Abstract
Emerging evidence has demonstrated that long noncoding RNAs (lncRNAs) play critical roles in the epigenetic and transcriptional regulation of mammalian circadian systems. Circadian rhythmicity regulates many aspects of our immune system, and perturbation of the circadian clock can augment the inflammatory response. However, knowledge of the precise functions of lncRNAs in the regulation of immune functions within the circadian system is relatively limited. In this study, differentially expressed lncRNAs induced by Clock knockdown were screened via mRNA/lncRNA microarray and bioinformatic prediction analysis. We identified a Clock-regulated lncRNA, AK028245, which was correlated with the activation of the immune response. The expression levels of AK028245 were decreased in the spleen of immunosuppressed mice and elevated in immune-activated mice treated with lipopolysaccharide (LPS). Further, Clock knockdown decreased the expression of OTUD7B and A20, 2 early immune response factors acting on the NF-κB signaling pathway. Interestingly, inhibition of AK028245 increased their expression, mitigating the effects of Clock knockdown. In addition, inhibition of AK028245 downregulated the expression of tumor necrosis factor–α and interleukin-6 in the late stages of LPS stimulation and the expression of interferon-γ and Cxcl12 in the peak stages. We conclude that this newly identified lncRNA plays a role in the crosstalk between Clock and immune response regulators, likely resulting in a proinflammatory response targeting OTUD7B and A20. The lncRNA AK028245 has revealed a new mechanism of the immune response and provided new targets for the treatment of immune disorders.
The regular 24-h environmental cycles generated by the Earth’s rotation have led to the evolution of daily circadian rhythms in almost all life forms on Earth (Bhadra et al., 2017). In mammals and other complex organisms, virtually all aspects of physiology are regulated by internal circadian rhythms, including behavior and locomotor activity, sleep-wake cycles, body temperature, endocrine systems, and metabolic and immune functions (Scheiermann et al., 2018). In mammals, the circadian clock is powered by the rhythmic activity of a number of clock proteins (Early and Curtis, 2016). At the center of the machinery lies the heterodimeric partnership of 2 transcription factors, Brain Muscle Arnt-Like Protein 1 (BMAL1) and Clock Locomotor Output Kaput (CLOCK). In their cooperative partnership, these proteins can bind to E-box sites throughout the genome to stimulate gene expression (Ripperger and Schibler, 2006). CLOCK exhibits profound control over the genome, with at least 10% of all transcripts being regulated in a circadian manner in most tissue types (Panda et al., 2002; Storch et al., 2002; Early and Curtis, 2016). CLOCK disturbances may therefore lead to a variety of diseases, as demonstrated by several studies previously (Roenneberg and Merrow, 2016; Hu et al., 2018; Kuhlman et al., 2018). The mechanisms by which circadian disorders are linked to different disease etiologies is not fully understood. It is worth noting that changes in immune function are common features of many diseases; thus, explorations of circadian rhythm biology and immunology may provide critical insights. Like most organ systems in the body, several components of the immune system appear to be regulated in a daily manner (Scheiermann et al., 2018). However, the cellular-level processes and underlying molecular mechanisms of this daily regulation are poorly understood (Nguyen et al., 2013).
Long noncoding RNAs (lncRNAs), a subclass of ncRNAs, are most commonly defined as transcripts (greater than 200 nucleotides) structurally resembling mRNAs, which have no protein-coding capacity (Mercer et al., 2009). Emerging research has found that lncRNAs play critical roles in biological processes such as transcriptional regulation, epigenetic modification, cell differentiation, as well as in some diseases (Rapicavoli et al., 2013; Zhan et al., 2016; Zhang et al., 2017; Xue et al., 2018). LncRNAs have been identified in virtually all immune cell types, although their precise functions remain an emerging topic in immune system research (Atianand and Fitzgerald, 2014). For example, linc00513 has been previously identified as a positive regulator of the type I interferon pathway, conferring functional susceptibility to systemic lupus erythematosus loci in systemic lupus erythematosus patients (Xue et al., 2018). Similarly, the lncRNA CRNDE has been shown to positively regulate inflammatory cytokines levels, subsequently activating the NF-κB and JAK/STAT pathways (Zhu-Ge et al., 2018). Thus, lncRNA, as a key regulatory factor, provides an additional regulatory layer for the regulatory network of immune genes.
Current research has outlined that the regulatory power of circadian clock systems on physiological processes is likely achieved by multisignal pathways and multilayer network regulation (Du et al., 2019). It is thereby necessary to explore the Clock-regulated immune response at the level of lncRNAs. Deubiquitinases are vital regulators of the NF-κB signaling pathway, functioning as molecular brakes of NF-κB signaling, helping to maintain immune homeostasis. Among these, the homologous deubiquitinases A20 and OTUD7B act on the canonical and noncannonical NF-κB pathway, respectively. In a previous study, Otud7b demonstrated Clock-controlled gene activity (Hou et al., 2017); however, the mechanisms remained unclear. In the present study, we set out to identify differentially expressed lncRNAs in Clock-knockdown cells, screening specifically for lncRNAs demonstrating Otud7b-related expression as determined via bioinformatic prediction. Further experiments were conducted to verify whether the expression of these candidate lncRNAs was sensitive to abnormal immunological conditions. This study identified the target, Clock-controlled lncRNA-AK028245, which demonstrated inhibitory functions on OTUD7B and A20 expression. Specifically, lncRNA-AK028245 was found to mitigate the effects of CLOCK on these deubiquitinases. Finally, the influence of AK028245 on the lipopolysaccharide (LPS)–induced immune response was examined to confirm the proinflammatory effect of AK028245.
Materials And Methods
Experimental Animals
Male ICR mice (6-8 weeks old, 20-22 g) were obtained from Dashuo Experimental Animals Co. Ltd. (Chengdu, China). These mice were housed in a rodent facility at 25 °C with a 12 h:12 h light-dark cycle for acclimatization. After being adapted to the experimental environment for 1 week, mice were randomly divided into groups of 5 mice each.
Immunosuppression Model Induced by Cyclophosphamide
The cyclophosphamide (CPA)–treated experiment had 2 groups (5 mice each). One group of healthy mice were used as a normal control group (NC). These mice were treated once daily with saline solution by intraperitoneal injection (i.p.) for 5 days at zeitgeber time (ZT) 04. The other group (CPA) were given 100 mg/kg/day CPA (i.p.) daily for 5 days. Body weights were recorded before the first i.p. administration and 24 h after the last dose. Animals were subsequently terminated, and the spleen was removed and weighed.
Acute Inflammation Model Induced by LPS
The LPS-stimulation experiment consisted of 3 groups (5 mice each). LPS was dissolved in 0.154 mol/L saline solution, and mice were injected i.p. with saline solution (naive) or with 1 mg/kg LPS in a 0.1-mL volume at ZT 04. One LPS subgroup (LPS-treated 6-h group) was terminated 6 h after administration, and the second LPS subgroup (LPS-treated 12-h group) was terminated 12 h after LPS administration. The naive group was terminated 12 h after saline administration (at ZT 16). The spleen was removed from each animal after termination. These animal procedures were approved by the Sichuan University Ethical Committee of Laboratory Animal Investigations. Experiments conformed to the Guide for the Care and Use of Laboratory Animals published by the National Institutes of Health (NIH publication No. 85-23, revised 1996).
