Abstract
Circadian clocks regulate multiple physiological processes in the eye, but their requirement for retinal health remains unclear. We previously showed that Drosophila homologs of spliceosome proteins implicated in human retinitis pigmentosa (RP), the most common genetically inherited cause of blindness, have a role in the brain circadian clock. In this study, we report circadian phenotypes in murine models of RP. We found that mice carrying a homozygous H2309P mutation in Pre-mRNA splicing factor 8 (Prpf8) display a lengthened period of the circadian wheel-running activity rhythm. We show also that the daily cycling of circadian gene expression is dampened in the retina of Prpf8-H2309P mice. Surprisingly, molecular rhythms are intact in the eye cup, which includes the retinal pigment epithelium (RPE), even though the RPE is thought to be the primary tissue affected in this form of RP. Downregulation of Prp31, another RNA splicing factor implicated in RP, leads to period lengthening in a human cell culture model. The period of circadian bioluminescence in primary fibroblasts of human RP patients is not significantly altered. Together, these studies link a prominent retinal disorder to circadian deficits, which could contribute to disease pathology.
The vertebrate eye is both a visual organ and a sensor that relays light inputs into the brain to entrain and reset the central circadian oscillator in the suprachiasmatic nucleus (SCN). Many genes are expressed with a 24-h rhythm in the eye, and while most are light driven, a sizeable number of transcripts (~300) cycle in constant darkness under the control of circadian clocks (Storch et al., 2007). Indeed, multiple tissues of the eye have cell-autonomous circadian clocks, which temporally compartmentalize function and allow physiological anticipation of daily light transitions associated with dusk and dawn (McMahon et al., 2014). Functions under circadian control in the retina include melatonin release (Tosini and Manaker, 1996), dopamine synthesis and secretion (Doyle et al., 2002; Ribelayga et al., 2004), gamma-aminobutyric acid turnover and release (Jaliffa et al., 2001), phagocytosis of photoreceptor outer segments (Goldman et al., 1980; Teirstein et al., 1980; LaVail, 1980; Grace et al., 1996), contrast sensitivity (Hwang et al., 2013), extracellular pH (Dmitriev and Mangel, 2001), intraocular pressure (Boyd and McLeod, 1964; Rowland et al., 1981), visual sensitivity (Walker and Olton, 1979; Bassi and Powers, 1986), and rod-cone electrical coupling (Ribelayga et al., 2008).
Despite the circadian regulation of multiple processes in the eye, the requirement of functional circadian clocks for retinal health remains uncertain (Felder-Schmittbuhl et al., 2018). Eukaryotic circadian clocks consist of transcription-translation loops, with the major loop in mammals consisting of a positive arm in which transcription factors BMAL1 and CLOCK drive the expression of Period (Per1-3) and Cryptochrome (Cry1-2) genes and a negative arm in which PER and CRY proteins negatively regulate CLOCK and BMAL1 (Partch et al., 2014). Negative feedback by PER-CRY not only drives rhythmic Per/Cry expression but also confers circadian expression on to hundreds of other genes downstream of BMAL1-CLOCK. Mice with mutations in core circadian factors display behavioral phenotypes and have also been extensively characterized for metabolic phenotypes and even for susceptibility to cancer. However, less is known about the impact of disrupted clocks in the eye. Loss of Per1/Per2 leads to subtle morphologic changes in the retina, largely attributed to developmental defects, while no phenotypic alterations are observed for Cry1/Cry2 double-knockout mutants (Ait-Hmyed et al., 2013; Selby et al., 2000). Developmental deletion of Bmal1 from retinal progenitors perturbs normal visual transduction, likely because of disrupted retinal neurogenesis (Sawant et al., 2019). Mice with adult-specific Bmal1 knockout display corneal neovascularization, inflammation, and keratinization (Yang et al., 2016), and Clock knockouts develop cataracts (Dubrovsky et al., 2010). Because both BMAL1 and CLOCK are transcriptional activators, whose loss produces reduced expression (in addition to disrupted cycling) of many target genes, it is unclear if the reported eye pathologies reflect consequences of clock disruption.
Retinitis pigmentosa (RP) is the most commonly inherited form of progressive retinal degeneration, affecting 1 in 3000 to 7000 people worldwide (Ferrari et al., 2011). While RP is a vastly heterogeneous disease with more than 80 genes currently implicated, the symptoms common to all RP types involve night blindness that first progresses to tunnel vision, often resulting in legal blindness, and then to complete blindness (Ferrari et al., 2011; Verbakel et al., 2018). The progression of disease is explained by the initial death of rods, which subsequently triggers the degeneration of cones, resulting in complete blindness. The onset and penetration of disease varies depending on the causal genetic lesion. However, even individuals with the same RP mutations can have different phenotypes, indicating an important role for modifier loci and/or epigenetic factors in this disease (Ferrari et al., 2011). Most RP mutations map to genes implicated in the visual cycle, such as Rhodopsin, but a class of RP is caused by mutations in U4/U6.U5 triple small nuclear ribonucleoprotein (tri-snRNP) components that form a well-conserved regulatory unit of the spliceosome (Verbakel et al., 2018). Interestingly, we recently showed that tri-snRNP spliceosome factors regulate circadian clocks in Drosophila melanogaster (Shakhmantsir et al., 2018). In addition, both Prpf4 and SM-like (LSM) genes, which associate with U6 snRNP, regulate circadian rhythms in cultured osteosarcoma cells (Zhang et al., 2009; Perez-Santangelo et al., 2014). Because of this emerging connection between the U4/U6.U5 tri-snRNP and circadian clocks, we sought to determine if human RP-causing mutations in U4/U6.U5 splicing factors are associated with circadian phenotypes, in particular in the eye.
