Abstract
Rhodopsin 7 (Rh7), a new invertebrate Rhodopsin gene, was discovered in the genome of Drosophila melanogaster in 2000, but its function has remained elusive. We generated an Rh7 null mutant (Rh70) by P element–mediated mutagenesis and found that an absence of Rh7 had significant effects on fly activity patterns during light-dark (LD) cycles: Rh70 mutants exhibited less morning activity and a longer siesta than wild-type controls. Consistent with these results, we found that Rh7 appears to be expressed in a few dorsal clock neurons that have been previously implicated in the control of the siesta. We also found putative Rh7 expression in R8 photoreceptor cells of the compound eyes and in the Hofbauer-Buchner eyelets, which have been shown to control the precise timing of locomotor activity. The absence of Rh7 alone impaired neither the flies’ responses to constant white light nor the ability to follow phase shifts of white LD cycles. However, in blue light (470 nm), Rh70 mutants needed significantly longer to synchronize than wild-type controls, suggesting that Rh7 is a blue light–sensitive photopigment with a minor contribution to circadian clock synchronization. In combination with mutants that lacked additionally cryptochrome-based and/or eye-based light input to the circadian clock, the absence of Rh7 provoked slightly stronger effects.
The adequate timing of daily activity is essential for all animals. Light is regarded as the most important factor that synchronizes the animal circadian clock with the 24-h solar day. This light synchronization is mediated by both visual and dedicated “circadian” photoreceptors. In Drosophila, both the visual photoreceptors in the compound eyes and ocelli, as well as the blue light photoreceptor cryptochrome (CRY) expressed within the brain clock neurons, contribute to clock synchronization by light (Rieger et al., 2003; Stanewsky et al., 1998; Emery et al., 2000; Fogle et al., 2015; Schlichting et al., 2014). In addition, specialized photoreceptive structures located between the retina and optic lobes, the Hofbauer-Buchner (H-B) eyelets, project to the clock neurons in the brain and participate in light synchronization (Helfrich-Förster et al., 2002; Malpel et al., 2002; Veleri et al., 2007; Schlichting et al., 2016). The photoreceptors of the compound eyes, ocelli, and H-B eyelets contain rhodopsins. Upon light excitation, the rhodopsins activate the canonical invertebrate phototransduction cascade in which the Phospholipase C-β (PLC-β) encoded by the norpA gene is a crucial enzyme (Bloomquist et al., 1988). Therefore, the combination of norpA loss-of-function mutants with cry mutants leads to severe decrements in light synchronization (Emery et al., 2000; Klarsfeld et al., 2004; Mealey-Ferrara et al., 2003; Stanewsky et al., 1998). Nevertheless, such mutants are still able to synchronize their activity to light-dark (LD) cycles, indicating that rhodopsins can additionally signal through norpA-independent pathways, as recently shown for Rh5 and Rh6 (Szular et al., 2012). In addition, the presence of an additional norpA-independent rhodopsin in the dorsal brain was predicted from studies showing that flies lacking compound eyes, ocelli, H-B eyelets, and CRY still show responses to light (Rieger et al., 2003; Helfrich-Förster et al., 2001; Veleri et al., 2003). The photopigment in question could be rhodopsin 7 (Rh7), the existence of which was predicted from the genome in 2000 (Adams et al., 2000; Terakita, 2005). Rh7 is present in most groups of arthropods, even in ancient ones such as Limulus polyphemus, but it is absent from arthropods living in dark environments (Senthilan and Helfrich-Förster, 2016). This suggests that Rh7 has a conserved function in light signaling. Nevertheless, Rh7 differs from other rhodopsins in several aspects such as unusually long extracellular and cytoplasmic tails and the lack of the G protein–binding QAKK motif (Senthilan and Helfrich-Förster, 2016). This suggests that it is rather involved in nonvisual light signaling; for example, it might play a role in basic insect circadian photoreception as melanopsin does in mammals.
The aim of the present study was to investigate a potential function of Rh7 in circadian photoreception. For this purpose, we investigated the expression of Rh7 in the brain and retina of the fly, generated an Rh7 knockout mutant, and monitored its activity pattern under different light intensities and qualities. We also generated double mutants that lacked Rh7 and CRY or Rh7 and PLC-β as well as triple mutants that lacked Rh7, CRY, and PLC-β and investigated their capability to synchronize to LD cycles and to follow phase shifts of the LD cycles. In addition, we determined the responses of some of these mutants to constant light (LL) that renders wild-type flies arrhythmic but does not disturb rhythmicity in cry and other mutants that block the light-dependent degradation of the clock protein Timeless (Stanewsky et al., 1998; Emery et al., 2000; Busza et al., 2004; Peschel et al., 2006; Chen et al., 2011; Buhl et al., 2016).
Materials and Methods
Generation and Characterization of an Rh7 Knockout Mutant
An Rh7 knockout mutant was generated by P element–mediated mutagenesis. The starting point was a transgenic line carrying a P-element insertion 147 bp upstream of the first noncoding exon of the Rh7 gene (Bellen et al., 2004). We obtained this insertion line, y− w−;;P{EPgy2}EY13118 (BL21406), from the Bloomington Drosophila Stock Center (Indiana University, Bloomington, IN, USA). Remobilization of the P-element by crossing these flies to a transposase strain (y− w−;+;KiΔ2-3) resulted in a precise excision (y− w− revertant) and a deletion line (y− w−;+;Rh70). The breakpoint was mapped by polymerase chain reaction (PCR) using a forward primer upstream of the P-element insertion (5′-GCCTTGGCAAACATGAGTCC-3′) in combination with different downstream reverse primers in exons 1, 2, 3, and 4, respectively. The breakpoint localized within a 1.36-kb genomic DNA fragment to the noncoding region of exon 4 (reverse primer: 5′-AAGGGGCGGCCATCACAATACTG-3′). Sequencing and alignment with the Rh7 sequence revealed that the deletion comprises 10.35 kb and extends over the entire Rh7 coding sequence, thereby providing an Rh7 knockout mutant (Fig. 1A). In flies homozygous for the deletion, no Rh7 transcript was detected by quantitative PCR using primers flanking the transcription start site (5′-TCATCAAATGCCCGATTG-3′ and 5′-GCACCACCACATTGTACCG-3′). Both the y− w−;+;Rh70 mutant and the revertant were crossed into a red-eyed (y− w+) background to exclude side effects on the synchronization of locomotor activity rhythms due to different pigmentation. Depending on the experiment, Rh70 mutants and revertant flies were used either in the white-eyed (y− w−) or red-eyed (y− w+) background.
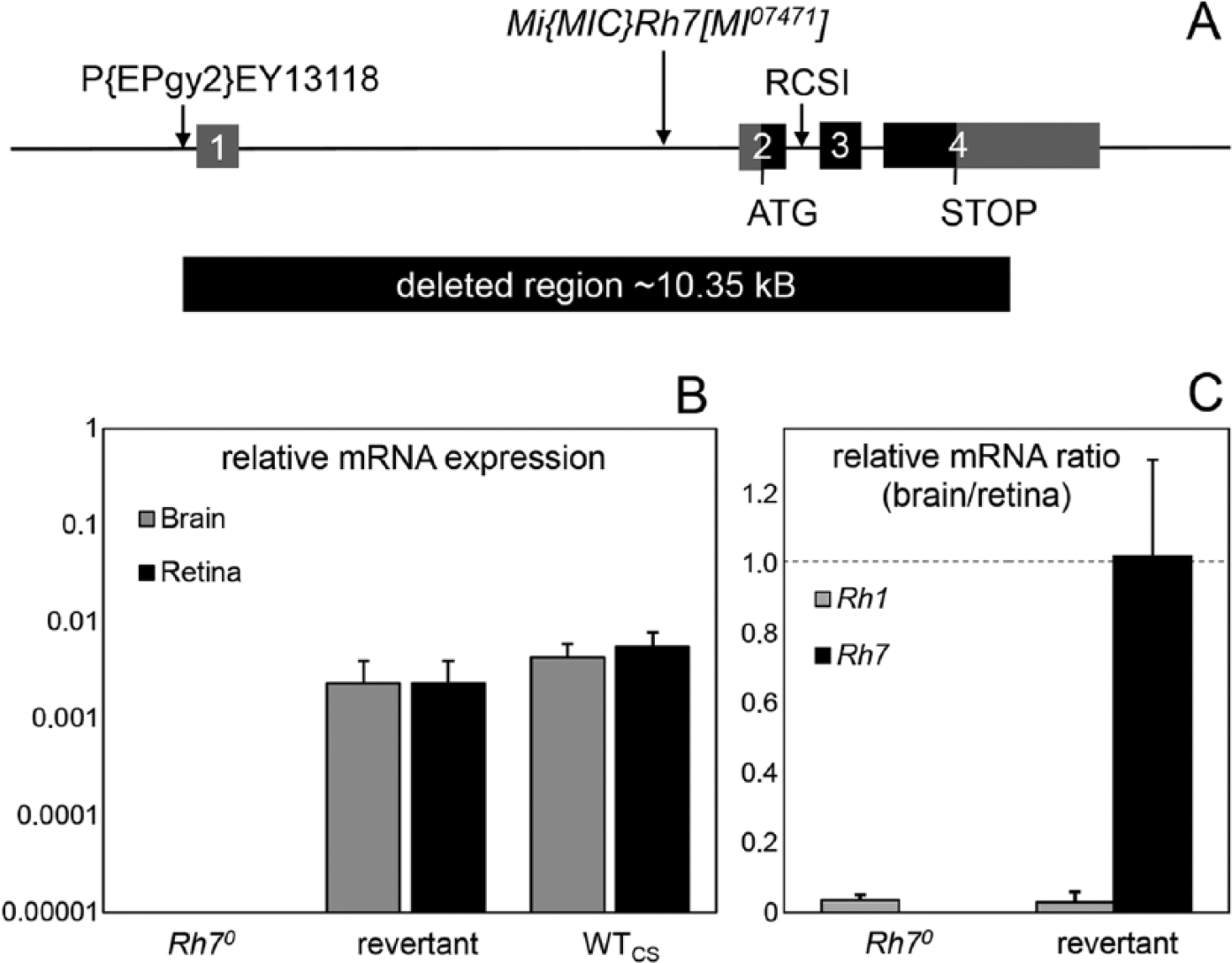
Genomic organization and expression levels of Rh7. (A) Genomic organization of the wild-type and mutant Rh7 locus. The Rh7 gene (CG5638) lies on the left arm of chromosome 3, spans 11.3 kb, and possesses 4 exons (boxes), of which the coding regions are shown in black and the noncoding regions in gray. The GFP insertion of the mimic line is located in the first intron, and the RCSI motif is located in the second intron. Rh70 mutants were generated by P element–mediated mutagenesis. The short arrow indicates the position of the P-element P(EPgy2)EY13118 in the 5′ untranslated region of the Rh7 gene prior to its excision. The approximately 10.35-kb deletion in the Rh7 gene after the imprecise excision of the P-element is indicated by the black box below. (B) Relative Rh7 mRNA expression levels (±SD) in retinas and brains of Rh70 mutants, revertants, and wild-type flies (WTCS) revealed by quantitative PCR. Rh7 is totally absent in Rh70 mutants and expressed at low levels in the brain and retina of revertants and wild-type flies. (C) Relative mRNA ratio (brain/retina) of Rh1 and Rh7 in Rh70 mutants and revertants revealed by quantitative PCR (±SD). Rh1 is predominantly expressed in the retina and marginally in the brain (brain/retina ratio of less than 0.1). In contrast, Rh7 is equally expressed in the brain and retina of revertants (brain/retina ratio close to 1), but it is absent in Rh70 mutants.
