Abstract
The kidney is one of the main targets for toxicity induced by xenobiotics. Sensitive detection of early impairment is critical to assess chemical-associated renal toxicity. The aim of this study was to identify potential nephrotoxic biomarkers in rat kidney tissues after exposure to mercury (Hg), a representative nephrotoxicant, and to evaluate these new biomarkers employing in vivo and in vitro systems. Mercuric chloride was administered orally to Sprague-Dawley rats for 2 weeks. Proteomic analysis revealed that aldo-keto reductase (AKR7A1) and glutathione S-transferase pi (GSTP1) were significantly elevated in kidney after Hg exposure. While the levels of conventional nephrotoxic clinical markers including blood urea nitrogen and serum creatinine were not elevated, the mRNA and protein levels of AKR7A1 and GSTP1 were increased upon Hg exposure in a dose-dependent manner. The increases in AKR7A1 and GSTP1 were also observed in rat kidneys after an extended exposure for 6 weeks to low-dose Hg. In in vitro rat kidney proximal tubular cells, changes in AKR7A1 and GSTP1 levels correlated well with the extent of cytotoxicity induced by Hg, cadmium, or cisplatin. AKR7A1 and GSTP1 were identified as new candidates for Hg-induced nephrotoxicity, suggesting that these biomarkers have potential for evaluating or predicting nephrotoxicity.
Introduction
Due to its intrinsic functions such as excretion, reabsorption, and accumulation of xenobiotics, the kidney is one of the main targets of xenobiotic-associated toxicity. 1 Environmental toxicants such as mercury (Hg) and cadmium (Cd), as well as pharmaceutical agents including cisplatin, gentamicin, and spironolactone, are known to be associated with impaired renal function. 2,3 Sensitive prediction of renal toxicity is an important issue in safety assessment of drug candidates or chemical compounds and also in clinical settings with acute/chronic renal impairment. Blood urea nitrogen (BUN) and serum creatinine (sCr) are the most commonly used indicators for renal dysfunction in clinical patients and in preclinical animal studies. 4,5 However, these parameters are not considered to be suitable biomarkers for assessment of early renal injury because they are elevated when the kidneys have lost more than 75% of their function. 6 In addition, since many other pathological factors including heart dysfunction, shock, and dehydration can increase and/or interfere with BUN or sCr levels, these markers lack sensitivity in detecting precise stages of renal damage. 7
To overcome these limitations, several alternative molecular markers have been identified, including kidney injury molecule-1 (KIM-1), secreted phosphoprotein 1 (Spp1), tissue inhibitor of metalloproteinases-1 (TIMP1), neutrophil gelatinase–associated lipocalin (Lcn2/NGAL), and cystatin C (CST3), 4,6,8 and the clinical usefulness of these markers has been actively studied. Most of these urinary biomarkers have been extensively studied in the context of acute kidney injury after exposure to high doses of xenobiotics. There are emerging attempt to explore the potential of these nephrotoxic biomarkers using in vivo and in vitro systems 9 ; however, it still remains unclear if these biomarkers are also useful for predicting chronic nephrotoxicity or potential impairment following low dose exposure to xenobiotics. Moreover, it needs to be evaluated if these molecular urinary biomarkers are useful as in vitro biomarkers for preclinical assessment and safety screening of chemicals.
Hg is a heavy metal that exists naturally in the environment. Environmental Hg is derived through naturally occurring and anthropogenic processes including evaporation, volcanic eruption, coal burning, and incineration. 10,11 Once released into the environment, Hg undergoes complicated transformations including biomethylation. Humans can be exposed to inorganic Hg and organic Hg through contaminated water, air, food, or soil. Of note, intake of methylmercury-contaminated food is a global concern and a regulatory issue in human health. 12 Hg potentially damages various organs because of its easy binding to low-molecular-weight proteins and tissue accumulation. Chronic exposure to low-dose Hg was reported to be associated with damages in nervous, renal, immune, and cardiovascular systems in humans. 13 –15 Upon the public health concerns, US Environmental Protection Agency (US EPA) has announced reference doses for mercuric chloride (HgCl2) of 0.3 μg/kg/day and for methylmercury 0.1 μg/kg/day. 16 In 2012, European Food Safety Authority has established tolerable weekly intakes (TWIs) for inorganic Hg of 4 μg/kg and for methylmercury of 1.3 μg/kg, and they mentioned that the mean dietary exposure did not exceed the TWI for methylmercury, while high fish consumers may exceed the TWI by up to approximately six-fold. 17 To cause acute damages in experimental animals, relatively high doses of Hg have been used; however, these situations may not exactly simulate human exposure scenarios of chronic exposure to low level Hg. Nephrotoxicity attributed to Hg has been well established in experimental animals and epidemiological studies. 2,18 –20 While elevated levels of BUN and sCr were detected after acute high-dose Hg exposure (up to 20 mg/kg, per oral) in animals, 20 the early stages of Hg-induced nephrotoxicity were difficult to detect due to the lack of sensitive markers. 21
The aims of this study are to (1) identify early nephrotoxic biomarkers in rat kidney samples after oral exposure of Hg for 2 weeks (up to 7.5 mg/kg) and for 6 weeks (0.15 mg/kg), (2) evaluate the upregulation of these new biomarkers employing in vivo and in vitro systems, and (3) expand the usefulness of the biomarkers by examining the responsiveness to different nephrotoxic compounds such as Hg, Cd, and cisplatin. The levels of the newly identified biomarkers were compared with that of other recently proposed nephrotoxic biomarkers.
