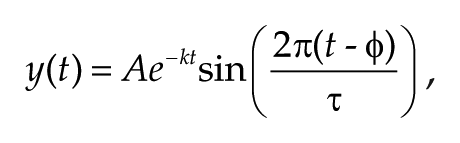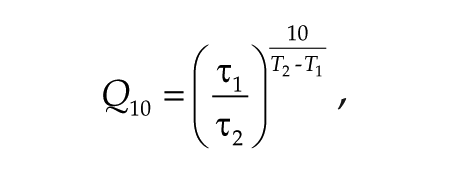Abstract
Most organisms have cell-autonomous circadian clocks to coordinate their activity and physiology according to 24-h environmental changes. Despite recent progress in circadian studies, it is not fully understood how the period length and the robustness of mammalian circadian rhythms are determined. In this study, we established a series of mouse embryonic stem cell (ESC) lines with single or multiplex clock gene ablations using the CRISPR/Cas9-based genome editing method. ESC-based in vitro circadian clock formation assay shows that the CRISPR-mediated clock gene disruption not only reproduces the intrinsic circadian molecular rhythms of previously reported mice tissues and cells lacking clock genes but also reveals that complexed mutations, such as CKIδm/m:CKIε+/m:Cry2m/m mutants, exhibit an additively lengthened circadian period. By using these mutant cells, we also investigated the relation between period length alteration and temperature compensation. Although CKIδ-deficient cells slightly affected the temperature insensitivity of period length, we demonstrated that the temperature compensation property is largely maintained in all mutants. These results show that the ESC-based assay system could offer a more systematic and comprehensive approach to the genotype-chronotype analysis of the intracellular circadian clockwork in mammals.
Circadian rhythms are intrinsic biological rhythms affecting diverse organisms, with a period of about 24 h. In mammals, the master pacemaker of the circadian clock resides in the suprachiasmatic nucleus (SCN) of the hypothalamus, and peripheral oscillators exist in most organs and tissues throughout the body, coordinating activity and physiological processes, including the sleep/wake cycle, feeding, hormonal secretion, and metabolism (Dunlap, 1999; Reppert and Weaver, 2002; Lowrey and Takahashi, 2011). In every cell, a set of core clock genes constitutes cell-autonomous transcriptional-translational feedback loops, in which the transcriptional activators, BMAL1 and CLOCK, heterodimerize and promote expressions of Period (Per1, Per2, and Per3) and Cryptochrome (Cry1 and Cry2) (Reppert and Weaver, 2002; Ko and Takahashi, 2006). The PER and CRY proteins repress BMAL1/CLOCK activity and thereby their own transcription, leading to the circadian oscillation of their messenger RNA (mRNA) expression. Despite recent progress in circadian studies, it is not fully understood how the period length and the robustness of mammalian circadian rhythms are controlled. In this study, we generated mutant clock gene mouse embryonic stem cell (ESC) lines using the clustered regularly interspaced short palindromic repeats (CRISPR)/Cas9-based genome editing method (Cong et al., 2013; Mali et al., 2013). We targeted Cry1, Cry2, CKIδ, and CKIε to address whether the CRISPR-mediated clock gene disruption can reproduce the intrinsic circadian molecular rhythms of previously reported mice tissues and cells lacking these genes, as disruption of Cry1/2 and CKIδ/ε is well known to result in significant change in the circadian period. Furthermore, we generated complexed mutants, such as the CKIδm/m:CKIε+/m:Cry2m/m triple gene-targeted mutant, expecting mutant cells with a much longer circadian period. Then, by using these mutant cells, we investigated temperature compensation properties of mammalian cells lacking clock genes. Temperature compensation is one of the most distinctive characteristics of circadian oscillators. The period length of rhythms is relatively constant over a wide range of physiological temperatures (Pittendrigh, 1954; Hastings and Sweeney, 1957; Zimmerman et al., 1968; Menaker and Wisner, 1983; Kondo et al., 1993). Previous studies have demonstrated that circadian gene expression rhythms are also temperature compensated in mammalian cultured cells (Izumo et al., 2003; Tsuchiya et al., 2003). Intriguingly, it has been reported that in Neurospora, several clock gene mutants such as frq3 and frq7 exhibit both abnormal period length and altered temperature compensation (Gardner and Feldman, 1981; Loros and Feldman, 1986; Liu et al., 1997; Ruoff et al., 2005). Also in Drosophila, clock gene mutants including perL have been found to show altered temperature compensation for the respective period (Huang et al., 1995; Rutila et al., 1996; Sawyer et al., 1997; Matsumoto et al., 1999). However, the relation between the circadian period and temperature compensation has yet to be extensively investigated in mammals. We generated mutant cells with altered period length to analyze temperature compensation properties of the mammalian circadian clock. We propose that an ESC-based assay system with CRISPR-mediated genome editing could offer a more systematic and comprehensive approach to the genotype-phenotype analysis of the intracellular circadian clockwork in mammals.
