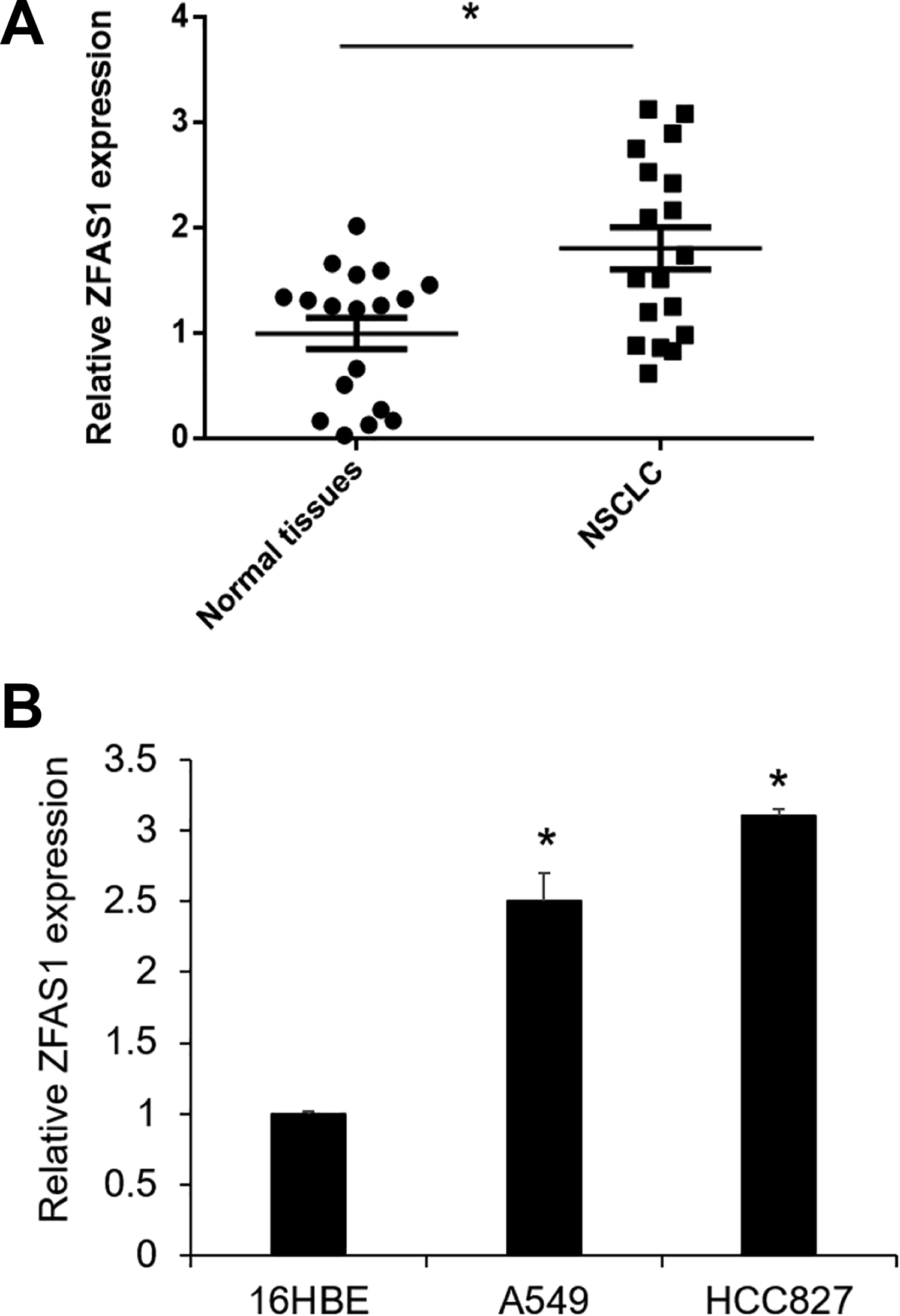
Expression level of ZFAS1 in NSCLC tissues and cell lines. (A) Relative expression levels of ZFAS1 in 18 pairs of human NSCLC and adjacent normal tissues by qRT-PCR. *P < 0.05 vs normal tissues. (B) Relative expression levels of ZFAS1 in 16HBE, A549, and HCC827 cell lines using qRT-PCR. *P < 0.05 vs 16HBE (n = 3). HCC is just the name of cell line. NSCLC: nonsmall cell lung cancer; HBE: human bronchial epithelial cell; qRT-PCR: quantitative real-time PCR: polymerase chain reaction; ZFAS1: zinc finger antisense1.