Cell Culture
MS-1, RAW264.7, MHS, and NIH-3T3 cells were obtained from the American Type Culture Collection cell bank (Manassas, VA, USA). They were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Hyclone; GE Healthcare Life Sciences, Logan, UT, USA), containing 10% inactivated fetal bovine serum (Gibco; Thermo Fisher Scientific, Waltham, MA, USA). Cells were maintained in cell incubators with 5% CO2 at a constant temperature of 37 °C.
Serum Shock Procedures
In the circadian serum shock experiments, cells were divided into 8 groups, each designated by the indicated time point of total mRNA extraction from cell lysates (Balsalobre et al., 1998). All experiments were performed in triplicate. Petri dishes measuring 5 × 105 cells/10 cm were used to plate cells 1 to 3 days prior to the experiment. Under these conditions, cells reached confluence after about 2 days, after which the medium was exchanged with serum-rich medium (DMEM, supplemented with 50% equinum serum [GIBCO]; designated as time point ST0). Two hours after ST0, the cell medium was replaced by serum-free DMEM (ST2). One hour later (ST3), Petri dishes were washed twice with ice-cold phosphate-buffered saline, and cells were collected for RNA extraction. In this manner, cells were collected every 3 h until cycle ST24 was reached (endpoints ST3-ST24, in equidistant intervals of 3 h).
mRNA and lncRNA Microarray
To investigate the expression of mRNAs and lncRNAs in relation to CLOCK, RNA from Clock-knockdown MS-1 cells and negative control cells were used for the mRNA and lncRNA microarray. The microarray was performed by RiboBio Co. Ltd. The fold change of each differentially expressed mRNA and lncRNA was obtained by log2 (normalized intensity of treat/normalized intensity of control). The quantile normalization method was used, and an average of repeated data was taken from each sample. The p values were calculated by the analysis of variance (ANOVA) method. Standard selection criteria used to identify differentially expressed lncRNAs/mRNAs were established at p ≤ 0.05 and |FoldChange| ≥ 1.5. The p values were calculated by the ANOVA method. If p values were less than or equal to the threshold (p ≤ 0.05), differential expression was presumed between the control and test sample. The software LncTAR was used for all lncRNA predictions.
Transfection of the Lentivirus-AK028245 and lncRNA Small Silencer
The transfection of lv-shRNA-Clock has been previously described (Hou et al., 2017). Briefly, AK028245 full-length sequence (https://www.ncbi.nlm.nih.gov/nuccore/AK028245) was synthesized and inserted into the gene transfer vector pLOV-EF1a-PuroR-CMV (H8888; Obio Technology Corp., Ltd., Shanghai, China). The titer of the virus was 1.34 × 109 TU/mL. Lv-Puro-AK028245 and lv-Puro-NC viruses were thawed and dissolved on ice. After the medium was replaced with fresh medium containing polybrene, the virus solutions were added into a 24-well plate configuration containing cells at a volume of 30 µL/well, according to the manufacturer’s instructions. The cells were cultured at a constant temperature of 37 °C and a saturation humidity of 5% CO2 for 24 h. Subsequently, the cells were transferred to cell culture bottles, where they were screened for successful transfection using puromycin (J&K Scientific Ltd., Beijing, China) at a concentration of 4 µg/mL (MS-1 cells and NIH-3T3 cells) and 2 µg/mL (RAW264.7 cells), according to the manufacturer’s instruction. The cycle length for transfection screening was 1 week. The overexpression of AK028245 was confirmed using reverse transcriptase polymerase chain reaction (RT-PCR) 48 h after the transfection.
LncRNA Smart Silencer, synthesized by RiboBio (Guangzhou, China), was used to knock down the expression of AK028245. AK028245 Smart Silencer solution contained a mixture of 3 small interfering RNAs (siRNAs) and 3 antisense oligonucleotides. The negative control (NC) did not contain domains homologous to humans, mice, or rats. LncRNA Smart Silencer transfection was performed with riboFECT CP Reagent (RiboBio) according to the manufacturer’s instructions. Approximately 5 × 103 cells were plated into each well of a 12-well plate at least 24 h before transfection to achieve 30% to 50% confluency. Cells were collected 24 h after transfection for RNA isolation.
Isolation of RNA and Quantitative RT-PCR Analysis
Total RNA was isolated with the Cell/Animal Total RNA Isolation Kit (Foregene, Chengdu, China) according to the manufacturer’s instructions. Samples were subjected to on-column deoxyribonuclease digestion. After quantification of RNA concentrations using NanoDrop 2000 (Thermo Fisher Scientific), RNAs were reverse-transcribed into cDNAs using the 5× All-in-One RT Master Mix (Vazyme, China). Quantitative PCR was performed using the AceQ qPCR SYBR Green Master Mix (Vazyme, China). The lncRNA primer assays were purchased from RiboBio, and the remaining primers were obtained from Sangon (Shanghai, China). The sequences were designed as follows: Clock: 5′ ATGGTGTTTACCGTAAGCTGTAG 3′ and 3′ CTCGCGTTACCAGGAAGCAT 5′, Gapdh: 5′ GCCTTCCGTGTTCCTACCC 3′ and 3′ GCCCTCAGATGCCTGCTTCA 5′, Otud7b: 5′TGTCCTGTCGGATTTTGTTCG 3′ and 3′ GTGGGGATAGGTTCCCAGC 5′, A20: 5′GAACAGCGATCAGGCCAGG 3′ and 3′ GGACAGTTGGGTGTCTCACATT 5′. Change in expression level was calculated by quantitative analysis (in triplicate) using the comparative cycle threshold method. The raw data of each target gene were normalized to Gapdh.
Western Blot Analysis
Spleen tissues and both treated and untreated cell lines were lysed with radioimmunoprecipitation assay buffer containing 1 mM phenylmethylsulfonyl fluoride, Halt protease inhibitor cocktail, and Halt phosphatase inhibitor (Beyotime Institute of Biotechnology, Haimen, China). The protein lysates underwent centrifugation at 14,000 × g for 20 min at 4 °C. Equal amounts of protein (0.5 mg/mL concentration; control and experimental) were loaded in each experiment. Protein separation was achieved by electrophoresis on 10% to 12% sodium dodecyl sulfate–polyacrylamide gel electrophoresis, followed by transfer to polyvinylidene difluoride membranes (Millipore, Billerica, MA, USA). After blocking with 5% skim milk at room temperature for 1 h, the membranes were incubated with primary antibodies at 4 °C overnight. The primary antibodies used included monoclonal anti-GAPDH (1:5000 dilution; cat No. EM1101; HuaBio Inc., Cambridge, MA, USA), polyclonal anti-CLOCK (1:10,000 dilution; cat No. ab3517; Abcam, Cambridge, MA, USA), polyclonal anti-OTUD7B (1:500 dilution; cat No. 16605-1-AP; ProteinTech Group, Inc., Chicago, IL, USA), and polyclonal anti-A20 (1:500 dilution; cat No. 23456-1-AP; ProteinTech Group, Inc.). The following day, membranes were incubated with a horseradish peroxidase–conjugated anti-rabbit IgG or anti-mouse IgG secondary antibody (1:2000 dilution; cat No. ZB-2301&ZB-2305; ZSGB-Bio Group, Inc., Beijing, China) at 37 °C for 2 h. Membranes were developed using an enhanced chemiluminescence reagent provided in the EasySee Western Blot kit (Beijing Transgen Biotech Co. Ltd., Beijing, China). Imaging was performed with a gel imager (GE Healthcare Life Sciences, Little Chalfont, UK). The images were further analyzed with ImageJ software (National Institutes of Health, Bethesda, MD, USA) to assess the differences in protein expression via protein band optical density. GAPDH was used as an internal control protein in all assays.
