Abstract
Various circular RNAs (circRNAs) are novel class of non-coding RNAs, which are pervasively transcribed in the genome. CircRNAs play important roles in human, animals and plants. Up to now, there was no report regarding circRNAs of cleft palate by 2,3,7,8-tetrachlorodibenzo-pdioxin (TCDD) induce. The present study screened identification and characterization of differential expressed-circRNAs in TCDD-induced cleft palate. 6903 circRNAs candidates came from cleft palates. Among them, 3525 circRNAs are up-regulation, and 3378 circRNAs are down-regulation by TCDD induce. The cluster and GO analysis found that circRNAs involved in biological process, cellular component, and molecular function. Through the analysis of KEGG Pathway, circRNAs made functions via classical signaling pathway in cleft palate, such as TGF-beta signaling pathway, BMP signal pathway, MAPK signaling pathway. In addition, we found down-regulated circRNA224, circRNA3302 and up-regulated circRNA5021 targeted tgfbr3, but up-regulated circRNA4451 targeted tgfbr2. circRNA4451 may make functions through TGF-beta signaling pathway. These results suggested that many different circRNAs may make important role in TCDD-induced cleft palate, which provided a theoretical basis for further research.
Introduction
Cleft palate is the most common congenital malformation in the oral and craniofacial region. The incidence of cleft palate is about one in 700 live births. 1 During the palatal development, environmental factors can affect palatal growth and development. As a persistent environmental pollutant, 2,3,7,8-tetrachlorodibenzo-pdioxin (TCDD) is widely distributed in every corner of people’s production and live, such as combustion of garbage, automobile exhaust, discharge of waste gas and water. TCDD is also a strong teratogen for cleft palate. 2 The incidence rate of cleft palate is 100% in pregnant mice (64 μg/kg body weight) by TCDD treated only once. 3 TCDD can alter the expression of non-protein-coding genes. For example, the disregulated expression of long non-coding RNA H19 mediated cleft palate by TCDD-treated. 4
Circular RNAs (circRNAs) are new class of non-coding RNAs and exist ubiquitously in eukaryotic cells. Because circRNAs have closed loop structures without 5′–3′ polarities, they can escape from deadenylation, decapping and degradation. 5 CircRNAs have great abundance, high stability, extensive functions, and disease-developmental-stage-specific expression patterns.6,7 The classifications of circRNAs are also not final verdict, but people can’t consistently conclude the function of circRNAs. Some studies have revealed that circRNAs are classified into two types, such as circular exonic RNAs (ecircRNAs) and circular intronic RNAs (ciRNAs). 8 Others are classified into three types, such as ecircRNAs, ciRNAs and exon-intron circRNAs (eiciRNAs). 9 But now the circRNAs were still unknown their functions in various species. CircRNAs were found to serve a critical role in embryonic development, and the dysfunctions of circRNAs were associated with different developmental disorders.10,11 For example, circRNAs involved in the development of many diseases, such as embryonic heart, 12 neurological disorders13,14 and male infertility.15,16 However, the roles of circRNAs in cleft palates by TCDD induce were still unknown. Therefore, identification of circRNAs can help us to explore the molecular mechanisms of cleft palate by TCDD induced.
Materials and methods
Construction of cleft palate model
18 pregnant C57BL/6 mice were supplied by Jiangxi Key Laboratory of Systems Biomedicine in Jiujiang University. All experiments were carried out in accordance with the Experimental Animal Center Guide for the care and use of laboratory animals and the Institutional Ethical Guidelines for experiments with animals. The mice were housed at constant temperature varying from to 20–24°C, and 50 ± 10% relative humidity with a light-dark 12 h/12 h cycle. Water and food were provided ad libitum. Mice were divided randomly into two groups, and nine mice were in each group. TCDD (Sigma, Saint Louis, MO, USA) was dissolved in dimethyl sulfoxide (DMSO, Sigma, Saint Louis, MO, USA). We prepared it for 100 μg/ml suspension with corn oil (Jinglongyu, Qinghuangdao, China), and administered orally to each pregnant mouse (64 μg/kg body weight) in the TCDD-treated group on embryo day 10 (E10). The same volume of DMSO and corn oil were administered to the control mice (control group).