Methods
Reagents
The following reagents were obtained from Sigma-Aldrich (St. Louis, Missouri, USA); HgCl2, cadmium chloride (CdCl2), cisplatin, cyano-4-hydroxycinamic acid, 3-(4,5-dimethylthiazol-z-yl)-2,5-diphenyltetrazotium bromide (MTT), ethylenediaminetetraacetic acid (EDTA), benzamidine, thiourea, agarose, glycine, and Coomassie brilliant blue. All other reagents used were in the highest quality available.
Animal treatment
All animal experiments were designed and performed according to the Guide for the Care and Use of Laboratory Animals from the NIH and approved by the Institutional Animal Care and Use Committee (IACUC) at Hanyang University (HY-IACUC-12-059A and HY-IACUC-13-041A). Male Sprague-Dawley rats (3 weeks old, 90–110 g; Koatech, Pyeongtaek, Korea) were given food and water ad libitum and kept under standard conditions for 3 days prior to the onset of administration. For 2-week experiment, rats were divided into four groups, each of which contained five animals. In the normal control group, rats were administered distilled water (DW) as a vehicle. To induce kidney damage with low-dose Hg, rats were orally administered 0.15, 1.5, and 7.5 mg/kg Hg2+ in the form of HgCl2 for 2 weeks. For 6-week treatment, rats were divided into two groups and orally administered DW or 0.15 mg/kg Hg2+ for 6 weeks. On the day after last administration, the animals were killed under isoflurane anesthesia. Blood samples were obtained from the abdominal aorta and kidney samples were immediately frozen in liquid nitrogen. The levels of BUN and sCr were analyzed by Neodin Vetlab (Seoul, Korea).
Proteomics
Proteomic analysis was conducted by Yonsei Proteome Research Center (Seoul, Korea). Briefly, kidney tissue samples were washed and grounded by liquid nitrogen. Protein extracts were prepared with extraction buffer (50 mM Tris, 5 mM EDTA, 7 M urea, 2 M thiourea, 4% CHAPS, and protease inhibitors). Proteins in sample buffer were resolved on an IPG strip (immobilized 4–7 pH gradient strips, 17 cm; Amersham Biosciences, Piscataway, New Jersey, USA) that had been rehydrated with a sample protein solution at 20°C for 14 h. The second dimension of electrophoresis was analyzed on vertical SDS gradient slab gels (9–17%, dimensions 180 × 200 × 1.5 mm) at a constant current of 40 mA for approximately 5 h. After protein fixation in 40% methanol and 5% phosphoric acid for 1 h, the gel was stained overnight with Coomassie Brilliant Blue G-250. After destaining the gel with H2O, the image was scanned using a GS-710 densitometer (Bio-Rad, Hercules, CA). The scanned data were converted to an image file and analyzed with Image Master Platinum 5.0 (Amersham Biosciences, Amersham, UK). 2-DE spots were excised from the gel with pipette tips and digested with trypsin solution (Promega, Madison, Wisconsin, USA). The tryptic peptide mixtures were concentrated and were analyzed by 4800 Proteomics Analyzer (Applied Biosystems, Foster City, California, USA) operated in MS and tandem mass spectrometry (MS/MS) modes. The MS/MS spectra were searched against the National Center for Biotechnology Information (NCBI) rat protein database using the Mascot algorithm (Matrix Science, Boston, Massachusetts, USA) for peptide and protein identification. Peptide mass fingerprinting was carried out using the Mascot search engine included in the GPS Explorer software (Version 3.6, Applied Biosystems). Mass spectra used for manual de novo sequencing were annotated with Data Explorer software (Applied Biosystems). A Mascot ion score >2.0 was used as the criterion for protein identification.
Histological analysis
For histological examination, kidney tissues were fixed in 4% paraformaldehyde, embedded in paraffin, and 5-μm cross sections were stained with hematoxylin and eosin. Histology was analyzed in a blinded fashion to analyze pathological alteration in cortex and medulla to examine cell death, infiltration, and cast formation.
Cell culture
Rat kidney proximal tubular epithelial cells lines (NRK-52E, CRL-1571) were purchased from American Type Culture Collection (ATCC) and cultured under standard conditions (37°C, 5% CO2) in Dulbecco’s modified Eagle medium containing 4.5 g/L glucose supplemented with 2 mM L-glutamine, penicillin/streptomycin (100 IU/mL and 100 µg/mL, respectively), and 5% (v/v) fetal bovine serum, described previously. 22 To examine cytotoxicity and the level of mRNA expression, cells were treated with HgCl2 (0, 0.1, 1, and 5 µM), CdCl2 (0, 0.1, 1, and 5 µM), or cisplatin (0, 5, and 25 µM) for 24 h.