For our studies, we used 2 previously characterized mouse models of human RP mutations: Prpf3-T494M and Prpf8-H2309P (Graziotto et al., 2011). We identified a long-period circadian phenotype in the wheel-running activity of Prpf8-H2309P mice. In addition, the circadian output of the retina of Prpf8-H2309P mice was dampened, as revealed by daily profiling of selected clock gene transcripts in the isolated retina. We propose that rhythm dampening in the retina underlies some of the previously reported phenotypes in Prpf8-H2309P mice, in particular the disrupted rhythm in phagocytosis of photoreceptor outer segments (Graziotto et al., 2011; Farkas et al., 2014). We show also that the knockdown of Pre-mRNA Splicing Factor 31 (PRPF31), another tri-snRNP spliceosome factor, lengthens circadian period in a human cell culture model. Finally, we measure circadian period in primary skin fibroblast cultures from RP patients with heterozygous mutations in PRPF31 and report no statistically significant differences in circadian period in these cells as compared with controls.
Methods
Mice Husbandry and Genotyping
Prpf3 knockin mice bearing the p.T494M missense mutation found in RP18 patients (Prpf3-T494M) and Prpf8 knockin mice bearing the p.H2309P missense mutation found in RP13 (Prpf8-H2309P) were obtained from Eric Pierce and have been characterized in Graziotto et al. (2011). Mouse lines used in this study were obtained from cryopreserved samples. Previously, Prpf3 and Prpf8 mutant mice were generated using embryonic stem cells from 129SvEvTac mice (Graziotto et al., 2011). The founders were crossed to C57BL/6J mice (without the Rd8 allele) and then all the heterozygous mice outcrossed into the C57BL/6J background for at least 8 generations prior to cryopreservation (Graziotto et al., 2011). For genotyping Prpf3-T494M mice, the forward primer was 5′-CTCATTCATTGGCATTAAAAATAAATAAACTCC-3′, and the reverse primer was 5′-CGTTGGCCTCTTCATGCGCTCTGTCGTGAC-3′. The polymerase chain reaction (PCR) product was digested with Xmn1, and the presence of a ~650-bp band signified the mutant allele. Prpf8-H2309P mice were genotyped with the forward primer 5′-CCAGACAAGCTGCTGACATTCAGCAGTC-3′ and the reverse primer 5′-TAGAGCTGTAAGTGGTCACTTCAGGC-3′. The mutant allele was identified by the presence of a band 400-bp larger than the wild-type one (Graziotto et al., 2011).
Mice were housed in group cages with water and food ad libitum in 12-h light:12-h dark cycles unless otherwise specified. The lights inside the cages in the mouse colony room provided ~20 to 50 lux intensity of white light. All procedures were approved by the University of Pennsylvania Institutional Animal Care and Use Committee.
Wheel-running Setup, Recording, and Data Analysis
To test SCN-driven circadian rhythms, we stably entrained Prpf3-T494M and Prpf8-H2309P mice in shared cages under 12-h light:12-h dark conditions for 3 weeks. Next, these mice were transferred to individual cages with running wheels and sensors that could track the wheel revolutions. Activity records of wheel-running activity were generated for 2 weeks’ worth of data, and analysis was performed using Clocklab/Matlab.
Eye Dissection and Tissue Collection
Mice were entrained in 12-h light:12-h dark conditions for 3 weeks and were euthanized by cervical dislocation at ZT1, ZT7, ZT13, and ZT19. Eyes were immediately removed (note: only 1 eye per mouse was used) and placed into a solution of 1X Hanks’ balanced salt solution with Ca2+ and Mg2+ (HBSS+; Invitrogen, Carlsbad, CA) kept at room temperature. The retina and the eye cup (RPE, choroid, Bruch’s membrane) were dissected in HBSS+ solution under a dissecting microscope. First, fat and muscle tissues around the eyeball were cleaned. Next, the iris and the lens were removed. The retina was detached from the eye cup, and the retina and eye cup samples were frozen immediately and separately on dry ice. All tissue samples were stored at −80 °C until the RNA isolation step.
RNA Isolation and Real-time Quantitative PCR
Mouse retina and eye cup samples were collected in individual RNAse-free tubes on dry ice at ZT1, ZT7, ZT13, and ZT19 and stored at −80 °C until all of the time points were collected (within 24 h). Within 1 week of their initial collection, the samples were homogenized with TRIzol (ThermoFischer, Waltham, MA) on ice with an RNAase-free pestle. Following the phase separation, the aqueous phase was transferred into a new tube, mixed with an equal volume of 70% ethanol, and loaded directly onto the RNeasy mini kit columns (Qiagen, Venlo, the Netherlands). The rest of the RNA isolation was done according to the manufacturer’s protocol. On-column DNase digestion (Qiagen) for 15 min at room temperature was included for all samples. cDNA was generated with Superscript II (Invitrogen), according to the manufacturer’s protocol. Quantitative real-time PCR reaction was performed in a ViiA7 Real-Time PCR system (Applied Biosystems, Foster City, CA) using SYBR Green Master Mix (Applied Biosystems) with gene-specific primers. Per2: F(5′-3′): GTCCACCTCCCTGCAGACAA, R(5′-3′): TCATTAGCCTTCACCTGCTTCAC; Rev-erb α: F(5′-3′): GCTCCATCGTTCGCATCAAT, R(5′-3′): CTAGAGGGCACAGGCTGCT; Cry1: F(5′-3′): GCCAGCTGATGTATTTCCCAG, R(5′-3′): CGCCAGCCTCAGTAGCCAG; Tim: F(5′-3′): GTTCCGAGACCAGACCCCTGA, R(5′-3′): CCAAATCGAGAGTGCCTGTTTC; Rorβ: F(5′-3′): CCTGGCTGATCGAACCAAG, R(5′-3′): TGCAGACTGCCGTGATAG; MerTK: F(5′-3′): TGACTCCTTGGAAGACTCTGAAG, R(5′-3′): CTTGAAGATTGCCTTGGTCAT; 36B4: F(5′-3′): TCCAGGCTTTGGGCATCA, R(5′-3′): CTTTATCAGCTGCACATCACTCAGA. The ΔΔCt method was used to calculate the relative gene expression across the day. 36B4 was used as a normalization control.