miR-590-3p Was Significantly Downregulated in NSCLC Tissues
The bioinformatic online tool Starbase v2.0 (http://starbase.sysu.edu.cn/) was used to predict the complementary region at the 3′-UTR of ZFAS1 and miRNAs. The miR-590-3p was found binding sites on ZFAS1(Fig. 2A). We subsequently examined the miR-590-3p expression in NSCLC tissues. We found that miR-590-3p was significantly decreased in NSCLC tissues compared with adjacent normal tissues (Fig. 2B).
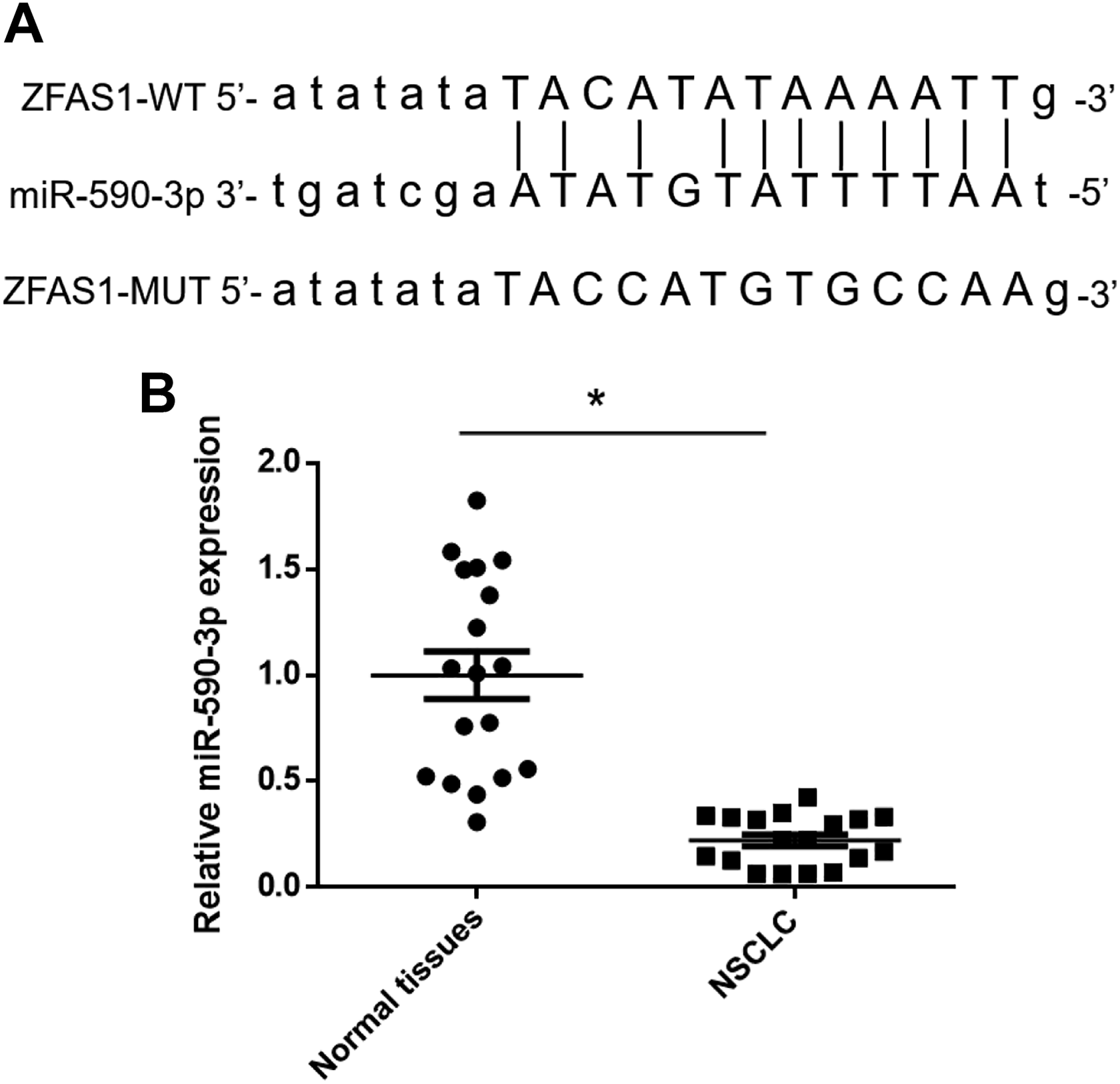
miR-590-3p was significantly downregulated in NSCLC tissues. (A) Prediction for miR-590-3p binding sites on ZFAS1 by bioinformatics analysis. (B) Relative expression levels of miR-590-3p in 18 pairs of human NSCLC and adjacent normal tissues by qRT-PCR. *P < 0.05 vs normal tissues. NSCLC: nonsmall cell lung cancer; MUT: mutant; qRT-PCR: quantitative real-time PCR; WT: wild type; ZFAS1: zinc finger antisense1.
miR-590-3p Was Directly Regulated by ZFAS1
To support these observations, luciferase assay was used to confirm that overexpression of miR-590-3p reduced the luciferase activity of ZFAS1-WT, but not ZFAS1-MUT in HBE16 cells (Fig. 3A). The reduced luciferase activity of ZFAS1-WT was not observed in miR-con as well (Fig. 3A). Results of qRT-PCR assay showed that si-ZFAS1 transfection significantly decreased the expression level of ZFAS1, while the pcDNA-ZFAS1 transfection significantly increased the expression level of ZFAS1 (Fig. 3B and C). Relatively, si-ZFAS1 