Double/Triple Mutants and Other Fly Strains
To generate Rh70 cry01 double mutants, recombination was necessary because both genes are located on the third chromosome. Mass crosses between y+ w−;+;P[w+]cry01 females (Dolezelova et al., 2007) and y− w−;+;Rh70 males were performed. The female offspring (including potential recombinants) were mated to w; Sp/CyO; MKRS/TM6B males, and 330 single crosses were carried out to recross P[w+]/TM6B progeny to double balancer flies. Successful recombination was confirmed by sequencing (LGC Genomics, Berlin, Germany). We obtained 2 recombinant lines, #39 and #112, that had a highly similar activity pattern under all investigated conditions. If not stated otherwise, we show here actograms of line #112. Line #112 was also used to generate norpAP41;+; Rh70 cry01 triple mutants using standard genetic crosses. norpAP41 is a loss-of-function allele that leads to the removal of the catalytic domain of the norpA-encoded PLC-β (Szular et al., 2012).
For investigating Rh7 gene expression in the brain and retina, we used a gfp reporter strain that carried gfp in the correct orientation with regard to the Rh7 gene in its first intron (Venken et al., 2011) (Fig. 1A). The relevant reporter strain, y− w−;+;Mi{MIC} Rh7MI07471 (BL43712), was obtained from the Bloomington Drosophila Stock Center.
Quantitative PCR
The relative mRNA levels of Rh1 and Rh7 in different fly strains were quantified via quantitative PCR, as described by Senthilan and Helfrich-Förster (2016). Total RNA was extracted from the brains and retinas of 3 to 4 flies per strain using the Quick-RNA MicroPrep Kit (Zymo Research, Irvine, CA, USA) and reversely transcribed using the QuantiTect Reverse Transcription Kit (Qiagen, Hilden, Germany). Quantitative PCR was then carried out with the SensiFAST SYBR No-ROX Kit (Bioline Reagents, London, UK) in combination with the Rotor-Gene Q (Qiagen) and 0.1-µM PCR primers. For each strain and tissue, 3 biological replicates were examined, and for each replicate, 2 PCRs were run. The relative mRNA levels were calculated using the ΔCt equation, and α-tubulin was used as the reference gene.
The following genes and primers (sequences 5′-3′) were used:
Rh1: GGAGTAGAAGATCAGGTATGAGCGTG and TGCCTACATCTGGTTCATGTCGAGC;
Rh7: CATCTGCGACTTTCTGATGCTCATC and GGATGCACCACCACATTGTACCGATC; and
α-tubulin: TCTGCGATTCGATGGTGCCCTTAAC and GGATCGCACTTGACCATCTGGTTGGC.
Recording of Locomotor Activity Rhythms
Locomotor activity recordings were performed at 3 different places with different systems. The majority of the experiments were conducted at the University of Regensburg using a homemade Drosophila recording system in which the flies were housed individually in photometer cuvettes (Helfrich-Förster, 1998) and illuminated with a light-emitting diode (LED)–based lighting system (Rieger et al., 2007). The following LEDs were used: white LEDs, 400-nm LEDs, and 470-nm LEDs (Lumitronix LED-Technik GmbH, Jungingen, Germany). The LL experiments were performed at the University of Würzburg using the Drosophila Activity Monitor (TriKinetics, Waltham, MA, USA) (Schlichting et al., 2015). The third set of experiments was performed at University College London (UCL) also using the Drosophila Activity Monitor and white LEDs (5050 surface-mount device cool white LED strips, 3000-4000 K).
The experiments at the Universities of Regensburg and Würzburg were performed at a constant 20 °C and the ones at UCL at 25 °C. In all experiments, flies of the different genotypes were recorded simultaneously to ensure comparability of the data. Each experiment was repeated at least twice.
LD cycles of different light intensities (University of Regensburg)
Previous studies have shown that the siesta of fruit flies increases with the light intensity (Rieger et al., 2007). In order to see whether Rh7 affects the light dependency of the siesta, Rh70 mutants and revertants were recorded at 10 lux (6 µW/cm2), 100 lux (70 µW/cm2), and 1000 lux (700 µW/cm2) (always 12-h:12-h LD cycles).
Phase shift experiments (University of Regensburg)
The flies were first recorded for 5 to 7 days in 12-h:12-h LD cycles (colored or white light) that had the same phase as the LD cycle under which the flies were reared (light intensity is indicated in each experiment). On day 8, the LD cycle was either advanced or delayed by 6 h, and the flies were recorded until they had re-entrained to the new light regime. Afterwards, they were transferred to constant darkness (DD) to assess their free-running behavior. The number of days needed for resynchronization was assessed by eye on coded actograms of individual flies by 7 members of the laboratory. An average was calculated out of these 7 values.
Phase shift experiments (UCL)
Flies were first recorded for 4 days in 12-h:12-h LD cycles (177-lux [62-µW/cm2] white light), during which the lights were ramped linearly for 2 h after lights-on and before lights-off, respectively. This LD cycle was in the same phase as the one in which the flies were reared. To ensure entrainment of potentially circadianly blind flies, a 25 °C:16 °C temperature cycle was superimposed on the initial LD cycle (warmer temperature during the light portion and cooler temperature during the dark portion of the LD cycle). The LD cycle was then delayed by 6 h, and flies were kept in this new cycle (same ramping, same intensity, constant 25 °C) for 6 days before being released to DD for another 4 days. The number of days required for resynchronization to the shifted LD cycle was determined, as described by Szular et al. (2012).
LL experiments (University of Würzburg)
Flies were first recorded for 5 to 7 days in 12-h:12-h LD cycles and then for 11 days in DD and finally in LL. The light intensity was 1000 lux (700 µW/cm2).
Data Analysis and Statistics
Actograms were plotted using the Fiji plugin ActogramJ (Schmid et al., 2011), and average actograms and activity profiles were calculated, as described by Schlichting et al. (2015) and Schlichting and Helfrich-Förster (2015). The offset of morning activity (M offset) and the onset of evening activity (E onset) were determined manually on the activity profiles of individual flies, as indicated in Supplementary Figure S1. Under DD and LL, the percentage of rhythmic and arrhythmic flies was determined, and in rhythmic flies, the free-running period was calculated in ActogramJ using the χ2 periodogram analysis of Sokolove and Bushell (1978). All results were expressed as mean ± SEM. Data were tested for normal distribution using the Kolmogorov-Smirnov test and further compared by analysis of variance, followed by a post hoc test with Bonferroni adaptation to determine significant differences (SYSTAT 11; Systat Software, Chicago, IL, USA). p < 0.05 was considered statistically significant. The Bonferroni multiple comparison test was applied for post hoc comparison.
Generation of Rh7 Antibodies
Three peptide antibodies against Rh7 were generated. The first one was raised against a 21-mer peptide in the intracellular domain (aa411-431: TRSSYMTRSRSSFTHRLRTST), the second one against an 18-mer peptide in the extracellular domain (aa54-71: TESSAVNVGKDHDKHVND), and the third one against a 25-mer peptide in the extracellular domain (aa86-110: YIHYRDKYDLSYIAKVNPFWLQFEP) of Rh7. A detailed description of antibody #1/#2 generation and characterization can be found in the work of Grebler et al. (2017). All 3 antibodies were generated in rabbits by different institutions/companies (antibody #1 [35-9185.82/977/99]: Regierungspräsidium Karlsruhe, Karlsruhe, Germany; antibodies #2 and #3: Pineda Antikörper-Service, Berlin, Germany). All 3 Rh7 antibodies were affinity purified using the original peptide bound to Sepharose 6B columns prior to use. Before immunization, serum samples were taken to obtain preimmune serum as a negative control for unspecific immunoreactivity. Dot blot analysis was used to confirm the selective binding of the Rh7 antibodies to the peptide used for immunization.