Cytotoxicity assay
Cytotoxicity was assessed by the MTT assay or by quantitating leakage of lactate dehydrogenase (LDH). Briefly, cells were seeded into 96-well cell culture dishes at a density of 5.0 × 10 4 cells/ml and allowed to grow for 24 h. Next, cells were treated with HgCl2, CdCl2, or cisplatin for 24 h. After treatment, 100 µl of a 5 mg/mL MTT solution was added to each well and the cells were further incubated for 2 h at 37°C. The medium was then replaced by 100 µl of dimethyl sulfoxide (DMSO) and mixed thoroughly to dissolve the formazan crystals, after which absorbance was read at 470 nm. The released amount of LDH was measured using a CytoTox 96® Non-Radioactive Cytotoxicity Assay kit (Promega). After incubation with the indicated chemicals, the conditioned medium was collected and centrifuged at 250 × g for 4 min. Next, 50 µl supernatant was transferred to a 96-well plate, and 50 µl LDH assay solution was added to each well. After 30 min of incubation in the dark at room temperature, 10 µl stop solution was added to each well and the absorbance was read at 490 nm. More than four independent experiments were carried out, each consisting of three technical replicates. Results are expressed as percentage, taking control as 100%.
Real-time polymerase chain reaction
Total RNA was extracted from frozen kidney tissue or NRK-52E cells using TRIzol reagent (Invitrogen, Carlsbad, California, USA) according to the manufacturer’s recommendations. RNA concentrations were quantified by measuring ultraviolet absorbance at 260 nm with a NanoDrop ND-2000 UV-Vis Spectrophotometer (NanoDrop Technologies, Wilmington, Delaware, USA). Complementary DNA (cDNA) was synthesized from 0.5 µg total RNA using iScript™ Reverse Transcription Supermix (Bio-Rad). The mRNA levels for nephrotoxic biomarkers were measured by quantitative real-time polymerase chain reaction. Reactions had a total volume of 20 µl containing 10 µl of iQ™ SYBR® Green Supermix (Bio-Rad), 0.5 µg cDNA, and 100 nM of each primer. cDNA amplification was performed using the following thermocycling parameters: 3-min enzyme activation at 95°C, followed by 45 cycles of 95°C for 10 s, 55°C for 30 s, and 72°C for 10 s. The primer sets for amplification of aldo-keto reductase (AKR7A1), glutathione S-transferase pi (GSTP1), and CST3 were designed by Primer-Blast at NCBI, while the primer sets for 18S ribosomal RNA (rRNA), KIM-1, Spp1, TIMP1, and Lcn2 were described previously. 23 –26 The primer sets are summarized in Table 1. Gene expression changes were determined using the 2−ΔΔCt method 27 and normalized to housekeeping gene 18S. Results are presented as mean fold change in mRNA expression level compared to the level in control group.
Primer sequences for amplification of nephrotoxic biomarkers.

Western blotting
Kidney samples were homogenized in radio-immunoprecipitation assay (RIPA) buffer (Thermo Scientific, San Jose, California, USA) and centrifuged at 15,000 × g for 15 min. The supernatant was separated, and the protein concentration of the lysate was determined with a BCA protein assay kit (Thermo Scientific). For Western blotting, equal amounts of protein were analyzed by sodium dodecyl sulfate–polyacrylamide gel electrophoresis. The protein was then transferred to a polyvinylidene fluoride (PVDF) membrane (EMD Millipore, Billerica, Massachusetts, USA). The membrane was blocked with PBS-T (phosphate buffered saline with 0.1% Tween-20) solution containing 5% bovine serum albumin and then incubated overnight at 4°C with primary antibodies against α-actinin (polyclonal, 1:5000; Santa Cruz Technology, Santa Cruz, California, USA), AKR7A1 (polyclonal, 1:1000; Santa Cruz Technology), or GSTP1 (monoclonal, 1:2000; Sigma-Aldrich). Subsequently, membranes were incubated for 1 h with horseradish peroxidase–conjugated secondary antibodies (1:5000). The blots were washed with PBS-T solution, and the chemiluminescent intensity of each band was quantitated with an Odyssey imaging system (LI-COR Biosciences, Lincoln, Nebraska, USA).
Statistical analysis
Data are presented as mean ± standard errors of the mean. Statistical analysis was performed by Student’s t-test or Mann–Whitney U test. One-way analysis of variance test was conducted for more than two groups followed by Dunnett’s post hoc analysis to compare each treatment group against the control group. Values significantly different from the control value are indicated as *p < 0.05 or **p < 0.01.
