Abstract
Work in recent years has provided strong evidence for the modulation of memory function and neuroplasticity mechanisms across circadian (daily), ultradian (shorter-than-daily), and infradian (longer-than-daily) timescales. Despite rapid progress, however, the field has yet to adopt a general framework to describe the overarching role of biological rhythms in memory. To this end, Iyer and colleagues introduced the term iterative metaplasticity, which they define as the “gating of receptivity to subsequent signals that repeats on a cyclic timebase.” The central concept is that the cyclic regulation of molecules involved in neuroplasticity may produce cycles in neuroplastic capacity—that is, the ability of neural cells to undergo activity-dependent change. Although Iyer and colleagues focus on the circadian timescale, we think their framework may be useful for understanding how biological rhythms influence memory more broadly. In this review, we provide examples and terminology to explain how the idea of iterative metaplasticity can be readily applied across circadian, ultradian, and infradian timescales. We suggest that iterative metaplasticity may not only support the temporal niching of neuroplasticity processes but also serve an essential role in the maintenance of memory function.
Keywords
Iterative Metaplasticity
Biological rhythms pervasively modulate memory function. Discourse in this domain tends to revolve around the circadian (daily) regulation of behavior and neuroplasticity (Krishnan and Lyons, 2015; Rawashdeh et al., 2018; Hartsock and Spencer, 2020). However, a growing body of work suggests that memory function is cyclically modulated also on ultradian (shorter-than-daily) and infradian (longer-than-daily) timescales. Studies across these 3 timescales are now beginning to converge, creating a need for language that can capture the complex phenomena they reveal.
Research in the memory field suggests that basal neural features may dictate neuroplastic capacity—that is, the ability of neural cells to undergo activity-dependent change. For instance, the basal excitability of the postsynaptic neuronal membrane determines the degree to which stimulation will induce postsynaptic neuroplasticity (Dong et al., 2006; Han, 2006; Zhou et al., 2009). Neuroplastic capacity can change over time, and this process is called metaplasticity (Abraham and Bear, 1996). In the context of circadian rhythms, circadian fluctuations in basal neural features may underlie circadian rhythms in neuroplastic capacity. We and others have described the circadian modulation of neuroplastic capacity as circadian metaplasticity (Gerstner and Yin, 2010; Gerstner, 2012; Hartsock and Spencer, 2020).
When we recently reviewed mechanisms of circadian metaplasticity (Hartsock and Spencer, 2020), we were unaware of another publication describing a related principle, iterative metaplasticity (Iyer et al., 2014). The authors define iterative metaplasticity as the “gating of receptivity to subsequent signals that repeats on a cyclic timebase.” In other words, iterative metaplasticity is the same as circadian metaplasticity but is not limited to the circadian timescale. Iyer and colleagues, however, provide examples drawn exclusively from the circadian literature. Here, we propose that iterative metaplasticity might be used as an umbrella term to encompass circadian, ultradian, and infradian forms. The literature we review spans a wide range of the phylogenetic scale, including studies from flies, crabs, birds, rodents, and humans. This body of work suggests that iterative metaplasticity across timescales may coordinate the adaptive timing of neuroplasticity processes and play a key role in the maintenance of optimal memory function.
Iterative Metaplasticity On A Daily Timescale: Circadian Metaplasticity
Circadian metaplasticity is a form of iterative metaplasticity that repeats on a daily timescale (Figure 1, middle panel). Circadian rhythms are generated and maintained endogenously by molecular clocks located in cells throughout the body (Reppert and Weaver, 2002). As first described in Drosophila (Bargiello et al., 1984; Zehring et al., 1984; Vosshall et al., 1994; Allada et al., 1998; Darlington, 1998; Price et al., 1998; Young and Kay, 2001) and subsequently found in many different vertebrate species, molecular clocks arise from daily transcriptional-translational feedback loops involving the 24-h oscillatory expression patterns of clock genes. The mammalian core clock genes include brain and muscle ARNT-like protein-1 (Bmal1), circadian locomotor output cycles kaput (Clock), period (Per1/2/3), and cryptochrome (Cry1/2) (Reppert and Weaver, 2002). Another clock gene, neuronal PAS domain protein 2 (Npas2), can replace Clock in certain circumstances (Reick, 2001; DeBruyne et al., 2006). Notably, many non-mammalian organisms exhibit a comparable molecular clock mechanism but with variations in the genetically encoded molecular players (Young and Kay, 2001). The existence of molecular clocks even in evolutionarily more ancient organisms, such as cyanobacteria (Swan et al., 2018), points to the phylogenetic conservation of molecular clocks and their importance for basic biological functions (Bhadra et al., 2017).