Primary Fibroblast Collection and Culture Establishment
Informed consent was obtained for a dermal skin biopsy after explanation of the nature of the study; procedures complied with the Declaration of Helsinki and were approved by the University of Pennsylvania institutional review board (#808828). Biopsy samples were processed according to the published protocol (Vangipuram et al., 2013). All samples are cryopreserved at passage 2. Experimental controls were obtained from Coriell Institute for Medical Research and were picked at random from the unaffected primary cell samples from male patients of similar age to the RP group.
Primary Fibroblast Maintenance
Tissue growth surface was pretreated with 0.1% v/v gelatin (G1393, Sigma, St. Louis, MO) for 30 min at room temperature. Cells were grown in complete culture media consisting of DMEM high glucose with GlutaMax (10569010, Invitrogen), 20% v/v fetal bovine serum (SH30071.03, Hyclone, Logan, UT), and penicillin/streptomycin 100 µg/mL (15140122, Invitrogen) at 37 °C under 5% CO2. Trypsin-EDTA 0.25% (25200056, Invitrogen) was used for routine cell splitting with a 1:4 ratio. Media were changed approximately every 3 days.
Virus Generation
HEK293T cells were grown on 145-mm plates using DMEM high glucose (4.5 g/L) with 1X glutamine and 5% fetal bovine serum. Cells were fed twice a week until they were 80% to 90% confluent. To generate the lentivirus, these HEK293T cells were transfected with 65.3 µg of pABpuro-BluF, which carries a BMal1-luciferase gene and was a gift from Steve Brown (plasmid 46824, Addgene, Cambridge, MA), 15 µg G protein of the vesicular stomatitis virus, 23 µg Hgpm2, 12 µg Tat, and 15 µg Rev using 2.5 M CaCl2 and 2X HEPES at pH 7.1. The supernatant was collected for 2 days following the transfection. This supernatant was then filtered through a 0.45-µM pore polyethersulfone filter (VWR international catalog #89126-828). The virus was concentrated by centrifugation at 49,000g for 90 min and stored at −80 °C.
Virus Infection and Luminescence Reporting of Circadian Gene Expression
Primary fibroblasts (passage #9-20) were seeded at ~80% confluence onto 35-mm dishes (3 replicate dishes per cell line). Cells were incubated with 8 µg/mL of polybrene (Sigma) diluted in complete media and treated with 10 µL of concentrated pABpuro-BluF virus for ~24 h. Next, the virus was removed, and cells were left in the complete media supplemented with 2 µg/mL puromycin dihydrochloride (Santa Cruz Biotechnology, Santa Cruz, CA) for 3 to 4 days. Following selection with puromycin, cells were synchronized with 0.1 µM dexamethasone (Sigma) diluted in the complete media for 1 h. Prior to placing the dishes into the luminometer, the medium with dexamethasone was changed to the lumicycle medium (no phenol red and 0.1 mM luciferin; Promega, Madison, WI), and the dishes were sealed tightly with high-vacuum silicone grease (Sigma). Real-time luminescence of the control and RP cells was monitored using a LumiCycle luminometer (Actimetrics, Wilmette, IL). The LumiCycle software program (Actimetrics) was used to calculate the period of the luminescence data. Period was determined by analyzing data recorded on days 2 to 5 only. To determine the period length of a given cell line, 3 to 4 independent viral infections were performed (in triplicate).
siRNA Knockdown
U2OS cells (ATCC, Manassas, VA) carrying a BMal1-luciferase reporter were maintained in Dulbecco’s Modified Eagle Medium (DMEM) with 20% v/v fetal bovine serum (SH30071.03, Hyclone) and penicillin/streptomycin 100 µg/mL (15140122, Invitrogen) at 37 °C under 5% CO2. Cells were seeded into 35-mm dishes at a density of ~3 × 105 cells/cm2, grown for 2 days in the antibiotic-free DMEM medium, and then transfected with small interfering RNAs (siRNAs) using 3 µL/plate DharmaFECT 4 Transfection Reagent (GE Life Sciences, Chicago, IL) and 2 µM of siGENOME siRNAs, as per the manufacturer’s instructions. In addition to the untreated control, the following siRNAs were used: siGENOME nontargeting siRNA pool #1 (GE Life Sciences, D-001206-13) as a negative control; SMARTpool: siGENOME CRY2 siRNA (GE Life Sciences, M-014151-01) as a positive control; and SMARTpool: siGENOME PRPF31 siRNA (GE Life Sciences, M-020525-01).
Cells were treated with siRNAs for 48 h and then processed for lumicycle recording as described in the section above or were collected for Prpf31 knockdown verification via western blotting as previously described (Garbe et al., 2013). Briefly, cells were collected in 1X passive lysis buffer (E1941, Promega), supplemented with cOmplete, EDTA-free Protease Inhibitor Cocktail (11873580001, Sigma). The following primary antibodies were used: PRPF31 A-6 antibody (sc-166792, Santa Cruz Biotechnology) and anti-beta actin antibody (mAbcam 8224).
Quantification and Statistical Analysis
The statistical parameters are included in the legends of each figure. JTK_CYCLEv3.1 (Hughes et al., 2010) was run in R for circadian statistical tests. GraphPad Prism was used for all other statistical analyses.
Results
Period of Circadian Wheel-running Activity Is Lengthened in Prpf8-H2309P Mice
Our previous findings implicated U4/U6.U5 spliceosome components, prp4 in particular, in the regulation of circadian rhythms of rest:activity in Drosophila (Shakhmantsir et al., 2018). To determine if the circadian behavioral role of U4/U6.U5 tri-snRNP is conserved in mammals, we sought to use mouse models mutant for these genes. Given that components of U4/U6.U5 tri-snRNP are essential genes required for development, we were not able to obtain viable homozygous mutants, and most heterozygous mutants are phenotypically normal (Graziotto et al., 2008; Bujakowska et al., 2009). Instead, we took advantage of the mouse models of human RP that were generated in the laboratory of Dr. Eric Pierce (Graziotto et al., 2011) to understand how Prpf3-T494M and Prpf8-H2309P knockin mutations affect circadian rhythms.
