Abstract
2,3,7,8-Tetrachlorodibenzo-p-dioxin (TCDD) is a ubiquitous environmental pollutant and also a strong teratogen for cleft palate (CP). But up to now, the underlying molecular mechanisms of TCDD-induced CP are largely unknown. More recently, accumulating evidences are revealing important roles of long noncoding RNAs (lncRNAs) in all kinds of diseases including CP. However, the role and molecular mechanism of lncRNAs in TCDD-induced CP are still largely unexplored. Thus, identification of differentially expressed lncRNA (DEL) might help figuring out the mechanism of CP induced by TCDD. In this study, a CP offspring model of C57BL/6 female mice was generated by TCDD (64 µg/kg body weight) induce on embryo day 10 (E10). The incidence rate of CP was 100% in the TCDD group (105) after cervical dislocation on E16. Then, the high-throughput RNA sequencing (RNA-seq) was established to search a comprehensive profile of the lncRNAs. In addition, a coexpression network of lncRNA and messenger RNA (mRNA) was performed to discern potential mechanism. The result showed that 26,246 novel lncRNAs and 9635 known lncRNAs were screened out, and 413 lncRNA transcripts and 65 mRNA transcripts were identified as being significantly different between the CP group and control group. Notably, we found that there are seven lncRNAs that can target Smad1 and Smad5, which are key molecules of bone morphogenetic protein (BMP) signaling pathway, which suggested that they may be concerned with BMP signaling in TCDD-induced CP. In addition, some lncRNAs targeted the important molecules of Hippo and Wnt signaling pathways. These results suggested that characteristic lncRNA alterations may play a critical role in TCDD-induced CP, which provided a theoretical basis for further research.
Introduction
Cleft palate (CP) and clefts of the lip are the most common craniofacial birth defects worldwide, with an approximate incidence of approximately 1/700. 1 The etiology of CP involves both genetic and environmental factors. 2 2,3,7,8-Tetrachlorodibenzo-p-dioxin (TCDD) is a widespread, persistent environmental contaminant, and comes from byproducts and wastes of industrial processes. 3 TCDD has been identified as a ubiquitous environmental pollutant with reproductive and developmental toxicity. 4 It is a well-known fact that TCDD can induce CP in pregnant C57BL6 mice. 5 However, the underlying mechanism remains largely unknown.
Long noncoding RNAs (lncRNAs) are a class of noncoding RNA molecules, which are defined as transcripts longer than 200 nucleotides. According to earlier reports, lncRNA played a key role in multiple diseases, such as coronary artery disease, 6 Alzheimer’s disease, 7 various cancers, 8 and deformities. 9 Recently, published works have provided evidence that lncRNAs are involved in development and progression of CP. 10 –12 However, it remains unclear whether lncRNAs have any role in TCDD-induced CP. Thus, identification of DELs between CP samples and normal specimens can help us to explore the pathogenesis of CP induced by TCDD.
In this study, high-throughput sequencing technology was utilized to investigate the DELs in TCDD-induced CP mice model. Then, gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) biological pathway analysis were performed through these DELs and showed that seven lncRNAs were enriched in regulation of bone morphogenetic protein (BMP) signaling pathway.
Materials and methods
Animals
Thirty pregnant C57BL/6 female mice (embryonic day0, E0) were obtained from Henan Laboratory Animal Center and housed in a controlled environment (22 ± 2°C constant temperature and 50 ± 10% relative humidity) with a 12-h light/12-h dark cycle. All experiments were carried out in accordance with the Experimental Animal Center Guide for the care and use of laboratory animals and the Institutional Ethical Guidelines for experiments with animals.
Construction of CP model and sample collection
Thirty pregnant mice were divided into two groups: the control group (n = 15) and TCDD-treated groups (n = 15); TCDD (≥98%, DD-2378-S, Sigma, St Louis, Missouri, USA) was dissolved in dimethyl sulfoxide (D2650, Sigma). Group division of mice was used by random number method. The above suspension (100 µg/ml) was diluted in corn oil (383756, Jinglongyu, Qinghuangdao, China) and administered orally to each pregnant mouse (64 µg/kg body weight) in the TCDD-treated group on embryo day 10 (E10), according to a previous report. 13 The equivalent volume of corn oil was administered to the control mice. On time points E16, all pregnant mice from both the control and TCDD-exposed group were euthanized by cervical dislocation. There were a total of 105 and 121 fetuses recovered from the terminated dams of the TCDD-treated and control groups, respectively. Under a stereomicroscope (SZ61TRC–SET, Olympus, Tokyo, Japan), fetal mice palates were quickly harvested by palate dissection. The isolated specimens were preserved in RNAlater solution (AM7020, Ambion, Austin, Texas, USA) at 4°C and stored at −80°C for further RNA isolation or fixed in 10% formalin for histological examination. All experiments were carried out three repetitions, such as TCDD group (MP-T1, MP-T2, and MP-T3) and control group (MP-C1, MP-C2, and MP-C3). We collected RNA from each of the 15 control and 15 TCDD-dosed animals three times. We always complied with experimental Animal Center Guide for the care and use of laboratory animals and the Institutional Ethical Guidelines for experiments with animals.
