Abstract
The AKT signaling pathway is important for circadian rhythms in mammals and flies (Drosophila). However, AKT signaling in mammals is more complicated since there are 3 isoforms of AKT, each performing slightly different functions. Here we study the most ubiquitous AKT isoform, Akt1, and its role at the organismal level in the central and vascular peripheral clocks. Akt1–/– mice exhibit relatively normal behavioral rhythms with only minor differences in circadian gene expression in the liver and heart. However, circadian gene expression in the Akt1–/– aorta, compared with control aorta, follows a distinct pattern. In the Akt1–/– aorta, positive regulators of circadian transcription have lower amplitude rhythms and peak earlier in the day, and negative circadian regulators are expressed at higher amplitudes and peak later in the day. In endothelial cells, negative circadian regulators exhibit an increased amplitude of expression, while the positive circadian regulators are arrhythmic with a decreased amplitude of expression. This indicates that Akt1 conditions the normal circadian rhythm in the vasculature more so than in other peripheral tissues where other AKT isoforms or kinases might be important for daily rhythms.
Circadian rhythms, or the innate 24-hour oscillations in behaviors and gene transcription, affect many aspects of life from the sleep cycle to metabolism. The central clock of the suprachiasmatic nucleus (SCN) in the hypothalamus controls central circadian rhythms of activity (Ibuka et al., 1977; Bartness et al., 2001; Dibner et al., 2010), while the peripheral clocks in various organs control tissue-specific physiologic circadian functions (Dibner et al., 2010). Although peripheral clocks receive input from the central clock (Scheer et al., 2003; Buijs et al., 2014), they also can be set, or entrained, separately (Stokkan et al., 2001). Peripheral clocks (e.g., the circadian clock mechanism) are present and oscillating in cardiac cells (Durgan et al., 2005) and vasculature (Brunet et al., 1999; McNamara et al., 2001; Nonaka et al., 2001; Rudic et al., 2005) where there the clock serves to control cyclic variation of heart rate (HR) and blood pressure (BP), respectively (Richards et al., 2014). Having a functional molecular clock and maintaining a consistent daily sleep schedule decrease the risk for adverse cardiovascular events as evidenced by studies on shift work and sleep dysfunction (Fujino et al., 2006; Wong et al., 2015; Morris et al., 2016; Vetter et al., 2016; Wang et al., 2016). These risk factors are related to deregulated rhythms of behavior, indicating the importance of maintaining a daily rhythm for cardiovascular health.
Proteins involved in cell signaling, such as kinases and phosphatases, play a vital role in regulating circadian rhythms. One of the kinases necessary for central behavioral rhythms in Drosophila is Akt, a prosurvival signaling kinase (Zheng and Sehgal, 2010). In flies, there is one gene for Akt (Staveley et al., 1998) which when overexpressed lengthens circadian period and when knocked down shortens circadian period (Zheng and Sehgal, 2010). In mammals, there are 3 isoforms of AKT, encoded by separate genes (Walker et al., 1998). AKT1 is the most ubiquitously expressed and generally phosphorylates substrates within the proliferative and cell motility pathways (Somanath and Byzova, 2009; Grabinski et al., 2011). In the vasculature, AKT1 is necessary for normal placental and retinal vascular development (Chen et al., 2001; Cho et al., 2001b; Lee et al., 2014) and endothelial cell behaviors such as migration, proliferation, and cell-cell barrier (Ackah et al., 2005; Di Lorenzo et al., 2009). The role of Akt1 in mammalian circadian rhythms has not yet been investigated.
Akt1 expression exhibits a circadian pattern in the liver, aorta, and heart (Rudic et al., 2005; Hughes et al., 2009). Not only are Akt1 transcripts regulated in a circadian manner, but AKT activation via phosphorylation by upstream kinases also oscillates. AKT phosphorylation at T308 in chick retina (Ko et al., 2010), guinea pig endocardial myocytes (Chen et al., 2016), and mouse aorta (Kunieda et al., 2008) is rhythmic, whereas phosphorylation at S473 is not rhythmic in the liver (Jouffe et al., 2013; Shavlakadze et al., 2013). Interestingly, the sleep hormone melatonin, which is secreted in a circadian pattern, can activate or inhibit AKT in the hypothalamus (Anhe et al., 2004; Faria et al., 2013), indicating that AKT activity in the SCN oscillates (Faria et al., 2013). In addition, the circadian transcription factor BMAL1 inhibits activation of the PI3K-AKT pathway in cancer cells (Jung et al., 2013). AKT activity is altered in mice with circadian gene mutations. Bmal1-knockout mice have decreased p-AKT in the vasculature (Anea et al., 2009), while Per2 mutant mice have increased p-AKT in response to vascular endothelial growth factor (VEGF) in vascular endothelial cells (Wang et al., 2008). Thus, circadian rhythms and core circadian genes regulate Akt1 expression and its activation.