Statistical Analyses
All experiments were independently repeated at least 3 times with each experiment performed in triplicate. Statistical analyses of the data were performed using the GraphPad Prism 6 software (GraphPad Software, La Jolla, CA, USA), and 1-way ANOVA was used to assess differences between groups. Bonferroni t tests were used for posttests. All data were reported as the mean ± SD, and p < 0.05 was considered statistically significant for all comparisons.
Results
Identification of Clock-responsive lncRNAs Targeted to
Otud7b
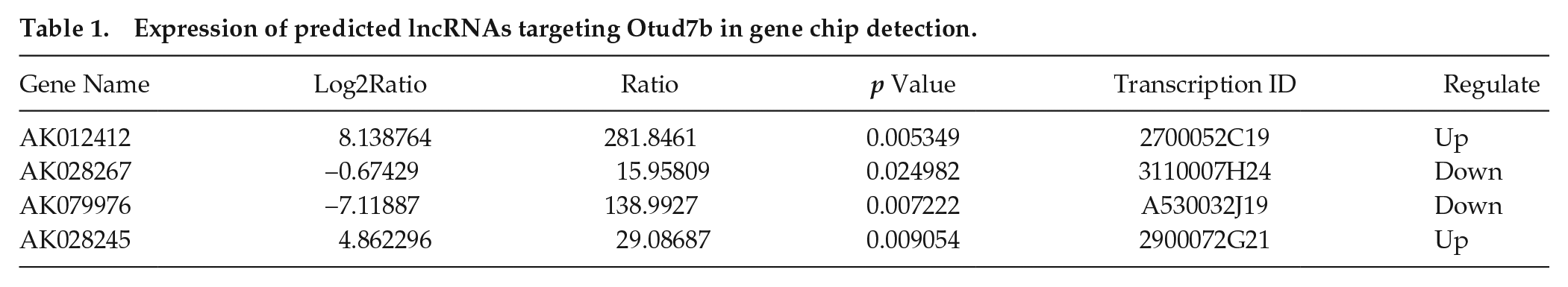
CLOCK is a core factor of the circadian system, acting as a transcription factor that forms a heterodimer with BMAL1. MS-1 cells are mouse islet endothelial cells that, similar to vascular endothelial cells, show significant circadian rhythm characteristics (Gao et al., 2014). Acting as structural cells, MS-1 cells are commonly used to study the interaction between circadian rhythm regulation and basic physiological activities (Gao et al., 2014). Here, changes in the expression of mRNAs and lncRNAs expressed in MS-1 cells were evaluated after Clock knockdown. The expression level of mRNAs and lncRNAs in Clock-knockdown MS-1 cells and negative control MS-1 cells was measured using microarray (Fig. 1A). Standard selection criteria were used to identify differentially expressed lncRNAs/mRNAs (p ≤ 0.05 and |FoldChange| ≥ 1.5). Microarray analysis revealed 2150 altered lncRNAs, with 1443 upregulated and 707 downregulated (Suppl. Fig. S1B). In addition, 1402 mRNAs were altered, with 935 upregulated and 467 downregulated (Suppl. Fig. S1A). Gene ontology clustering analysis was subsequently used to evaluate the biological functions of genes that were affected by CLOCK particularly. Results suggested that there were 152 differentially expressed genes involved in immune system processes (Fig. 1B; P < 0.01). Among them, the deubiquitination enzyme OTUD7B, an inhibitor of the NF-κB signaling pathway, was identified as a Clock-controlled gene in a previous study (Hou et al., 2017). However, the specific regulatory mechanisms of this gene remained unclear. In vitro promoter assays have previously shown that CACGNG, CACGTT, and CATG (T/C) G are functional Clock-binding motifs (Yoshitane et al., 2014). In this study, we did not find any potential motifs 1000 bp upstream of the Otud7b gene transcription initiation site (data not shown). Thus, we intended to identify lncRNAs that may be nodes related to Otud7b. An Otud7b-associated lncRNA gene cluster was identified and summarized using LncTar software. In total, there were 4 lncRNAs—AK012412, AK028267, AK079976, AK028245—that were altered after Clock knockdown and simultaneously related to Otud7b (Table 1; Fig. 1C).
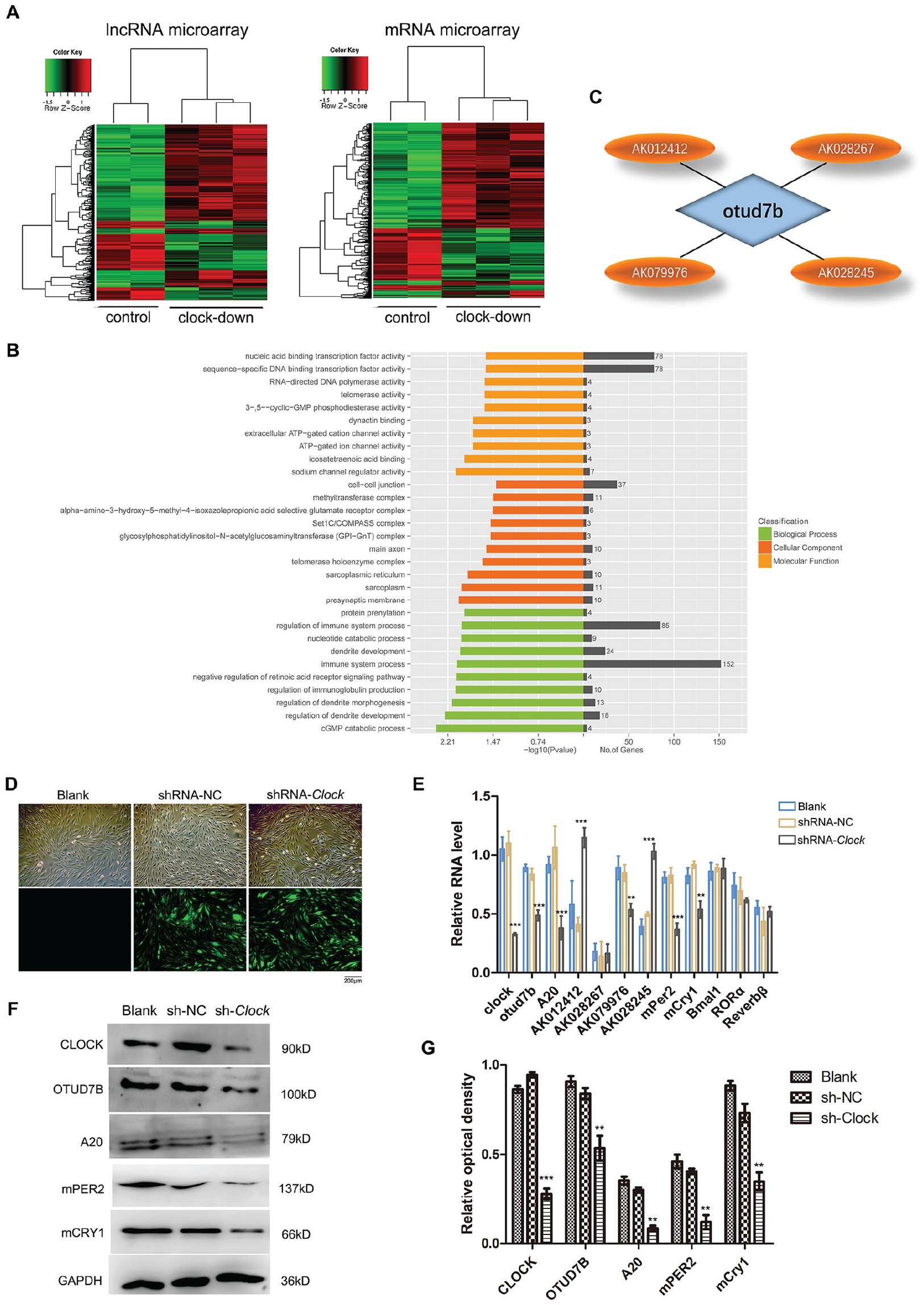
Identification of Clock-responsive lncRNAs targeting Otud7b. (A) Heat map of mRNA and lncRNA microarrays after Clock was downregulated by short hairpin RNA (sh-RNA) in MS-1 cells. Standard selection criteria were used to identify deferentially expressed mRNAs and lncRNAs established at p ≤ 0.05 and /FoldChange/ >1.5. (B) The Gene Oncology (GO) analysis in relation to differential expression of mRNA in a MS-1 cell line in response to Clock gene was knockdown (Hou et al., 2017). (C) Predicted lncRNAs with Otud7b as the target factor obtained by using LncTar. (D) Green fluorescence represents the transfection efficiency of the virus in the shRNA-NC and shRNA-Clock groups (magnification, ×100). (E) Quantitative reverse-transcriptase polymerase chain reaction analysis in blank, shRNA-NC, or shRNA-Clock cells reveals that the expression levels of mPer2, mCry1, and 3 lncRNAs were altered. (F, G) The protein expression was determined by Western blot analysis. Clock-knockdown downregulated mPER2, mCRY1, OTUD7B, and A20 protein expression levels. Error bars indicate mean ± standard errors of the mean. *p < 0.05, **p < 0.01. shRNA-NC: negative control (MS-1 cells transfected with lv-GFP-Puro NC virus); shRNA-Clock: transfected with lv-shRNA-Clock virus. GAPDH was used as an internal control. Data represents the mean ± SD from 3 independent experiments.