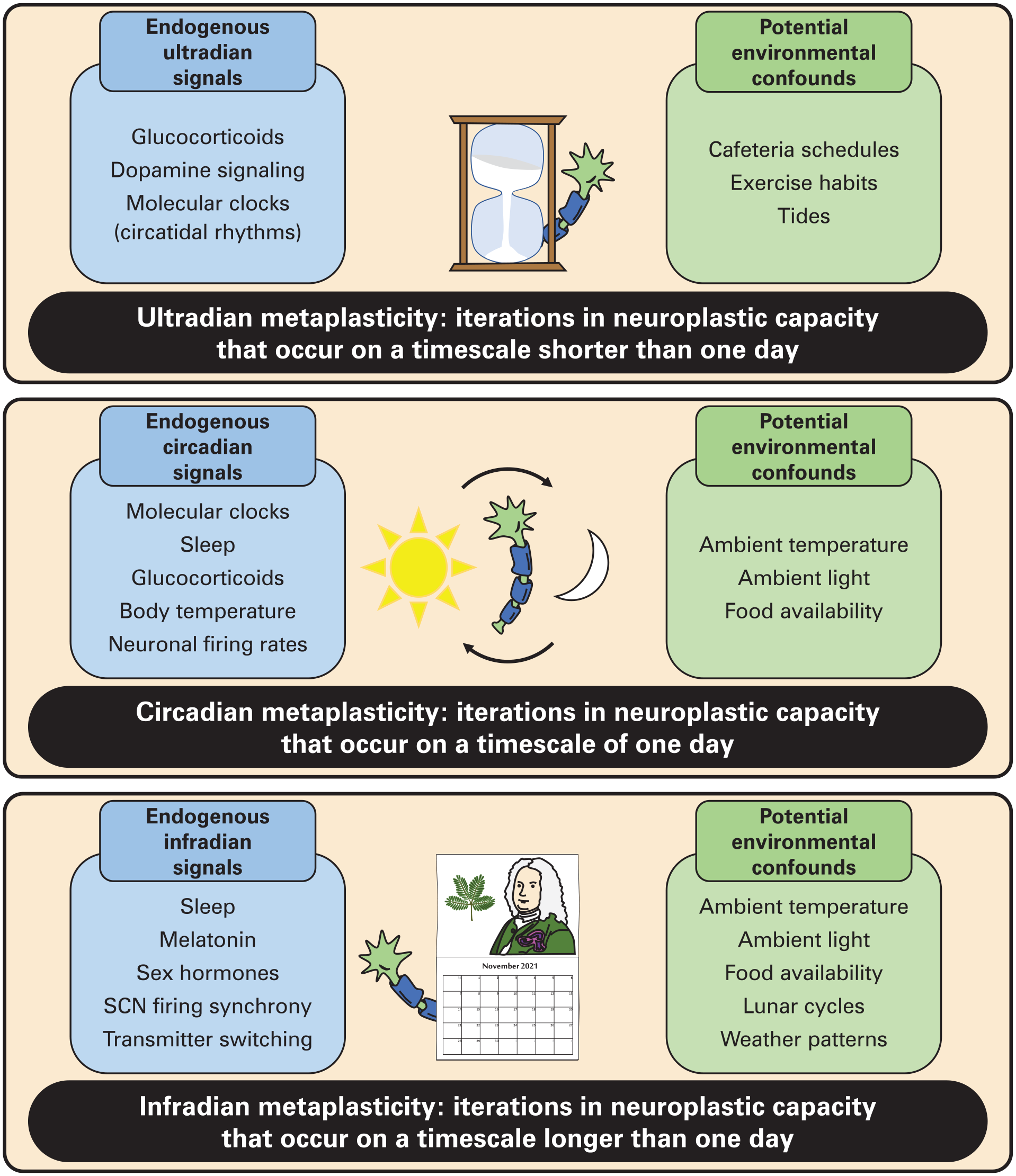
Sources of iterative metaplasticity across timescales. Iterations in neuroplastic capacity may occur on ultradian (hourglass), circadian (solar cycle), and infradian (calendar) timescales. On each timescale, iterations generated by endogenous cyclic signals must be distinguished from iterations generated by environmental cyclic signals. Abbreviation: SCN = suprachiasmatic nucleus.
Although mammalian molecular clocks are expressed in cells throughout the body, they are organized into a hierarchy. At the top of this hierarchy sits the suprachiasmatic nucleus (SCN) of the hypothalamus. The SCN receives direct input from intrinsically photosensitive retinal ganglion cells, which support the time-of-day-dependent induction of Per1 and Per2 gene expression in SCN cells in response to ambient light exposure (Reppert and Weaver, 2001; Hattar, 2002; Güler et al., 2008). Thus, molecular clocks in the SCN are able to align their oscillations to the solar cycle (Colwell, 2011). Complex interactions among SCN neurons and astrocytes generate tissue-level synchrony of circadian rhythms in SCN firing rates (Green and Gillette, 1982; Herzog et al., 1998; Albus et al., 2002; Kudo et al., 2015), which convey solar timing information to nearby brain regions (Abrahamson and Moore, 2001). In turn, these downstream sites control a variety of signals—including hormones, body temperature, and neuronal firing—that relay solar timing information to molecular clocks local to other tissues (Koronowski and Sassone-Corsi, 2021). By synchronizing to these SCN-orchestrated signals, molecular clocks in tissues outside the SCN can coordinate their rhythms to the rhythm of the SCN master pacemaker.