Flies with reduced levels of U4/U6.U5 tri-snRNP components had prolonged or disrupted rest:activity rhythms in free-running conditions (constant darkness), and so we tested behavioral rhythms of Prpf3-T494M and Prpf8-H2309P knockin mutants under similar conditions. Mouse strains were entrained to light:dark cycles for 3 weeks and then monitored wheel-running activity in constant darkness. Homozygous Prpf8-H2309P mice exhibited longer periods of circadian wheel-running behavior compared with their littermate controls without the mutation (p = 0.01; Fig. 1A, C). The circadian period of the activity rhythm in homozygous Prpf3-T494M mice was not significantly different from that of littermate controls (p = 0.08; Fig. 1B). Importantly, Prpf3-T494M and Prpf8-H2309P mice begin to display early signs of retinal degeneration only at ~2 years of age (Graziotto et al., 2011). Because we tested circadian behavior in these knockin mice at the age of 2 to 4 months, it allows us to exclude the blindness of RP mice as a potential cause of the circadian behavioral phenotype we observed (Lockley et al., 2007). Overall, our data indicate that the role of Prpf8, and perhaps Prpf3, in circadian behavior is conserved from fruit flies to mammals.
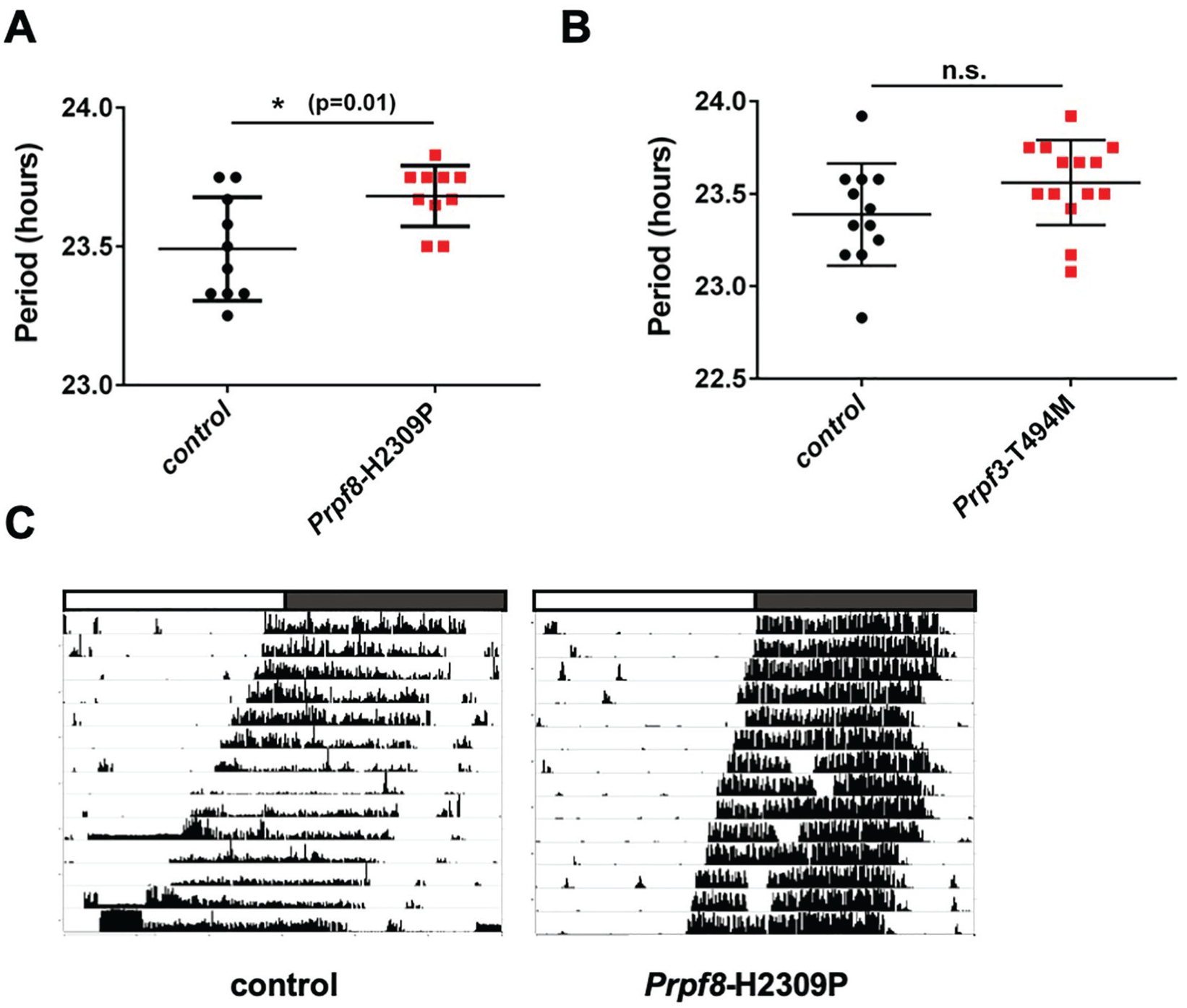
The period of circadian wheel-running activity is lengthened in Prpf8-H2309P mice. (A) Circadian wheel-running activity in Prpf8-H2309P homozygous mice is significantly longer than in their littermate controls (+/+). (B) No significant difference in period length between Prpf3-T494M homozygous mice and their littermate controls (+/+) was observed. n.s., not significant at the 0.05 level, *p = 0.01 to control (+/+) as determined by Student t test, n = 10-14. Error bars = +/− SEM. (C) Representative activity records of free-running behavior of a Prpf8-H2309P homozygous mouse and a littermate control. The white and the gray bars on top of the records indicate subjective day and night, respectively. Fourteen days of activity are displayed.