Materials and Methods
Cell Culture
Mouse ESCs derived from PER2Luc knock-in mice were kindly gifted by Dr. Joseph Takahashi’s group (Yoo et al., 2004). In the PER2Luc ESCs, PER2::luciferase fusion reporter is expressed from the endogenous Per2 locus, in which an SV40 polyadenylation site is inserted after luciferase. ESCs were maintained on MEF feeder cells in an ES medium (Glasgow minimum essential medium [MEM] supplemented with 15% fetal bovine serum [FBS; HyClone, Logan, UT], 0.1 mM MEM nonessential amino acids, 0.1 mM 2-mercaptoethanol, 1000 U/mL of leukemia inhibitory factor [LIF], and 100 U/mL penicillin-streptomycin).
Single Guide RNA Design
The single guide RNA (sgRNA) target site was determined by seeking sequences corresponding to N19NGG in the coding region of the target genes. For each sequence, 13 bases preceding the protospacer-adjacent motif (PAM) sequence with AGG, TGG, GGG, or CGG were aligned to the mouse genome (mm9) using bowtie software; any sequence that had more than a single match was excluded from consideration (Mali et al., 2013).
Plasmids
A human codon-optimized Cas9 expression plasmid was obtained from Addgene (Cambridge, MA; #41815). For the construction of the sgRNA expression plasmid, sgRNA plasmid (Addgene #41817) was polymerase chain reaction (PCR) amplified with a pair of oligos containing 2 BbsI sites. PCR products were self-ligated. The oligos used were as follows: BbsI-f, 5′-GAAGACCTGTTTTAGAGCTAGAAATAG-3′; BbsI-r, 5′-TCGAAGACCCGGTGTTTCGTCCTTTCC-3′. A pair of oligos for the sgRNA targeting site was annealed and ligated into the introduced BbsI sites of the sgRNA plasmid.
Transfection
Mouse ESCs were cotransfected with an hCas9 expression vector, sgRNA expression vectors, and a plasmid with a puromycin selection marker using FuGENE HD (Promega, Madison, WI). The cells were selected with 2 µg/mL puromycin for 2 days and then passaged for mutant cloning. ESC colonies were picked and cultured to establish mutant cell lines.
Sequence Analysis of Genomic DNA
Genomic DNA samples were extracted from mouse ESCs grown under feeder-free conditions. PCR was performed with PrimeSTAR MAX DNA Polymerase (TAKARA, Japan) under the following conditions: 98 °C for 1 min; 35 cycles of 98 °C for 10 sec, 60 °C for 5 sec, 72 °C for 10 sec; 72 °C for 20 sec; and hold at 4 °C. PCR products were treated with Exonuclease I (New England Biolabs, Ipswich, MA) and shrimp alkaline phosphatase (TAKARA) at 37 °C for 30 min followed by inactivation at 95 °C for 10 min and used for sequencing analysis. The primers for PCR are listed in Supplemental Table S2.
Off-Target Analysis
All potential off-target sites with homology to the 11 bases preceding the PAM sequence with NGG were aligned to the mouse genome (mm9). In addition, each of the 19 bases preceding the PAM sequence with NGG was also analyzed, allowing for ungapped alignments with up to 3 mismatches. The potential off-target sites were amplified by PCR and sequenced. The primers for PCR are listed in Supplemental Table S2.
In Vitro Differentiation of Mouse ESCs
The in vitro differentiation of ESCs was performed as described previously (Umemura et al., 2013; Umemura et al, 2014). Briefly, to form embryoid bodies (EBs), 2 × 103 of the dissociated ESCs were seeded in low-attachment 96-well plates in a differentiation medium (Dulbecco’s modified Eagle’s medium supplemented with 10% FBS, 1 mM sodium pyruvate, 0.1 mM nonessential amino acids, GlutaMax-I [Invitrogen, Carlsbad, CA], 0.1 mM 2-mercaptoethanol, and 100 U/mL penicillin-streptomycin). Two days later, the EBs were plated onto gelatin-coated 24-well plates and cultured for 26 days in a differentiation medium, which was exchanged every other day.