Sample collection and RNA extraction
On time points 16.5 (E16.5), the samplings of TCDD-treated groups (MP1-T, MP2-T, and MP3-T) and control group (MP1-C, MP2-C, and MP3-C) were quickly collected by palate dissection. There were three biological replicates in per group, and each made up of five different individual palatal tissues. Subsequently, the palates of fetuses were harvested from both groups under a stereomicroscope (Olympus SZ-X12, Olympus, Tokyo, Japan) and immediately preserved in RNAlater solution (Ambion, Austin, TX, USA) at 4°C, and then stored at −80°C for further RNA isolation. Total RNA was extracted by a standard protocol using Trizol reagent (Invitrogen, Carlsbad, CA, USA). For quantitation of circRNAs, RNase R (Epicentre, Madison, WI, USA) was added to degrade linear RNAs. RNA quality and concentration were measured and analyzed by Bioanalyzer 2100 and RNA 6000 Nano LabChip Kit (Agilent, Carlsbad, CA, USA) with RIN number >7.0. Then, we performed the paired-end sequencing on an Illumina Hiseq 4000 at the LC-BIO (LC-Bio Technology Co., Ltd, Hangzhou, China) following the vendor’s recommended protocol.
Sequence and primary analysis
Using the Illumina paired-end RNA-seq approach, we sequenced the transcriptome of palates and generated a total of million paired-end reads of from under 100 nt to 4000 nt length. This yielded gig abases (15Gb) of sequence, representing approximately seven times the size of the genome (13Gb) Prior to assembly, the low-quality reads (1, reads containing sequencing adaptors; 2, reads containing sequencing primer; 3, nucleotide with q quality score lower than 20) were removed. After a total of 13Gbp of cleaned, paired-end reads were produced. The raw sequence data were submitted to the NCBI Short Read Archive with accession number. We aligned reads of control and TCDD-treated group to the reference genome from NCBI, Ensemble or NCBI database using Tophat package, which initially removed a portion of the reads basing on quality information accompanying each read, and then mapped the reads to the reference genome from NCBI. Tophat allowed multiple alignments per read (up to 20 by default) and a maximum of two mismatches when mapped the reads to the reference. Tophat built a database of potential splice junctions and confirmed these by comparing the previously unmapped reads against the database of putative junctions. The aligned read files were processed by scripts in house, which used the normalized RNA-seq fragments and counted to measure the relative abundances of the transcripts. The unit of measurement was Fragment per kilobase of exon per million fragments mapped (FPKM). The reference gene transfer format (GTF) annotation file was downloaded from NCBI, Ensemble or NCBI database. CIRCExplorer was used to de novo assemble the mapped reads to circRNAs firstly. Then, these assembled circRNAs from all samples generated unique circRNAs. The expression of circRNAs in different samples or groups were calculated by scripts. Only the comparisons with “p value” less than 0.05 were regarded as showing deferential expression RNA-seq reads mapping. And circRNA sequences were predicted using the protocol by Memczak et al. 17
Analysis and annotation of circRNAs
The expression level of each transcript was measured as the number of clean reads mapped to its sequence. The mapped clean read number was normalized to RPKM. We used DESeq to analyze the expression of circRNAs. p-value >0.05 and | log2 foldchange | ≥2 were considered to indicate significant expression abundance. KEGG was the major public pathway-related database for helping to further understand the biological functions of genes. KEGG pathway enrichment analysis identified significantly signal transduction pathways using the corrected p-value <0.05, which was as a threshold to find significantly enriched KEGG terms in significant differential expressed genes comparing them to the whole genome background. The calculation formula of p-value was as follows:
N represented the total number of KEGG Pathway annotated genes in cleft palate, and n represented the number of differentially expressed genes in N, M represented the number of KEGG Pathway annotated genes, and m represented the number of KEGG Pathway annotated genes in M. After correction for multiple testing, we chose pathways with a P-value <0.05 to represent those significantly enriched in differential expressed genes.