Results
Identification of AKR7A1 and GSTP1 as nephrotoxic biomarkers in rat kidney after 2-week Hg exposure
In order to identify protein biomarkers of early nephrotoxicity, rat kidney tissues were examined using proteomics. Rats were treated with Hg, a representative nephrotoxicant, for 2 weeks via the oral route. Compared to the control group, rats exposed to 0.15, 1.5, or 7.5 mg/kg Hg did not show statistically significant difference in body weight, food uptake, and water consumption (data not shown). The Hg levels tested did not induce changes in the levels of BUN or sCr, the most commonly used end-stage clinical markers for renal functional damage (Figure 1(a)). Histological examination revealed that there were slight morphological changes in kidney pathology such as cell death, infiltration, and cast formation in Hg-exposed rats compared to control (Figure 1(b)). To identify potential protein biomarkers, a proteomic analysis was conducted with rat kidney samples isolated after 2-week Hg exposure. The protein levels of AKR7A1 and GSTP1 were highly increased in rat kidneys after exposure to 1.5 or 7.5 mg/kg Hg (Figure 2(a)). The increased protein levels of AKR7A1 and GSTP1 in rat kidneys after 2-week Hg exposure were further confirmed by Western blot analysis (Figure 2(b)). To evaluate the potential of AKR7A1 and GSTP1 as new nephrotoxic biomarkers, the mRNA expression levels in rat kidney were compared with those of recently suggested molecular biomarkers of KIM-1, Spp1, TIMP1, Lcn2/NGAL, and CST3. As shown in Figure 2(c), the mRNA levels of AKR7A1 and GSTP1 were increased upon Hg exposure in a dose-dependent manner compared to traditional biomarkers, suggesting that these two proteins are potential biomarker candidates for Hg-induced nephrotoxicity.

Effects of 2-week Hg exposure on rat kidney. Rats were treated with Hg (0, 0.15, 1.5, or 7.5 mg/kg) for 2 weeks. (a) The levels of BUN and sCr were assessed after Hg exposure for 2 weeks. (b) Histological changes in rat kidney following 2-week Hg exposure were examined with H&E staining. Histopathological examination demonstrates cell death (black arrow), infiltration (white arrowhead), and cast formation (black arrowhead). Scale bar = 200 μm. Representative images are shown. n = 5. Hg: mercury; H&E: hematoxylin and eosin.

Identification of nephrotoxic protein markers in rat kidney after 2-week Hg exposure. Rats were treated with Hg (0, 0.15, 1.5, or 7.5 mg/kg) for 2 weeks. (a) Two-dimensional proteomic analysis revealed that the levels of AKR7A1 and GSTP1 were increased in kidneys isolated from Hg-exposed rats after 2-week treatment. Representative images from control and 1.5 mg/kg Hg-exposed rats were shown. (b) The protein levels of AKR7A1 and GSTP1 in kidney were measured by Western blot analysis. (c) The mRNA levels of the newly identified biomarkers (AKR7A1 and GSTP1) and previously suggested nephrotoxic biomarkers (KIM-1, Spp1, TIMP1, Lcn2, and CST3) were analyzed by RT-qPCR. n =5. Values are mean ± SEM. *p < 0.05 or **p < 0.01 vs. control. Hg: mercury; AKR7A1: aldo-keto reductase 7A1; GSTP1: glutathione S-transferase pi; mRNA: messenger RNA; KIM-1: kidney injury molecule-1; Spp1: secreted phosphoprotein 1; TIMP1: tissue inhibitor of metalloproteinases-1; Lcn2: neutrophil gelatinase–associated lipocalin; CST3: cystatin C; RT-qPCR: quantitative real-time PCR; SEM: standard error of the mean.
Upregulation of AKR7A1 and GSTP1 in rat kidney after 6-week Hg exposure
Next, rats were exposed to low-dose Hg (0.15 mg/kg Hg) for 6 weeks, thereby simulating long-term low-level exposure in humans, and the expression levels of these biomarkers in rat kidneys were analyzed. The levels of AKR7A1 and GSTP1 protein were significantly increased by Hg exposure for the extended period (Figure 3(a)). The mRNA levels of AKR7A1 and GSTP1 in Hg-exposed kidney tissue correlated well with the protein levels, while the mRNA levels of KIM-1, Spp1, TIMP1, and Lcn2 were not affected by Hg exposure for 6 weeks (Figure 3(b)). The clinical levels of BUN and sCr were not statistically significantly different compared to those of the control group, indicating that there was no severe functional damage to the kidneys by 6-week exposure to 0.15 mg/kg Hg (data not shown). The increases in the AKR7A1 and GSTP1 mRNA and protein levels in rat kidneys were consistent after Hg exposure for 2 weeks (0.15, 1.5, and 7.5 mg/kg Hg) and for 6 weeks (0.15 mg/kg Hg), suggesting that these two protein markers may be useful to identify early renal toxicity caused by low-level exposure to nephrotoxicants such as Hg.