RNA library construction and sequencing
Total RNA was extracted using Trizol reagent (Invitrogen, Carlsbad, California, USA) according to the manufacturer’s procedure. The total RNA quantity and purity were analyzed using NanoDrop 2000 (Thermo Fisher Scientific, Waltham, MA, USA). Approximately 10 µg of total RNA representing a specific adipose type was used to deplete ribosomal RNA according to the manuscript of the Epicentre Ribo-Zero Gold Kit (Illumina, San Diego, USA). For purification, divalent cations were used to fragment the poly(A)− or poly(A)+ RNA fractions into small pieces under elevated temperature. Then, the cleaved RNA fragments were reverse transcribed to create the final cDNA library according to the manufacture’s procedure (Illumina), the average insert size for the paired-end libraries was 300 ± 50 bp. Then, the paired-end sequencing was performed using Illumina Hiseq 4000 (LC Bio, China) according to the vendor’s recommended protocol.
Identification of LncRNA
Transcripts with a length shorter than 200 bp and transcripts that overlapped with known mRNAs were discarded. Then, we utilized coding potential calculator 2(CPC2) and coding-noncoding index (CNCI) to predict transcripts with coding potential. 14,15 After the transcripts with CPC score <−1 and CNCI score <0 were deleted, lncRNAs were acquired.
Identification of DELs and mRNAs
StringTie was used to perform the expression level for mRNAs and lncRNAs by calculating the number of fragments per kilobase of transcript sequence per millions of base pairs sequence. 16,17 The differentially expressed mRNAs and lncRNAs with log2(fold change) > 1 or log2(fold change) < −1 were screened out by R package (version 3.4.3, R Development Core Team, Vienna, Austria), Ballgown (version 3.7; www.bioconductor.org/packages/release/bioc/html/ballgown.html). 18
Screening of downstream signaling pathway and target genes of lncRNAs
The cis-/trans-target genes of lncRNAs containing 100,000 upstream and downstream genes were screened out by python script and then analyzed using the BLAST2GO (version 2.5; http://www.blast2go.de). 19
Quantitative real-time polymerase chain reaction
At least three biological repeats for each strain and treatment were analyzed. Total RNAs were isolated using Trizol reagent (Invitrogen) according to the manufacturer’s instructions. To measure the expression of lncRNA MSTRG.56310.1, MSTRG.51656.1, MSTRG.40182.1, MSTRG.38487.1, MSTRG.1991.1, MSTRG.13399.1, and ENSMUST00000193657, first-strand cDNA was generated by SYBR PrimesScript RT-PCR kit (TaKaRa Biotechnology, Kyoto, Japan) using 500 ng total RNA. Quantitative real-time polymerase chain reaction (qRT-PCR) was performed to detect the expression of the above lncRNAs using the Applied Biosystems 7500 real-time PCR system (Foster City, California, USA). Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) served as an internal control. The qRT-PCR conditions were as follows: polymerase activation 30 s at 95°C, 40 cycles of 95°C for 5 s, 60°C for 30 s, and 72°C for 20 s. PCR products were identified by melting curve analysis. The data were calculated using the 2–ΔΔCt method. All primers were designed and synthesized by Invitrogen Company. The primer sets used in lncRNAs were as follows: MSTRG.56310.1-(forward, F): 5′-CTCTCGGTTGTGGTGCATCT-3′ and MSTRG.56310.1-(reverse, R): 5′-GCCGGCTGCCATATAGAACT-3′; MSTRG.51656.1-F: 5′-TAGGCAGGGGGAGTCTTGTT-3′ and MSTRG.51656.1-R: 5′-GTGAGCCATCTCCCTAGCAC-3′; MSTRG.40182.1-F: 5′-AATAGCCCTCACTCCACAGG-3′ and MSTRG.40182.1-R: 5′-CCTCCAGTTACAGTAGGGGC-3′; MSTRG.38487.1-F: 5′-AGATGGCTCAGCGGTTAAGA-3’ and MSTRG.38487.1-R: 5′-CAAACACATCACGTATCTTGAGG-3′; MSTRG.1991.1-F: 5′-ACTCACCAGACTCTCTCCCT-3′ and MSTRG.1991.1-R: 5′-AGACACAGTGCTTGCCTAGT-3′; MSTRG.13399.1-F: 5′-TGCCCACAACCAAACTCTGT-3′ and MSTRG.13399.1-R: 5′-TCCTGGCTGGTATGCACAAG-3′; ENSMUST00000193657-F: 5′-GCACAAAGCCTTGGGTATGG-3′ and ENSMUST00000193657: 5′-ACAGGCTGGCAAGATGATCC-3′; GAPDH-F: 5′-TGACGTGCCGCCTGGAGAAAC-3′ and GAPDH-R: 5′-CCGGCATCGAAGGTGGAAGAG-3.