Upstream of AKT, PI3K phosphorylates PIP2 to PIP3, which recruits AKT to the plasma membrane through its PH domain. At the plasma membrane, AKT is phosphorylated by PDK1 on T308 and mTORC2 on S473, resulting in its activation (Alessi et al., 1997; Sarbassov et al., 2005). Inhibition of PI3K in human osteosarcoma (US2O) cells leads to longer circadian period (Zhang et al., 2009), and chemical inhibition of PI3K delays phase of circadian gene expression (Zhang et al., 2009). When PTEN, a negative regulator of the PI3K-AKT signaling pathway, is knocked down, period length and BMAL1 protein rhythms are altered (Matsumoto et al., 2016).
Substrates downstream of PI3K-AKT pathway are also important for circadian biology. In fact, AKT phosphorylates and inhibits FOXO transcription factors (Brunet et al., 1999). This pathway is required for Clock transcription in the liver (Chaves et al., 2014), and dFOXO–/– flies have decreased sleep-wake rhythms (Zheng et al., 2007). AKT also rhythmically inhibits GSK through phosphorylation of GSK3β on serine 9 (S9) and GSK3α on serine 21 (S21) (Iitaka et al., 2005). Gsk3αS21A/βS9A double mutant mice expressing nonphosphorylatable residues, rendering GSK constitutively active, have decreased rhythmic amplitude of activity and overall decreased activity (Paul et al., 2012). GSK3β, in turn, phosphorylates many core circadian proteins, including the positive feedback loop transcription factors CLOCK (Spengler et al., 2009) and BMAL1 (Sahar et al., 2010) and the negative regulators PER2 (Iitaka et al., 2005), CRY2 (Harada et al., 2005), and REVERBα (Yin et al., 2006).
Because AKT is regulated over a 24-hour period and several components of the PI3K-AKT signaling axis are important for circadian rhythms, we examined the role of Akt1 in mammalian central and peripheral circadian rhythms. Here we show that the loss of Akt1 dampens the amplitude of circadian wheel running but does not affect circadian period length. Analysis of circadian gene expression in the aorta and endothelial cells isolated from Akt1–/– mice is characterized into 2 distinct groups. The negative circadian regulators (Per2, Reverbα, and Dbp), whose expression is controlled most strongly by the E- and E′-boxes in their promotors (Ripperger et al., 2000; Ueda et al., 2005; Nakahata et al., 2008), exhibit heightened and later expression in Akt1–/– mice. The positive regulators (Bmal1, Clock, and Npas2), whose expression is controlled by Rev response elements (Preitner et al., 2002; Ueda et al., 2005), are expressed earlier with decreased amplitude compared with control mice. Collectively, these data elucidate a new role for Akt1 in vascular circadian gene expression.
Materials and Methods
Mouse Strains and Care
Akt1–/– mice were generated as previously described (Cho et al., 2001b). Due to poor survival of the Akt1 homozygous animals, Akt1+/– mice were bred to Akt1–/– mice to generate Akt1–/– and Akt1+/– control littermates. All mice were congenic (Fernandez-Hernando et al., 2009) and backcrossed more than 10 generations on a C57BL/J6 background. Male mice aged 8 to 16 weeks were used for all experiments. Mice were housed in standard conditions at 25 °C, under 12 hours of light per day, from 0700 h to 1900 h. Zeitgeber time (ZT) 0 corresponds to 0700 h. Circadian time (CT) 0 corresponds to the time at which the lights would have turned on if the mice were not under complete darkness. All procedures were approved by the Institutional Animal Care and Use Committee of Yale University.
Circadian Wheel Running
Male mice, 8 to 10 weeks old, were housed in a light-tight enclosed system (Actimetrics, Coulbourn Instruments, Holliston, MA) at 25 °C. Mice were housed individually in cages with running wheels for 14 days under the same 12-h:12-h light-dark schedule as the mouse facility (lights on 0700 h to 1900 h, LD). After 14 days, the lights were turned off and remained off (DD) for 14 days. Wheel revolutions per minute were counted, and data were analyzed and plotted using Clocklab software (Actimetrics, Wimette, IL) in MATLAB (MathWorks, Natick, MA).
Circadian Gene Expression
Male mice, 12 to 14 weeks old, were entrained to a 12-h:12-h light-dark cycle (LD) in light-controlled cages for 14 days. At the end of the 14th day, the lights were turned off, and mice were sacrificed under red light starting the following day, 16 hours after lights-off. After decapitation, the lights were turned on, and the aorta, heart, and liver were snap frozen and stored at −80 °C. RNA was extracted using Trizol and the RNeasy kit (Qiagen, Germantown, MD). RNA was reverse transcribed to cDNA using the TaqMan Reverse Transcription kit (Applied Biosystems, Foster City, CA). qPCR was conducted with Sybr Green on the CFX96 Touch Real-Time PCR machine (BioRad, Hercules, CA). Primers for qPCR are listed in Supplementary Table S5.