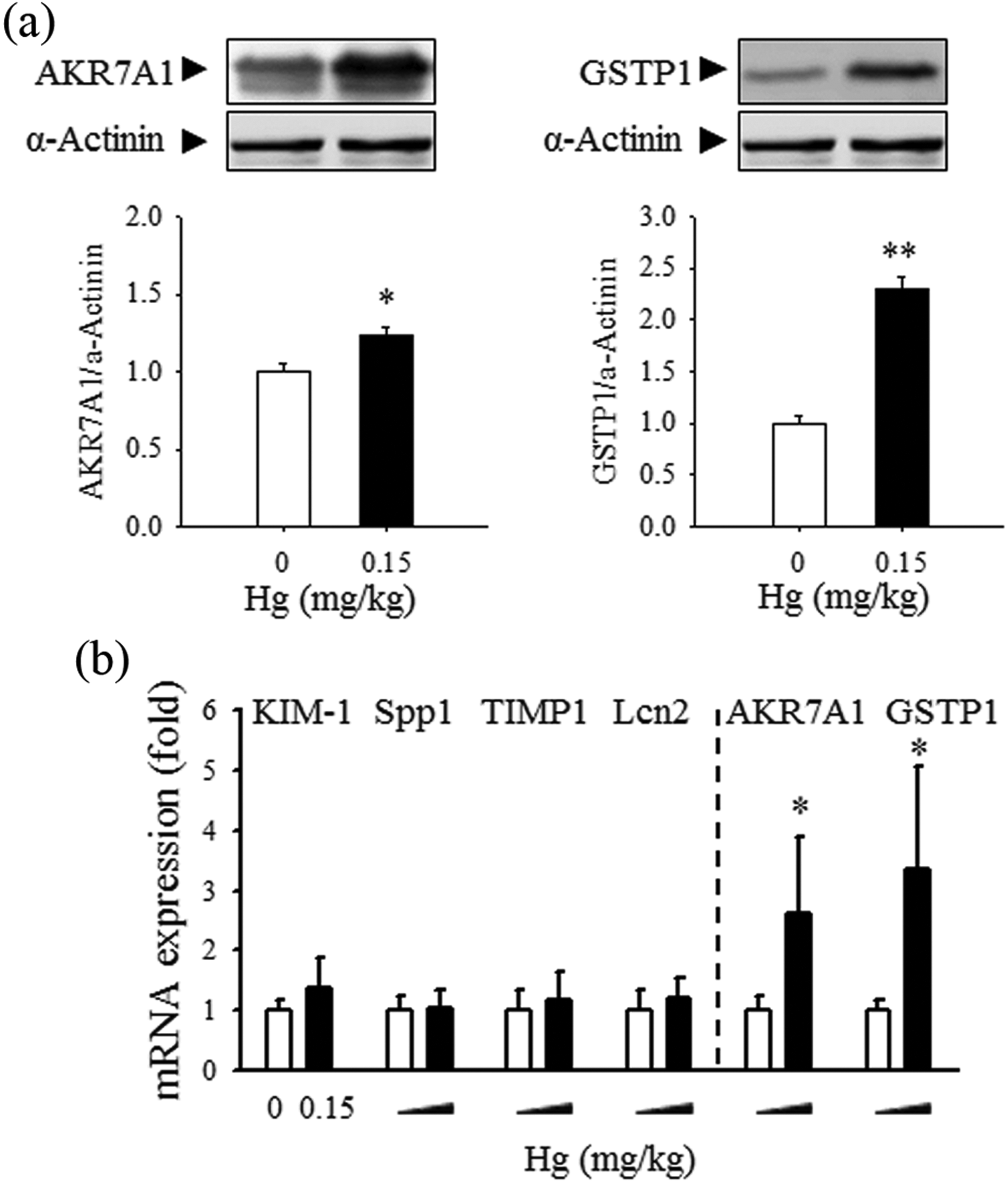
Effects of 6-week Hg exposure on protein markers in rat kidneys. Rats were treated with Hg (0 or 0.15 mg/kg) for 6 weeks. (a) The protein levels of AKR7A1 and GSTP1 in kidney were measured by Western blot analysis. (b) The mRNA levels of the newly identified biomarkers (AKR7A1 and GSTP1) and previously suggested nephrotoxic biomarkers (KIM-1, Spp1, TIMP1, and Lcn2) were analyzed by RT-qPCR. n = 5. Values are mean ± SEM. *p < 0.05 or **p < 0.01 vs. control. Hg: mercury; AKR7A1: aldo-keto reductase 7A1; GSTP1: glutathione S-transferase pi; mRNA: messenger RNA; KIM-1: kidney injury molecule-1; Spp1: secreted phosphoprotein 1; TIMP1: tissue inhibitor of metalloproteinases-1; Lcn2: neutrophil gelatinase–associated lipocalin; CST3: cystatin C; RT-qPCR: quantitative real-time PCR; SEM: standard error of the mean.