Integrated analysis of circRNAs
According research reports, circRNAs can influence the post transcriptional regulation function of mRNA, and mRNAs were obtained from small RNASeq in TCDD-treated groups (MP1-T, MP2-T, and MP3-T) and control group (MP1-C, MP2-C, and MP3-C) on E16.5. We can analyze function of them deeply. In order to define all the possible circRNA-mRNA interactions, two computational target prediction algorithms were used to identify their binding sites. Targetscan 7.0 (http://www.targetscan.org/) was constructed the regulatory network of circRNA-mRNA.
Impact TGFB/smad pathway molecules in TCDD induced-MEPM cells
Mouse embryonic palatal mesenchymal (MEPM) cells were derived from palatal tissue at 13th day of C57BL/6 mice embryos. The method of MEPM cell culture was according to Wang’ report. 18 The MEPM cells were cultured in flasks with DMEM (Dulbecco’s Modified Eagle’s medium)/F12 medium (Hyclon, Logan, Utah, USA) supplemented with 10% fetal calf serum (Sijiqing, Hangzhou, China). The MEPM cells were placed in a humidified incubator at 37°C with 5% CO2 atmosphere with media replaced every other day. To investigate the biological function of circRNA4451, 10 nM TCDD exposed to MEPM cells, and MEPM cells were transfected with 40 nM each of siRNA targeting circRNA4451 or negative control (siRNA NC) (GenePharma, Shanghai, China) using Lipofectamine 2000 transfection reagent (Invitrogen, Carlsbad, CA, USA), according to the manufacturer’s instructions. After 48 h, total lysates from different MEPM cells treated 5X SDS-lysis buffer supplemented with proteases inhibitors (M250, Amresco, Ohio, USA) were determined. Protein concentration was determined using a standard BSA protein assay (Dingguo, Beijing, China). A total of 40 mg proteins were fractionated on 12% SDS-PAGE, and transferred to nitrocellulose membranes. After blocking with 5% nonfat milk, the membranes were immunoblotted with the primary antibodies: p-Smad2 (sc101801, Santa Cruz, California, USA), Tgfr2 (BA0526, Boster Biotech, Wuhan, China), p-Smad3 (GD-CZ5616 R, Santa Cruz, California, USA). Actin (WD03056, Weiao Biotech, Shanghai, China) was probed as a loading control. Then membranes were washed and incubated with HRP-conjugated secondary antibody (sc-2004 or sc-2005, Santa Cruz, California, USA), Western blot analysis was performed using the Odyssey Infrared Imaging System (Li-Cor, Lincoln, NE, USA).
Statistical analysis
The data of tests was made automatically using SPSS software version 18.0 (SPSS lnc., Chicago, IL, USA). Double-sided Student’st test was used for comparison of two groups. Multiple group comparisons were assessed using one-way analysis of variance, and Turkey’s post hoc test was used to determine which groups differ from each other. All data were presented as mean ± standard deviation (SD). The differences were considered statistically significant when *p < 0.05, or **p < 0.01.