Cell Culture
Mouse lung endothelial cells (MLECs) were cultured in EGM-2 Endothelial Cell Growth Medium-2 BulletKit (Lonza, Basal, Switzerland) with 15% fetal bovine serum, 4 mM L-glutamine, and penicillin-streptomycin at 37 °C and 5% CO2. MLECs were immortalized by the polyoma middle T-antigen as previously described (Ackah et al., 2005; Lee et al., 2014) and propagated to the same extent by serial passaging.
Circadian Gene Expression in Cells
MLECs were grown to confluence and shocked with 100 nM dexamethasone in EGM-2 media (Lonza). After 12 hours, cells were collected in Trizol every 2 hours and frozen at −80 °C. RNA was extracted using the RNeasy kit (Qiagen). Reverse transcription and qPCR were carried out as described above.
Statistics
Data are expressed as mean ± SEM. Comparisons between 2 single data points were made using Student t test. Comparisons between groups of data over time were compared using 2-way analysis of variance (ANOVA) with Bonferroni posttest. Complete statistics are included in the supplementary online material. To determine circadian gene expression parameters, statistics were calculated using JTK_CYCLE, a nonparametric algorithm (Hughes et al., 2010).
Results
Wheel Running Rhythms of Akt1–/– Mice
The central circadian rhythm set by the SCN in the hypothalamus can be measured by recording activity of light-entrained animals subjected to complete darkness. Akt1+/– and Akt1–/– offspring of Akt1+/– mice bred to Akt1–/– mice were used due to the reduced number, size, and survival of Akt1–/– from breeding heterozygous mice as experienced in our laboratory and as previously published (Chen et al., 2001; Cho et al., 2001b). Akt1+/– and Akt1–/– littermates were acclimated to light-tight cages with running wheels and entrained to a 12-h:12-h light-dark (LD) schedule for 14 days; then the mice were subjected to complete darkness (DD) for 14 days. During the LD cycle, Akt1–/– and Akt1+/– mice were entrained with tau equal to 24 hours and exhibited normal phase angle of entrainment, alpha or duration of activity, and rhythm amplitude represented as number of wheel revolutions. The only differences measured were the known differences in body weight (Cho et al., 2001b), indicating that there are no effects of Akt1 deletion on voluntary activity under LD (Suppl. Fig. S1). In DD, Akt1–/– mice had a similar circadian period, tau, compared with both Akt1+/– controls (Suppl. Fig. S2A) and published data from WT C57Bl/J6 mice (23.77 ± 0.02 h) (Schwartz and Zimmerman, 1990). Although Akt1–/– mice had comparable alpha (Suppl. Fig. S2B), they exhibited some minor defects in lower total activity during the subjective dark phase (Suppl. Fig. S2E, p < 0.01), as well as a significantly diminished precision of activity onset in the second week of DD (Suppl. Fig. S2C, p < 0.05). In general, Akt1–/– mice have relatively intact wheel running rhythms with a slight decrease in rhythm amplitude.
Circadian Gene Expression in Akt1–/– Hearts, Livers, and Aortae
To discern the role of Akt1 in the molecular circadian clock, peripheral circadian gene expression in the heart, liver, and aorta was examined. After entraining under LD for 14 days, mice were sacrificed at 4-hour intervals across the circadian cycle after at least 16 hours in complete darkness, and circadian gene expression was analyzed. In all tissues, the mRNA levels of negative regulators (Per2, Reverbα, and Dbp) and positive regulators (Bmal1, Npas2, and Clock) of the circadian feedback loop were assessed. There was no shift in the timing of peak expression of circadian regulators in the heart. However, the magnitude of Reverbα expression, but not other genes, was attenuated in Akt1–/– hearts. There was a less than 2-fold difference in amplitude of expression for each of the 6 genes analyzed (Fig. 1, Suppl. Table S1).