Immunohistochemistry
Preparation of whole-mount brains and immunostainings were performed, as described by Hermann et al. (2012). Primary antibodies were applied for 2 days at room temperature: polyclonal guinea pig anti-Vrille (VRI) (Glossop et al., 2003) at 1:3000, rabbit anti-Rh7 at 1:100, and chicken anti-GFP (ab13970; Abcam, Cambridge, UK) at 1:2000 for enhancing green fluorescent protein (GFP) in the y−w−;+;Mi{MIC} Rh7MI07471 line. Rabbit anti-CRY (Yoshii et al., 2004) and monoclonal mouse anti–pigment-dispersing factor (PDF) (Developmental Studies Hybridoma Bank, University of Iowa, Iowa City, IA, USA; depositor: Justin Blau) were applied at 1:1000 and 1:5000, respectively, on the y−w−;+;Mi{MIC} Rh7MI07471 line. Furthermore, rabbit anti–diuretic hormone 31 (DH31) (Kunst et al., 2014) was applied at 1:1000, together with mouse anti-PDF, on the y− w−;+;Mi{MIC} Rh7MI07471 line. Preparation of retinas was performed, as described by Hsiao et al. (2012), with the same incubation time of primary antibodies as for the whole-mount brains (mouse anti-Rh1 [Developmental Studies Hybridoma Bank, University of Iowa] at 1:100, mouse anti-Rh5 from Steve Britt [University of Colorado Denver, Denver, CO, USA] at 1:100, rabbit anti-Rh6 from Claude Desplan [New York University, New York, NY, USA] at 1:500, and chicken anti-GFP [Abcam] at 1:1000). In brains and retinas, fluorescent secondary antibodies (Alexa Fluor 488, 555, 635; Invitrogen, Carlsbad, CA, USA) were applied overnight. Immunostainings were visualized by confocal microscopy (TCS SPE; Leica, Wetzlar, Germany).
Results
Rh7 Gene Structure and Rh7 Knockout Mutants
The Rh7 gene comprises 11.3 kb and possesses 4 exons, of which 1 is fully and 2 are partly coding (Fig. 1A). As with other Drosophila opsin genes, Rh7 possesses a P3 (palindromic binding site for paired-class homeodomain transcription factors)/rhodopsin core sequence I (RCSI) motif that is important for transcription (Papatsenko et al., 2001). The conserved P3/RCSI motif of Rh7 is located within Rh7’s second intron (Fig. 1A). Rh7 is expressed at relatively low levels in the retina and brain of wild-type flies (Fig. 1B, C).
To investigate the function of Rh7, we generated an Rh7 knockout mutant by P element–mediated mutagenesis. Remobilization of the P-element resulted in a fly line with precise excision of the P-element (revertant) as well as a fly line with imprecise excision (y−w−;+;Rh7 mutant). In homozygous mutant flies, no Rh7 transcripts were detected by quantitative PCR, neither in the brain nor in the retina (Fig. 1B), whereas the expression of the downstream-located gene, CG9760, remained unaffected (not shown). The deletion was further characterized by the determination of the DNA breakpoint, revealing that it comprises approximately 10.35 kb and extends over the entire Rh7 coding sequence (Fig. 1A). Therefore, this mutant is a true knockout and in the following will be referred to as Rh70. The revertant line with precise excision of the P-element was used as a genetic control for the Rh70 knockout mutant.
Spatial Expression of Rh7
As shown by quantitative PCR, Rh7 seems to be expressed in the fly brain and retina (Fig. 1B). In order to reveal the spatial distribution of Rh7, we performed anti-Rh7 immunohistochemistry on whole-mount brains and retinas. Furthermore, we tested gfp expression in a transgenic fly line that carries the gfp reporter gene within the Rh7 gene (MIMIC line: y− w−; +;Mi{MIC} Rh7MI07471). All 3 Rh7 antibodies detected the peptide used for immunization on dot blots and in retinas that overexpressed Rh7 in the outer photoreceptor cells of the retina (Grebler et al., 2017). However, no Rh7 labeling was found in the brain (not shown) probably because Rh7 expression levels are below the detection limit of the antibodies.
In whole-mount brains of the MIMIC line, we found GFP staining in several small cells throughout the brain and particularly the optic lobes (Fig. 2). In addition, we found prominent staining in a few dorsal brain cell bodies that resembled the dorsal clock neurons of the fruit fly. Double labeling with an antibody against the clock protein VRI revealed that some of these cells are indeed clock neurons (Fig. 2A, B). Per hemisphere, the 2 dorsal clock neurons 1a (DN1a) and a mean 4.34 (±0.18) cells of the approximately 15 dorsal clock neurons 1p (DN1p) expressed GFP, whereas none of the lateral clock neurons were labeled. Double labeling with an antibody against CRY confirmed previous results, showing that the 2 GFP-positive DN1a cells were also CRY positive (Yoshii et al., 2004) (Fig. 2C). However, out of the 4 to 5 GFP-positive DN1p, only 1 contained CRY (Fig. 2D). Since it was shown recently that 4 to 5 DN1p express the peptide DH31, we also tested whether the Rh7-positive DN1p are DH31 positive. However, we found that this was not the case. There was no overlap between DH31-positive and Rh7-positive neurons (not shown). Thus, we conclude that the Rh7-positive DN1p are DH31 negative and that most of them are also CRY negative.
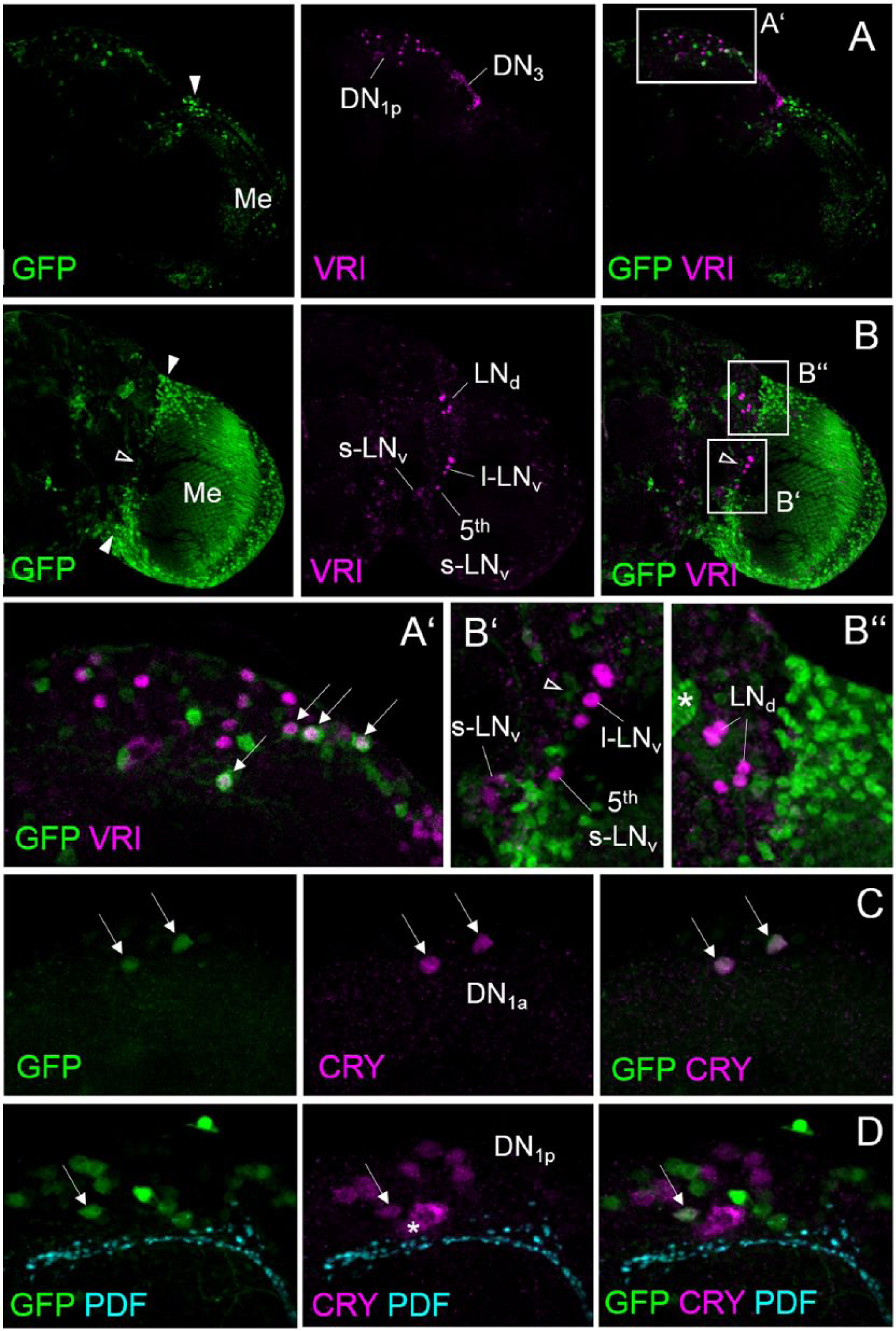
Rh7 is expressed in the DN1a and 4 to 5 DN1p. Whole-mount stainings of the right brain hemisphere of 2 brains carrying a gfp reporter insertion in the Rh7 gene (y− w−;;Mi{MIC}Rh7MI07471) are shown. In all images, Rh7 (=GFP) is visualized in green. The first brain is double labeled by an antibody against the clock protein VRI (in magenta) and depicted from the posterior (A) and anterior (B). The second one is triple labeled by antibodies against CRY (in magenta) and PDF (in cyan). (A) In the posterior dorsal brain, several cells are labeled by GFP, of which 4 belong to the DN1p. GFP staining was also seen in the posterior medulla (Me) and in several cells between the medulla and dorsal brain (arrowhead). The latter are close but not identical to the DN3. (A’) Magnification of the dorsal brain with the DN1p. Arrows point to 4 neurons that are double labeled by GFP and VRI. (B) In the anterior brain, many cells on the surface of the medulla and between the medulla and central brain (arrowheads) are labeled by GFP. Some of these cells are close to the lateral clock neurons (LNd, s-LNv, l-LNv, and 5th s-LNv), but there was no colabeling (the open arrowhead in B and B’ points to GFP-positive cells close to the l-LNv). (B’) Magnification of the area containing the s-LNv, l-LNv, and 5th s-LNv. The s-LNv and l-LNv were identified by anti-PDF staining (not shown). (B’’) Magnification of the area containing the LNd. The asterisk points to a labeled fiber tract of unknown origin. (C, D) Triple labeling of the dorsal neurons with Rh7 (=GFP), CRY, and PDF. Confocal sections of the same brain are shown; C is more anterior than D and shows the DN1a that are CRY and GFP positive (arrows), and D is more posterior and depicts the DN1p and the PDF terminals of the s-LNv. Only 1 DN1p contains GFP and CRY (arrow). The asterisk marks unspecific cytoplasmic CRY staining of a cell that does not belong to the DN1p.