Statistical analysis
The χ2 test and the Student’s t-test were applied for statistical comparisons using SPSS 18.0 software (SPSS, Inc., Chicago, Illinois, USA). All p values were two-sided, and p values <0.05 were taken as statistically significant.
Results
Establishment and identification of cleft palatal model
First, we compared the incidence rate of CP in pregnant C57BL/6 mice treated with or without TCDD. The fetuses were removed from the uterus on E16. We found that the CP incidence rate of fetal offspring was 100% in the TCDD-treated group (105) but was 0% in the control group (121) in all 30 female mice (Figure 1(a)). Furthermore, morphological observation revealed that the palatal shelves of fetal mice had palatal wrinkles and fused together in the control group but not in the TCDD-treated group (Figure 1(b) and (c)). Histology analysis revealed that the palate shelves contacted each other in the control group, while failed to contact each other and led to CP in the TCDD-treated group (Figure 1(d) and (e)).

Morphology and histology identified cleft palates. On E16.5, the number of fetuses was 121 and 105 in control and TCDD group, respectively. These data were the mean ± SD of two independent experiments. *p < 0.05 (a). The palate shelves of fetuses were no palatal wrinkles and fused together in control group (b), whereas the palatal wrinkles were obvious and failure to fuse in TCDD-treated group (c). Histology analysis of palatal shelves in the control and TCDD-treated group: the palate shelves of fetuses contacted each other in control group (d). The palate shelves of fetuses failed to contact each other and led to cleft palate in TCDD-treated group (e). HE staining ×40 in both group. TCDD: 2,3,7,8-tetrachlorodibenzo-p-dioxin; HE: hematoxylin and eosin.
Transcriptome sequencing analysis of TCDD-induced CP
To obtain a comprehensive understanding about the transcriptome analysis of TCDD-induced CP, total RNAs of palatal tissues were isolated from the TCDD group and the control group. RNA-seq libraries were constructed using total RNA with rRNA depletion and sequenced by the paired-end method (2 × 100 bp) platform (Online Supplemental Figure S1). Approximately 18 GB (Mouse Palatal Control, MP_C and TCDD-treated palate, MP_T) of sequences were obtained. After discarding the low-quality sequences below the Q20 threshold, N base ratio less than 5% of reads, and joint contamination sequence (15 bp overlap), the residual rRNA and high-quality reads that having at least 90% similarity were successfully aligned against mice reference genome (Tables 1 and 2). To identify unannounced RNA transcripts, we aligned reads of TCDD-treated and control groups to their ensemble number using NCBI database. 20 Next, we removed known mRNAs and transcripts that are smaller than 200 bp and then predicted the remaining transcripts of lncRNA using CPC and CNCI prediction software. Subsequently, we defined and filtered the remaining transcripts, which have the potential to encode protein, and then, lncRNA sequences were obtained. These lncRNAs generate 48.23% exonic, 50.1% intronic, and 1.67% intergenic region in the control group and 50.52% exonic, 47.85% intronic, and 1.63% intergenic region in the TCDD-treated group (Figure 2 and Online Supplemental Figure S2).
Summary of sequence data generated for cleft palate transcriptome and quality filtering.

MP_T: TCDD group; MP_C: corn oil-treated group, control group.
Overview of two total RNA-seq datasets.
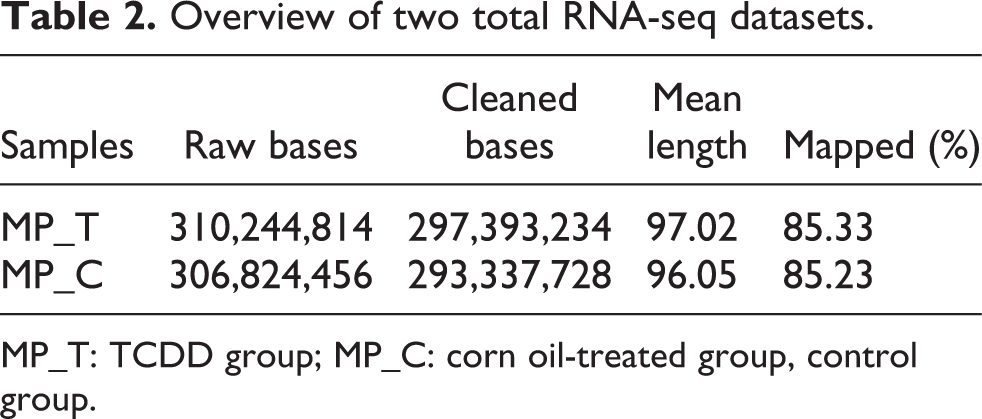
MP_T: TCDD group; MP_C: corn oil-treated group, control group.

The lncRNA from all samples generated exonic, intronic, and intergenic region were on control and TCDD-treated group. TCDD: 2,3,7,8-tetrachlorodibenzo-p-dioxin; lncRNA: long noncoding RNA.