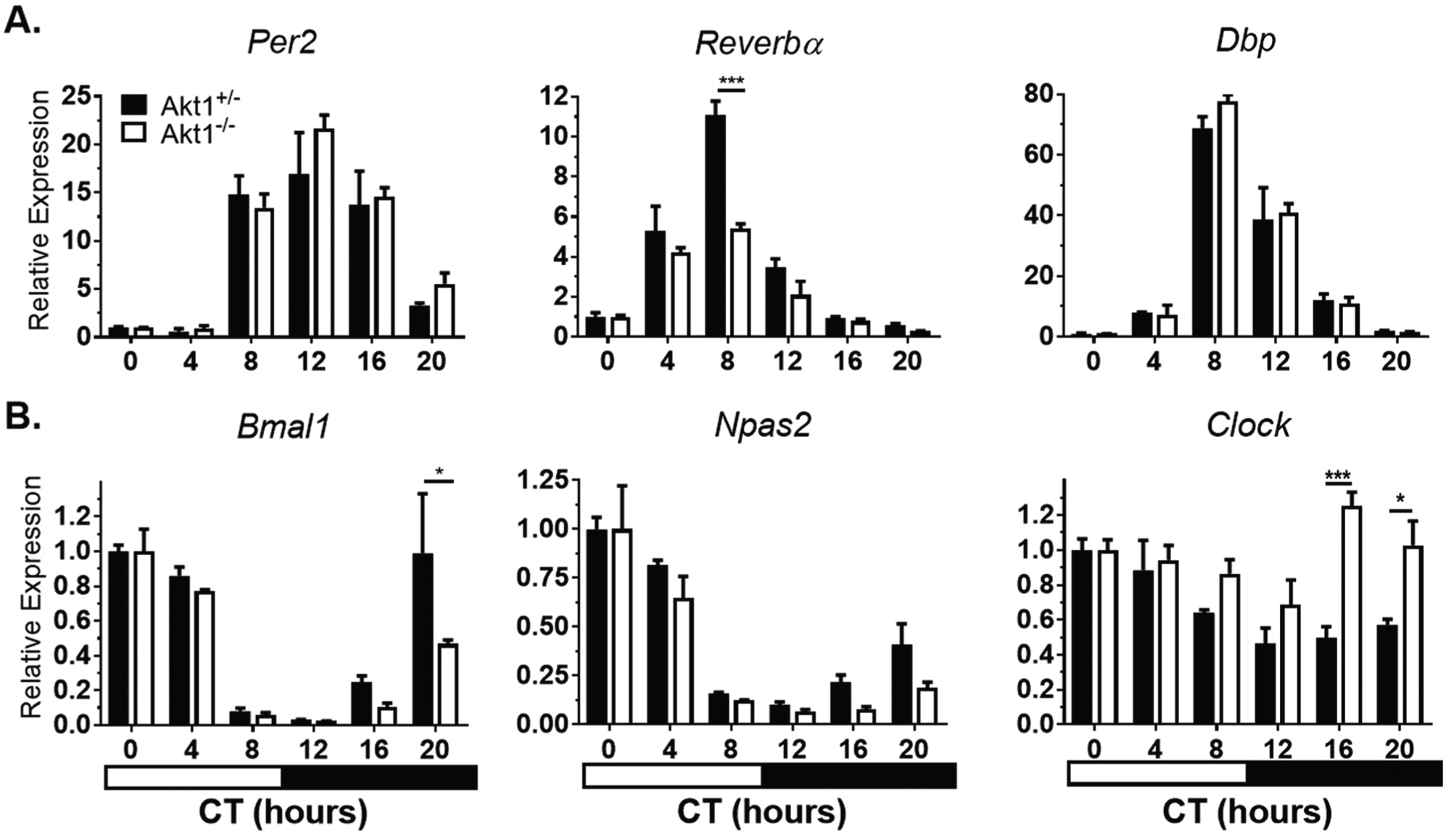
Circadian gene expression in Akt1–/– hearts. Mice were sacrificed at the given times in complete darkness. Negative regulators (A) and positive regulators (B) of circadian rhythms were quantified by qPCR and gene expression normalized to Gapdh expression and CT0 (0700 h). Black bars, Akt1+/–. White bars, Akt1–/–. Horizontal black and white bar shows subjective dark and light phases of circadian time. Data are represented by mean ± SEM. n = 3; ***p < 0.001, *p < 0.05.
The liver, which is one of the strongest and most well-studied peripheral clocks, was examined next. In the liver, Akt1 plays a less significant role in function and metabolism compared with Akt2 (Cho et al., 2001a; Cho et al., 2001b). The loss of Akt1 in the liver had little effect on circadian gene expression (Fig. 2, Suppl. Table S2). However, the magnitude of Reverbα expression was reduced in Akt1–/– livers compared with Akt1+/– livers. Clock was not rhythmically expressed in either Akt1+/– or Akt1–/– livers (Suppl. Table S2).