Daily Cycling of Retinal Transcripts Is Dampened in Prpf8-H2309P Mice
Based on the wheel-running phenotype of Prpf8-H2309P mice, we decided to probe circadian function in peripheral tissues. Given the connection between Prpf8 mutations and retinal degeneration (McKie et al., 2001; Graziotto et al., 2011), and a previous report of defective phagocytosis of photoreceptor outer segment discs in Prpf3-T494M and Prpf8-H2309P mice (Farkas et al., 2014), we chose to evaluate the daily rhythmicity of gene expression in the eye. The shedding of photoreceptor outer segment discs, which are then phagocytosed by the retinal pigment epithelium (RPE), is one of the most robust circadian processes in the vertebrate eye (Goldman et al., 1980; Teirstein et al., 1980; LaVail, 1980; Grace et al., 1996). Because RP caused by mutations in splicing factors is often associated with abnormalities in the RPE, we hypothesized that circadian rhythms of gene expression would be disrupted in the RPE, which could potentially account for the disrupted rhythm of outer segment phagocytosis (Graziotto et al., 2011; Farkas et al., 2014).
The traditional method for isolation of the RPE requires multiple enzymatic and temperature-dependent steps. With this method, we saw weak cycling of clock gene transcripts in control animals, possibly because the process interferes with the regulation of rhythms (data not shown). As a result, we switched to dissecting the whole eye cup (consisting of RPE, choroid, sclera, and Bruch’s membrane, with RPE constituting roughly ~1/6 of the entire prep), which provides a faster, manual method expected to better preserve endogenous cycling. We assayed daily gene expression in the isolated retina and the eye cup across 4 daily time points. Contrary to our hypothesis, the daily cycling of gene expression in the retina, but not in the RPE, was compromised in Prpf8-H2309P mice (Fig. 2). The transcripts of core clock genes, such as Per2 and Rev-erbα, did not display significant cycling in Prpf8-H2309P mice, although they did so in sibling controls (Fig. 2A). Our recent findings show that spliceosome factors regulate circadian clock cycling in Drosophila, at least partially via splicing of the timeless (tim) clock RNA (Shakhmantsir et al., 2018). Thus, we assayed the cycling of Tim mRNA in the murine retina and identified a significant cycle in sibling controls but a dampened daily rhythm in the retinas of Prpf8-H2309P mice (Fig. 2A). Not all of the genes we tested displayed the same differences between the control and the Prpf8-H2309P mice. Cry1 cycling was not detected in either the control or Prpf8-H2309P mice, while Rorβ, a circadian clock regulator previously implicated in retinal degeneration (Andre et al., 1998), cycled robustly in both genotypes. These results indicate that daily expression rhythms of some of the core circadian genes in the retina are sensitive to the Prpf8-H2309P mutation.
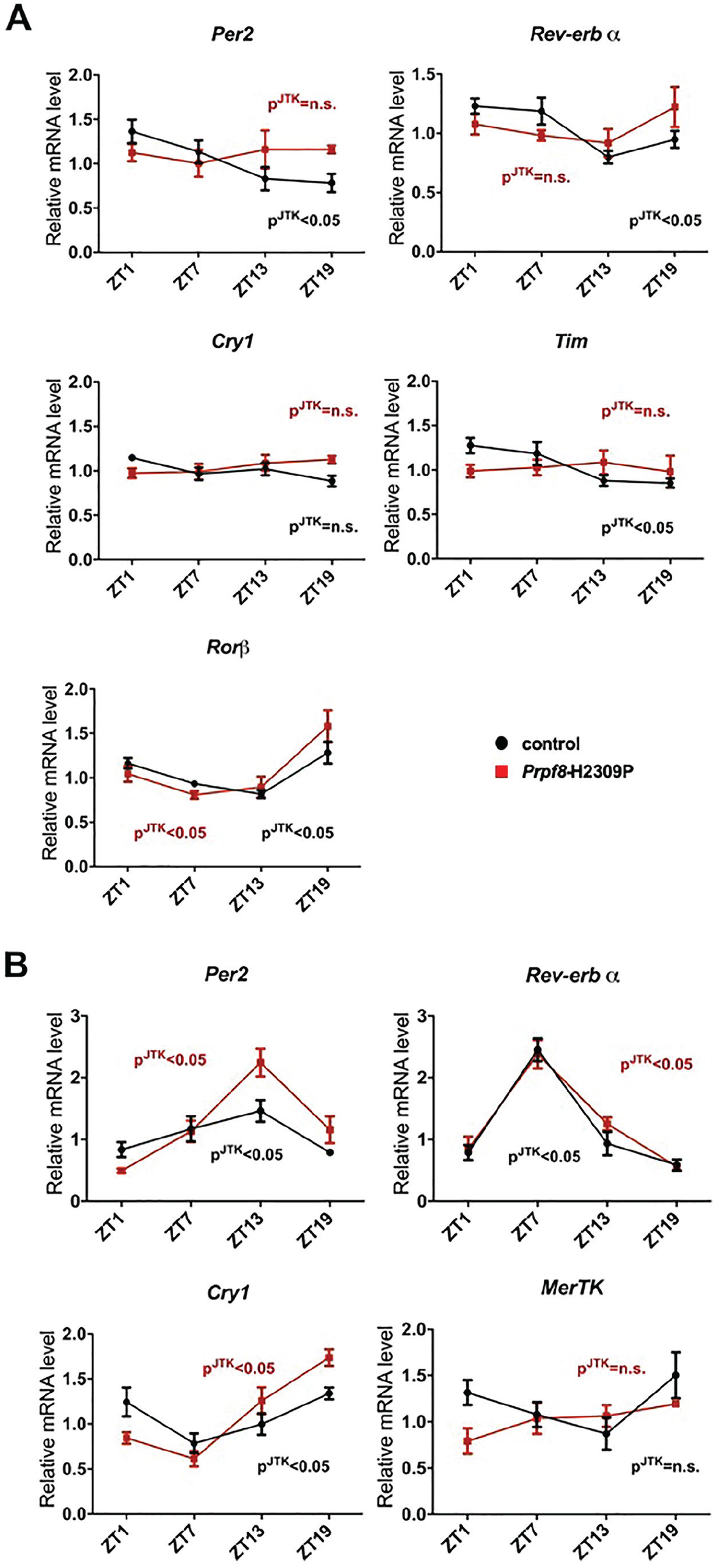
Daily cycling of retinal transcripts is dampened in Prpf8-H2309P mice. Mice were entrained for 3 weeks in 12-h light:12-h dark conditions, and then retinas (A) and eye cups (B) were harvested at indicated ZT time points. Quantitative polymerase chain reaction experiments were performed in triplicate, normalized to 36B4, and analyzed using the ΔΔCt method. pJTK indicates cycling as assessed by JTK cycle analysis for control mice (black) and homozygous Prpf8-H2309P mutants (red). Each time point represents data collected from 3 to 7 eyes across 2 experimental runs. JTK output files are available in Supplemental File 1.