In whole-mount retinas of the MIMIC line, we found strong GFP labeling in the fenestrated layer that is located between the retina and lamina, directly underneath the retina (Fig. 3A). The H-B eyelet that lies in the fenestrated layer at the median posterior margin of the retina was also labeled by GFP (Fig. 3A). Furthermore, weak GFP staining was present in R8 photoreceptor cells of the retina, predominantly in the Rh6-expressing R8 cells that comprise the majority of R8 cells (Fig. 3A-D).
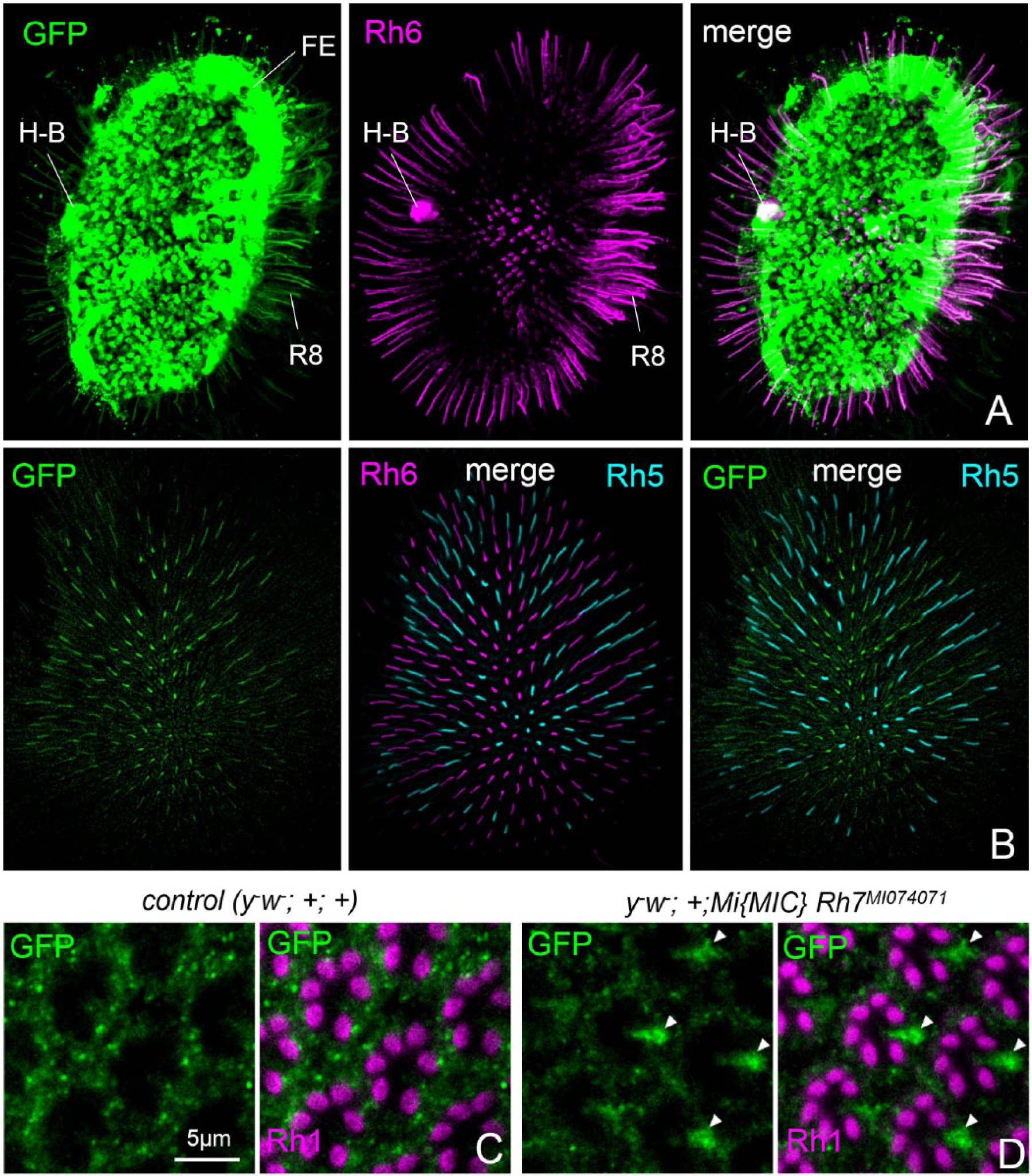
Rh7 appears present in the fenestrated layer (FE) underneath the retina, in the H-B eyelets, and weakly in R8 photoreceptor cells. (A) Rh6 and putative Rh7 expression in a whole-mount retina. Rh6 is visualized in magenta and Rh7 (=GFP) in green by a gfp reporter insertion in the Rh7 gene (y− w−;;Mi{MIC}Rh7MI07471). (B) Rh6 (in magenta), Rh5 (in cyan), and putative Rh7 (in green) expression in a whole-mount retina. Rh6 is present in about 70% of the ommatidia, whereas Rh5 is present in the remaining 30%. Rh7 appears to be colocalized with Rh6. (C) Higher magnification of the retina of y− w−;;Mi{MIC}Rh7MI07471 and control flies (y− w−;+;+) labeled for GFP as a reporter for Rh7 and with anti-Rh1 that marks the rhabdomeres of photoreceptor cells 1 to 6 in each ommatidium. About 7 ommatidia are depicted in each image. Arrowheads point to GFP staining in most R8 photoreceptor cells of y− w−;;Mi{MIC}Rh7MI07471 flies that is absent in the control flies.
Rh7 and CRY Suppress the Siesta with Additive Effects on the M Offset but Nonadditive Effects on the E Onset
Wild-type fruit flies show a bimodal activity pattern, meaning that they are mainly active in the morning and evening. Between morning and evening activity peaks, they keep a siesta, and the duration of this siesta increases with environmental temperature and light intensity (Majercak et al., 1999; Rieger et al., 2007). Based on our finding that Rh7 is expressed in a subset of the DN1p, which are implicated in regulating the siesta (Guo et al., 2016), as well as in R8 photoreceptor cells and H-B eyelets, which control the timing of morning and evening activity (Schlichting et al., 2015, 2016), we analyzed the activity profiles of Rh70 mutants under 12-h:12-h white LD cycles. We found that revertants and Rh70 mutants increased the siesta with increasing light intensity but that Rh70 mutants always kept a significantly longer siesta than the revertants (p < 0.01) (Fig. 4A). The longer siesta of Rh70 mutants was due to an earlier M offset, while the E onset was not changed in the mutants (Fig. 4A). cry0 mutants were reported to also show a longer siesta than wild-type flies (Rieger et al., 2003; Yoshii et al., 2004), but in contrast to Rh70 mutants, this seemed to be mainly due to a later E onset (Schlichting et al., 2015). In order to see how CRY and Rh7 interact in the timing of morning and evening activity, we recorded cry01 and Rh70 cry01 mutants under white light (700 µW/cm2) (Fig. 4B, C). We found that Rh70, cry01, and Rh70 cry01 mutants all had highly significant earlier M offsets than the controls (revertants) (p < 0.00001). The M offsets of Rh70 and cry01 mutants were not significantly different from each other (p = 0.306), whereas M offsets of Rh70 cry01 double mutants were significantly earlier than M offsets of both single mutants (p < 0.001 for Rh70 and cry01). Thus, CRY and Rh7 seemed to have additive effects on the M offset (Fig. 4C). For the E onset, this was different. As already found in the first experiment, E onsets of Rh70 mutants and controls were not different from each other (p = 1.000), but cry01 mutants had a significantly later E onset than all the other lines (p < 0.001) (Fig. 4C). Most interestingly, the E onset of the Rh70 cry01 double mutants was significantly earlier than that of all the other lines (p < 0.01). Thus, there were clearly no additive effects of CRY and Rh7 on the E onset. On the contrary, the lack of CRY and Rh7 advanced the E onset instead of delaying it (Fig. 4C). The differential effects of CRY and Rh7 on the M offset and E onset are also reflected in the length of the siesta. cry01 mutants exhibited the longest siesta that was significantly different from the siesta of all the other lines (p < 0.001). The siesta of Rh70 mutants was slightly longer than that of the controls (p = 0.049) but not different from the Rh70 cry01 double mutants (p = 1.000). The Rh70 cry01 double mutants also showed a significantly longer siesta than the controls (p = 0.012) and a significantly shorter one than cry01 mutants (p = 0.00001) (Fig. 4C). In spite of these significant differences in the activity pattern, all fly strains were normally rhythmic under DD with no significant differences in the period length.