Result
Model identification of cleft palate
Cleft palatal model was generated by pregnant C57BL/6 mice. 18 mice were randomly divided into TCDD-treated and control group averagely. On the E16.5, the fetuses were removed from the uterus. We found that the rate was 100% of cleft palate in TCDD-treated group. Histology analysis of palatal shelves contacted each other in control group (Figure 1(a)). But the palatal shelves failed to contact each other and led to cleft palate in TCDD-treated group (Figure 1(b)). Morphological and histological analysis of palatal shelves. On E16.5, histology analysis of palatal shelves in the control and TCDD-treated group. The palate shelves of fetuses contacted each other in control group (a). The palate shelves of fetuses failed to contact each other and resulted in cleft palate in TCDD-treated group (b). TCDD, 2,3,7,8-tetrachlorodibenzo-p-dioxin. HE staining X 40 in both group.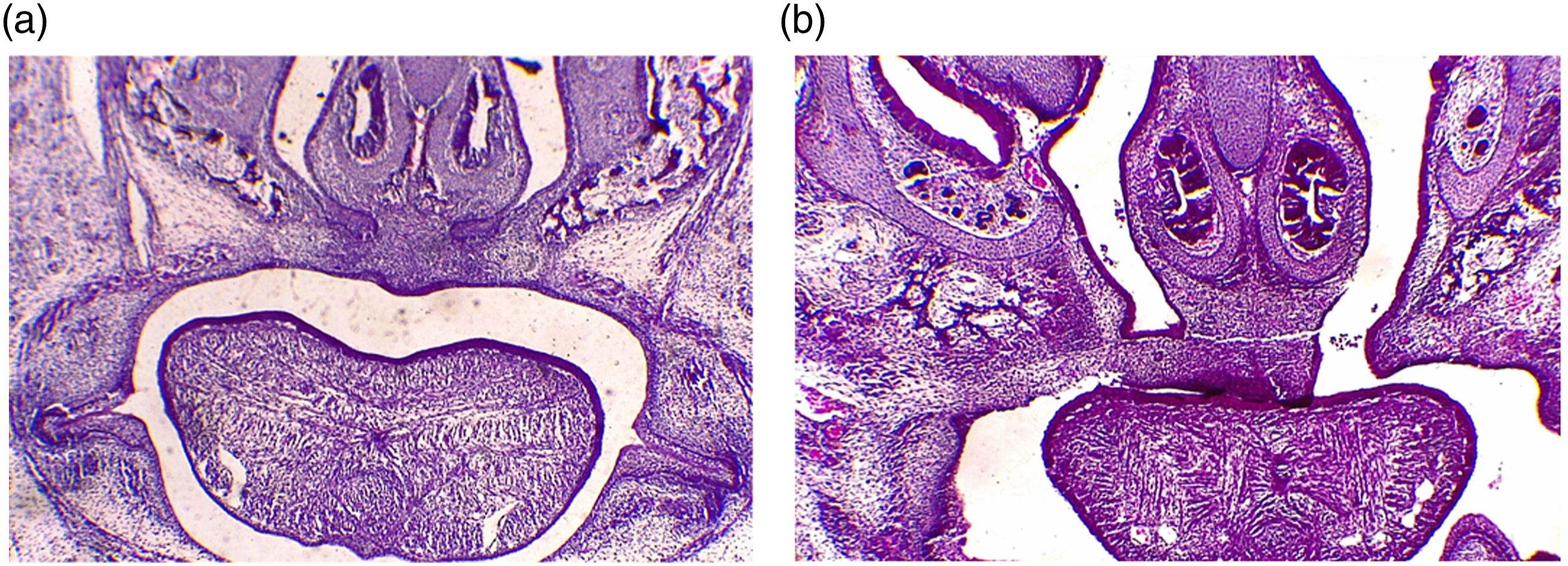
Analysis transcriptome sequencing of cleft palate by TCDD induced
To obtain a comprehensive understanding transcriptome by TCDD-induced palate, the palatal tissues of TCDD-treated group and control group were subjected to NGS on E16.5. CircRNA libraries were constructed from the total RNA with rRNA depletion plus RNase R, and sequenced by the paired-end method (300 bp ± 500 bp) using the Illumina Hiseq 4000 platform. Those residual rRNAs, about 85.23% control group and 85.39% TCDD-treated group of high-quality reads were successfully aligned against the mice genomes, which were showed in supplementary table S1 (Table S1). After discarding the low-quality raw reads, 590,730,962 clean reads remained, the reads statistics of sequencing data were compared with reference genome (Table S2).
To identify unannotated RNA transcripts, we aligned reads of TCDD-treated and control group again to NCBI, Ensemble or NCBI database using Tophat. 19 The results based on the sequence reads, totally 24,076 transcripts candidates were identified by the circRNA identifier (CIRI) software in cleft palates. In control group, 48.23% transcripts were completely matched to exons of genome, and 50.1% transcripts were located entirely within reference intronic regions, even 1.67% transcripts were located entirely within reference intergenic regions. In TCDD-treated group, 50.52% transcripts were completely matched to exons of genome, and 47.85% transcripts were located entirely within a reference intronic regions, and 1.63% transcripts were located entirely within a reference intergenic region, all of which showed in supplementary figure 1 (Figure S1). The results indicated that most transcripts were between the exonic and intronic regions, and total circRNA-sequence (circRNA-Seq) strategies could cover the RNAs.