transfection significantly increased the expression level of miR-590-3p in A549 and HCC827 cells and pcDNA-ZFAS1 transfection decreased the expression level of miR-590-3p in A549 and HCC827 cells (Fig. 3D and E). RNA immunoprecipitation experiments also confirmed that both miR-590-3p and ZFAS1 were present in the Ago2-pulled down pellet (Fig. 3F, G). Collectively, these results suggested that miR-590-3p targeted the 3′-UTR of ZFAS1 and was negatively correlated with ZFAS1 expression.

miR-590-3p was directly regulated by ZFAS1. (A) Luciferase reporter assay was performed to confirm that miR-590-3p bound to specific regions in the ZFAS1 3′-UTR region. *P < 0.05 vs miR-con (n = 3). (B and C) The relative mRNA expression of ZFAS1 was detected by transfecting with empty vector, si-ZFAS1, or pcDNA-ZFAS1 in NSCLC cells. *P < 0.05 vs si-NC or Ctl-OE (n = 3). (D and E) The relative mRNA expression of miR-590-3p was detected by transfecting with empty vector, si-ZFAS1, or pcDNA-ZFAS1 in NSCLC cells. *P < 0.05 vs si-NC or Ctl-OE (n = 3). (F and G) Association of ZFAS1 and miR-590-3p with Ago2 in NSCLC cell lines. *P < 0.05 vs IgG (n = 3). pcDNA-ZFAS1 means that the ZFAS1 sequences were cloned into pcDNA3.1 vector. ZFAS1: zinc finger antisense1; UTR: untranslated region; miR-con: miRNA control; mRNA: messenger RNA; si-ZFAS1: ZFAS1-small interfering RNA; NSCLC: nonsmall cell lung cancer; si-NC: negative control siRNA; Ctl-OE: control (empty pcDNA3.1 vector); IgG: immunoglobulin G.