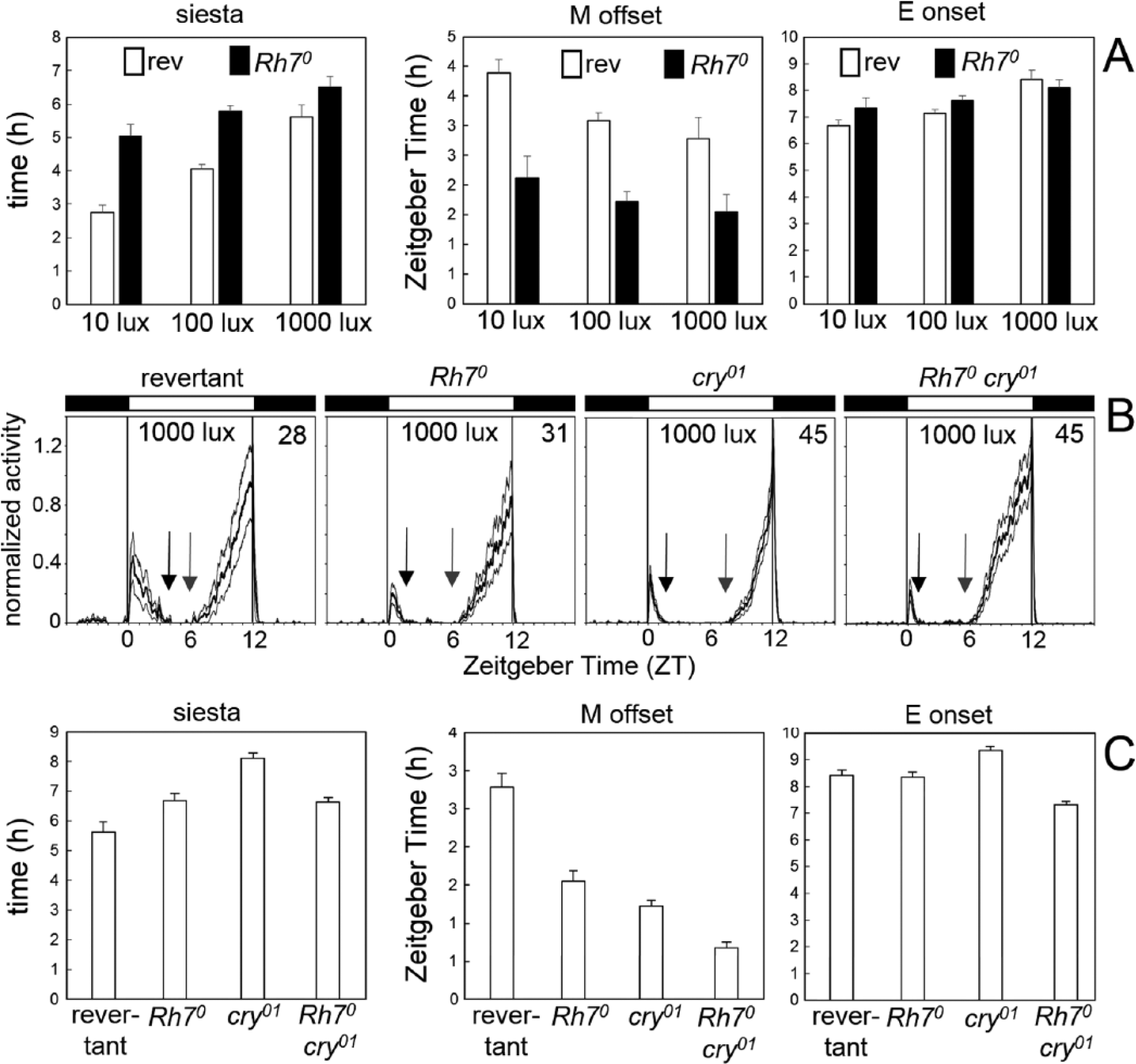
Rh7 affects morning activity and the siesta. (A) For revertants and Rh70 mutants, the length of the siesta is calculated together with the timing of M offset and E onset, dependent on the light intensity. Two-way analysis of variance revealed that the length of the siesta depended on the light intensity (F2,162 = 33.49; p < 0.000001) and genotype (F2,162 = 59.54; p < 0.000001). The same was true for the M offset (light: F2,162 = 14.63; p < 0.000001 and genotype: F2,162 = 139.81; p < 0.000001). In contrast, the E onset depended only on the light intensity (F2,162 = 23.01; p < 0.000001) and not on the genotype (F2,162 = 3.59; p = 0.06). (B) Average activity profiles of revertants and Rh70 mutants in LD (white light of 1000 lux) (number of included flies in the upper right corner). The gray lines above and below the mean (black) represent the SEM. The arrows point to the M offset and E onset, respectively, with the distance between the 2 arrows indicating the length of the siesta. (C) Calculated length of the siesta and the timing of M offset and E onset for revertants, Rh70, cry01, and Rh70 cry01 double mutants under white light of 1000 lux. For details and statistics, see the text.
Rh7 Does Not Contribute to Clock Synchronization to White Light
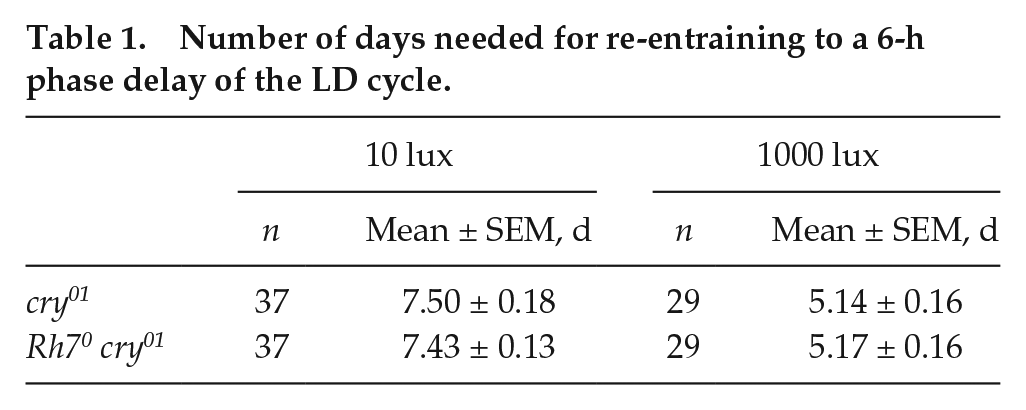
Photoreceptor mutants need longer to resynchronize to a shift of the LD cycle than wild-type flies. Therefore, phase shift experiments, also called jet lag experiments, are conventionally used to test whether a mutation affects photosensitivity of the circadian clock. Wild-type fruit flies are very light sensitive and usually resynchronize within 1 to 2 days to a 6- to 8-h advance or delay of the LD cycle, whereby the flies take a bit longer to resynchronize to phase delays (Kistenpfennig et al., 2012; Saint-Charles et al., 2016). In order to test if a photoreceptor contributes to circadian photosensitivity, one has to either use very low light intensities or combine the relevant photoreceptor mutant with others that are known to affect circadian photosensitivity such as cry mutants (Helfrich-Förster et al., 2001). Here, we did both: first, we measured the ability of Rh70 mutants and revertants to resynchronize to a 6-h delay of a dim LD cycle (0.001 lux [0.0005 µW/cm2]) (Fig. 5), and second, we crossed the Rh70 mutation into the cry01 mutant background and tested resynchronization to the same delay of the LD cycle but using higher light intensities (10 lux [6 µW/cm2] and 1000 lux [700 µW/cm2]). We found that the flies needed 7 to 8 days to resynchronize to the lower irradiance and approximately 5 days to resynchronize to the higher irradiance (Table 1 and Fig. 5). Nevertheless, there was no significant difference in the speed of resynchronization between the flies with and without Rh7, indicating that Rh7 does not or only marginally contributes to circadian photosensitivity. To further sensitize the system, we removed Rh7 function in the background of flies lacking visual system function and Cry (norpAP41; Rh70 cry0). Despite the lack of canonical rhodopsin signaling and Cry function, norpAP41; cryb double mutants slowly resynchronize their behavior to shifted LD cycles presumably because of norpA-independent light signaling to the clock by Rh5 and Rh6 (Szular et al., 2012). The norpAP41; Rh70 cry0 triple mutants required the same amount of time for resynchronization to 6-h delayed LD cycles (177 lux [62 µW/cm2]), demonstrating that Rh7 does not function in this norpA- and cry-independent pathway (Fig. 6).
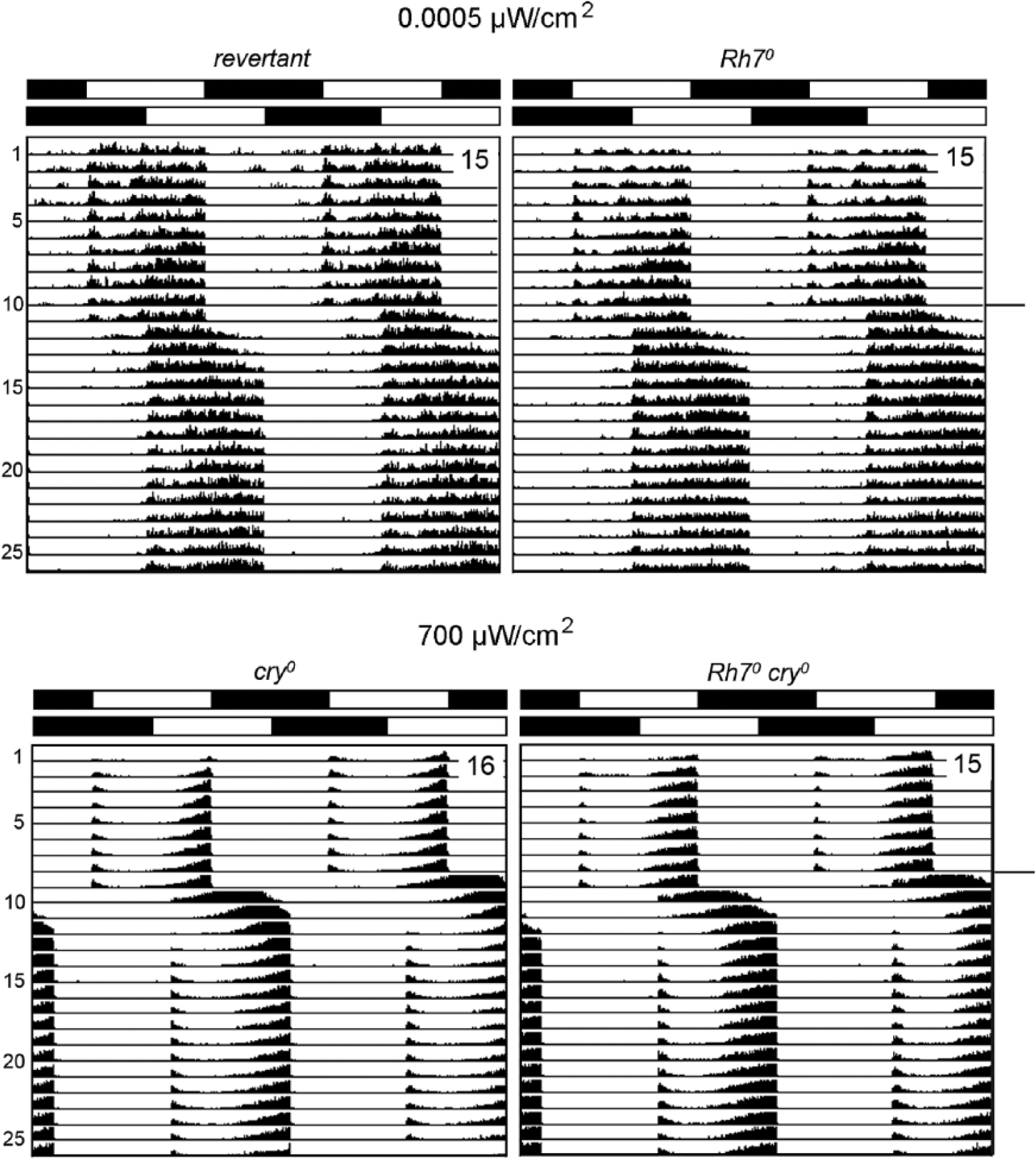
Resynchronization of revertants, Rh70, cry01, and Rh70 cry01 mutants to 6-h phase delays of white light. Double plots of average actograms are shown with the number of included flies indicated in the upper right corner of each plot. Only flies that lived until the end of the experiment are included. The LD cycle and its phase delay on day 12 are depicted with white-black bars on the top of each average actogram. Revertants and Rh70 mutants were exposed to white light of 0.001 lux (0.0005 µW/cm2), whereas cry01 and Rh7 cry01 mutants were exposed to 1000-lux (700-µW/cm2) white light. All fly strains resynchronized to the new LD cycle within approximately 5 days; no significant differences occurred between the lines: revertants needed 5.1 ± 0.4 days to resynchronize, and Rh70 mutants needed 5.0 ± 0.4 days. For the timing of resynchronization of cry01 mutants and Rh7 cry01 double mutants, see Table 1.