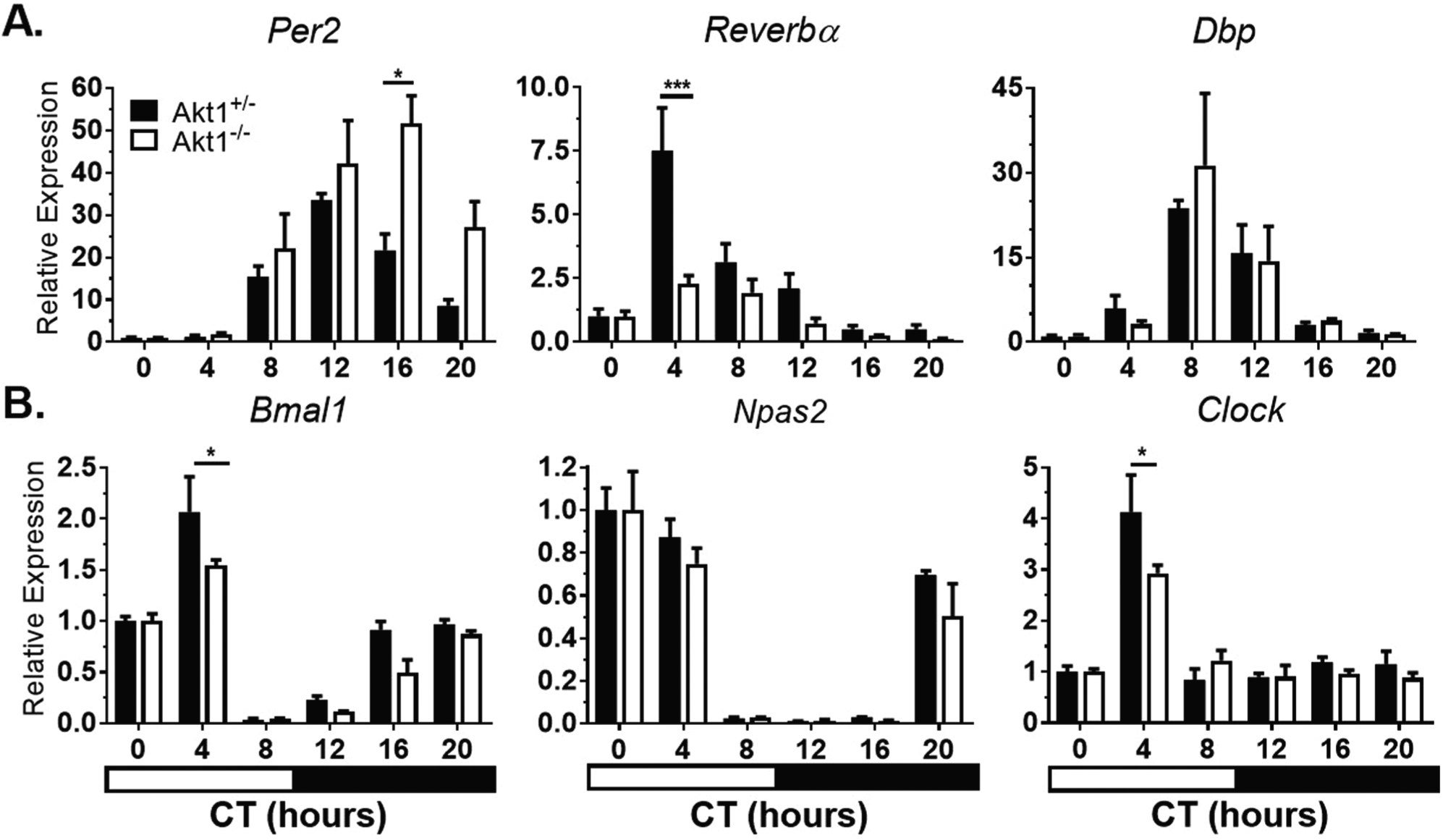
Circadian gene expression in Akt1–/– livers. Mice were sacrificed at the given times in complete darkness. Negative regulators (A) and positive regulators (B) of circadian rhythms were quantified by qPCR and gene expression normalized to Gapdh expression and CT0 (0700 h). Black bars, Akt1+/– . White bars, Akt1–/– . Horizontal black and white bar shows subjective dark and light phases of circadian time. Data are represented by mean ± SEM. n = 3; ***p < 0.001, *p < 0.05.
In the aorta, negative regulators of circadian rhythms (Per2, Reverbα, Dbp) showed increased levels of expression (peak to trough; 2.95- to 21.50-fold greater depending on the gene) and later peak expression by 15° to 90° in Akt1–/– aortae compared with Akt1+/– aortae (Fig. 3A, Suppl. Table S3). Interestingly, circadian genes in the positive feedback loop (Bmal1, Npas2) showed earlier peak expression in Akt1–/– aortae with decreased amplitude (1.35- to 2.65-fold, depending on the gene). The expression of Clock lost its rhythmicity in Akt1–/– aortae (Fig. 3B, Suppl. Table S3).