Overview of RNA-seq of lncRNAs and identification of mRNAs
The genome databases of NCBI and ensemble served as the annotation references for lncRNA and mRNA analyses, respectively. LncRNA and mRNA were identified based on their sequences. The normalized RNA sequencing (RNA-seq) fragment was used to measure the relative abundances of the transcripts. Two hundred six down- and 207 upregulated lncRNA transcripts were selected (Figure 3(a) and (b)), and 19 down- and 46 upregulated mRNA transcripts were identified (Figure 3(c) and (d)). Then, hierarchical clustering analysis was performed to further screen DELs and mRNA between the TCDD-treated group and control group (Figure 4(a) and (b)). The top 10 DELs and mRNA were shown in Tables 3 and 4, respectively. Next, the full-length and open-reading frame length distribution of lncRNAs were compared with that of mRNAs. We found that lncRNAs were less conserved than mRNAs in sequences (Figure 5(a) and (b)).

Bar and volcano plots of palatal transcripts. The difference of lncRNAs expression profiles (a and b) and mRNAs expression profiles (c and d) were labelled in the all distribution of the transcripts. The green points indicated downregulated transcripts, and the red points indicated upregulated transcripts in the all plots.
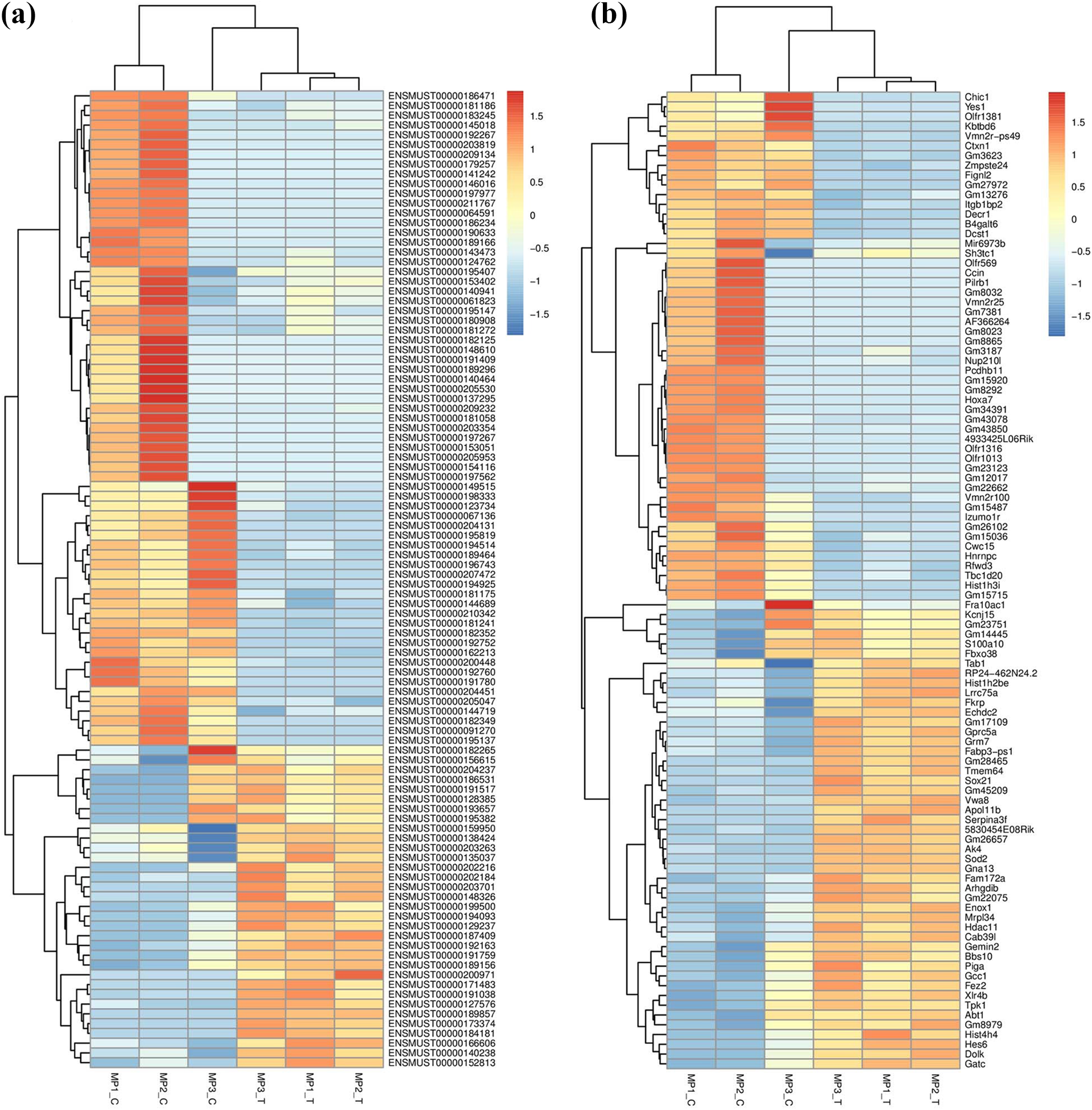
Hierarchical heat maps of the differentially expressed profiles between the TCDD-induced and control groups. Hierarchical heat maps of the differentially expressed profiles between the TCDD-induced and control group (a and b). Each column represents a sample and each row represents a transcript. “Red” indicates higher expression, and “blue” indicates lower expression. MP_T (TCDD-induced group) and MP_C (control group). TCDD: 2,3,7,8-tetrachlorodibenzo-p-dioxin.