In vitro evaluation of AKR7A1 and GSTP1 increase using proximal tubular epithelial cells
Identification of sensitive in vitro markers is critical for preclinical assessment and safety screening of chemicals and drug candidates. Therefore, the potential of AKR7A1 and GSTP1, the newly identified nephrotoxic biomarkers, as in vitro predictive markers was investigated in rat kidney tubular cells (NRK-52E). Rat tubular epithelial cells were treated with HgCl2 (0, 0.1, 1, and 5 µM), after which, Hg-induced cytotoxicity was evaluated by quantitating MTT conversion and the released level of LDH. As shown in Figure 4(a), treatment with Hg (1 and 5 µM) for 24 h significantly decreased cell viability as observed by MTT assay. The leakage of LDH showed a similar pattern. The expression levels of several nephrotoxic biomarkers including KIM-1, Spp1, TIMP1, and Lcn2, along with the two new candidate biomarkers of AKR7A1 and GSTP1 were examined. Interestingly, the mRNA level of the previously proposed biomarkers was not affected by Hg exposure, whereas the mRNA expression levels of AKR7A1 and GSTP1 in tubular cells were significantly elevated by Hg (Figure 4(a)). Low-level Hg exposure (0.1 µM), which did not show significant cytotoxicity in tubular cells, induced increase in mRNA levels of AKR7A1 and GSTP1 although the statistical significance was not achieved. This finding suggests that the changes in these biomarkers can be upregulated in the absence of the cytotoxicity observed by MTT and LDH assays, suggesting that AKR7A1 and GSTP1are more sensitive.
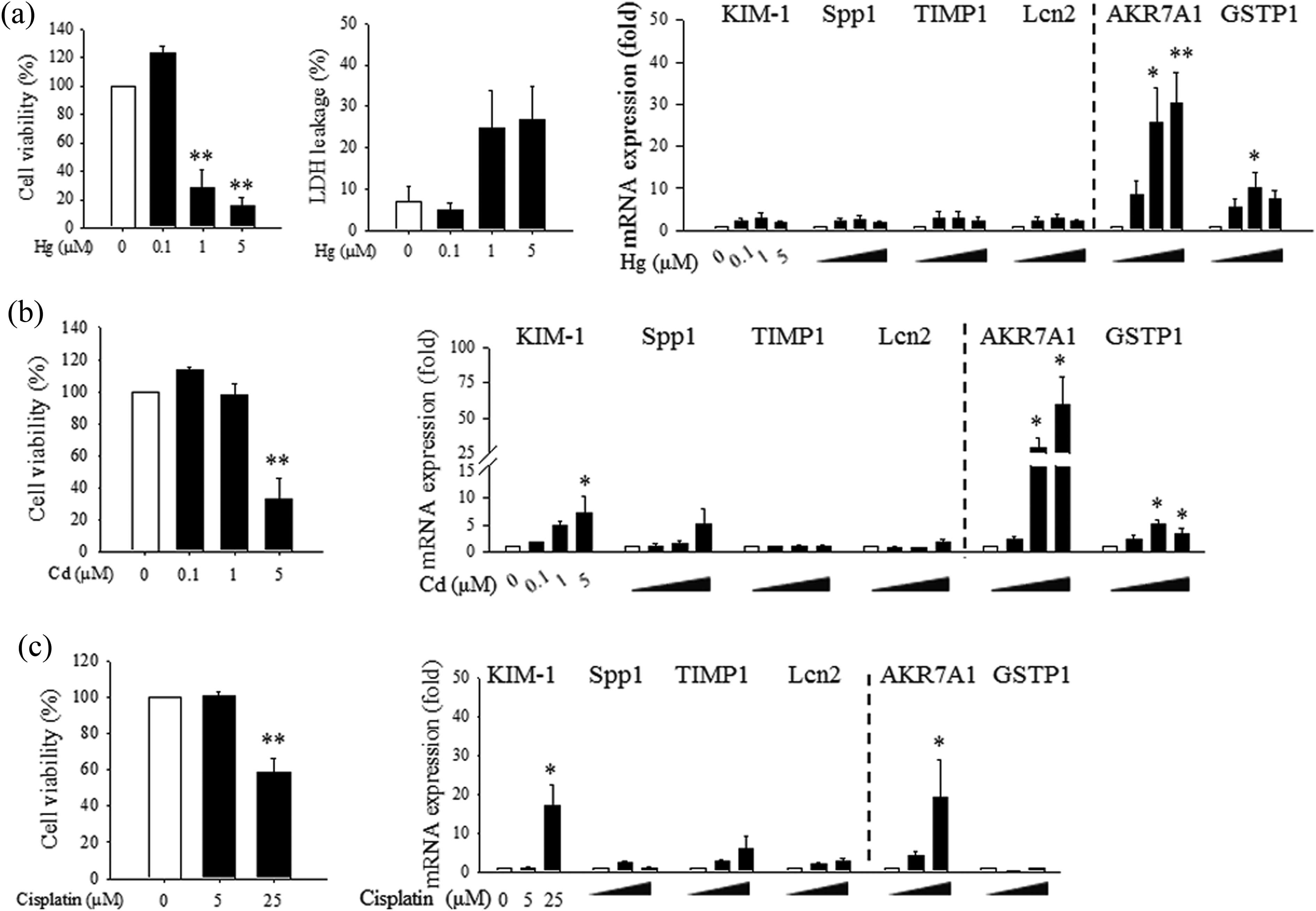
In vitro evaluation of nephrotoxic biomarkers in rat kidney tubular epithelial cells after exposure to Hg, Cd, or cisplatin. Rat tubular epithelial cells (NRK52E) were treated with Hg (0, 0.1, 1, and 5 µM HgCl2), Cd (0, 0.1, 1, and 5 µM CdCl2) or cisplatin (0, 5, and 25 µM) for 24 h. (a) Cell viability and cytotoxicity were measured by the MTT and LDH leakage assays in NRK52E cells after Hg treatment for 24 h. The mRNA levels of KIM-1, Spp1, TIMP1, and Lcn2, in addition to those of the newly identified biomarkers (AKR7A1 and GSTP1), in Hg-treated NRK52E cells were determined by RT-qPCR. (b) and (c) The changes in cell viability and mRNA expression in NRK52E cells exposed to cadmium (b) or cisplatin (c) were analyzed. n = 4–5. Values are mean ± SEM. *p < 0.05 or **p < 0.01 vs. control. Hg: mercury; AKR7A1: aldo-keto reductase 7A1; GSTP1: glutathione S-transferase pi; mRNA: messenger RNA; KIM-1: kidney injury molecule-1; Spp1: secreted phosphoprotein 1; HgCl2: mercuric chloride; CdCl2: cadmium chloride; TIMP1: tissue inhibitor of metalloproteinases-1; Lcn2: neutrophil gelatinase–associated lipocalin; CST3: cystatin C; Cd: cadmium; MTT: -(4,5-dimethylthiazol-z-yl)-2,5-diphenyltetrazotium bromide; LDH: lactate dehydrogenase; RT-qPCR: quantitative real-time PCR; SEM: standard error of the mean.