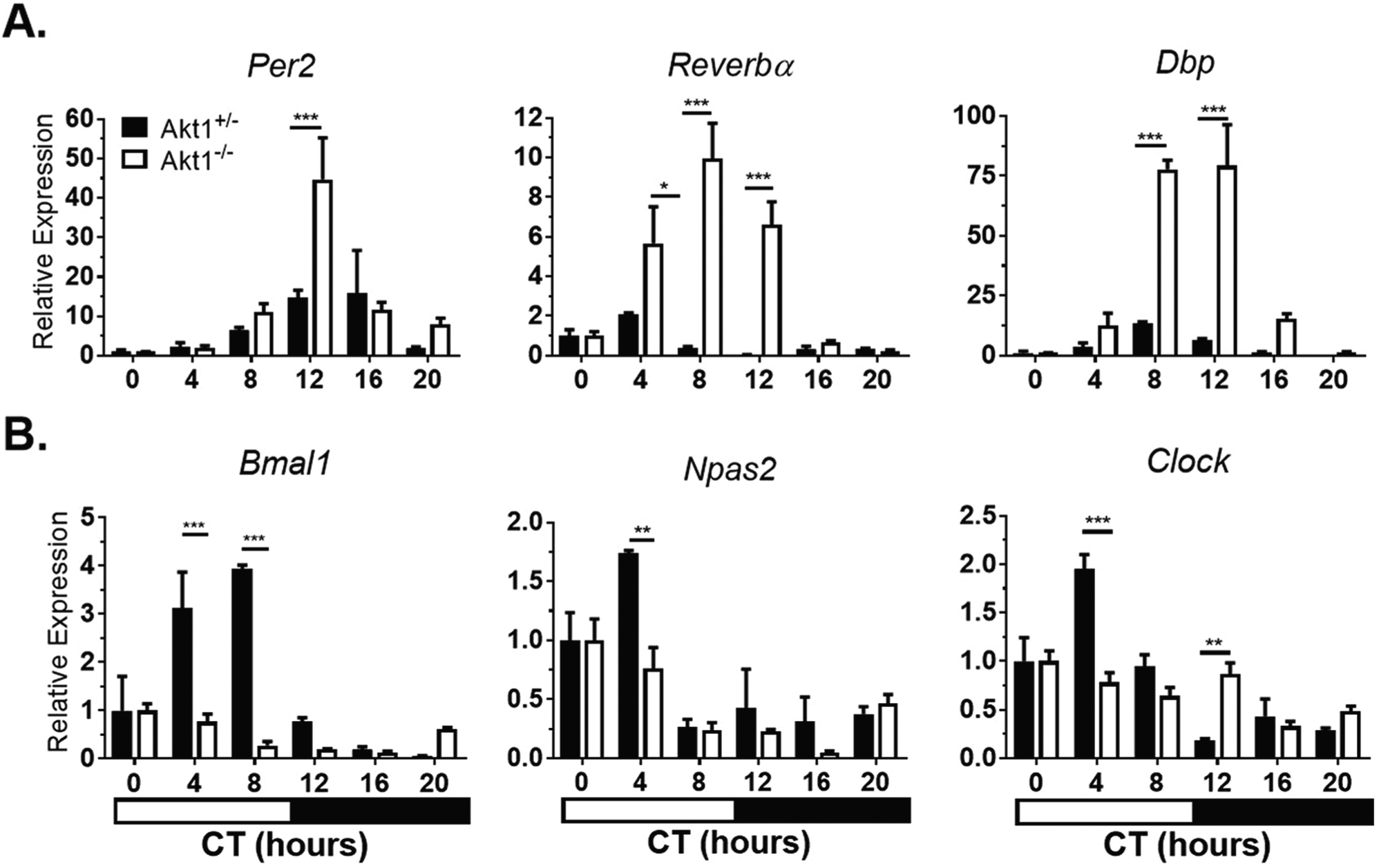
Akt1 affects timing and amplitude of circadian gene expression in the aorta. Mice were sacrificed at the given times in complete darkness. Gene expression was normalized to Gapdh expression and CT0 (0700 h). (A) Negative regulators show larger amplitude and later peak of expression. (B) Positive regulators generally show smaller amplitude and earlier peak of expression. Clock in the Akt1–/– aortae was arrhythmic using JTK_CYCLE analysis. Black bars, Akt1+/– . White bars, Akt1–/–. Horizontal black and white bar shows subjective dark and light phases of circadian time. Data represented as mean ± SEM. n = 3; ***p < 0.001, **p < 0.01, *p < 0.05.
Abnormal Circadian Gene Expression in Akt1–/– MLECs
The abnormal regulation of circadian gene expression in whole aortae of Akt1–/– mice implied that either, or both, the endothelial cells and smooth muscle cells of the vessel wall exhibit impaired circadian gene expression. Previous work from our laboratory and others (Ackah et al., 2005; Somanath et al., 2008; Di Lorenzo et al., 2009; Lee et al., 2014) has shown a critical role of Akt1 in endothelial cell function; therefore, we examined the role of Akt1 in isolated mouse lung endothelial cells (MLECs). In these experiments, unlike the in vivo experiments in mice either heterozygous or homozygous for the Akt1 allele, MLECs were isolated from littermate WT and Akt1–/– mice and immortalized as described (Lee et al., 2014). In preliminary experiments, 3 different treatments (50% horse serum, 10 µM forskolin, or 100 nM dexamethasone) were used to induce circadian gene expression as described in other cell types (Balsalobre et al., 1998; Nagoshi et al., 2004; Chalmers et al., 2008; Hughes et al., 2009). While all 3 treatments induced circadian gene expression, dexamethasone produced the most robust stimulation of daily rhythms in MLECs. Therefore, circadian gene expression in Akt1+/+ and Akt1–/– MLECs after dexamethasone shock was evaluated. As seen in Figure 4A (and quantified in Suppl. Table S4), negative circadian regulators (Per2, Reverbα, and Dbp) exhibited an increase in amplitude of expression (3.8- to 4.8-fold, p < 0.001) and more rapid induction. The positive circadian regulators, Bmal1 and Npas2, exhibited circadian rhythmicity in Akt1+/+ MLECs; however, Clock was not calculated to be significantly rhythmic in these cells (p = 0.36, Suppl. Table S4). The positive regulators were arrhythmic in Akt1–/– cells (Fig. 4B), indicating that Akt1 regulates both their rhythmicity and amplitude in MLECs. Generally, positive regulators in Akt1–/– MLECs were less rhythmic with smaller amplitude (−3.23 and −3.06 for Bmal1 and Npas2, respectively, p < 0.01, Suppl. Table S4) than in Akt1+/+ cells. Similar to the negative regulators, the positive regulators also were expressed earlier although the changes were less than that seen for the negative regulators (22.5°-82.5°, Suppl. Table S4).