Our data suggest that Prpf8 has a tissue-specific effect on daily gene cycling in the eye as cycling in the eye cup was just as robust in the RP mutant mice as in their sibling controls. In fact, some transcripts, such as Per2 and Cry1, displayed an even more robust cycling in the RP mutant mice compared with the controls (Fig. 2B). Expression of MerTK, a gene implicated in the phagocytosis of photoreceptor outer segments (Law et al., 2015), did not cycle in either Prpf8-H2309P or control mice (Fig. 2B).
Human Prpf31 Sets Circadian Period
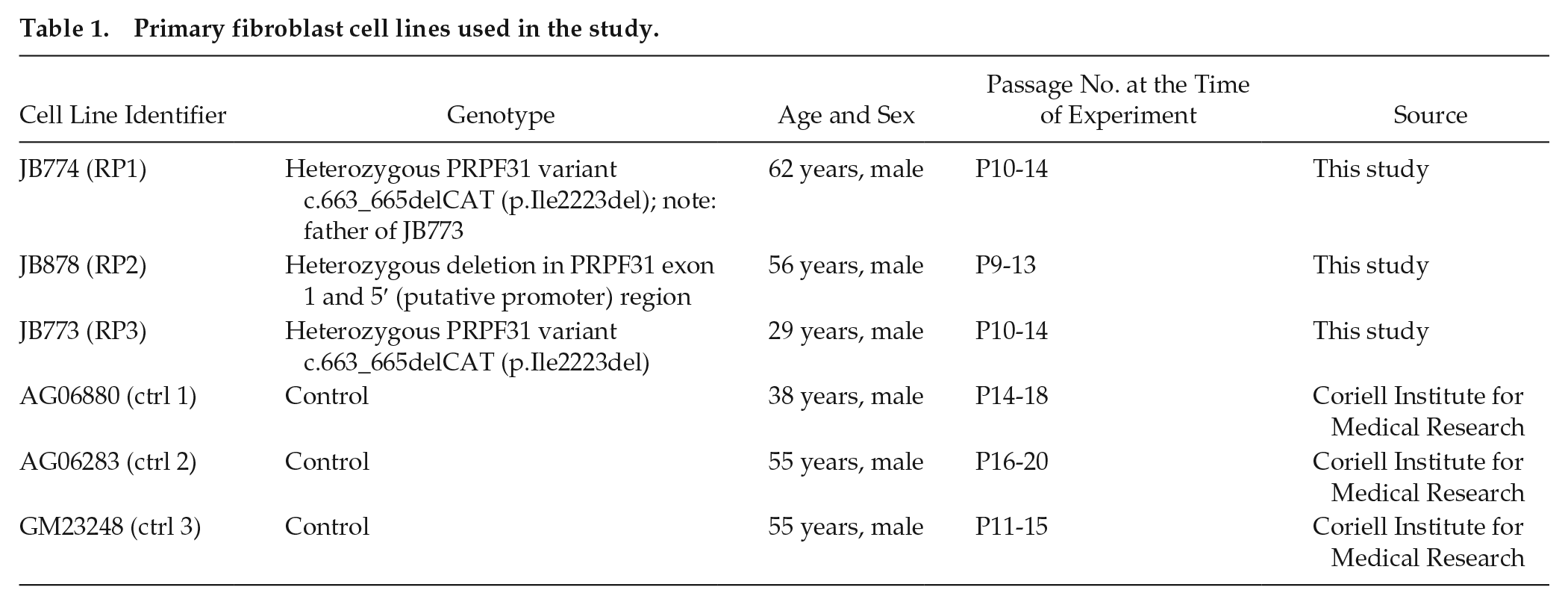
Because mouse models of RP do not reflect all the intricacies of the human disease (Graziotto et al., 2011), we sought to use human models to address the role of Prp factors implicated in RP in circadian rhythms. Our initial goal was to obtain cells from patients with RP and develop primary cultures to assay rhythms. Unfortunately, recruitment of RP patients with verified mutations in splicing factor genes is not trivial, because these individuals represent a small subset of the RP-affected population. As mutations in PRPF31 are some of the most represented among this small group (Bowne et al., 2008; Van Cauwenbergh et al., 2017), we were able to identify 3 patients heterozygous for a PRPF31 mutation (Table 1) and obtain skin samples from which we cultured fibroblasts. We infected primary fibroblast cultures with a recombinant lentivirus carrying a Bmal1::luciferase construct (Brown et al., 2005) and assayed circadian expression of the luciferase reporter. We report that the period of RP samples was not significantly different from that of the control primary fibroblast lines derived from skin biopsy (Suppl. Fig. S1).
Primary fibroblast cell lines used in the study.
The patient study was limited by the small numbers we were able to obtain, and so we considered other approaches toward manipulating Prp in human cells. Human osteosarcoma cells (U2OS) stably expressing a pBmal1-luciferase (pBmal1-dLuc) reporter (Ramanathan et al., 2012) provide an effective model to examine rhythms via cycling of luciferase activity. Given that haploinsuffiency is a proposed mechanism for PRPF31 involvement in RP (reviewed in Rose and Bhattacharya, 2016), we chose to downregulate PRPF31 in U2OS cells and determine the resulting effect on circadian parameters. First, we verified that siRNA against PRPF31 knocked down levels of PRPF31 protein by conducting western blot analysis (Fig. 3C). Next, we assayed rhythms of the luciferase reporter and found that siRNA knockdown of PRPF31 lengthens the bioluminescence rhythm as compared with the treatment with scrambled siRNA control (Fig. 3A, B). The knockdown of CRY2 served as a positive control and, as expected, significantly lengthened the period length (Fig. 3A, B; e.g., as used in Zhang et al., 2009). Together, these data indicate that PRPF31 is an important modulator of circadian period in humans.