The 10 upregulated and downregulated lncRNAs in cleft palatal transcriptome.

MP_T: TCDD group; MP_C: corn oil-treated group, control group; lncRNA: long noncoding RNA.
The 10 upregulated and downregulated mRNAs in cleft palatal transcriptome.

MP_T: TCDD group; MP_C: corn oil-treated group, control group; mRNA: messenger RNA.
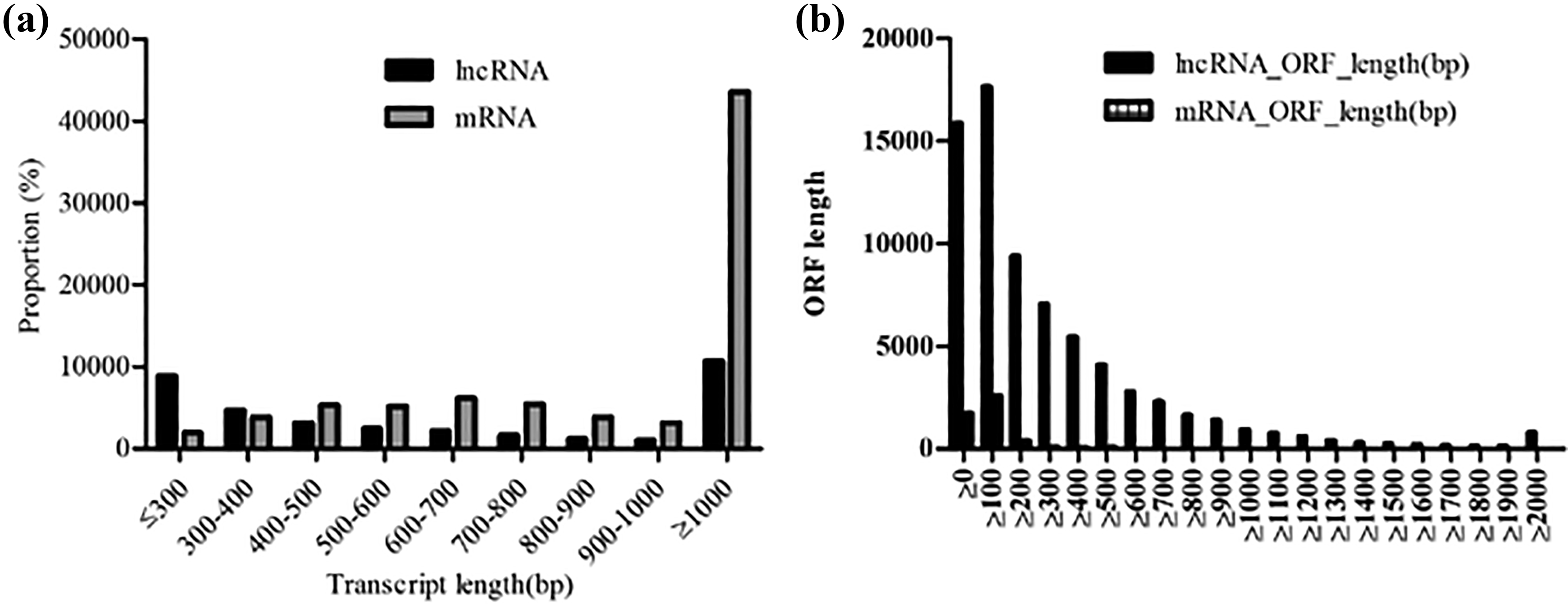
Genomic features of lncRNAs and mRNAs. The different length distribution of lncRNAs and mRNAs (a). The different ORF length distribution of lncRNAs and mRNAs. The ORF length of lncRNAs was always shorter than that of mRNAs (b). mRNA: messenger RNA; lncRNA: long noncoding RNA.
GO and KEGG pathway analysis
To explore the potential biological properties of these lncRNAs in TCDD-induced CP, GO and KEGG pathway enrichment analyses were established. The biological process, cellular component, and molecular function that were associated with these lncRNAs have been shown in Figures 6 and 7. Notably, the results of KEGG pathway analysis revealed that these lncRNAs were involved in regulation of Hippo signaling pathway, Wnt signaling pathway, and BMP signaling pathway (Figure 8). As the BMP signaling pathway plays an important role in the regulation of CP, 21,22 therefore, we speculated that lncRNA might regulate palatoschisis via BMP signaling pathway.