Expression of predicted lncRNAs targeting Otud7b in gene chip detection.
To confirm the altered expression of the 4 candidate lncRNAs, their expression levels were measured by qRT-PCR in MS-1 cells transfected with lv-shRNA-Clock and lv-shRNA-NC viruses. Observation of green fluorescence indicated that the number of cells successfully transfected was >85% (Fig. 1D). Clock-knockdown downregulated the expression of mPer2 and mCry1, which were core clock genes, whereas others were not unaffected (Fig. 1E, F). As shown in Figure 1E, 3 lncRNAs (AK012412, AK079976, AK028245) were altered significantly compared with blank and shRNA-NC cells (p < 0.05). Consistent with our previous results, Clock knockdown downregulated Otud7b mRNA (Fig. 1E) and protein (Fig. 1F, G) expression. In addition, because OTUD7B belongs to the A20 deubiquitination family (and A20 exhibits the same inhibitory effect on NF-κB), the expression of A20 was also evaluated. The results showed that A20 was downregulated in Clock-knockdown cells (Fig. 1E-G).
LncRNAs Exhibit Circadian-related Variable Expression
Genes regulated by the circadian system are characterized by rhythmic changes in expression within a 24-h cycle. Thus, the gene expression changes of 3 candidate lncRNAs were monitored for time-dependent changes in vitro (Halberg, 1969; Cornelissen, 2014). After equinum serum shock induction, the expression of circadian genes and circadian clock control genes should exhibit synchronization. The results of real-time PCR analysis in this study demonstrated that the expression of core circadian genes including Clock, Bmal1, mPer2, and mCry1 exhibited circadian rhythmic fluctuations, and Clock knockdown decreased amplitudes of Clock, mPer2, and mCry1 (Fig. 2A-D). Of the 3 verified lncRNAs demonstrating altered gene expression after Clock knockdown, AK079976 and AK028245 exhibited detectable time-of-day differences in expression (Fig. 2F, G), and Clock knockdown decreased their amplitudes too, whereas AK012412 did not exhibit circadian rhythmic characteristics (Fig. 2E).
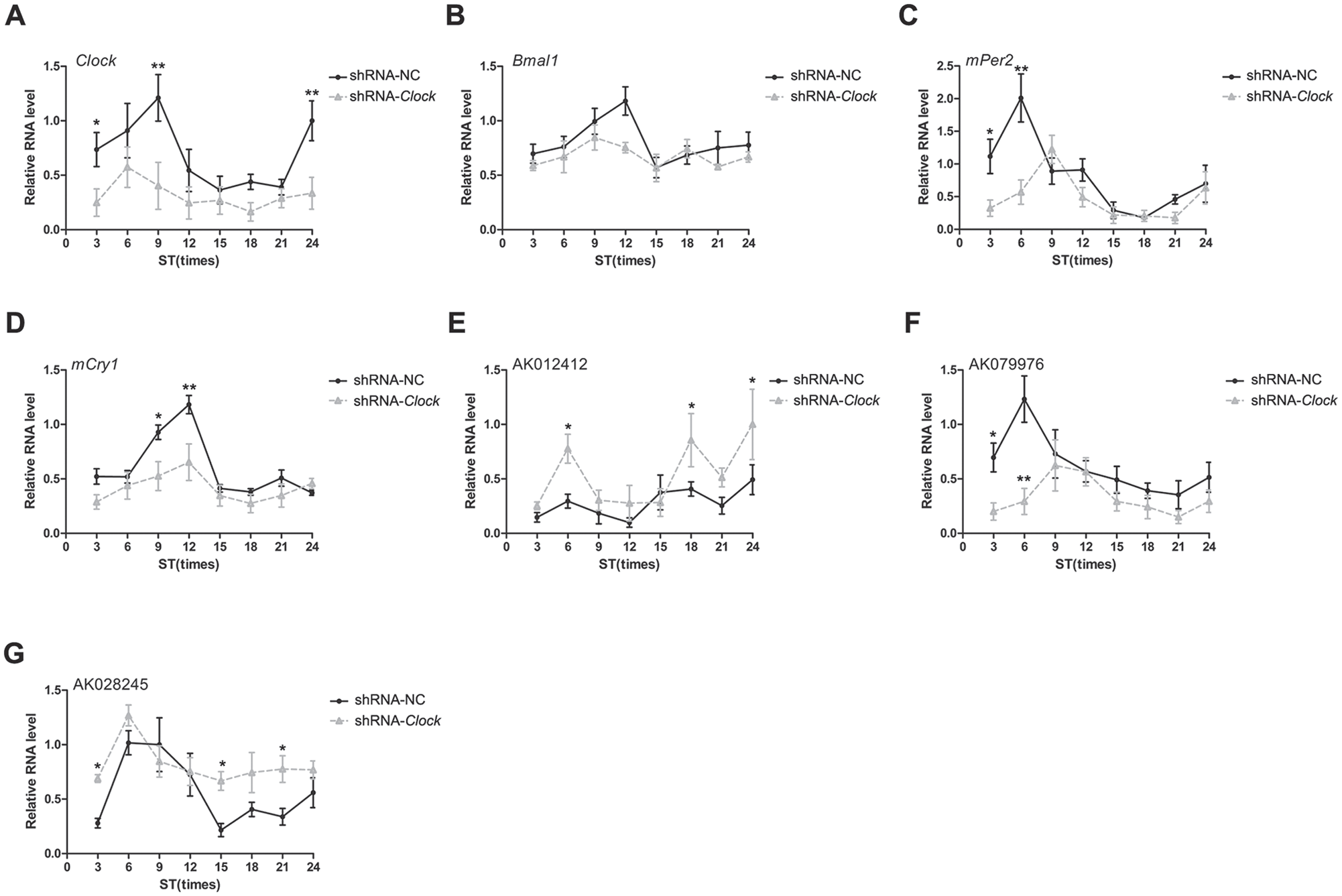
LncRNA circadian variation detected by quantitative reverse-transcriptase polymerase chain reaction. Synchronization induction was performed in short hairpin negative control (shRNA-NC) cells and shRNA-Clock cells. (A) Clock. (B) Bmal1. (C) mPer2. (D) mCry1. (E) AK012412. (F) AK079976. (G) AK028245. Error bars indicate means ± standard errors of the mean. *p < 0.05, **p < 0.01. Data represent the mean ± SD from 3 independent experiments.