Number of days needed for re-entraining to a 6-h phase delay of the LD cycle.
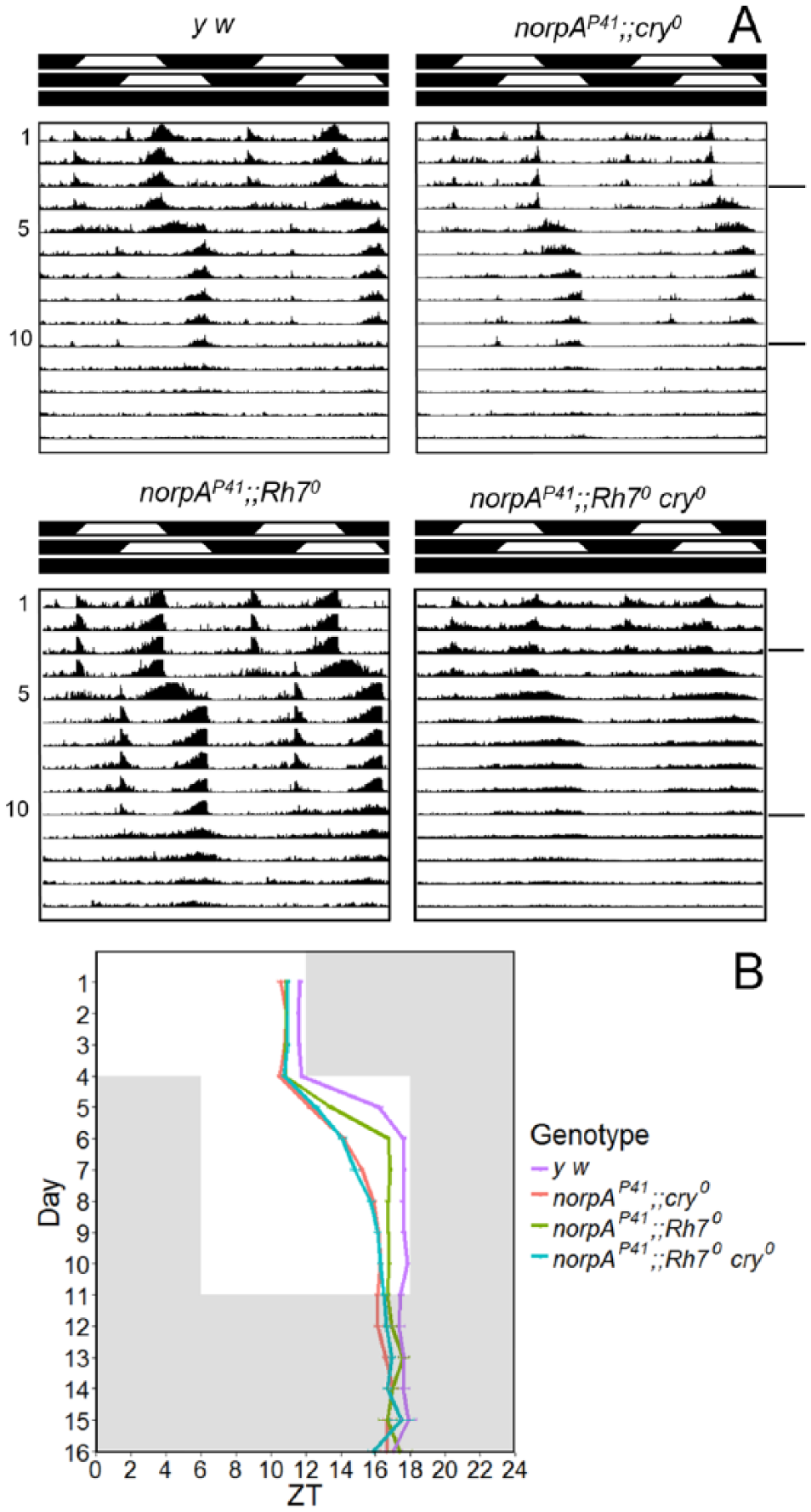
Resynchronization of Rh70 and Rh70 cry01 mutants in a norpAP41 background to 6-h phase delays of ramping white light. (A) Double plot of average actograms of 16 flies per genotype of y w, norpAP41;;cry0, norpAP41;;Rh70, and norpAP41;;Rh70 cry0. The 2 different LD cycles and the DD cycle are indicated with white-black bars on the top of each actogram. (B) Determination of the evening activity peak for the 4 tested genotypes from 3 independent experiments. While y w flies (n = 26) moved their evening peak by 4.8 ± 0.3 h on the first day after the phase shift and 1.1 ± 0.2 h on the second day, the various mutants needed longer. norpAP41;;Rh70 mutants (n = 27) delayed their evening peak by 2.6 ± 0.2 h on the first day and 3.3 ± 0.1 h on the second day. norpAP41;;cry0 (n = 42) and norpAP41;;Rh70 cry0 (n = 30) needed between 4 to 5 days to shift their evening peak to the new conditions, without any significant differences between them (p > 0.1 for all days after shift; norpAP41;;cry0: 1.7 ± 0.1 h, 2.0 ± 0.1 h, 1.0 ± 0.1 h, 0.7 ± 0.1 h, and 0.3 ± 0.1 h; norpAP41;;Rh70 cry0: 1.8 ± 0.2 h, 1.5 ± 0.2 h, 0.8 ± 0.2 h, 1.0 ± 0.2 h, and 0.4 ± 0.2 h). Error bars represent ±SEM.
Rh7 Contributes to Clock Synchronization under Blue Light
The results above show that Rh7 is neither required nor substantially contributing to circadian clock synchronization using white light. Yet, at the same time, this does not rule out Rh7’s contribution to circadian clock photosensitivity at specific wavelengths. The circadian system of wild-type fruit flies is most sensitive in blue green, with sensitivity maxima at 420 nm and 500 nm, and the action spectrum of eyeless mutants is shifted to blue, with the second maximum occurring at 470 nm (Helfrich-Förster et al., 2002). We therefore chose 470-nm blue light as an adequate wavelength to test for the potential contribution of Rh7 to light resetting. In addition, we tested 400-nm violet light. The flies were subjected to the same phase shift experiments as described above, with the exception that we applied colored light and tested phase advances of the LD cycle in addition to phase delays. Moreover, we monitored activity at the end of the experiment in DD in order to see whether the starting point of free-running activity coincided with that of the light period of the phase-shifted second LD cycle, which would indicate stable clock synchronization (Fig. 7). We found that all flies stably resynchronized to the shifted colored LD cycles and that mutant and revertant flies free-ran with a similar period (23.5 ± 0.1 h for revertants and 23.6 ± 0.1 h for Rh70 mutants). Rh70 mutants needed slightly longer to resynchronize to the shifted LD cycle than the revertants, but these differences were only significant for blue light, both during advances and delays (Fig. 7).