GO analysis. The lncRNAs networks were mainly associated with biological process, cellular component, and molecular function. GO: gene ontology; lncRNA: long noncoding RNA.

KEGG pathway analysis. KEGG analysis identified the top 20 categories of the biological processes that associate with these lncRNAs. KEGG: Kyoto Encyclopedia of Genes and Genome; lncRNA: long noncoding RNA.

KEGG pathway analysis. KEGG pathway analysis can deduce the systematic biological behavior (BMP signaling pathway). KEGG: Kyoto Encyclopedia of Genes and Genome; BMP: bone morphogenetic protein.
DELs are correlated with BMP signaling pathway
We next decided to ascertain the association between DELs and BMP signaling pathway. Through bioinformatics prediction approach, we found that lncRNA MSTRG.56310.1, MSTRG.51656.1, MSTRG.40182.1, MSTRG.38487.1, MSTRG.1991.1, MSTRG.13399.1, and ENSMUST00000193657 could targeted Smad1 and Smad5, which are target genes of BMP signaling. Therefore, we chose them for further analysis. The BMP signaling pathway-related genes, Smad1 and Smad5, were predicted as a target gene of these lncRNAs (Figure 9). Then, the expression levels of these DELs were verified in the TCDD-treated group and control group by qRT-PCR. As shown in Figure 10, the qRT-PCR results were consistent with the RNA-seq data. The expression of ENSMUST00000193657, MSTRG.13399.1, and MSTRG.56310.1 transcripts was increased (2.64 ± 0.42 fold), (2.68 ± 0.23 fold), and (2.8 ± 0.22 fold) in the TCDD-induced group compared with the control group, respectively (Figure 10(a)). The expression of MSTRG.1991.1, MSTRG.38487.1, MSTRG.40182.1, and MSTRG.51656.1 was decreased to (0.36 ± 0.01 fold), (0.34 ± 0.01 fold), (0.29 ± 0.01 fold), and (0.36 ± 0.06 fold) compared with the control group, respectively (Figure 10(b)). In addition, we found that one lncRNA only targeted one coding protein gene (relative molecules of the pathway) in both Hippo and Wnt signaling pathways. For example, lncRNA MSTRG.29999 targeted Zeb2 of Hippo signaling pathway (Table 5). LncRNA MSTRG.40988 targeted Dvl1 of Wnt signaling pathway (Table 6).

Coexpression network of lncRNA and mRNA. mRNA: messenger RNA; lncRNA: long noncoding RNA.

The relative level of lncRNAs was measured by qRT-PCR. The expression of ENSMUST00000193657, MSTRG.13399.1, and MSTRG.56310.1 transcripts was increased in TCDD-induced group compared with the control group, respectively (a). The expression of MSTRG.1991.1, MSTRG.38487.1, MSTRG.40182.1, and MSTRG.51656.1 was decreased compared with the control group, respectively (b). qRT-PCR: quantitative real-time polymerase chain reaction; lncRNA: long noncoding RNA.
LncRNA only targeted coding protein gene in Hippo signaling pathways.
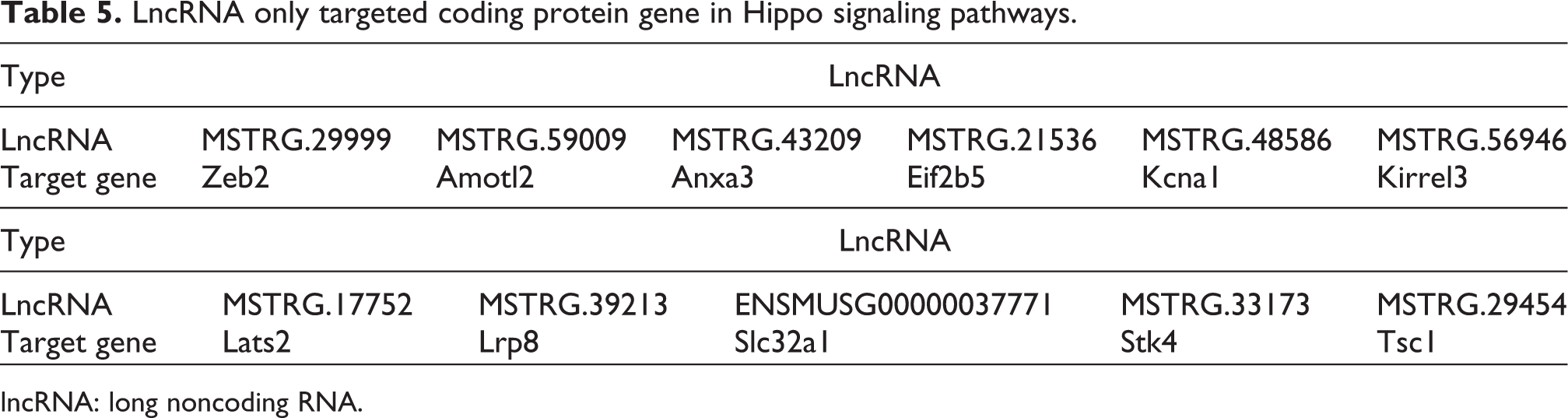
lncRNA: long noncoding RNA.