Bioluminescence Recording
For bioluminescence recording, the medium was replaced with a differentiation medium containing 0.2 mM luciferin and 100 nM dexamethasone. Bioluminescence was measured and integrated for 1 min at 20-min intervals with PMT-based equipment at 35.6 °C. For the analysis of temperature compensation, EBs were plated in a gelatin-coated 35-mm dish (5 EBs/dish), and the dishes were placed in a custom-made 4-channel PMT device with 4 wells in line and temperature-controllable heat blocks at both ends, generating a temperature gradient among the 4 dishes.
Data Analysis
The bioluminescence data recorded by PMT were analyzed using a sine wave fitting. A linear baseline was subtracted from the raw data. The detrended data from between 36 and 108 h was then used for analysis. Sine wave fitting was performed using the following equation:
where A = amplitude, k = damping constant, t = time, τ = period, and φ = phase. For statistical analyses, 1-way analysis of variance (ANOVA) followed by Bonferroni’s post hoc comparisons tests were performed. For the calculation of Q10 value, the period values were assigned to the following equation:
where τ1 and τ2 are the period at temperature T1 and T2, respectively. T1 = 33.5 °C and T2 = 36.0 °C.
Results
In Vitro Differentiation Culture of Mouse ESCs Lacking Cry1 and/or Cry2 Reproduces Genetically Determined Period Changes of Mammalian Circadian Clock
To generate mouse ESCs carrying mutations that disrupt clock gene function, we designed sgRNAs targeting Cry1 and Cry2 (Suppl. Fig. S1A). We used mouse PER2Luc knock-in ESCs that harbored the Per2 gene fused with a luciferase reporter (Welsh et al., 2004; Yoo et al., 2004). For the generation of Cry1 and Cry2 mutant ESCs, we transfected PER2Luc ESCs with a Cas9 expression vector and Cry1- or Cry2-targeting sgRNA expression vector and established a number of cell lines, each derived from a single ESC colony. The genotyping analysis demonstrated that cloned Cry1 #11, #13, #14 and Cry2 #5, #15, #18 only have alleles with a frame-shift mutation, resulting in a premature stop codon downstream of the targeted site (Suppl. Fig. S1A,B). For a Cry1/Cry2 double mutant cell line, PER2Luc ESCs were transfected with sgRNA expression vectors for Cry1 and Cry2 simultaneously (Suppl. Fig. S1C). We analyzed the PER2Luc bioluminescence rhythms of differentiated ESCs. The Cry1 mutant cells exhibited dampened bioluminescence rhythms with a shorter