Upregulation of AKR7A1 and GSTP1 by Cd and cisplatin in tubular epithelial cells
In order to investigate if the levels of AKR7A1 and GSTP1 were affected by other nephrotoxicants, tubular cells were treated with cadmium (CdCl2; 0, 0.1, 1, and 5 µM) or cisplatin (0, 5, and 25 µM). Treatment with cadmium for 24 h decreased cell viability in a dose-dependent manner (Figure 4(b)). The mRNA expression levels of AKR7A1 and GSTP1 in tubular cells were significantly increased by cadmium treatment, showing superior performance compared to traditional molecular markers. These trends were also observed in drug-induced nephrotoxicity. Cisplatin, a well-established nephrotoxic chemotherapeutic agent, induced significant cytotoxicity in tubular epithelial cells, and the mRNA level of AKR7A1 was significantly increased by cisplatin (Figure 4(c)).
Discussion
In this study, two new protein markers of AKR7A1 and GSTP1 were identified using a proteomic analysis of Hg-exposed rat kidneys. This observation is in line with our recent report regarding protein alteration in rat kidney following exposure to Hg. 21 These markers were upregulated in the absence of functional damage, as measured via BUN and sCr levels in our experimental condition. The upregulation of AKR7A1 and GSTP1 by Hg was validated using in vivo kidney tissues and in vitro tubular epithelial cells. Tubular epithelial cells were treated with Cd and cisplatin to examine if these markers are upregulated in respond to other nephrotoxicants.
Drug- or chemical-associated renal impairment is one of the most serious target organ toxicities. 28 However, early and accurate detection of renal damage is still limited due to a lack of sensitive biomarkers. Here, it is demonstrated that the new markers AKR7A1 and GSTP1 maybe more sensitively upregulated than the previously suggested nephrotoxic markers, meaning that they might be useful for in vitro safety screening in kidney tubular epithelial cells. The consistent results in our in vivo and in vitro experimental systems are interesting since there are extensive movements to develop in vitro alternative toxicity testing methods relevant to in vivo toxicity. 29 Due to the target organ–selective nature of many nephrotoxic xenobiotics, in vitro methods have become invaluable tools for obtaining predictive toxicity information related to kidney damage. 30,31 The mRNA levels of AKR7A1 and GSTP1 were statistically significantly upregulated after Hg treatment compared with the levels in the control group. Moreover, the mRNA upregulation of AKR7A1 was greater than that of other markers, even at the lowest concentration of Hg (0.1 µM), which did not show cytotoxicity. This finding indicates that these biomarkers might be good candidates for predicting nephrotoxicity.