Changes in lncRNA Expression in Immunocompromised Mice
We identified 2 clock-controlled lncRNAs in this study, AK079976 and AK028245, both of which are predicted Otud7b gene targets. Both OTUD7B and A20 are inhibitors of the NF-κB signaling pathway and play a key role in immune activity. Therefore, the expression of lncRNA in the spleen, an important immune organ, was evaluated under different immune states. The volume of the spleen tissues of mice injected with CPA was significantly smaller than that of the control group (Fig. 3A) 5 days later (Fig. 3B), and the spleen index was significantly reduced; in addition, the expression of interleukin (IL)–2 and IL-6 was reduced compared with the control group (Fig. 3C). In contrast, the expression of IL-10 was significantly higher than that of the control group. AK028245 expression was decreased after CPA treatment (p < 0.05), while AK079976 expression was not significantly affected (Fig. 3C). In a separate experiment, LPS stimulation was used to simulate an acute immune injury model, verified by severe spleen injury apparent by hematoxylin and eosin staining. In this model, the splenic nodules adhered to each other, with a small amount of lymphocyte necrosis, severe dilation, and congestion of the red pulp. In addition, a large number of disseminated extramedullary hematopoietic cells (Fig. 3D) were visible. The expression of IL-6 and tumor necrosis factor-α (TNF-α) was increased, and lncRNA-AK028245 showed a similar significant increase (Fig. 3E). Therefore, AK028245 was identified as a potential element affecting spleen immune activity by acting on OTUD7B.
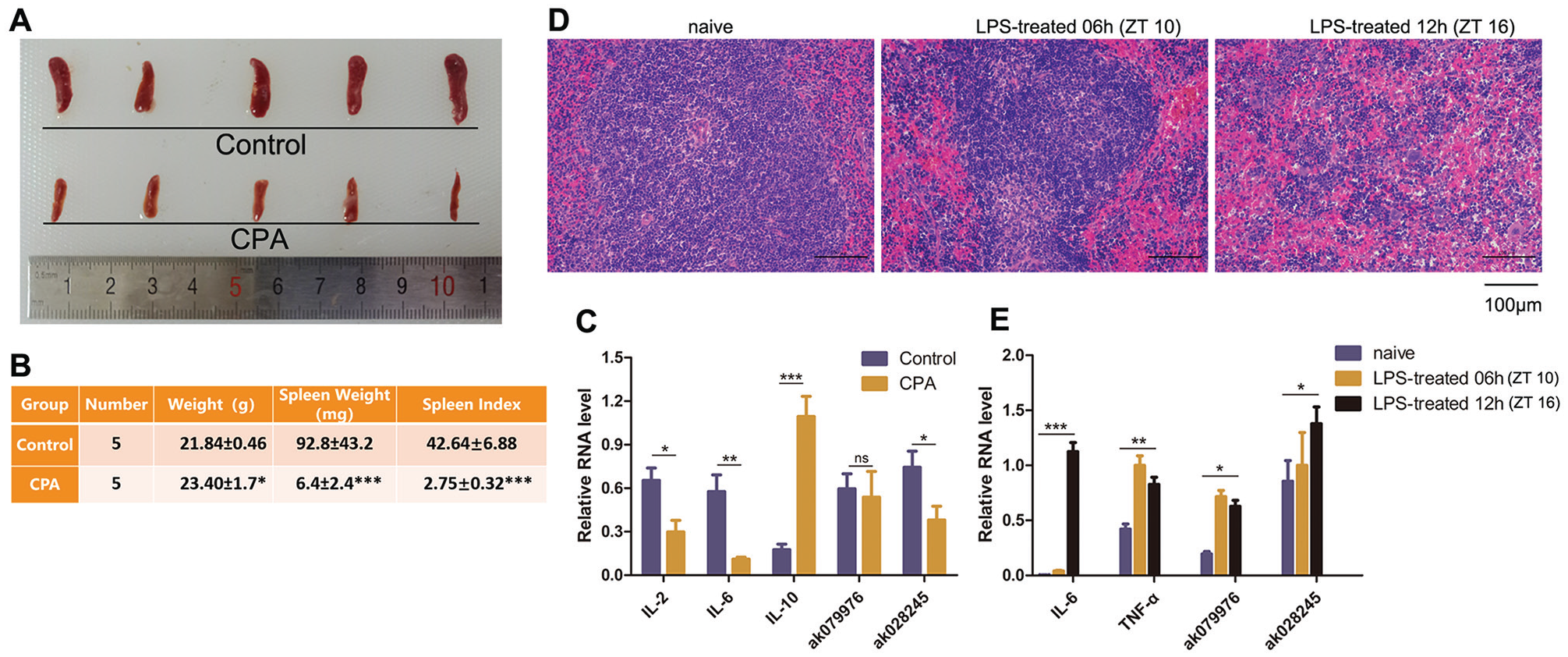
Changes in lncRNA expression in immunocompromised mice. (A) Shape and size of spleen tissues in cyclophosphamide-treated mice for 5 days. CPA, cyclophosphamide. (B) Spleen index in CPA-treated mice. Spleen index = (spleen weight [mg] /body weight [g]) × 10. (C) LncRNAs and cytokines expression in CPA-treated mice by quantitative reverse-transcriptase polymerase chain reaction. (D) Spleen tissues in the hematoxylin and eosin–stained histopathological images (objective: 200×) in lipopolysaccharide (LPS)–treated mice. Naive: normal control group, injected with saline. (E) LncRNAs and cytokine expression in LPS-treated mice. ZT, zeitgeber time. n = 5 mice per group. *p < 0.05, **p < 0.01. Data represent the mean ± SD from 3 independent experiments.
lncRNA AK028245 Regulates the Expression of OTUD7B and A20
According to the NCBI, AK028245 is located on chromosome chr5:122377407-122377720:+, with a full length of 314 bp, as shown in Supplemental Figure 2. In this experiment, AK028245 was both knocked down and overexpressed using an lncRNA smart silencer and lentiviral transfection. Three cell lines were used: NIH-3T3, MHS, and RAW264.7. MHS cells are mouse alveolar macrophages, and RAW264.7 cells are mouse mononuclear macrophages, which are commonly used in molecular immunology. NIH-3T3 cells are mouse embryonic fibroblasts with stable morphological properties, often used for transfection and gene expression studies. As shown in Figure 4A and B, both AK028245 silencing and overexpression were effectively detected by qPCR. After AK028245 silencing, the mRNA expression of Otud7b and A20 was increased in MHS cells and RAW26.7 cells, although Otud7b was not significantly changed in NIH-3T3 cells (Fig. 4C). Western blot results showed that protein expression levels exhibited similar trends as mRNA expression (Fig. 4E, F). After AK028245 was overexpressed, the mRNA and protein expression of both Otud7b and A20 were significantly decreased (Fig. 4D, G, H).