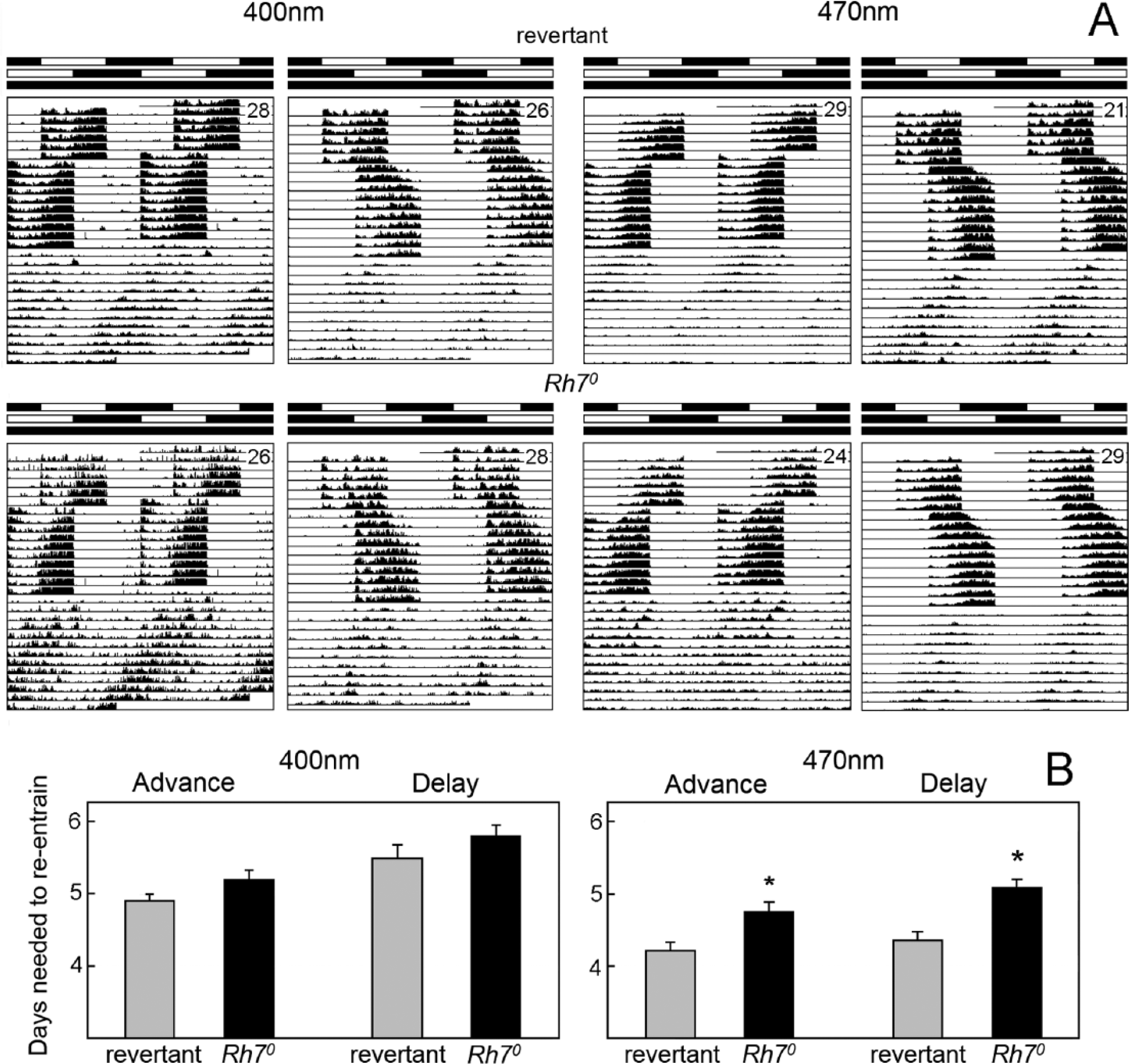
Resynchronization of revertants and Rh70 mutants to 6-h phase shifts of colored light (400-nm violet and 470-nm blue). (A) Double plot of average actograms of revertants and Rh70 mutants under 6-h phase advances and delays of the blue and violet 12-h:12-h LD cycles. The intensity of the 470-nm blue light was 0.0006 μW/cm2, and that of the 400-nm violet light was 0.0004 μW/cm2. Labeling is as in Figure 5. (B) Revertants and Rh70 mutants required a different number of days to resynchronize their locomotor activity rhythm to the 6-h shifts. Under blue light (470 nm), resynchronization took significantly longer in Rh70 mutants (p < 0.01), independent from the shifting direction. Under violet light (400 nm), the same tendency was present but did not reach significance. Error bars represent ±SEM.
Locomotor Activity Rhythms under LL and DD
Wild-type fruit flies show robust free-running locomotor activity rhythms under DD but become arrhythmic under LL (Konopka et al., 1989) due to light- and CRY-dependent degradation of the clock protein Timeless (Ceriani et al., 1999; Busza et al., 2004). The latter is impaired in several photoreceptor mutants, especially when the relevant photopigments interact directly or indirectly with Timeless (Peschel et al., 2006, 2009; Chen et al., 2011). Since Rh7 is expressed in the brain and more specifically in a small subset of clock neurons, it could theoretically interact with Timeless and promote its light-dependent degradation. If true, flies without Rh7 should remain more rhythmic under LL conditions than the controls.
We tested revertants, Rh70, cry01, and Rh70 cry01 mutants under DD and LL conditions. As expected, all flies free-ran under DD with a period slightly shorter than 24 h (Fig. 8 and Table 2). Only line #39 of the Rh70 cry01 double mutants showed a slightly higher number of arrhythmic flies than the other strains, presumably caused by the genetic background (Table 2). Under LL, revertants and Rh70 mutants became arrhythmic after a few days, whereas cry01 and Rh70 cry01 mutants remained rhythmic (Fig. 8). cry01 mutants exhibited the typical split into a prominent long-period component and a weak short-period component (Yoshii et al., 2004; Rieger et al., 2006; Dolezelova et al., 2007) (Table 2 and Fig. 8). All Rh70 cry01 double mutants showed strong free-running rhythms with a long period (Table 2). The short-period component was only clearly visible in 3 of the 16 recorded flies and did not appear in the average actogram (Fig. 8). Most interestingly, the power of the long-period component in Rh70 cry01 double mutants was significantly higher than that of all the other strains (p < 0.001); it was even significantly higher than the power of the rhythms in DD (p < 0.001) (Table 2), although the activity level of the Rh70 cry01 double mutants was not different from the other strains except cry01 mutants, which showed, on average, rather low activity under DD and LL (Table 2).

The lack of Rh7 and CRY made the flies strongly rhythmic under LL of 1000 lux. (A) Average actograms (double plots) of flies under LD, DD, and LL. Only flies that survived until the end of the experiment are included (the number of included flies is given at the top; the actual number of recorded flies is higher and indicated in Table 2). Whereas revertants and Rh70 mutants became arrhythmic after approximately 3 days in LL, cry01 and Rh70 cry01 mutants remained rhythmic in LL. Both mutants free-ran with a long period. In cry01 mutants, a second weak free-running component with a short period is visible (broken line in the right half of the average actogram). This component was absent in Rh70 cry01 double mutants. Note the robust rhythmicity of Rh70 cry01 double mutants under LL (only flies of line #112 are included in the average actogram). (B) Actograms of individual flies. The upper row shows examples of flies with weaker rhythms, and the lower row shows examples of flies with more robust rhythms under LL. Gray arrows point to the weak short-period component of cry01 mutants and to the indication of it in Rh70 cry01 double mutants under LL.
Locomotor activity rhythms under DD and LL.
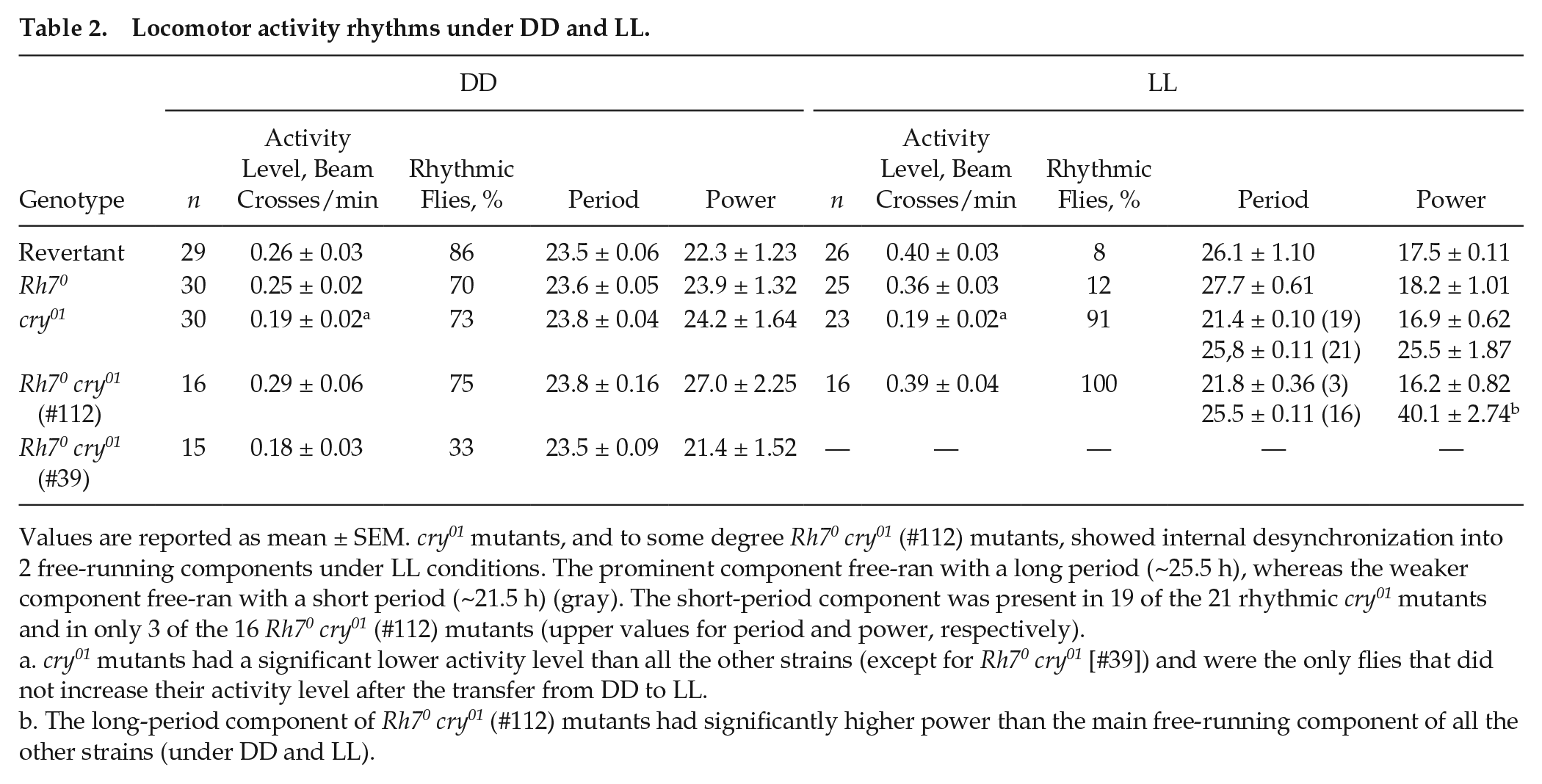
Values are reported as mean ± SEM. cry01 mutants, and to some degree Rh70 cry01 (#112) mutants, showed internal desynchronization into 2 free-running components under LL conditions. The prominent component free-ran with a long period (~25.5 h), whereas the weaker component free-ran with a short period (~21.5 h) (gray). The short-period component was present in 19 of the 21 rhythmic cry01 mutants and in only 3 of the 16 Rh70 cry01 (#112) mutants (upper values for period and power, respectively).
cry01 mutants had a significant lower activity level than all the other strains (except for Rh70 cry01 [#39]) and were the only flies that did not increase their activity level after the transfer from DD to LL.