Classification and functional annotation of circRNAs
Potential circRNA candidates were identified through their sequences. The normalized-RNA-seq fragements were counted to measure the relative abundances of transcripts. The unit of measurement was RPKM. The GTF annotation file was downloaded from NCBI, Ensemble or NCBI database. CIRCExplorer was used to de novo assembly for the mapped reads at first. These assembled circRNAs from all samples were generating unique circRNAs. 97.84% exonic circRNAs (eciRNAs) and 2.16% intronic circRNAs (ciRNAs) were on control group. 97.15% eciRNAs and 2.85% ciRNAs were on TCDD-treated group (Figure S2). The size of cleft palate-related circRNA candidates ranged from under 100 nt to 4000 nt. About the length of 98.74% circRNAs were less than 2000 nt, among which 27.90% length were less than 500 nt. and the length from 500 nt to 1000 nt were 70.84% (Figure S3). Venn diagram showed the number of overlaps circRNAs in control and TCDD-treated group (Figure S4). From the result, it suggested that the samples of repeatability were perfect and stable.
The candidate circRNAs were analyzed by fusions TopHat-Fusion packages (http://uvacse.virginia.edu/resources/rivanna/rivanna). All circRNAs were referenced against chromosome, genes, transcripts, average T/G, KEGG and GO databases, with the number of chromosome (21), genes (48,672), transcripts (118,888), average T/G (2.40), KEGG (5475) and GO (20,927), respectively (Table S3). According to GO classification system, circRNAs were classified into three major functional categories (biological process, cellular component, molecular function, and Figure 2). 5058 circRNAs were notably represented in the biological process category. 879 circRNAs involved in the cellular components. 1413 circRNAs involved in the category of molecular function. CircRNAs involved in “biological process”, “transcription, DNA-dependent” and “positive regulation of transcription, DNA-templated” groups were notably represented in the biological process category. “Cytoplasm” and “nucleus” were the most represented in the cellular components. A significant proportion of clusters were assigned to “protein binding” in the category of molecular function. List of major functional categories were in cleft palate (Table S4). GO enrichment analysis of differentially expressed circRNAs. The circRNAs networks were mainly associated with biological process, cellular component and molecular function by GO analysis in cleft palates.
The Kyoto Encyclopedia of Genes and Genomes (KEGG) found some important different pathways in 3962 circRNAs (Figure 3). The important pathways were involved in TGF-beta signaling pathway, MAPK signal pathway, Insulin signaling pathway, B cell receptor signaling pathway and Insulin signaling pathway in cleft palate. In addition, we found that 2746 circRNAs were significantly up-regulated, while 2308 circRNAs were significantly down-regulated in response to TCDD (Figure S5). We then made a volcano plot and a hierarchical clustering of the circRNAs basing on the six samples’ log10 (RPKM+1), with the results indicated on TCDD treatments (Figure 4). KEGG pathway analysis of differentially expressed circRNAs. KEGG pathway analysis can deduce the systematic biological behavior, such as MAPK and TGF-beta signaling pathway in palatal shelves by TCDD induced. TCDD, 2,3,7,8-tetrachlorodibenzo-p-dioxin. The vocalno plot and Hierarchical clustering of differentially expressed circRNAs. The vocalno plot of differentially expressed circRNAs (a). Heatmap of circRNAs libraries was for the differentially expressed circRNAs between control group and TCDD-treated group (b). TCDD, 2,3,7,8-tetrachlorodibenzo-p-dioxin.