The original reviews suggesting that circadian rhythms in neuroplasticity represent a form of metaplasticity focus on similarities between the SCN and hippocampus (Gerstner and Yin, 2010; Gerstner, 2012; Iyer et al., 2014). For example, daily solar resetting of molecular clocks in the SCN depends on key intracellular signals, particularly the cAMP-MAPK-CREB cascade (Ginty et al., 1993; Obrietan et al., 1998; Tischkau et al., 2003), that also mediate neuroplasticity in the hippocampus (Frey et al., 1993; Impey et al., 1998; Thomas and Huganir, 2004). In addition, the capacity of neurons to undergo long-term potentiation, an electrophysiological phenomenon important for many forms of activity-dependent neuroplasticity, is modulated in a time-of-day-dependent fashion in both the SCN (Nishikawa et al., 1995) and hippocampus (Harris and Teyler, 1983; Dana and Martinez, 1984; Raghavan et al., 1999). Thus, the circadian modulation of basal neural features underlies circadian solar entrainment in the SCN and the circadian modulation of activity-dependent neuroplasticity in the hippocampus.
In the hippocampus and other regions outside the SCN, circadian oscillations in neuroplastic capacity can arise from 3 main sources: molecular clocks, SCN-orchestrated intercellular signals, and circadian-regulated processes that influence neuroplasticity. Molecular clocks act largely through the control of gene transcription. The BMAL1:CLOCK/NPAS2 heterodimer acts as a transcription factor at E-box response elements that are associated with the promoter regions of not only some other core clock genes (Per and Cry) but also many non-clock genes (R. Zhang et al., 2014; Mure et al., 2018). Work in Drosophila suggests that the molecular clock belies rhythmic expression of many neuroplasticity-related transcripts (Figure 2, middle panel) (Claridge-Chang et al., 2001; Cirelli et al., 2004), including NCA localization factor 1 (Nlf-1), which controls the membrane localization of leak channels and basal membrane excitability in neurons (Flourakis et al., 2015). In the rodent ventral tegmental area, the dopamine synthetic enzyme tyrosine hydroxylase, whose promoter contains an E-box, exhibits greater expression during the active phase than the inactive phase even in constant darkness, pointing to a mechanism for the circadian regulation of dopaminergic neuromodulation (Chung et al., 2014; Logan et al., 2019). Similarly, the dopamine degradative enzyme monoamine oxidase A has an E-box in its promoter that binds BMAL1:NPAS2 and modulates nucleus accumbens dopamine concentrations in a time-of-day-dependent