LncRNA only targeted coding protein gene in Wnt signaling pathways.
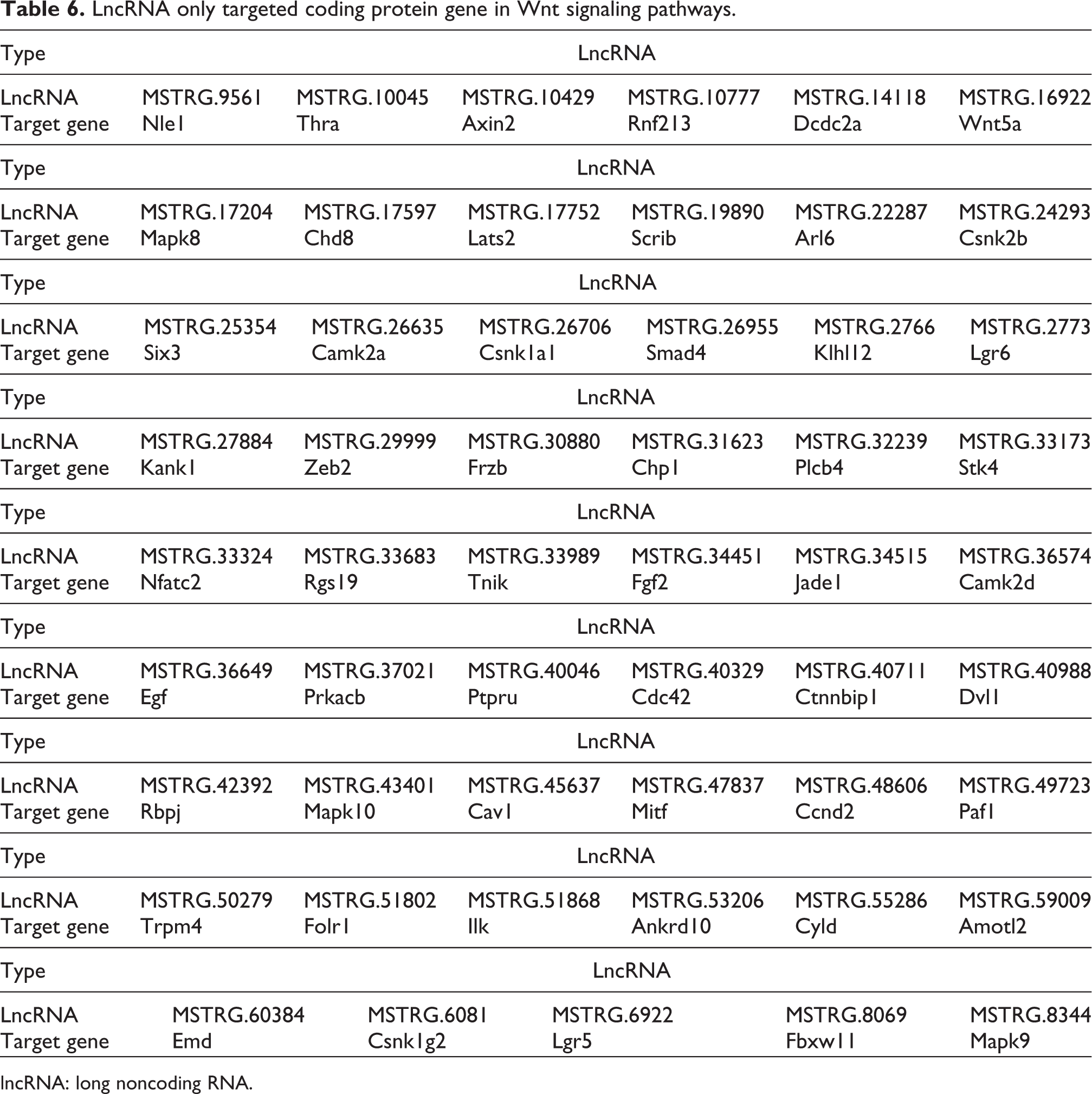
lncRNA: long noncoding RNA.
Discussion
TCDD is considered as a wide range and a highly toxic environmental contaminant, and a strong teratogen for CP (single low dose teratogenesis). 11 However, the systematic collection and characterization of lncRNAs in TCDD-induced CP were still not be reported. In this study, 26,246 novel lncRNAs and 9635 known lncRNAs have been screened out in TCDD-induced CP by RNA-seq, and among which 413 lncRNA transcripts and 65 mRNA transcripts were identified as being significantly different. The cluster and GO analysis revealed that lncRNAs were associated with multiple biological process, cellular component, and molecular function. Meanwhile, many lncRNAs were highly enriched in BMP signaling pathway by bioinformation analysis. For example, seven lncRNAs were targeted together with Smad1 and Smad5 in BMP signaling pathway, which suggested that they may play a significant role in TCDD-induced CP. Subsequently, BMP signaling pathway-related lncRNAs were verified by qRT-PCR analysis, and the results were consistent with RNA-seq.