Coexpression network of circRNAs
Many circRNAs particularly were responsed to cleft palate by TCDD induced, but the function of circRNAs remained largely unknown. In order to reveal the co-expression pattern of circRNA-miRNA and circRNA-miRNA-mRNA, the circRNA-miRNA and circRNA-miRNA-mRNA co-expression networks were constructed by the high-throughput RNA sequencing results, and mainly analyzed by Targetscan 7.0 (http://www.targetscan.org/) and miRanda (http://www.microrna.org/microrna/home.do). A constructed network map contained circRNAs, miRNAs and mRNAs, such as endogenous RNA (ceRNA). These circRNAs were chosen from the higher or lower expression of circRNA-Seq data (Table S5 or S6). The expression of circRNA5021 and circRNA4451 were up-regulation, but circRNA224, circRNA3302 were low-regulation in cleft palate by TCDD induced (Figure 5(a) and Figure 5(b)). The relationship between circRNAs and important pathways. The down-regulated circRNA224, circRNA3302 and up-regulated circRNA5021 targeted tgfbr3 (a), but up-regulated circRNA4451 targeted tgfbr2 (b). Red indicates circRNAs were high expression, and green indicates circRNAs were low expression in TCDD-treated group. TCDD, 2,3,7,8-tetrachlorodibenzo-p-dioxin.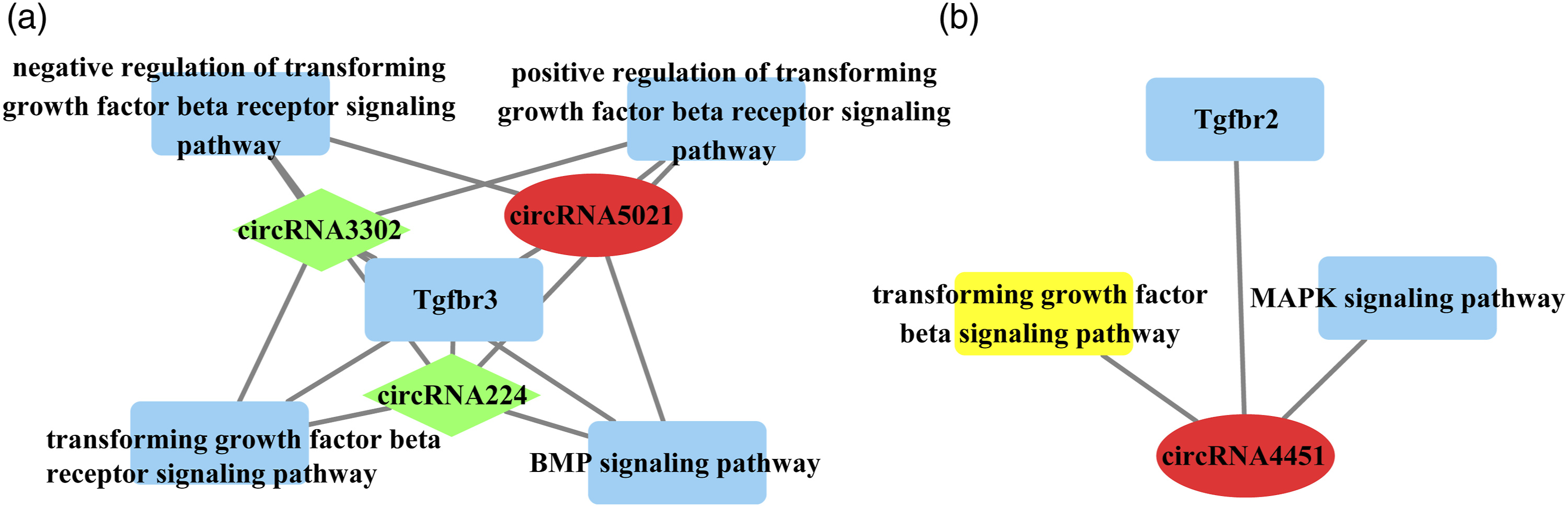
Predicted function of circRNAs
The gene expression was regulated by non-coding RNAs (ncRNAs) and revealed the full transcriptome level through the regulatory network of the ceRNAs.