manner (Hampp et al., 2008). Many studies have shown that interactions between the molecular clock machinery and histone-modifying enzymes may also contribute to circadian rhythms in gene expression and memory-related behavior (Nakahata et al., 2008; Michan et al., 2010; Chung et al., 2014; Perreau-Lenz et al., 2017; Kwapis et al., 2018). Through transcriptional control, molecular clocks in the brain can generate rhythms in the availability of transcripts important for activity-dependent plasticity.
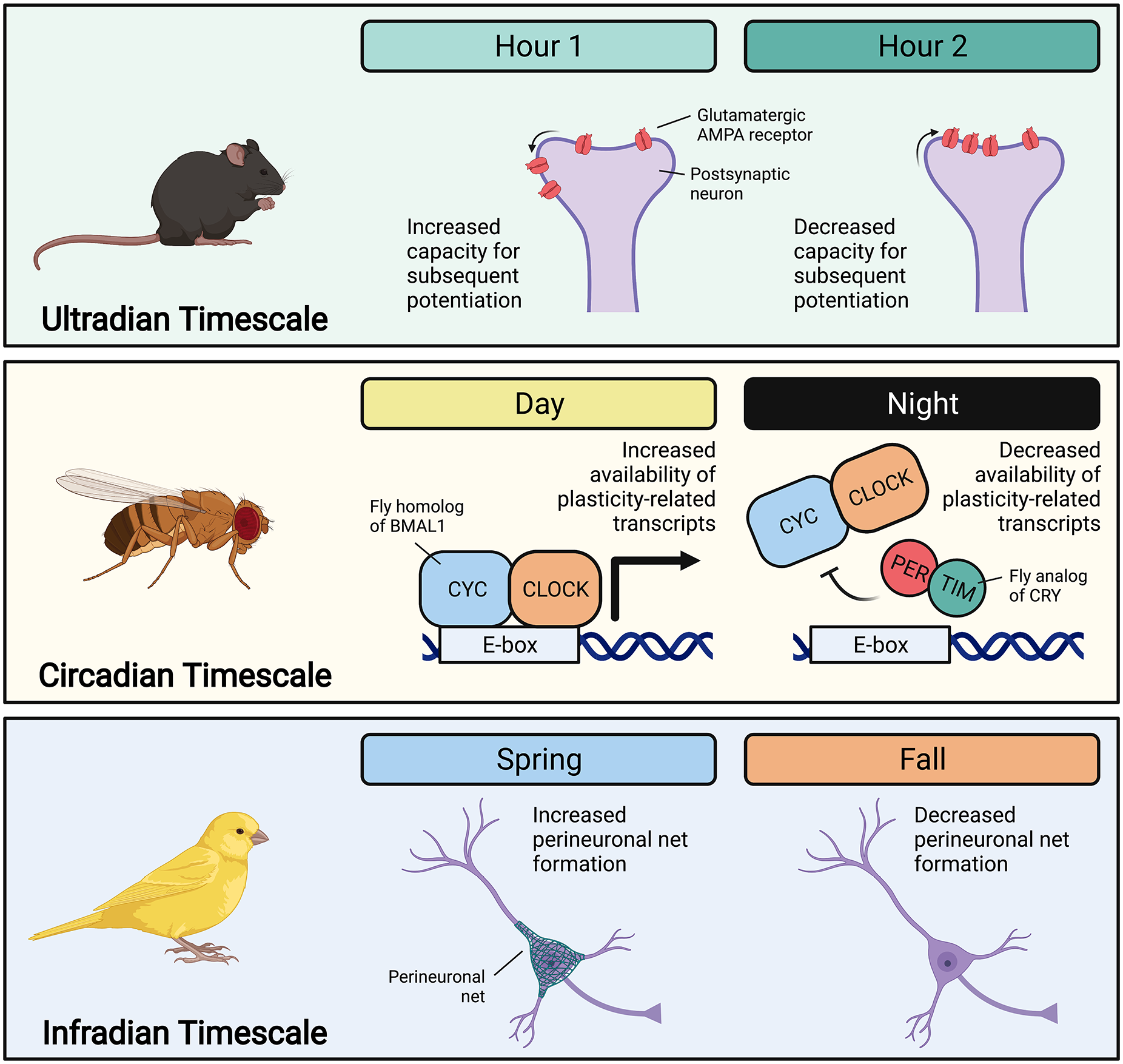
Examples of iterative metaplasticity across timescales. On the ultradian timescale, work in mouse hippocampal neurons suggests glucocorticoid-dependent oscillations in AMPA receptor membrane localization that create oscillations in the capacity for long-term potentiation (Sarabdjitsingh et al., 2014). On the circadian timescale, studies of Drosophila demonstrate clock-regulated rhythms in the transcription of plasticity-related genes (Claridge-Chang et al., 2001; Cirelli et al., 2004). On the infradian timescale, canary research illustrates seasonal differences in the formation of perineuronal nets, which stabilize synapses (Cornez, Collignon, et al., 2020). Across timescale and organism, iterations in such basal neural features generate iterations in the capacity for activity-dependent plasticity and memory. Abbreviations: BMAL1 = brain and muscle ARNT-like protein-1; CLOCK = circadian locomotor output cycles kaput; PER = period; CRY = cryptochrome; CYC = cycle; TIM = timeless
Molecular clocks can also influence neuroplastic capacity through non-transcriptional means. For instance, the molecular clock protein PER1 acts as a nuclear chaperone for ribosomal s6 kinase (RSK) in the mouse hippocampus, modulating long-term potentiation and hippocampus-dependent memory (Rawashdeh et al., 2014; Rawashdeh et al., 2016). In addition, CRY proteins can bind directly to Gs proteins, and this binding appears to be a mechanism by which the presence of CRY proteins in the mouse nucleus accumbens regulates susceptibility to stress-induced impairments in escape learning (Porcu et al., 2020). Moreover, the quantity of many transcripts oscillates in synapses but not the cell cytoplasm, suggesting that the molecular clock influences transcript trafficking from the nucleus to the synapse (Noya et al., 2019). These findings are just a few among now abundant examples of how the direct regulation of neuroplasticity mechanisms by molecular clocks contributes to circadian metaplasticity (Krishnan and Lyons, 2015; Rawashdeh et al., 2018; Hartsock and Spencer, 2020).