Numerous studies show that BMP signaling plays important roles in embryonic development, including palatogenesis. 23 –25 BMP signaling is propagated through BMP type I receptors (ALK1, ACVR1/ALK2, BMPRIA/ALK3, and BMPRIB/ALK6) and type II receptors. BMP type I receptor can phosphorylate the intracellular proteins Smad1/5/8. The phosphorylated Smad1/5/8 also can bind Smad4 and translocate to the nucleus to regulate target genes. Simultaneously, all BMP receptors are expressed in dynamic and differential patterns along the anterior–posterior axis of palatal shelves during palatal development, and deregulation of BMP signaling leads to CP formations. 26,27 Inactivation of Alk3 bound with BMP2 and BMP4 with high affinity, which led to CP. 28 Similarly, inactivation of Alk3 decreased mesenchymal cell proliferation and caused to CP. 29 In addition, neural crest-specific ablation of Alk2 resulted in CP due to palatal shelf elevation failure. 30 People also found that the CP happened in BMP7-deficient mice. 31 These findings illustrated that BMP signaling has a decisive influence in palate formation.
In this study, seven lncRNAs were found to be enriched in BMP signaling pathway through biological information prediction. Due to the fact that Smad played a vital role in BMP signaling pathway, we speculated whether these lncRNAs can target Smads. Consistently, the predicted results showed that lncRNA MSTRG.56310.1, MSTRG.51656.1, MSTRG.40182.1, MSTRG.38487.1, MSTRG.1991.1, MSTRG.13399.1, and ENSMUST00000193657 may induce the CP through suppressing the BMP signaling pathway by targeting Smad1 and Smad5. Notably, this hypothesis should be empirically tested, and further studies are required to address this aim. In addition, someone lncRNA only targeted one coding protein gene. For example, lncRNA MSTRG.29999 targeted Zeb2 of Hippo signaling pathway. LncRNA MSTRG.40988 targeted Dvl1 of Wnt signaling pathway. The Hippo and Wnt signaling pathways are also involved in CP. 32 However, the precise mechanisms of lncRNAs in TCDD-induced CP require further investigation.
Conclusions
In summary, we obtained most of DELs in TCDD-induced CP, and 26,246 novel lncRNAs and 9635 known lncRNAs were found. Subsequently, we found that seven lncRNAs can target Smad1 and Smad5 via inhibiting BMP signaling pathway in TCDD-induced CP. In addition, some lncRNAs targeted the important molecules through Hippo and Wnt signaling pathways. These data may provide the basis for further investigation of the function and molecular mechanism of lncRNAs in TCDD-induced CP.
Supplemental material
Supplemental Material, Figure_S1 - Identification and characterization of differentially expressed lncRNA in 2,3,7,8-tetrachlorodibenzo-p-dioxin-induced cleft palate
Supplemental Material, Figure_S1 for Identification and characterization of differentially expressed lncRNA in 2,3,7,8-tetrachlorodibenzo-p-dioxin-induced cleft palate by L-Y Gao, X-L Hao, L Zhang, T Wan, J-Y Liu and J Cao in Human & Experimental Toxicology
Supplemental material
Supplemental Material, Figure_S2 - Identification and characterization of differentially expressed lncRNA in 2,3,7,8-tetrachlorodibenzo-p-dioxin-induced cleft palate
Supplemental Material, Figure_S2 for Identification and characterization of differentially expressed lncRNA in 2,3,7,8-tetrachlorodibenzo-p-dioxin-induced cleft palate by L-Y Gao, X-L Hao, L Zhang, T Wan, J-Y Liu and J Cao in Human & Experimental Toxicology
Supplemental material
supplementary_material - Identification and characterization of differentially expressed lncRNA in 2,3,7,8-tetrachlorodibenzo-p-dioxin-induced cleft palate
supplementary_material for Identification and characterization of differentially expressed lncRNA in 2,3,7,8-tetrachlorodibenzo-p-dioxin-induced cleft palate by L-Y Gao, X-L Hao, L Zhang, T Wan, J-Y Liu and J Cao in Human & Experimental Toxicology
Footnotes
Author contributions
L-YG, J-YL, and JC were responsible for the experimental design. L-YG and X-LH contributed to the execution of experiments, data statistics, and manuscript composition. L-YG, LZ, and TW participated in the discussion and interpretation of data. J-YL and X-LH conceived the study and revised the manuscript. L-YG and JC were responsible for the funding application, and the supervision and management of the project. All authors have contributed to and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the grants of the National Natural Science Foundation of China [81502843].
Statement of Ethics
The Experimental Animal Center Guide for the Care and Use of Laboratory Animals in Henan (SCXK).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