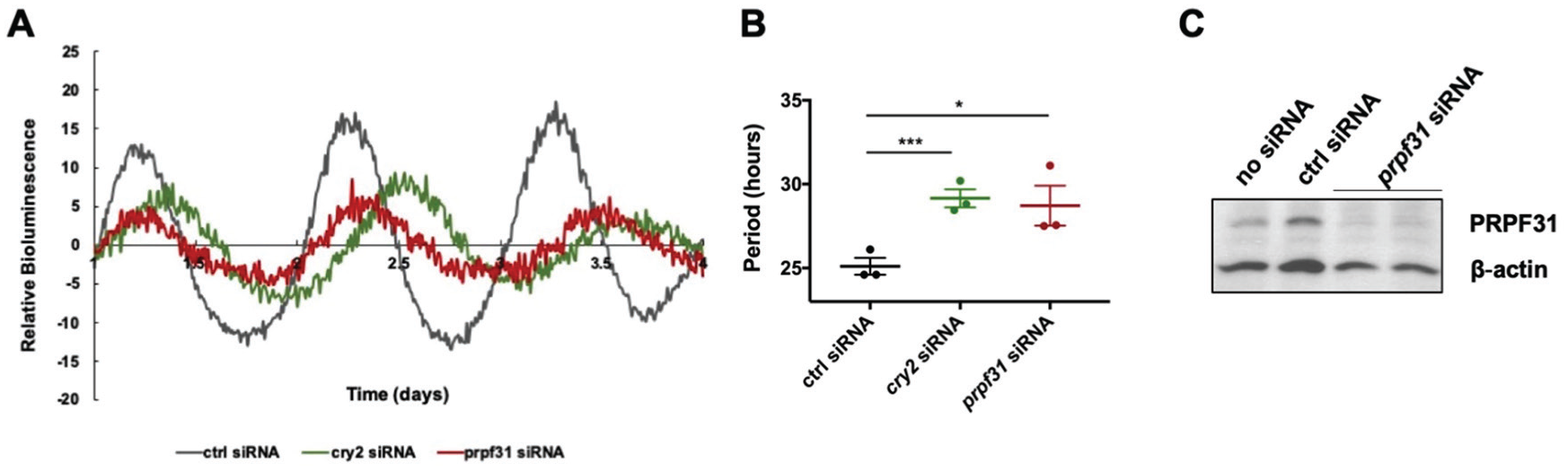
Human Prpf31 sets circadian period. Prpf31 downregulation in human U2OS cells lengthens the circadian period. U2OS cells stably expressing a BMAL1 promoter-driven destabilized luciferase (pBmal1-dLuc) were reverse transfected with targeted siRNAs, synchronized with dexamethasone (dex) 24 h posttransfection, and bioluminescence was monitored in real time. Relative bioluminescence signal (baseline subtracted) is plotted over days 2 to 5 (first day after dexamethasone treatment was excluded). *p < 0.05, ***p < 0.001 to ctrl siRNA treatment as calculated by a paired Student t test. Error bars = ±SEM, n = 2-3 biological replicates.
Discussion
In this study, we reveal a novel circadian phenotype of Prpf8-H2309P mice, which is characterized by lengthened period of the circadian wheel-running activity rhythm and dampening of daily gene expression in the retina (Figs. 1-2). Prpf8 is a key component of U5 snRNP and U4/U6.U5 tri-snRNPs that serve to assemble the active spliceosome (Grainger and Beggs, 2005). The circadian role of mammalian Prpf8 is intriguing in light of our recent report on the circadian role of U4/U6.U5 tri-snRNP complex in Drosophila melanogaster (Shakhmantsir et al., 2018). We propose that the U4/U6.U5 tri-snRNP components are evolutionarily conserved regulators of circadian clocks.
Human mutations in Prpf8 lead to autosomal dominant RP (McKie et al., 2001; Towns et al., 2010). Since Prpf8 is ubiquitously expressed, it remains unclear why the RP-causing mutations seem to exclusively affect retinal health (Ruzickova and Stanek, 2017). Multiple ideas exist about the role of U4/U6.U5 tri-snRNPs in the retina. First, some studies indicated that the expression of U4/U6.U5 components is particularly high in the retina (Tanackovic et al., 2011; Cao et al., 2011), suggesting a particularly heavy load of spliceosomal processing. Second, there could be unique eye-specific targets of U4/U6.U5 tri-snRNPs that are sensitive to decreases in splicing efficiency tolerated in the rest of the body (Tanackovic et al., 2011; Linder et al., 2011). Third, some tri-snRNP components, including Prpf8, are associated with ciliary dysfunction, which is implicated in RP pathogenesis (Buskin et al., 2018; Wheway et al., 2015).
Our study reveals that an RP-causing H2309P mutation in Prpf8 disrupts daily gene expression in the retina, suggesting circadian dysregulation as a novel retinal phenotype of RP. Previous studies of homozygous Prpf3-T494M and Prpf8-H2309P and heterozygous Prpf31+/− mice focused on the RPE as the primary site of initial RP pathology (Graziotto et al., 2011). While the late-onset degenerative RPE phenotype is first recorded in mice at only ~2 years of age, functional changes, in particular affecting the daily rhythm in phagocytosis, are already present in RPE isolated from 2-week-old homozygous Prpf8-H2309P mice (Farkas et al., 2014). Because RP caused by mutations in splicing factors is associated with RPE abnormalities (Graziotto et al., 2011; Farkas et al., 2014), we hypothesized that disruption of the local circadian rhythm in the RPE would underlie the disrupted rhythm of outer segment phagocytosis. Contrary to our hypothesis, we found that the cycling of gene expression in the retina, and not the RPE, was compromised in Prpf8-H2309P mice (Fig. 2A). We propose that the retinal clock coordinates the timing of photoreceptor outer segment shedding, which drives the rhythm of RPE phagocytosis. It is possible that our method of assaying RPE rhythms as part of the entire eye cup provides insufficient RPE enrichment, and so dysregulated rhythms in the RPE are masked by intact rhythms in other parts of the eye cup. In the future, it may be worth assaying rhythms specifically in the RPE of Prpf8 mutants, perhaps with histological methods. In addition, in this study, we measured transcript cycling in Prpf8-H2309P mice only under light:dark conditions and not in constant darkness. Under these conditions, it is possible that the dampened cycling is caused by deficits in circadian entrainment rather than in free-running cycling. It is unlikely, though, that the deficits trace to the visual system, as these mice are visually normal (Graziotto et al., 2011).