SCN-orchestrated intercellular signals serve primarily to coordinate extra-SCN molecular clocks, but these signals can also generate circadian metaplasticity through direct actions on neuroplasticity processes. For instance, the secretion of glucocorticoid hormones (e.g. cortisol in humans, corticosterone in rodents) exhibits a circadian rhythm that is SCN-dependent (Moore and Eichler, 1972). Although glucocorticoids are believed to play an important role in circadian entrainment of extra-SCN molecular clocks in mammals (Oster et al., 2017; Spencer et al., 2018), they can also exert direct effects on mechanisms of neuroplasticity. Accordingly, the circadian peak in corticosterone secretion has been shown to enhance learning-induced spine formation in the murine motor cortex (Liston et al., 2013). In that study, corticosterone injection immediately after training was sufficient to increase spine formation at both the circadian peak and trough of endogenous glucocorticoid release, suggesting that the effect was mediated directly by glucocorticoids rather than through glucocorticoid entrainment of cortical molecular clocks. In addition, constant high corticosterone was disruptive to new spine formation, pointing to a need for circadian peaks and troughs in corticosterone secretion for optimal spine turnover. Furthermore, corticosterone injection enhanced new spine formation only when learning occurred, indicating that glucocorticoids did not directly alter spines but rather the capacity for learning to do so. Taken together, these findings suggest that circadian rhythms in glucocorticoid release can generate circadian rhythms in neuroplastic capacity.
Sleep is a clear example of how circadian-regulated processes can influence neuroplastic capacity. The timing of sleep is organized in part by the circadian system (Borbély et al., 2016), but sleep and the circadian system have discriminable effects on the regulation of neuroplasticity. For instance, circadian rhythms in messenger RNA (mRNA) abundance have been identified for 67% of synaptic mRNAs (Noya et al., 2019). Of those mRNAs, 93% exhibit circadian oscillation only in synaptic elements. Although 75% of the proteins associated with cycling transcripts are also rhythmic, sleep deprivation can eliminate oscillations in protein abundance. These data suggest that sleep may regulate the translation of synaptic mRNAs into proteins (Noya et al., 2019). Hence, circadian rhythms in sleep timing can contribute to circadian rhythms in neuroplastic capacity through sleep-dependent mechanisms.
Collectively, such findings illustrate multiple and likely interacting mechanisms by which the endogenous circadian system can produce circadian metaplasticity. In some cases, however, daily rhythms in neuroplastic capacity may emerge also in response to daily environmental signals independent of the circadian system. Such daily rhythms are described as diurnal rhythms to distinguish them from endogenously generated circadian rhythms. Retinal ganglion cells, including those carrying solar entrainment information to the SCN, also project to many other brain areas (Hattar et al., 2006; LeGates et al., 2014). In turn, significantly shortened light cycles impair hippocampal long-term potentiation and hippocampus-dependent memory without disrupting SCN clock gene expression (LeGates et al., 2012; Fernandez et al., 2018). The most straightforward explanation is that periodic light exposure can generate time-of-day differences in neuroplasticity processes not only through its effects on the circadian system but also through non-circadian modulatory influences (LeGates et al., 2014). Continued studies of metaplasticity on the daily timescale must identify whether iterations are generated through endogenous versus environmental cyclic signals.
Iterative Metaplasticity On A Shorter-Than-Daily Timescale: Ultradian Metaplasticity
Ultradian metaplasticity is a form of iterative metaplasticity repeating on a timescale shorter than 1 day (Figure 1, upper panel). In many mammals, basal glucocorticoid secretion exhibits not only a circadian rhythm but also a superimposed ultradian oscillation with a period of approximately 60 min (Lightman and Conway-Campbell, 2010; Spencer and Deak, 2017). The ultradian oscillation results primarily from glucocorticoid-mediated negative feedback on adrenocorticotrophic hormone release in the anterior pituitary gland (J. J. Walker et al., 2010). Pulsatile glucocorticoid administration is sufficient to generate pulsatile gene expression in the rat hippocampus (Conway-Campbell et al., 2010). In mouse hippocampal neurons, consecutive pulses of glucocorticoid have been shown to generate a decrease and rebound in the capacity for long-term potentiation, an effect that appears to depend on the trafficking of glutamatergic AMPA receptors in and out of the postsynaptic membrane (Sarabdjitsingh et al., 2014). This latter finding suggests that consecutive hourly pulses of glucocorticoid may toggle neurons back and forth between states of increased and decreased neuroplastic capacity (Figure 2, upper panel). Ultradian pulses in glucocorticoid secretion have been hypothesized to keep the glucocorticoid system dynamically engaged throughout the day (Lightman and Conway-Campbell, 2010). If true, downstream effects on neuroplasticity may represent either an incidental consequence or, conceivably, a mechanism serving the parallel function of maintaining neuroplastic capacity.