Elevation of BUN or sCr reflects collapse of renal function, 6 and exposure to high dose of Hg (20 mg/kg) or Cd (25 mg/kg) is known to result in significant increase in BUN level. 20,32 In our results, the clinical functional parameters of BUN or sCr were not affected (Figure 1), while the mRNA and protein levels of AKR7A1 and GSTP1 in rat kidney were increased both after 2-week- and 6-week Hg exposure. Although there were no apparent functional nephrotoxicity observed by BUN or sCr levels by Hg (up to 7.5 mg/kg) in our in vivo experiments, Hg-induced alterations in AKR7A1 and GSTP1 in kidney were observed in dose-dependent and exposure duration–dependent manner. Interestingly, in an adverse outcome pathway approach, a recent concept to explain toxicological phenomena, xenobiotic-induced toxicity is a complicated sequential process from a molecular initiating event, which refers to the direct interaction of chemical with its biological target molecule, to subsequent responses at cellular, tissue, organ, and individual level through the relationship between key events. 33 It should be noted that identification of KEs in molecular and cellular levels is critical to predict or explain toxicity (i.e. adverse outcome) in organ and organism level. 29,34 In this regard, our results demonstrate that Hg-induced biomarker changes in the absence of functional alteration may represent early effect pathways. Histopathological changes in Hg-exposed kidneys support that there are minute adverse reactions in tissue level in the Hg-dose range tested. This is in line with the results from in vitro cell system, where upregulation of AKR7A1 or GSTP1 was observed at the lower concentration of Hg (0.1 μM), Cd (1 μM), or cisplatin (5 μM) in the absence of cytotoxicity. Further characterization of the role of these biomarkers warrants a future study that might include a relationship between these biomarkers and functional damage.
Extensive efforts have focused on developing urinary biomarkers for kidney damage. 35 –37 In particular, attention has been paid to the representative candidates of CST3, b2-microglobulin, total protein, KIM-1, albumin, clusterin, and trefoil factor 3. 38 –40 However, these urinary markers can be detected when they are excreted into the urine following functional disorder of the kidney, including destructed renal cell membrane or glomerulus disorder. AKR7A1 is a member of aldo-keto reductase superfamily. These proteins can detoxify toxic metabolites such as dialdehyde metabolite of aflatoxin B1. AKR7A1 expression can be induced by exposure to several xenobiotics such as ethoxyquin or coumarin. 41 GSTP1, a member of GST superfamily, is a classical detoxification enzyme that catalyzes the conjugation of glutathione (GSH) to a numerous xenobiotics and protects cells from oxidative stress. 42 The main goal of this study is to identify and validate new nephrotoxic biomarker candidates, and the detailed mechanisms how these markers are related with other intracellular toxic phenomena may be beyond the scope of the current study. However, in order to get a clue for possible relationship between disrupted redox balance by Hg and the increase of these enzyme biomarkers in renal tubular cells, we measured oxidative stress markers in our in vitro cell system, as shown in Supplemental Figure 1. The generation of reactive oxygen species as measured by dichlorofluorescein (DCF) was increased by Hg in a dose-dependent manner (Supplemental Figure 1A), supporting that oxidative stress might be an important contributing factor to Hg-induced renal cell damage. The effects of Hg on thiol status including glutathione system (GHS/GSSG) and related enzyme systems are known as complicated networks. 43 –45 While the levels of total GSH were increased by Hg in a dose-dependent manner, which is consistent to previous reports, 43,45 there was no obvious alteration in the levels of reduced GSH (Supplemental Figure 1B and 1C), possibly due to the dynamics of thiol status. Since AKR7A1 and GSTP1 are detoxification enzymes, the increased oxidative stress and alterations of thiol status might be related to compensatory upregulation of these enzyme markers. Further studies on the details for the expression of these two enzymes, such as the time course and the signaling pathways for their induction, would be meaningful to understand the upregulation of these enzymes in response to the renal cell damage.
It is likely that upregulation of AKR7A1 and GSTP1 in kidney tissue is xenobiotic-associated metabolic induction, and these can be explained as mechanism-based biomarkers in view of predictive toxicology. Supporting our observation, there have been several previous reports on the potential of GSTs as putative markers for kidney damage. 46,47 Based on our results, AKR7A1 and GSTP1 may serve as potential biomarkers for the early stage of nephrotoxicity, which can be detected in the absence of late kidney dysfunction. It is not clear that these two biomarkers can serve as noninvasive clinical markers that should be detected in human urine or blood; however, here we suggest that these markers might be reflecting xenobiotic-associated renal damage relevantly both in vivo rats and in vitro renal tubular cells, supporting their potential as nephrotoxic markers for in vitro safety assessment of chemicals. It would be necessary to determine whether these enzymes can be found in blood or serum in appropriate time- and severity-window, to expand and validate the clinical potential of these two enzyme markers.
Collectively, our results support that AKR7A1 and GSTP1 may be biomarker candidates for early stages of nephrotoxicity and be suitable for screening chemical-associated nephrotoxicity in in vitro safety assessment.
Supplemental material
Supplementary_material - Identification of aldo-keto reductase (AKR7A1) and glutathione S-transferase pi (GSTP1) as novel renal damage biomarkers following exposure to mercury
Supplementary_material for Identification of aldo-keto reductase (AKR7A1) and glutathione S-transferase pi (GSTP1) as novel renal damage biomarkers following exposure to mercury by Y-J Shin, K-A Kim, E-S Kim, J-H Kim, H-S Kim, M Ha, and O-N Bae in Human & Experimental Toxicology
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Ministry of Food and Drug Safety, Republic of Korea (grant nos. 12162MFDS736 and 15162MFDS045).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