18
Many circRNAs mediated their target protein-coding genes. CircRNA4451 may be regarded as centers and targeted tgfbr2 through TGF-beta signaling pathway. We examined whether knockdown of circRNA4451 would affect the TGF-beta signaling pathway. After Inhibition of circRNA4451 expression by siRNAs, the expression of tgfbr2, p-Smad2 and p-Smad3 were increased (Figure 6). As above statement, the regulatory roles of circRNAs in cleft palatal pathogenesis might be very complicated, which need study deeply in the future. The effects of TGF-b/Smad signaling molecules by siRNA circRNA4451 and TCDD induced MEPM cells. After siRNA circRNA4451, the expression of TGFBR2, phosphorylation of Smad2 and Smad3 proteins were measured by Western blot in MEPM cells of TCDD-treated and control group, respectively. B-actin (Action) was as loading controls, and band fluorescence intensity was analyzed and quantified using Odyssey Infrared Imaging System. The relative expression level of TGFBR2 and phosphorylation of Smad2/3 proteins were calculated after normalization with the loading control b-actin. The data are the mean values ± standard deviation of three replicate experiments. *p < 0.05, **p < 0.01 vs. the corresponding control values. TCDD, 2,3,7,8-tetrachlorodibenzo-p-dioxin; TGFBR2, transforming growth factor beta-receptor type II.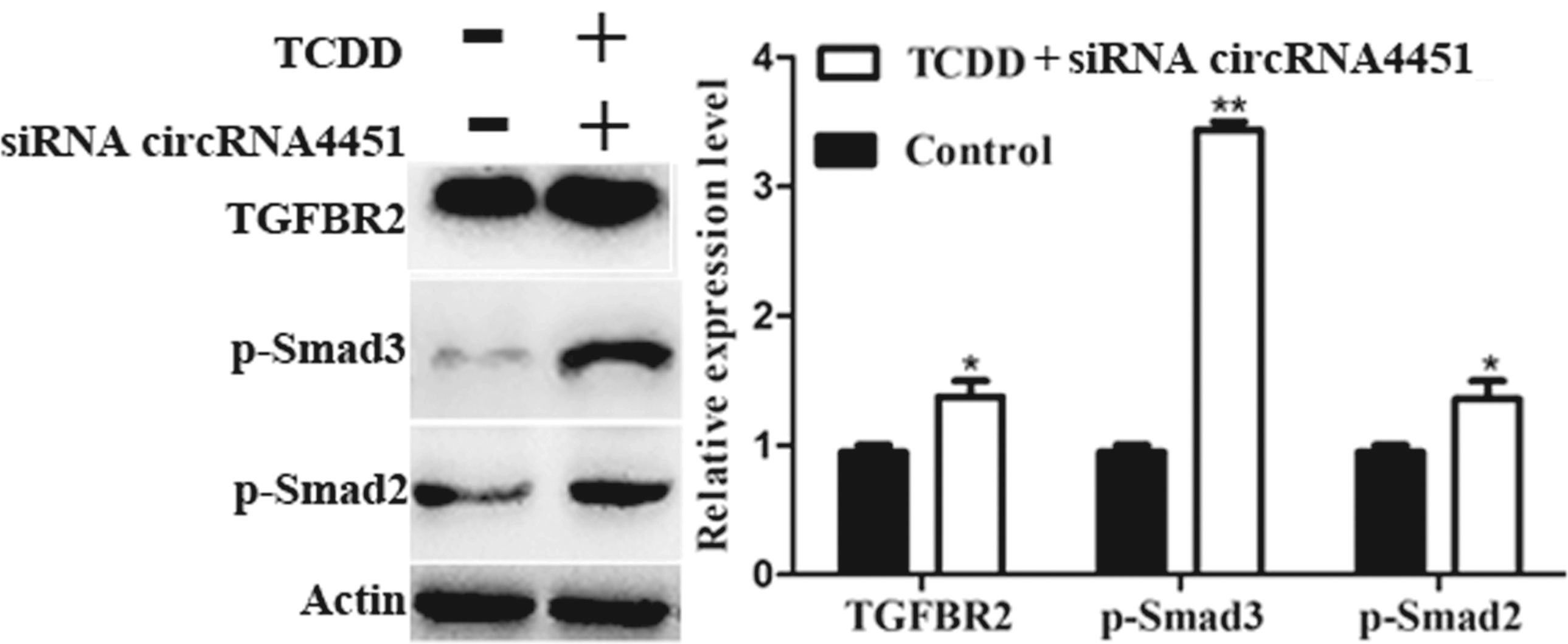
Discussion
CircRNAs were thought to be ‘junk’ just as other non-coding RNA in the genome. 20 The circRNAs were first identified over 30 years ago. 21 However, these circRNAs were found that they played important roles in the regulation of multiple diseases at recent years, such as different cancers, 22 nervous system diseases,23,24 coronary artery disease, 25 diabete diseases,26,27 cardiovascular disease. 28 However, the functions of circRNAs are still not clear. The biogenesis, specific expression, turnover, localization, and degradation of circRNAs still need us to deeply investigate.