Ultradian rhythms in the activity of ventral tegmental area dopamine neurons may represent another mechanism of ultradian metaplasticity. Dopamine projections from the ventral tegmental area to the limbic forebrain play a key role in the neuroplasticity processes underlying motivated behavior (Baik, 2013; Morales and Margolis, 2017). For instance, dopamine receptor activation can modulate the activity of multiple intracellular cascades involved in neuroplasticity, including extracellular signal–regulated kinase (ERK) (Choi et al., 1999; Chen et al., 2004), protein kinase A (PKA) (Bateup et al., 2008), and calcium/calmodulin-dependent protein kinase II (CaMKII) (Anderson et al., 2008). In the mouse striatum, which receives strong innervation from the ventral tegmental area, extracellular dopamine concentrations fluctuate with a period of approximately 4 h (Blum et al., 2014). These ultradian dopamine rhythms correspond with ultradian rhythms in locomotion, which are disrupted by manipulations of dopamine (Blum et al., 2014). Ultradian rhythms in dopamine release have not yet been connected to ultradian rhythms in neuroplasticity, but the ability of dopamine to modulate numerous neuroplasticity processes suggests that dopamine is likely to mediate ultradian rhythms in neuroplastic capacity, especially for motivated behavior.
Circatidal (approximately one tidal cycle) rhythms, which exhibit a period of approximately 12.4 h, may represent another set of mechanisms for ultradian metaplasticity. Circatidal rhythms have been observed in multiple aquatic species, including crabs (Reid and Naylor, 1990), hooded shrimp (Akiyama, 2004), mussels (Zaldibar et al., 2004), leeches (L. Zhang et al., 2013), and mangrove crickets (Takekata et al., 2012; Satoh and Terai, 2019). To our knowledge, no studies to date have directly examined circatidal contributions to ultradian metaplasticity. One investigation, however, has demonstrated that transcripts relevant for cellular metabolism and endoplasmic reticulum function exhibit circatidal expression rhythms in the mangrove cricket head (Satoh and Terai, 2019), indicating the potential for circatidal influences on molecular processes supporting neuroplasticity. Circatidal rhythms in the expression of genes relevant for neuroplasticity have also been shown in the central nervous system of the horseshoe crab (Simpson et al., 2017). In addition, parallel and in some cases overlapping mechanisms appear to mediate circadian and circatidal rhythms (L. Zhang et al., 2013; O’Neill et al., 2015; Tran et al., 2020), suggesting that certain molecular processes underlying circadian metaplasticity may support circatidal rhythms in aquatic organisms. For these reasons, we expect that the characterization of circatidal contributions to ultradian metaplasticity will prove fruitful in future investigations.
Metaplasticity on the ultradian timescale may be generated also by periodic environmental signals. For instance, metabolic signals induced after food intake may regulate neuroplasticity processes (Mandal et al., 2018). Ghrelin, for example, is a gastrointestinal peptide regulated by food intake. Ghrelin can cross the blood-brain barrier and enhance inhibitory avoidance memory retention in a dose-dependent manner (Carlini et al., 2004; Rhea et al., 2018). Similarly, glucagon-like peptide-1 (GLP-1) is released after feeding, crosses the blood-brain barrier, and improves passive avoidance learning (During et al., 2003; Marathe et al., 2013). Thus, ultradian rhythms in food availability (e.g., breakfast-lunch-dinner schedules on a college campus) may be sufficient to generate rhythms in neuroplastic capacity. When periodic environmental signals give rise to oscillations in neuroplastic capacity, the oscillations themselves may or may not serve an important purpose in overall organism function. Such oscillations could simply reflect the ability of an organism to modulate neuroplastic capacity as environmental events demand. In any event, as for circadian metaplasticity, future studies on ultradian metaplasticity will need to discriminate whether oscillations are generated by endogenous versus environmental signals.
Iterative Metaplasticity On A Longer-Than-Daily Timescale: Infradian Metaplasticity
Infradian metaplasticity is a form of iterative metaplasticity repeating on a longer-than-daily timescale (Figure 1, lower panel). Circalunar (approximately one lunar cycle) rhythms, including moon-phase-dependent fluctuations in sleep quality (Casiraghi et al., 2021) and the human ovulatory cycle (Helfrich-Förster et al., 2021), may contribute to infradian metaplasticity. We have already discussed an example of how sleep may directly influence neuroplastic capacity (Noya et al., 2019). It follows that circalunar rhythms in sleep quality could potentially generate circalunar rhythms in neuroplastic capacity, but we have no knowledge of studies that have explicitly demonstrated this link.
In contrast, effects of the ovulatory cycle on neuroplasticity have been well documented. In rodents, which exhibit a 4-to-5-day ovulatory cycle similar in many respects to the human ovulatory cycle (Becker et al., 2005), ovarian steroid hormones directly influence processes such as neuromodulatory neurotransmission (Barth et al., 2015), cytoskeletal remodeling (Parducz et al., 2006; Hansberg-Pastor et al., 2015), and transcriptional regulation (Marino et al., 2006). Moreover, this hormonal modulation of neuroplasticity results in a fluctuation in hippocampal CA1 dendritic spine density throughout the ovulatory cycle (Woolley et al., 1990). In humans, performance in implicit memory tasks exhibits a fluctuation across the ovulatory cycle in a task- and estrogen-dependent manner (Maki et al., 2002). Similarly, declarative and motor memory consolidation have been shown to correlate with levels of estrogen and progesterone, respectively (Genzel et al., 2012). These data suggest a steroid-dependent fluctuation in basal neural physiology that sets the stage for circalunar rhythms in activity-dependent plasticity.