ZFAS1 Enhanced NSCLC Cell Proliferation by Targeting miR-590-3p In Vitro
In this study, the Cyquant assays showed that overexpressing ZFAS1 enhanced cell proliferation in A549 and HCC827 cells compared to si-NC (Fig. 4A, B). These findings suggested that ZFAS1 influenced tumor growth in NSCLC progression.

ZFAS1 enhanced NSCLC cell proliferation by targeting miR-590-3p in vitro. (A and B) Cyquant assay was used to determine the cell proliferation after ZFAS1 overexpression in NSCLC cell lines. *P < 0.05 vs cell without transduction, **P < 0.05 vs Ctl-OE (n = 3). (C and D) Cyquant assay was used to determine the cell proliferation after ZFAS1 knockdown in NSCLC cell lines. *P < 0.05 vs cell without transduction, **P < 0.05 vs si-ZFAS1 (n = 3). (E and F) Cyquant assay was used to determine the cell proliferation by transfecting si-ZFAS1, si-ZFAS1 + miR-con, or si-ZFAS1+ miR-590-3p inhibitor into NSCLC cell lines. *P < 0.05 vs si-ZFAS1, **P < 0.05 vs si-ZFAS1 + miR-con (n = 3). (G and H) Cyquant assay was used to determine the cell proliferation by transfecting ZFAS1-OE, ZFAS1-OE + miR-con, or ZFAS1-OE + miR-590-3p-OE into NSCLC cell lines. *P < 0.05 vs ZFAS1-OE, **P < 0.05 vs ZFAS1-OE + miR-con (n = 3). (I) The protein levels of Cdc42 as determined by western blotting analysis in NSCLC cell lines transfected with si-NC, si-ZFAS1, or si-ZFAS1 + miR-590-3p inhibitor. (J) The protein levels of Cdc42 as determined by western blotting analysis in NSCLC cell lines transfected with Ctl-OE, ZFAS1-OE, or ZFAS1-OE + miR-590-3p-OE. HCC is just the name of cell line. ZFAS1: zinc finger antisense1; NSCLC: nonsmall cell lung cancer; Ctl-OE: control (empty pcDNA3.1 vector); si-ZFAS1: ZFAS1-small interfering RNA; miR-con: miRNA control; ZFAS1-OE: ZNFX1 Antisense RNA 1-over expression; Cdc42: cell division cycle 42; si-NC: negative control siRNA; IgG: immunoglobulin G; GAPDH: glyceraldehyde 3-phosphate dehydrogenase.
To determine whether the effects of ZFAS1 were mediated by miR-590-3p, ZFAS1 was knocked down in A549 and HCC827 cells (Fig. 4C, D). The results demonstrated that knockdown of ZFAS1 decreased A549 and HCC827 cell proliferation, while cotransfection of miR-590-3p inhibitor and si-ZFAS1 abolished the effects caused by si-ZFAS1 (Fig. 4E, F). Similarly, cotransfection of miR-590-3p-mimics and pcDNA-ZFAS1 eliminated the increase in proliferation of A549 and HCC827 cells caused by pcDNA-ZFAS1 (Fig. 4G, H).
In addition, protein expression of the cell proliferation-associated marker Cdc42 was decreased in cells with si-ZFAS1 overexpression compared with si-NC, while cotransfection of miR-590-3p inhibitors and si-ZFAS1 abolished the effects caused by si-ZFAS1 (Fig. 4I). Similarly, cotransfection of miR-590-3p-mimics and pcDNA-ZFAS1 eliminated the increase in the expression of Cdc42 of A549 and HCC827 cells caused by pcDNA-ZFAS1(Fig. 4J). Together, these findings implied that ZFAS1 promoted the progression of NSCLC cells by regulating miR-590-3p.
Discussion
Primary features of the NSCLC are migration and invasion of neoplasms, which are responsible for the high mortality rate 22 . Although the treatments have been advanced, finding new molecular targets for its treatment has the potential to improve the clinical strategies and outcomes of this disease. Recently, more and more functional studies have indicated that lncRNA expression is aberrant in human cancer 23 . However, lncRNAs that are involved in the progression of NSCLC and their clinical application values still remain incomplete.