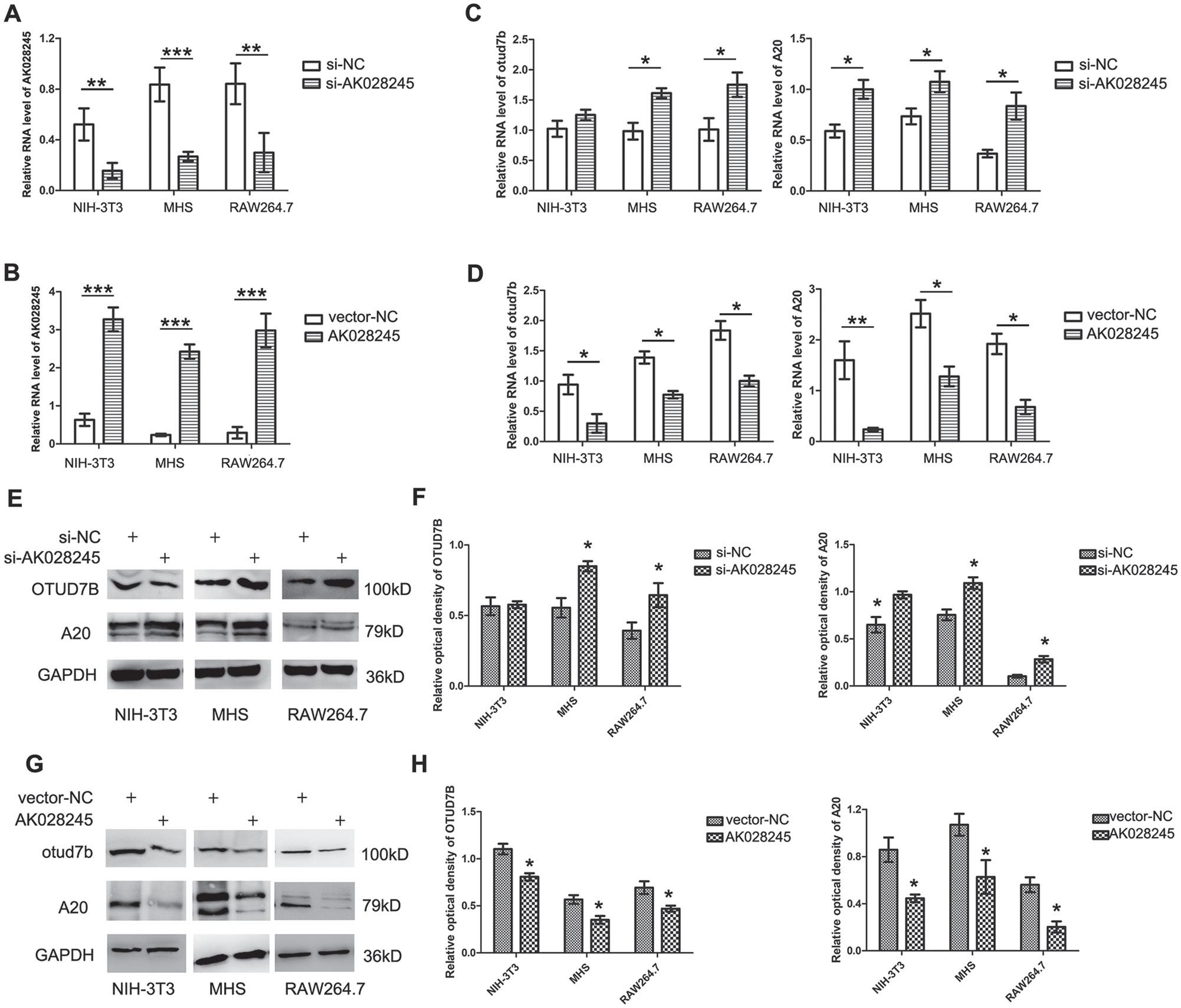
lncRNA AK028245 regulates the expression of OTUD7B and A20. (A) AK028245 smart silencer was performed in NIH-3T3, MHS, and raw264.7 cell lines. (B) NIH-3T3, MHS, and raw264.7 stably overexpressed AK028245 or a control vector. (C) Otud7b and A20 mRNA levels were identified in AK028245-silenced cells by quantitative reverse transcriptase polymerase chain reaction. (D) Otud7b and A20 mRNA levels were identified in overexpressing AK028245 cells. (E, F) OTUD7B and A20 protein levels were identified in AK028245-silenced cells by Western blot. (G, H) OTUD7B and A20 protein levels were identified in overexpressing AK028245 cells by Western blot. *p < 0.05, **p < 0.01, ***p < 0.005. Data represent mean ± SD from 3 independent experiments.
Inhibition of AK028245 Mitigates the Clock-induced Decrease of Otud7b and A20
The above results confirmed that the lnc-RNA AK028245 is a negative regulator of OTUD7B and A20. Subsequently, we hypothesized that CLOCK regulates OTUD7B and A20 by using AK028245 as a signaling molecule. To explore whether AK028245 plays a role in the regulation of CLOCK on OTUD7B and A20, AK028245 was silenced in a Clock-knockdown model of RAW264.7 cells. Clock knockdown was correlated with Otud7b and A20 mRNA downregulation, and when AK028245 was silenced simultaneously with Clock, the effect of CLOCK was entirely lost (Fig. 5A, B).
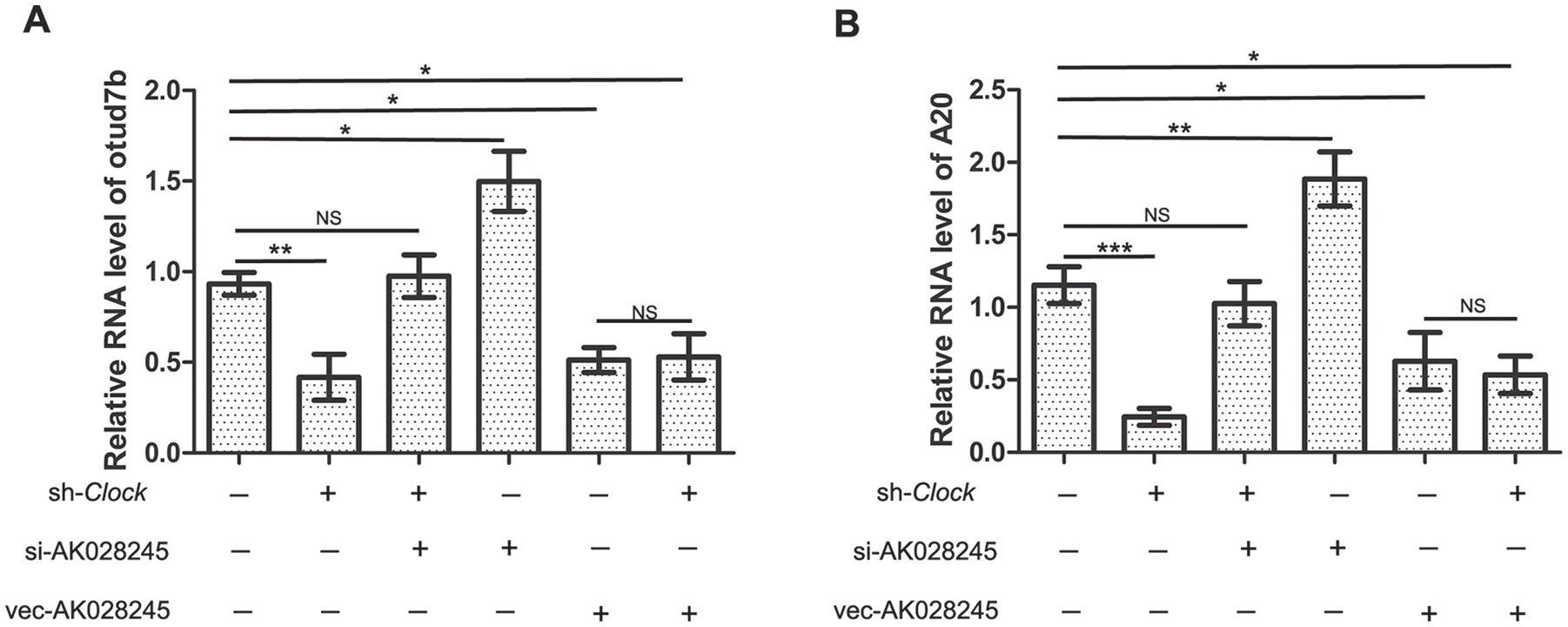
Changes in Clock and AK028245 together effect the expression of Otud7b and A20. (A) Otud7b expression levels detected by quantitative reverse transcriptase polymerase chain reaction (RT-qPCR). (B) A20 expression levels detected by RT-qPCR. Sh-Clock: cells transfected with lv-shRNA-Clock virus; si-AK028245: smart silencer AK028245; Vec-AK028245: transfected with overexpressed AK028245 virus. Values are expressed as mean ± SD. *p < 0.05, **p < 0.01.