Circannual (approximately 1 year) rhythms are also likely to support infradian cycles in neuroplastic capacity. Circannual variations in songbird song learning have received a great deal of attention. For instance, brain nuclei important for song learning grow in volume during breeding season in a variety of species (Tramontin and Brenowitz, 2000; Van Meir et al., 2006; Surbhi et al., 2016), owing in part to neurogenesis (Nottebohm, 1989). In addition, the maintenance of new neural cells is enhanced at the beginning of the breeding season (Larson et al., 2019). In adult male canaries, seasonal elevations in song production have also been shown to correlate with increases in the formation of perineuronal nets, which stabilize synapses (Figure 2, lower panel) (Cornez, Collignon, et al., 2020). Some of these changes depend on testosterone (Nottebohm, 1980), the release of which fluctuates in a seasonal fashion in songbirds (Marler et al., 1988; Schlinger and Brenowitz, 2002). In captive female starlings, the volume of song control nuclei and the rate of song learning are elevated after testosterone implantation and return to baseline after implant removal (Orije et al., 2020). Testosterone also augments perineuronal net formation and song production in male canaries (Cornez, Shevchouk, et al., 2020). Together, these studies point to numerous mechanisms underlying circannual modulation of neuroplastic capacity in avian species.
In mammals, provocative evidence suggests effects of photoperiod on neurotransmission. In the mouse SCN, where gamma-aminobutyric acid (GABA) can exert both inhibitory and excitatory effects (Choi et al., 1999; De Jeu and Pennartz, 2002; Irwin and Allen, 2009), the ratio of inhibitory to excitatory GABAergic synapses increases with short photoperiods and decreases with long photoperiods (Farajnia et al., 2014). In rats, in hypothalamic regions outside the SCN, some neurons can even switch between dopamine and somatostatin expression based on the length of the photoperiod (Dulcis et al., 2013). One report in hamsters has shown that differences in day length produce differences in SCN spine density (Mendoza-Viveros et al., 2017). Similarly, a report in mice has shown that long days enhance novel and spatial object recognition (Dellapolla et al., 2017). Another study illustrates that short photoperiods disrupt hippocampal spine density and spatial memory in mice (Pyter et al., 2005). These findings suggest several mechanisms of metaplasticity on the infradian timescale in mammals.
Endogenous pathways supporting circannual timing are not yet well understood. In mammals, individual SCN neurons tend to increase their spontaneous firing activity for only a few hours each day (Schaap et al., 2003; Brown et al., 2006). In nocturnal and diurnal species alike, SCN neurons tend to be active during the daytime, but individual SCN neurons exhibit different phases in their peak firing (VanderLeest et al., 2007). During short photoperiods, as in winter, the phases of peak firing tend to be closely aligned; the opposite is true during long photoperiods. Consequently, short days give way to a short window of spontaneous SCN activity at the tissue level, and long days give way to a long window (VanderLeest et al., 2007; Coomans et al., 2015). The SCN is therefore well positioned to coordinate circannual timing information for the rest of the body. Pineal melatonin may represent one important signaling molecule used by the SCN for this purpose. The release of melatonin is controlled by a well-characterized polysynaptic relay from the SCN to the pineal gland (Larsen et al., 1998; Møller and Baeres, 2002; Coomans et al., 2015). In optimal conditions, melatonin is secreted for the duration of the daily dark period (Turek and Gillette, 2004). A melatonin sensor in the pituitary gland modulates the neuroendocrine output of the hypothalamus in a manner dependent on the length of the photoperiod (Wood and Loudon, 2014; Wood et al., 2020). How circannual photoentrainment information is then communicated throughout the rest of the brain remains an area of active inquiry.
Future studies will need to assess whether cycles in neuroplastic capacity on the infradian timescale emerge from endogenous versus environmental signals. In many geographical locations, for example, the passage of seasons brings about changes in ambient temperature, photoperiod, and weather patterns. These environmental fluctuations may function as both circannual entrainment signals and direct causes of metaplasticity, as is observed with environmental cues on the circadian timescale. Although mechanisms of infradian metaplasticity have yet to be fully elucidated, these findings indicate the potential for circalunar and circannual rhythms in neuroplastic capacity, highlighting an important avenue for continued investigation.