The disruption of tri-snRNP spliceosome factors tends to prolong the circadian period in flies (multiple tri-snRNP components; Shakhmantsir et al., 2018), human osteosarcoma lines (Prpf4, Prpf31; Zhang et al., 2009; this study), and mice (Prpf8; this study). Thus, we conclude that Prp factors have broad and well-conserved roles in circadian rhythms. Our patient fibroblast study suggests that this role may extend to an effect on periodicity in a disease context, but we are aware of the limitations of this study (Suppl. Fig. S1). Ideally, we would have liked to use samples from the unaffected noncarrier family members as controls, but such arrangements were impossible at the time of our experiment, and so we used unaffected age-matched controls from the publicly accessible tissue collections. Circadian period varies considerably in human fibroblasts (Brown et al., 2005), which we noticed also in our study and which may have contributed to the lack of statistical significance. Interestingly, the period was quite tightly clustered in the 3 patient-derived cell cultures and was affected in the opposite direction from PRPF31 knockdown. PRPF31 mutations are generally thought to be loss of function (reviewed in Rose and Bhattacharya, 2016), but the lesions studied here have not been completely characterized and so could affect the circadian clock in a specific way. While the number of patients available is small, we note that this is often a limitation of translational studies, and so validation of laboratory findings in human subjects sometimes relies on only 1 to 2 patients (for example, Han et al., 2013). Whether or not circadian rhythms in the fibroblasts of RP patients are consistently affected, circadian rhythms in the retina might still be impaired. In that case, one could use fibroblasts to derive induced pluripotent stem cells, which could then be differentiated into ocular tissues, transfected with clock luciferase reporters, and tested for circadian rhythm defects (Reichman et al., 2014; Rathod et al., 2019).
In mammals, photoreceptors and RPE cells are postmitotic, which means their functions need to remain efficient throughout the life of an organism for retinal function to remain optimal. Aging causes dampening of circadian cycles in the eye, particularly in the retina (Baba and Tosini, 2018). Interestingly, many retinal diseases, including RP, arise later in life, when circadian function might be compromised (Baba et al., 2018). For example, mice with global developmental deletion of Bmal have normal photoreceptors at 2 to 3 months of age (Storch et al., 2007), but a recent study demonstrates that photoreceptor viability is impaired in older 8-month-old Bmal-/- mice (Baba et al., 2018). Clock/Npas2 knockout mice, similarly to Bmal-/- mice, display photoreceptor deficits as mice age (Baba et al., 2018). Some of the deficits reported in this study are consistent with the aberrant refractive development described recently for clock mutants in Drosophila and mice (Stone et al., 2019). The specific defect in refractive development is indicative of myopia, implicating circadian function in visual acuity. The importance of cycling in the inner retina was also demonstrated by another recent study, which identified circadian and BMAL1 regulation of a major inner blood-retina barrier (iBRB) tight junction component (Hudson et al., 2019). The authors speculate that the circadian permeability of the iBRB contributes to the daily clearance of spent photoreceptors as well as their overall health and substrate availability. Aberrant cycling of claudin-5 in the course of aging could contribute to age-related macular degeneration characteristics, such as drusen accumulation and RPE atrophy.
It is clear that circadian rhythms are important for retinal health. While perturbations of core clock components often do not lead to drastic retinal phenotypes in healthy animals (Ait-Hmyed et al., 2013; Selby et al., 2000), it is possible that circadian misregulation synergizes with other factors known to modify retinal health, such as aging, bright light, and genetic retinal diseases. We speculate that circadian dysregulation in the eye, which arises in the Prpf8 mice well before any signs of retinal degeneration, could contribute to disease progression and pathology.
Supplemental Material
JBR887876_Supplementary_figure_01_CLN – Supplemental material for RNA Splicing Factor Mutations That Cause Retinitis Pigmentosa Result in Circadian Dysregulation
Supplemental material, JBR887876_Supplementary_figure_01_CLN for RNA Splicing Factor Mutations That Cause Retinitis Pigmentosa Result in Circadian Dysregulation by Iryna Shakhmantsir, Scott J. Dooley, Siddharth Kishore, Dechun Chen, Eric Pierce, Jean Bennett and Amita Sehgal in Journal of Biological Rhythms
Supplemental Material
JTK.outputs – Supplemental material for RNA Splicing Factor Mutations That Cause Retinitis Pigmentosa Result in Circadian Dysregulation
Supplemental material, JTK.outputs for RNA Splicing Factor Mutations That Cause Retinitis Pigmentosa Result in Circadian Dysregulation by Iryna Shakhmantsir, Scott J. Dooley, Siddharth Kishore, Dechun Chen, Eric Pierce, Jean Bennett and Amita Sehgal in Journal of Biological Rhythms
Footnotes
Acknowledgements
We would like to acknowledge the Neurobehavior Testing Core at the University of Pennsylvania, in particular Dr. W. Timothy O’Brien and Brianna Ciesielski, whose help with mouse colony maintenance was indispensable to the success of this project. We thank Dr. Joseph Bedont for his assistance with the assembly of the wheel-running setup. We thank Dr. Steven Brown for his pABpuro-BluF construct and Dr. Andrew Liu for pBmal1-dLuc construct. Finally, we thank Dr. Joshua Dunaief and his lab for their advice on mouse eye dissections. Support was provided by HHMI, NIH grant R37NS048471-14, RO1 EY020902, Brenda and Matthew Shapiro Stewardship, and the Robert and Susan Heidenberg Investigative Research Fund for Ocular Gene Therapy.
Conflict of Interest Statement
The authors have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