ZFAS1 is located on the antisense strand of the Znfx1 (zinc finger nuclear transcription factor, X-Box binding (NFX)-1-type containing) promoter region and is a host to three small nucleolar RNAs 24 . Recent studies have demonstrated that lncRNA ZFAS1 plays important functional roles in the progression of various tumors, such as colorectal cancer and melanoma, in which lncRNA ZFAS1 functions as oncogene. For instance, lncRNA ZFAS1 has been verified to contribute to colorectal cancer progression via the miR-150p-5p/vascular endothelial growth factor A (VEGFA) axis 25 . LncRNA ZFAS1 has also been shown to promote tumorigenesis through regulation of miR-150-5p/member RAS oncogene family, ras-related protein rab-9A (RAB9A) in melanoma 26 . Interestingly, although lncRNA ZFAS1 functions as oncogene in colorectal cancer and melanoma and so on, lncRNA ZFAS1 has been found to be downregulated in breast cancer, which functions as tumor suppressor gene 27 . Despite these findings, the underlying molecular mechanism of lncRNA ZFAS1 in NSCLC still remains largely unknown.
Recently, a lot of reports reveal that lncRNAs interact with miRNAs in various cancers. LncRNAs regulate miRNAs by targeting mRNA binding during tumorigenesis. Among those miRNAs, miR-590-3p has been validated to play important roles in the progression of different types of cancers by the regulation of lncRNAs. miR-590-3p has been detected to promote progression of papillary thyroid carcinoma with lncRNA ZFAS1 18 . miR-590-3p has also been found to contribute to cell proliferation and invasion of gastric cancer with urothelial cancer associated 1 (UCA1) and CAMP responsive element binding protein 1 (CREB1) 28 . However, little is known about the functional significance of lncRNA and miR-590-3p in NSCLC.
In this study, we present a novel mechanism of ZFAS1 in the tumorigenesis of NSCLC. We demonstrate that ZFAS1 is increased in NSCLC tissues and cell lines. ZFAS1 promoted NSCLC cell proliferation in vitro. Furthermore, ZFAS1 has also been verified to directly interact with miR-590-3p to influence NSCLC cell proliferation in vitro. In addition, the protein expression of the cell proliferation–associated marker Cdc42 was increased in cells overexpressing lncRNA ZFAS1, while Cdc42 protein expression was decreased in cells following ZFAS1 knockdown. However, it is worthy to verify this model in vivo and investigate the possible triple axis involved in NSCLC. And the change of impeded effect may be considered when miR-590-3p inhibitor and si-ZFAS1 are co-transfected.
In summary, our data provides new insights into the underlying mechanism of NSCLC progression and implied that ZFAS1 may serve as a potential promising therapeutic target for NSCLC.
Footnotes
Acknowledgments
The authors would like to thank the Pathology Division of the Affiliated Hospital of Inner Mongolia Medical University for the pathologic diagnosis.
Authors’ Contributions
YS and BA conceived the study. YS and YZ designed the experiments and wrote the manuscript. YZ and LS collected the samples and completed the experimental part of the study. YZ, ZL, LS, and BA analyzed the data and revised the manuscript.
Ethical Approval
Ethical approval was obtained for all experimental procedures by the Ethics Committee of the Affiliated Hospital of Inner Mongolia Medical University, Inner Mongolia Autonomous Region, China.
Statement of Human and Animal Rights
All procedures with human subjects in this study were conducted in accordance with the Human Ethics Committee of the Affiliated Hospital of Inner Mongolia Medical University. This article does not contain any studies with animals.
Statement of Informed Consent
Verbal informed consent was obtained from the patients for their anonymized information to be published in this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by the Natural Science Foundation of Inner Mongolia (no.2019BS08002) and the Million Technology Projects of Inner Mongolia Medical University (no. YKD2018KJBW020).