Up to now, circRNAs are potentially involved in multiple biological and pathological processes, but there was no report about circRNAs in cleft palates by TCDD induced. In this study, we conducted a genome-wide identification of circRNAs in cleft palate by TCDD induced. Our results demonstrated that there were many circRNAs in cleft palates, which provided the new resources of circRNAs for further research to cleft palates, which also firstly revealed common and distinct properties of circRNAs in cleft palates.
We found that circRNAs were located in exon, intron and intergenic in genome, which were consistent with previous report. 29 We also found that there were two-classification circRNAs (ecircRNAs and ciRNAs). EcircRNAs were 97.84% and 97.15% circRNAs in control group and TCDD-treated group, respectively. And ciRNAs were seldom (12.16% and 12.85%) in two groups, respectively. We firstly found the result in cleft palates. The result suggested that the circRNAs had important functions. The lengths of circRNAs were from 0 bp to 4000 bp. In performing KEGG pathway analyses, “protein binding” and “cytoplasm” were found to be the two most frequently represented subclasses. In addition, we used the pathway analysis to illuminate the functional clusters of circRNAs.
Many reports revealed that TGF-β/Smad signaling was very important in palate development.30,31 We also found that TGF-β/Smad signaling occurred in cleft palates by TCDD induced. In the meantime, we found circRNA224, circRNA3302 and circRNA5021 targeted tgfbr3, but circRNA4451 targeted tgfbr2 and made functions through TGF-beta signaling pathway. Obviously, the precise mechanism needed to further study.
In the present study, we found that 2746 circRNAs were significantly up-regulation, and 2308 circRNAs were significantly down-regulation comparing with corresponding control group. Up to now, the data of circRNAs in cleft palates by TCDD induce are not reported. Recently, some researchers discovered that some circRNAs can act as microRNA (miRNA) sponges, and other circRNAs could also directly target mRNAs by partial base pairing.32,33 In our study, we also found that circRNAs directly regulated the expression of mRNA through signal pathway, which was associated with previous findings that circRNAs were directly involved in pathways.34,35
Conclusion
In summary, our study was firstly identifying 3378 up-regulated and 3525 down-regulated circRNAs in cleft palate by TCDD induced. GO analysis found that these circRNAs involved in diverse biological functions. The expression levels of circRNA4451 and circRNA5021 were significantly up-regulated in cleft palate and they might be associated with very important functions. These circRNAs were also very closely relative with important pathways (such as TGF-beta signaling pathway, BMP signal pathway and so on). The several key pathways might be providing some information to help us articulate the different mechanisms in cleft palate by TCDD induced. These circRNAs may be potentially involve in the development of cleft palates and served as potential diagnosis or/and prognostic biomarkers for cleft palates. However, further studies need to validate. Moreover, the functions and mechanisms of the circRNAs need to require further investigation.
Supplemental Material
Supplemental Material - Identification and characterization of differentially expressed circRNA in 2,3,7,8-tetrachlorodibenzo-p-dioxin-induced cleft palate
Supplemental Material for Identification and characterization of differentially expressed circRNA in 2,3,7,8-tetrachlorodibenzo-p-dioxin-induced cleft palate by Liyun Gao1, Jingwen Tan, Chunhua Han, Junfei Fan, Jiayin He, Ting Luo, Shiqun Yu, Xiangxin Che, Lin Zhang and Xin Wang in Human & Experimental Toxicology
Footnotes
Author contributions
Liyun Gao contributed to the study conception and design. Materials were prepared by Jingwen Tan. Chunhua Han did experiments. Data collection and analysis were performed by Junfei Fan and Jiayin He. The first draft of the manuscript was written by Shiqun Yu, Ting Luo, Xiangxin Che and Lin Zhang. Xin Wang modified the manuscript. All authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the grants of the National Natural Science Foundation of China (81,502,843, 81,360,447), Jiangxi Natural Science Foundation Project (20202BAB206067), Jiujiang base and talent plan - high level scientific and technological innovation talent project (S2020QNZZ011 and 202311158), Future project of Jiujiang administration of traditional Chinese medicine (2021b697), Science and Technology Program of Jiangxi Provincial Health Commission (202,311,158), Special Research Project of Jiangxi Cognitive Science and Interdisciplinary Research Center (RZYB202206), Science and Technology Plan Project of Jiangxi Provincial Health Commission (SKJP220225877).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