The Functions Of Iterative Metaplasticity
The circadian system is widely regarded as a mechanism for predictive homeostasis, the physiological ability to anticipate and adapt to periodic environmental challenges (Moore-Ede, 1986). We and others view circadian metaplasticity to serve this function by organizing learning processes into adaptive temporal domains (Gerstner, 2012; Krishnan and Lyons, 2015; Rawashdeh et al., 2018; Hartsock and Spencer, 2020). For instance, enhanced fear learning and recall during the daily sleep period may promote safety behaviors when a prey animal would otherwise be quite vulnerable to danger (Chaudhury and Colwell, 2002; Eckel-Mahan et al., 2008). Non-circadian forms of iterative metaplasticity that are also endogenously generated may be viewed to niche memory function on non-daily timescales. That songbirds are best at song learning during mating season is a fitting illustration (Van Meir et al., 2006; Surbhi et al., 2016; Cornez, Collignon, et al., 2020). Whereas endogenously generated forms of iterative metaplasticity support predictive homeostasis, environmentally generated forms of iterative metaplasticity may instead support reactionary homeostasis, the better known type of homeostasis (Moore-Ede, 1986). Thus, endogenously generated iterations in neuroplastic capacity may help an organism to predict environmental change, whereas iterations generated only in response to environmental signals may enable the organism to adjust its behavior in real time as conditions demand.
But conceiving of iterative metaplasticity strictly as a mechanism for homeostasis may miss something more profound about its functional importance. Liston and colleagues (2013) have shown that the daily peak in glucocorticoid concentrations is necessary for spine formation, while the daily trough in glucocorticoid concentrations is necessary for spine stabilization. Similarly, another report demonstrates that, whereas hippocampal adenylyl cyclase activity is necessary for optimal recall of contextual fear in mice, increasing hippocampal adenylyl cyclase activity during its daily trough impairs recall (Eckel-Mahan et al., 2008). A third study has shown that constitutively high expression of the normally oscillatory transcript microRNA-132 in the mouse forebrain can disrupt expression of the hippocampal plasticity proteins MeCP2 and SIRT1 as well as hippocampus-dependent memory (Aten et al., 2018). These data suggest that neuroplastic capacity that is oscillatory may better serve memory function than neuroplastic capacity that is always high. It is also noteworthy that iterative metaplasticity is observed virtually every time it is measured, which is especially clear in the growing body of work on circadian metaplasticity (Krishnan and Lyons, 2015; Rawashdeh et al., 2018; Hartsock and Spencer, 2020). Together, these findings lead us to consider the possibility that iterative metaplasticity may be necessary for maintaining neuroplastic capacity more generally.
We have discussed previously that sleep might be seen as one mechanism by which the circadian system regulates neuroplasticity (Hartsock and Spencer, 2020). From this perspective, sleep can help us to think about why iterations may be essential. Sleep maintains neuroplastic capacity ultimately by counteracting processes that occur during wakefulness. For instance, wakefulness tends to promote long-term potentiation, and sleep tends to promote long-term depression (Tononi and Cirelli, 2014). Wakefulness tends to promote neurotoxic metabolite accumulation, and sleep tends to promote neurotoxic metabolite clearance (Xie et al., 2013). Wakefulness tends to promote synaptic transmission, and sleep tends to promote metabolic recovery (Noya et al., 2019). This pattern is even seen in Drosophila, with dopamine neurons promoting active forgetting of olfactory memories during wakefulness but being suppressed during sleep (Berry et al., 2015). Sleep-driven iterations in neuroplastic capacity support optimal memory function in a variety of memory tasks (M. P. Walker and Stickgold, 2004; Rasch and Born, 2013; Tononi and Cirelli, 2014). Other forms of iterative metaplasticity may work the same way, maintaining memory function by toggling neural cells between states of cellular expenditure (increased neuroplastic capacity) and subsequent restoration (decreased neuroplastic capacity).
Further exploration of this hypothesis will require convergent evidence from many research areas. We will need to clarify exactly how basal neural parameters, such as membrane potential, kinase activity, and DNA methylation status, help to set the stage for activity-dependent neuroplasticity (Josselyn and Tonegawa, 2020). We can then begin to understand the functional impact of oscillations in these parameters. To this end, we will be challenged to develop techniques that can precisely alter oscillatory function with minimal off-target disturbance, as Aten and colleagues (2018) have achieved with a quadruple transgenic mouse in which the oscillatory expression profile of microRNA-132 is replaced by a constitutive expression profile. In addition, we will need to understand how oscillations in neuroplasticity may differ among species, with a particular focus on how environmental factors such as sunlight or tides may produce particular oscillatory signatures. Through these pursuits, we may gain a deeper appreciation of how iterative metaplasticity across timescales can modulate the capacity of an organism to learn and remember at any given moment in time.
Footnotes
Acknowledgements
This work was supported by a National Science Foundation predoctoral fellowship to M.J.H. (grant number DGE144083), a National Institutes of Health predoctoral fellowship to H.K.S. (grant number T32HL149646), and a National Institutes of Health research program grant to R.L.S. (grant number MH115947). Figure 2 was created with ![]() .
.
Conflict Of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