The long-period component of Rh70 cry01 (#112) mutants had significantly higher power than the main free-running component of all the other strains (under DD and LL).
Discussion
The aim of the present study was to investigate the potential function of Rh7 in the light-dependent regulation of daily activity patterns. We found that the presence of Rh7 is important for a normal duration of morning activity and the siesta. In Rh70 mutants, morning activity ended significantly earlier than in wild-type controls, leading to a long siesta. Despite these clear effects of Rh7 on activity patterns, we found only weak effects of Rh7 on flies’ resynchronization to phase shifts of the LD cycle. Rh7 alone also barely influenced rhythmicity under LL. In this section, we discuss the role of Rh7 in more detail, particularly with regard to its expression pattern in the dorsal brain and the compound eyes.
To determine the site of action of Rh7, we used an Rh7-GFP reporter line carrying the gfp gene in the first intron of Rh7. Such reporter lines are known to faithfully report the expression of the relevant gene (Venken et al., 2011). We found Rh7 expression in the 2 DN1a, in 4 to 5 DN1p, and in several neurons in the optic lobes. The putative expression of Rh7 in the 2 DN1a and the subset of the DN1p is consistent with its observed role in determining the length of the siesta because some CRY-positive dorsal clock neurons have been previously implicated in the control of the siesta (Guo et al., 2016). The 2 DN1a as well as about 6 CRY-positive DN1p signal via glutamate and metabotropic-inhibiting glutamate receptors to the lateral clock neurons (s-LNv) that control morning activity (Collins et al., 2012, 2014; Hamasaka et al., 2007) and at least 6 CRY-positive DN1p also to the LNd that control evening activity (Guo et al., 2016). In the late morning, they inhibit electrical activity of the s-LNv, leading to the M offset and the start of the siesta. Similarly, the DN1p seem to inhibit the E onset and prolong the siesta (Guo et al., 2016). Our data suggest that Rh7-mediated light input into the 2 DN1a, 1 of the CRY-positive DN1p, and 3 to 4 CRY-negative DN1p also affects their electrical activity and consequently the M offset. CRY, which is present in different and more DN1p than Rh7, additionally affects the E onset. Future work has to show whether Rh7 in the DN1a and different subsets of the DN1p can rescue aspects of morning activity and the siesta in Rh70 mutants.
In addition, Rh7 seems to be expressed in the retina, most strongly in the fenestrated layer beneath the retina and more weakly in R8 photoreceptor cells and H-B eyelets. Very little is known about the fenestrated layer except that it is a layer of glia and pigment cells, providing optical insulation of the retina and preventing extraneous light rays from inappropriately activating the photoreceptors (Tomlinson, 2012). It is called the “fenestrated layer” because it has exit holes for the axons of the photoreceptors that leave the retina. It is unclear why Rh7 is expressed in the fenestrated layer. Nevertheless, the expression of Rh7 in R8 photoreceptor cells is in concert with the structure of the Rh7 gene: From sequence alignment, Rh7 is closely related to Drosophila Rh3, Rh4, and Rh5 genes (Senthilan and Helfrich-Förster, 2016), suggesting that it is rather expressed in the inner (R7 and R8) than outer (R1-R6) receptor cells. Furthermore, the ortholog of Drosophila Rh7, the mosquito Op10, is expressed in the inner R7 receptor cell of the mosquito, together with Op8 (Hu et al., 2014). The expression of Rh7 in R8 photoreceptor cells and the H-B eyelet is also interesting from a functional perspective because Rh6, the major rhodopsin of these cells, plays a key role in the timing of morning and evening activity under dim light and in resynchronizing rhythmic activity to phase shifts of dim LD cycles (Schlichting et al., 2015; Saint-Charles et al., 2016). Thus, Rh7 might interact with Rh6 in a still unknown manner in phasing activity. Furthermore, the H-B eyelets have recently been shown to be involved in the adaptation of flies to long days, which encompasses a lengthening of the siesta (Schlichting et al., 2016). Again, Rh7 might interfere with photoreception in the H-B eyelets and thereby modulate the length of the siesta. As stated above for the DN1p, rescue experiments of Rh7 in different structures of the eyes will show whether Rh7 in R8 photoreceptor cells and/or the H-B eyelet does indeed affect the siesta.
When comparing the activity patterns of control flies, Rh70 and cry01 single mutants, and Rh70 cry01 double mutants under LD cycles with each other, we found that Rh7 and CRY affected the M offset and the length of the siesta in a similar manner. Both proteins appear to delay the M offset; consequently, their absence provokes an earlier M offset and a longer siesta. CRY additionally advances the E onset and consequently has a stronger impact on the siesta than Rh7. In Rh70 cry01 mutants, the effects of both mutations were additive on the M offset but not on the E onset. Consequently, the siesta of Rh70 cry01 double mutants was as large as that of Rh70 single mutants. It is still unclear why the absence of CRY and Rh7 leads to an earlier E onset than the lack of Rh7 alone, whereas the absence of CRY alone provokes a late E onset.
A physical interaction of Rh7 and CRY is possible, but it appears unlikely that such an interaction in the dorsal neurons can cause the observed effects on the E onset and the siesta because the 2 proteins are only coexpressed in 3 dorsal clock neurons (2 DN1a and 1 DN1p). Alternatively, Rh7 and CRY could interact in the photoreceptor cells of the compound eyes and H-B eyelets. CRY has previously been shown to be associated with the phototransduction cascade in the eyes and to affect their photosensitivity (Mazzotta et al., 2013). Thus, Rh7 and CRY may both influence light signaling to the clock either by interacting with each other or by independent parallel pathways. The second observation that suggests an interaction of Rh7 and CRY is the high power rhythmicity of Rh70 cry01 double mutants under LL as compared to cry01 mutants; yet again, this could be explained by parallel pathways. For example, Rh7 may remove some of the light input remaining in cry01 mutants, and this results in very robust LL rhythms.
In contrast to the significant effects of Rh7 on activity patterns and (together with CRY) on LL rhythmicity, we found that Rh7 contributes only marginally to flies’ phase-shifting capabilities. Only under blue light did Rh70 mutants need significantly longer than control flies for reaching their previous phase. This indicates that Rh7 is blue light sensitive. At present, it is unknown to which wavelengths Rh7 responds. One study suggests that it is sensitive to visible light because it contains a tyrosine at position 191 that is also present in all other visible light-detecting rhodopsins such as Rh1, Rh2, Rh5, and Rh6 but changes into a phenylalanine in ultraviolet (UV)–sensitive Rh3 and Rh4 (Chou et al., 1996) (Suppl. Fig. S2). Another study suggests that Rh7 is rather UV sensitive because it carries a lysine at position 168, as do the UV-sensitive pigments Rh3 and Rh4 (Salcedo et al., 2003). We conducted no phase shifts with UV light but used violet light (400 nm) instead, which is rather close to the UV light absorption maxima of Rh3 and Rh4 (~330 nm and ~370 nm, respectively) (Salcedo et al., 2003). Under these conditions, we did not see significant phase-shifting differences between Rh70 mutants and control flies, which is consistent with electroretinogram recordings under 370-nm light that revealed no differences between Rh70 mutants and control flies (Grebler et al., 2017). Thus, our results suggest a role for Rh7 as a blue-light photopigment rather than a UV-sensitive one.
In summary, we show here that Rh7 is a promising photopigment that might interact with circadian photoreception of the fruit fly in the DN1a, DN1p, inner R8 photoreceptor cells, and putatively also H-B eyelets and, in this way, modulate activity patterns. Future experiments will be needed to determine the way(s) in which Rh7 acts.
Footnotes
Acknowledgements
This work was started in 2003 at the University of Regensburg and continued from 2009 onward at the University of Würzburg, with some experiments carried out at UCL. The authors thank Wolfgang Bachleitner for help with generating the Rh7 knockout mutant; Patrick Weidner for help with dissecting retinas; Dirk Rieger, Enrico Bertolini, and Heike Wecklein for help with locomotor activity recordings; and Janina Kempf for performing quantitative PCR. They are grateful to Paul Hardin (Texas A&M University, College Station, TX, USA) for anti-VRI, Michael Nitabach (Yale School of Medicine, New Haven, CT, USA) for anti-DH31, Steve Britt (University of Colorado Denver) for anti-Rh5, Claude Desplan (New York University) for anti-Rh6, and Joachim Bentrop (KIT, Karlsruhe, Germany) for anti-Rh7 #1. This study was funded by the German Research Foundation (GK 640: “Natural and artificial photoreceptors”; SFB 1047: “Insect timing” [project A2]) and EuClock (6th Framework Programme no. 018741). R.G. was supported by a grant from the Excellence Initiative of the German Research Foundation to the Graduate School of Life Sciences, University of Würzburg. M.O. was supported by a grant from the Biotechnology and Biological Sciences Research Council given to R.S. (BB/J018589/1). M.S. was supported by a grant from the Hanns Seidel Foundation funded by the Federal Ministry of Education and Research.
Conflict of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