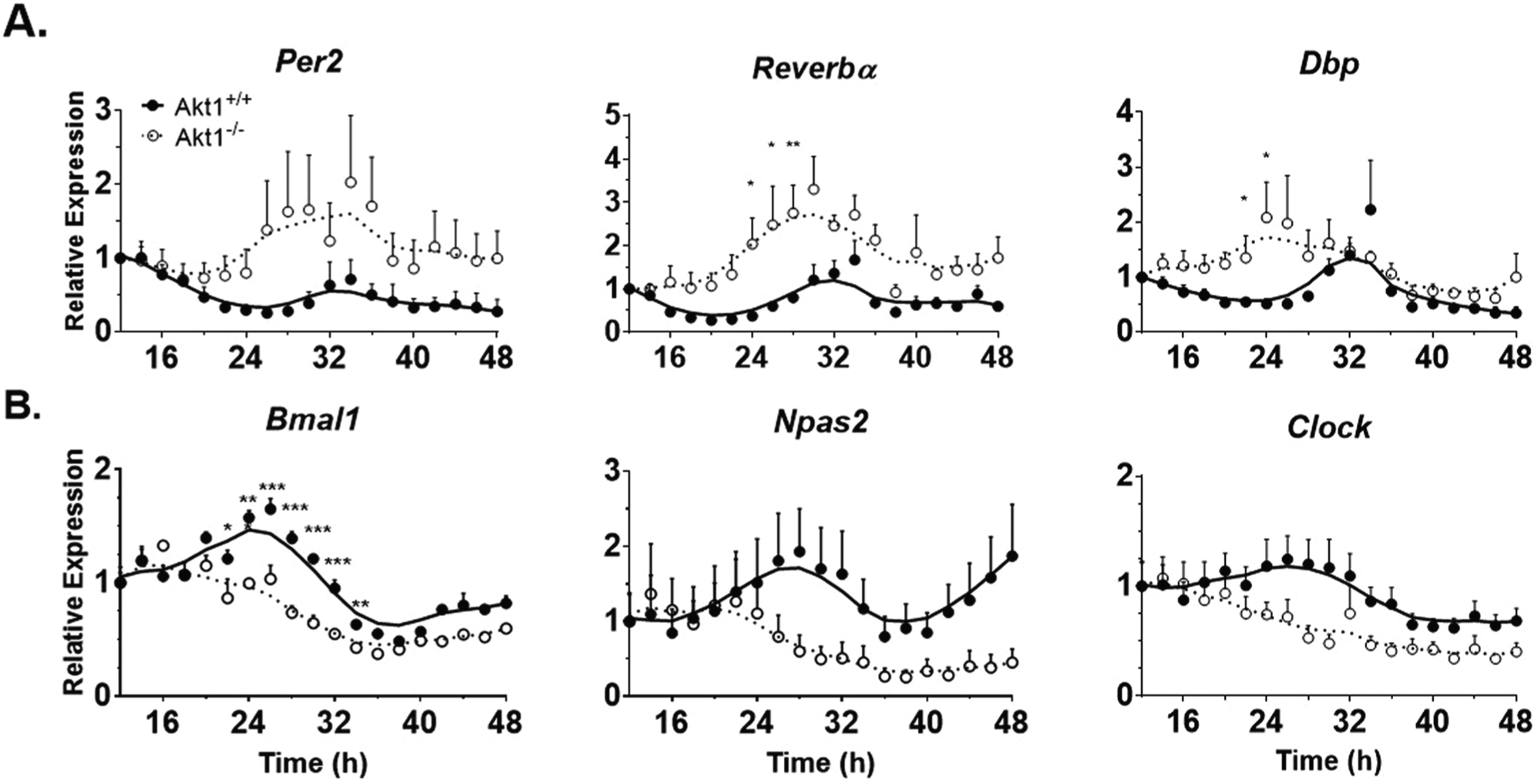
Altered circadian gene expression in Akt1–/– mouse lung endothelial cells. Expression at the indicated times post shock was normalized to Gapdh expression and the first time point at 12 hours post shock. (A) Negative regulators have higher amplitude of rhythms and earlier peak expression in Akt1–/– cells. (B) Positive regulators in Akt1–/– cells have lower amplitude rhythms than Akt1+/+ transcripts and earlier peak expression. Neither the negative regulators nor Dbp in Akt1–/– nor Clock in either cell type are significantly rhythmic according to JTK_CYCLE (Suppl. Table S4). Data are represented as mean ± SEM. A Lowess curve is fitted to the data. Akt1+/+, closed circles and solid line. Akt1–/–, open circles and dotted line. n = 5; *p < 0.05, **p < 0.01, ***p < 0.001.