AK028245 Regulates LPS-induced Activation of the NF-κB Signal Pathway
Given that A20 and OTUD7B are inhibitory signals upstream of the NF-κB signaling pathway and are involved in immune disease and inflammatory processes, we validated whether AK028245 had an effect on the activity of important immune inflammatory signals. In normal RAW264.7 cells, LPS stimulation (100 ng/mL) leads to elevated AK028245 expression (Fig. 6A), which is consistent with the results of our animal experiments. LPS was thus used to induce an inflammatory response in AK028245 silenced cells. Using qPCR, the mRNA abundance of 4 proinflammatory cytokines was evaluated. Among them, TNF-α and IL-6 were affected by the activation of the classical NF-κB signal pathway, whereas IFN-γ and Cxcl12 were affected by the nonclassical pathway. IL-6 expression increased rapidly after LPS treatment, peaking at 4 h. Subsequent expression demonstrated a gradual decline. The silenced of AK028245 had no significant effect on IL-6 expression before 4 h (Fig. 6B); however at 8 h, silencing of AK028245 significantly reduced IL-6 expression compared with control siRNA (si-NC). The expression of TNF-α increased rapidly, peaking at 2 h. Between 4 h and 8 h, the expression of TNF-α in the si-AK028245 group was lower than in the si-NC group (Fig. 6C). Interestingly, we found that both IL-6 and TNF-α increased significantly at peak levels when AK028245 was overexpressed (Fig. 6F, G). With regard to the nonclassical pathway, we saw that both IFN-γ and Cxcl12 reached peak levels at 4 h after stimulation. Their expression was significantly lower at the peak within AK028245 silenced (Fig. 6D, E), whereas it increased within AK028245 overexpressed (Fig. 6H, I). It can be seen that AK028245 promotes the expression of IL-6, TNF-α, IFN-γ, and Cxcl12. These results suggest that AK028245 is indeed a promoter of the NF-κB signal pathway, both classical and nonclassical.
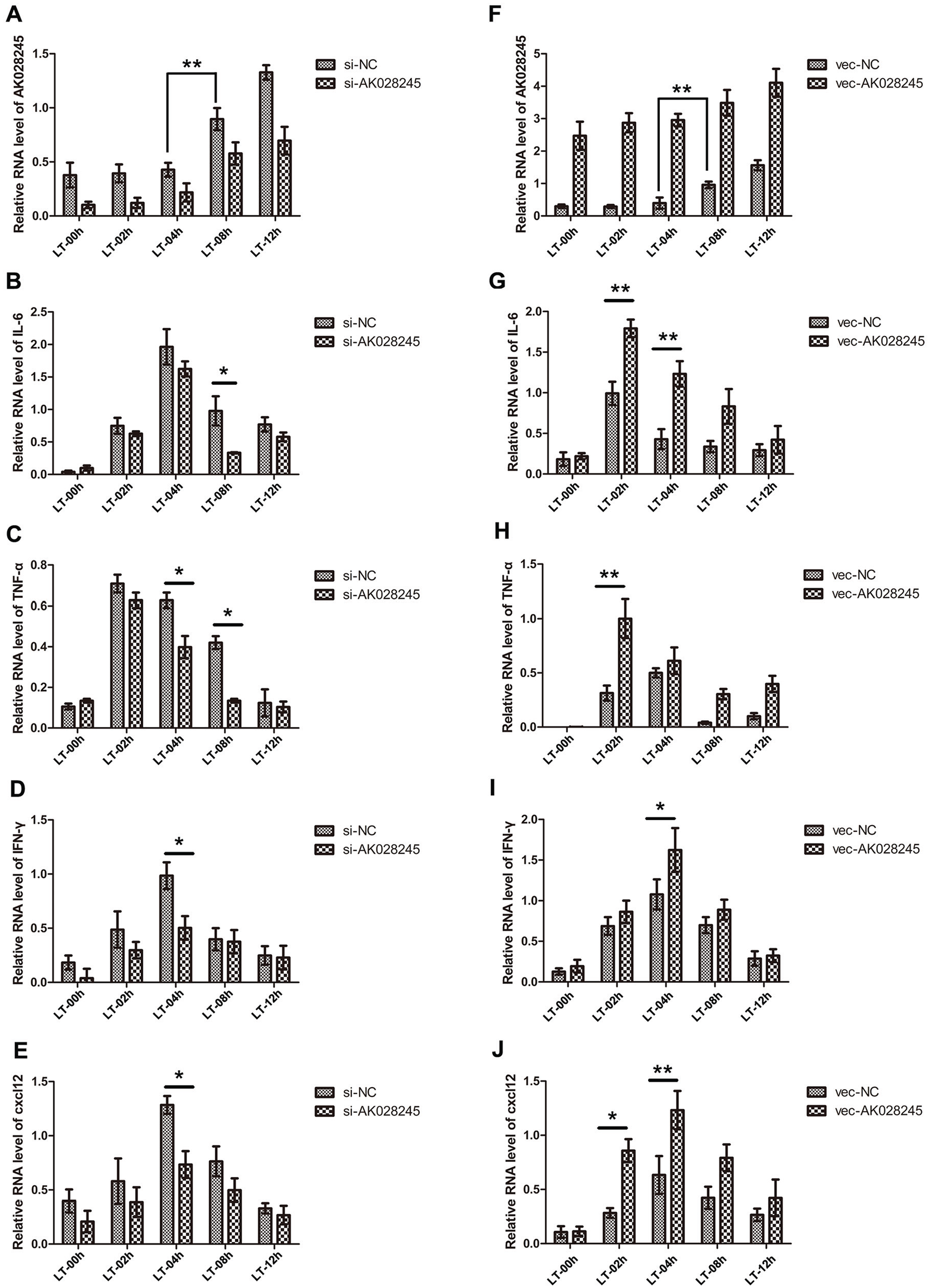
AK028245 affects the expression of interleukin-6 (IL-6), tumor necrosis factor–α (TNF-α), interferon-γ (IFN-γ), and Cxcl12. (A) Lipopolysaccharide (LPS) stimulation (100 ng/mL) increased AK028245 expression levels in RAW264.7 cells, as detected by reverse transcriptase polymerase chain reaction. (B-E) The expression level of IL-6, TNF-α, IFN-γ, and Cxcl12 at consecutive time points after LPS stimulation (100 ng/mL) in AK028245-silenced RAW264.7 cells. (F-I) The expression level of IL-6, TNF-α, IFN-γ, and Cxcl12 at consecutive time points after LPS stimulation (100 ng/mL) in AK028245-overexpressed RAW264.7 cells. LT, LPS treated. *p < 0.05, **p < 0.01. Data represent the mean ± SD from 3 independent experiments.