Discussion
Previous work in lower organisms has suggested that the PI3K-AKT pathway is important for regulating circadian rhythms. Here we show that Akt1 has little effect on the central clock but modulates rhythms of circadian gene expression in mice. Akt1–/– mice are less active during the dark phase of the day but have normal period length, which is different from the circadian phenotype in Drosophila where Akt maintains period length. In addition, the loss of Akt1 regulates the expression of circadian genes in intact blood vessels and isolated endothelial cells, implying that AKT1 signaling influences key inputs into peripheral control of circadian gene expression.
Akt1 phosphorylates multiple substrates critical for vascular function and regulates angiogenesis and vascular remodeling (Ackah et al., 2005; Chen et al., 2005; Shiojima et al., 2005; Somanath et al., 2008; Lee et al., 2014; Kerr et al., 2016). Akt1–/– aortae generally express negative circadian regulators with higher amplitude and positive circadian regulators with lower amplitude (Fig. 3). Interestingly, circadian gene expression in hearts and livers is less affected by the loss of Akt1, indicating that Akt1 does not play a vital role in the daily rhythms of these tissues and uniquely regulates gene expression in vascular tissue and endothelial cells. Mechanistically, the effects of Akt1 deletion on circadian rhythms may be partially explained by AKT1 phosphorylating key substrates, GSK3β and FOXO. GSK3β +/– mice have a smaller phase angle and longer period than WT controls (Lavoie et al., 2013). When GSK3β is chemically inhibited with LiCl, NIH3T3 cells exhibit a phase delay of Per2 expression (Iitaka et al., 2005). Overexpression of GSK3β, which is functionally similar to activation of GSK3β by Akt1 deletion, shifts Per2 transcription later (Iitaka et al., 2005), similar to the effect on Per2 expression in the present experiments in aortae. However, in Akt1–/– aortae, the expression of Dbp and Reverbα is also affected, indicating that Akt1 likely has a broader effect in regulating upstream E-box transcription factors that activate transcription of the negative regulators (Ueda et al., 2005). In fact, BMAL1 is phosphorylated by AKT in livers, resulting in its inactivation (Dang et al., 2016). In addition to GSK3β, FOXO transcription factors are required for Clock transcription in the liver (Chaves et al., 2014). FOXO phosphorylation, and likely activity, is altered in Akt1–/– mice (Lee et al., 2014), which may explain why Clock is not rhythmically expressed in this tissue.
In summary, these data indicate that Akt1 plays a role in the amplitude and timing of expression of circadian genes. In the vasculature and isolated endothelial cells, Akt1 appears to decrease amplitude of expression of positive circadian regulators that are activated by Rev response elements (RREs) in their promoters (Ueda et al., 2005). At the same time, there is increased amplitude of expression of genes activated by E-box elements (Ueda et al., 2005), indicating that AKT1 affects transcription of groups of circadian genes upstream through activating or inhibiting specific DNA response elements. However, whether Akt1 has direct effects by phosphorylating circadian substrates or only indirect effects through intersecting pathways is unknown. It would be interesting to examine whether growth factor-dependent activation of AKT and phosphorylation of AKT motifs in CLOCK, TRIP12, and other circadian proteins might be important for the daily oscillations regulated by PI3K-AKT signaling. Ongoing work will focus on investigating the vascular effects of Akt1 and the mechanism(s) behind the observed circadian effects. When considering targeting the PI3K-AKT axis for treatment of diseases such as cancer or metabolic syndrome, the present data indicate that there may be off-target effects on vascular circadian rhythms.
Footnotes
Acknowledgements
We thank Roger Babbitt for his invaluable technical assistance. This study was supported by the NIH (R01 HL64793, R01 HL61371, and P01 HL1070295), the American Heart Association MERIT Grant to W.C.S., an NIH training grant (T32 GM007324 to PI W.C.S. and A.K.L. supported), a predoctoral AHA grant (15PRE21670000) to A.K.L., a Sarnoff Cardiovascular Research Foundation fellowship to J.M.S., and the George M. O’Brien Kidney Center Grant from the NIH (P30 DK079310) to H.V. and for use of the circadian wheel running system and light-tight cages.
American Heart Association 10.13039/100000968, 16MERIT27950008
Conflict of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