Discussion
With the development of lncRNA detection techniques, accumulating evidence indicates that lncRNAs participate in a wide variety of important biological phenomena (Yang et al., 2017). In this study, a new Clock-controlled lncRNA was found to inhibit the expression of A20 and OTUD7B, inhibitory factors upstream of the NF-κB pathway. AK028245 also inhibited the impact of CLOCK on A20 and OTUD7B. In addition, AK028245 accelerated the expression of IL-6, TNF-α, IFN-γ, and Cxcl12 in the LPS-induced immune response.
In recent years, the number of studies on the regulatory role of lncRNAs in circadian rhythmicity has been increasing gradually (Chen et al., 2017; Henriques et al., 2017; Yang et al., 2017; Bhadra et al., 2018; Park and Belden, 2018). However, there have been no reports to date regarding the regulatory functions of lncRNAs in relation to Clock gene expression. As a core factor of the circadian rhythm system, CLOCK plays a key role in the function of the circadian system and participates in the regulation of many related physiological functions and disease developments (Cui et al., 2015; Cao, 2018; Masri and Sassone-Corsi, 2018; Stenvers et al., 2019). The aims of this study were focused on lncRNA expression during Clock knockdown to elucidate the molecular mechanisms implicated in biological processes related to circadian rhythmicity, particularly the immune response. Microarray analysis revealed 2150 altered lncRNAs, with 1443 upregulated and 707 downregulated (data not shown). LncRNAs exhibiting the spectrum of expression change observed herein likely represent multiple levels of regulation of the circadian system from epigenetic levels, genetic, and so forth (Bhadra et al., 2018; Park and Belden, 2018). They also provide valuable targets for physiological or disease research, demonstrating circadian influence.
Over the past decade, the roles of some lncRNAs in immune regulation have been reported (Carpenter et al., 2013). For example, Carpenter et al. showed that lncRNA-Cox2 regulates a large set of inflammatory response genes and that knockdown of this lncRNA resulted in increased expression of a gene set including Ccl5 and Cx3cl1 (Carpenter et al., 2013). LncRNA-Lethe physically associates with the NF-κB p65 (RelA) subunit, directly or indirectly blocking its recruitment to the promoters of immune genes such as IL-6 and IL-8 (Rapicavoli et al., 2013). In this study, we demonstrated that lncRNA-AK028245 regulates expression of TNF-α, IL-6, IFN-γ, and Cxcl12 in LPS-stimulated cells. The previous study provided compelling evidence for the possible functions of lncRNA-AK028245 in the immune system. In addition, other studies have shown that CLOCK functions as a positive regulator of NF-κB–responsive promoters by the direct binding of CLOCK protein to the p65 subunit of NF-κB, enhancing the immune response (Spengler et al., 2012). The results of this study showed that CLOCK may act as an inhibitor of AK028245 since Clock knockdown promoted the expression of proinflammatory lncRNA-AK028245. This phenomenon appears to be consistent with previous studies. First, the role of CLOCK as an enhancer of NF-κB signaling occurs independently of specific transactivator functions, which are required of CLOCK in BMAL1-dependent regulation of circadian genes. The lncRNA discovered herein displayed time-of-day differences in expression and appeared to be regulated by the circadian system, which is itself governed by the regulation of A20 and OTUD7B. As a whole, the circadian system entails a sophisticated and complex regulatory network. The effect of Clock inhibition on AK028245 is, on one hand, evidence of a link between the circadian system’s involvement in immune regulation. On the other hand, it can also be taken as a compensatory effect of Clock knockdown to coordinate its strong proinflammatory effect. Such regulation would be more conducive to maintaining the homeostasis between the circadian rhythm and the immune system.
Previous studies have shown that the circadian system is involved in immune regulation through multiple pathways (Bellet et al., 2012; Nguyen et al., 2013; Sanchez and Kay, 2016; Scheiermann et al., 2018). In this study, the expression of immune response–related mRNA was most obvious after the downregulation of Clock. The deubiquitination enzyme family member Otud7b was shown to be the Clock-controlled gene most prevalent in our previous studies. Therefore, we hypothesized that there is significant cross-talk between OTUD7B and A20, communicating changes between CLOCK and NF-κB. Regulation involving noncoding RNA may be an important and missing link to describe the interaction between circadian systems and physiological responses. This study identified a novel Clock-controlled lncRNA that showed significant changes in the presence of immune abnormalities. The expression of both A20 and OTUD7B increased when AK028245 was silenced, and the results of AK028245 overexpression revealed similarly consistent conclusions. Therefore, AK028245 appears to be the signal source of the 2 upstream inhibitory factors of the NF-κB signaling pathway, affecting their functions. In contrast to the action of micro RNAs (miRNAs), lncRNAs are involved in the regulation of gene expression on multiple levels, including transcriptional regulation by the recruitment of chromatin-modifying complexes and posttranscriptional regulation by interaction with miRNAs or proteins (Qu et al., 2016). Although we found changes in mRNA levels of A20 and Otud7b due to AK028245, the specific mechanism remains to be further studied.
The inconsistency of NIH-3T3 with MHS and RAW264.7 cell lines may be due to differences in cellular immunity. MHS and RAW264.7 are immune cells, which are more sensitive to stimulation of immune-related signals, whereas NIH-3T3 cells have relatively weak immunity. In addition, the inhibitory effect of A20 on the NF-κB pathway is about 4 to 5 times that of OTUD7B. In NIH-3T3 cells, lnc-AK028245 does not affect the expression of Otud7b, but the expression of A20 is meaningful. This result may demonstrate the effective role of lnc-AK028245 in the immune activities of the A20 family.
In summary, we elucidated lncRNA profiles of wild-type and Clock-knockdown cells in mice by RNA microarray analysis, identifying and characterizing lncRNA-AK028245, which promotes the immune response. LncRNA-AK028245 therefore acts as an important component of the circadian system, participating in immune activation. In future studies, further exploration of AK028245 should be undertaken to expand on its molecular mechanisms of action in the regulatory network governing circadian-sensitive immune processes.
Supplemental Material
Supplemental_material_JBR – Supplemental material for The Clock-Controlled lncRNA-AK028245 Participates in the Immune Response via Immune Response Factors OTUD7B and A20
Supplemental material, Supplemental_material_JBR for The Clock-Controlled lncRNA-AK028245 Participates in the Immune Response via Immune Response Factors OTUD7B and A20 by Fang Qi, Zhou Jiang, Wang Hou, Bo Peng, Shuting Cheng, Xiaolong Zhang, Zhihan Luo, Zeyong Dai, Yumeng Wang, Yanyou Liu, Yuhui Wang and Zhengrong Wang in Journal of Biological Rhythms
Footnotes
Acknowledgements
This research was supported by the National Natural Science Foundation of China (grant 31500935).
Conflict of Interest Statement
The authors have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
NOTE
Supplemental material is available for this article online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