period, and Cry2 mutant cells showed a longer period (Fig. 1A,B,D). Cry1/Cry2 double mutant cells exhibited arrhythmic bioluminescence (Fig. 1C). These data reproduce the circadian phenotypes observed in Cry1 and/or Cry2 knockout mice and cells (van der Horst et al., 1999; Vitaterna et al., 1999; Liu et al., 2007; Ruan et al., 2012), demonstrating that the combined approach of CRISPR-mediated gene targeting and differentiation-dependent circadian clock formation in ESCs provides a useful assay system to investigate genetically determined properties of the intracellular circadian clock in mammals.
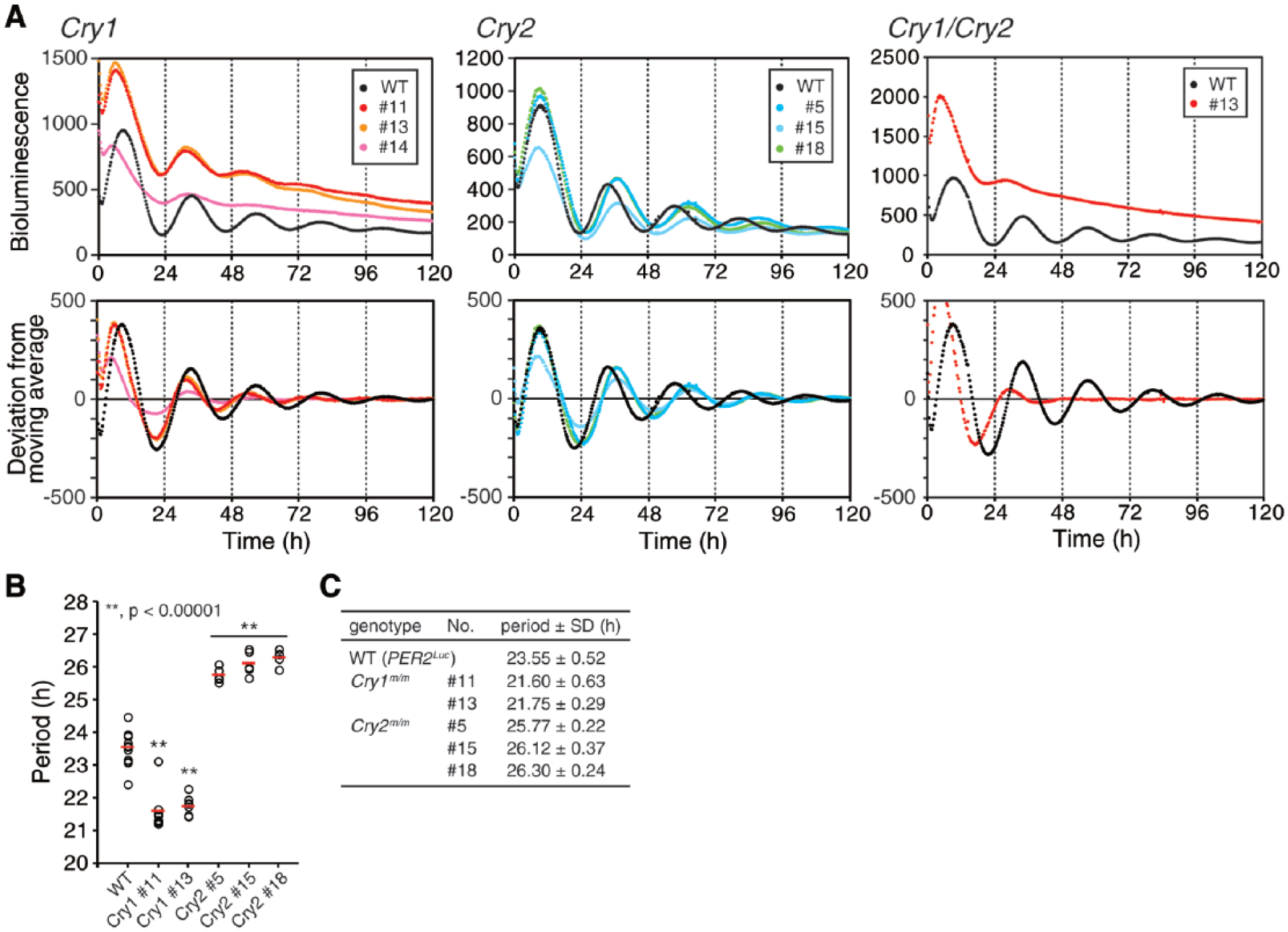
Clock gene mutant embryonic stem cells develop circadian phenotypes compatible with corresponding Cry1 and/or Cry2 knockout tissues and cells. (A–C) Cry1 (A) and Cry2 (B) mutant cells exhibit rhythms with a shorter and longer period, respectively, and Cry1/Cry2 double mutant cells (C) show attenuated rhythms. The average of the raw (upper) and detrended (bottom) data is plotted (n = 5). (D) The period length of Cry1 or Cry2 mutant cells is analyzed in A and B. Each open circle indicates the value calculated from the data of 1 embryoid body, and red bars indicate the mean (**p < 0.00001).
Overlapping Function of CKIδ and CKIε in Regulation of the Circadian Clock
Next, we applied the CRISPR-mediated gene targeting approach to other clock genes (i.e., CKIδ and CKIε ), which play an important role in setting the speed of the circadian clock (Meng et al., 2008; Lee et al., 2009; Lee et al., 2011). We first obtained 1-allele targeted (CKIδ+/m) (#10) and 2-allele targeted (CKIδm/m) (#2) CKIδ mutants as well as a 2-allele targeted (CKIεm/m) (#109) CKIε mutant using single gene targeting (Suppl. Fig. S2A,B,D). To generate the CKIδ/ε double mutant cell lines, we targeted CKIε in CKIδ+/m and CKIδm/m backgrounds. As a result, both CKIε+/m (#206, #209) and CKIεm/m (#204, #211) mutant lines were obtained in a CKIδ+/m background, but only CKIε+/m (#312, #315) mutant lines were obtained in a CKIδm/m background (Suppl. Fig. S2B). These results suggest that the complete loss of both CKIδ and CKIε compromises the viability and/or proliferation of mouse ESCs. Notably, the targeting of CKIδ could cause occasional off-target mutations in the CKIε locus, which has 1 base mismatch at 7 bases upstream of the protospacer adjacent motif (PAM) (Suppl. Table S1). Indeed, the CKIδ mutant #9 was found to possess a mutation at the CKIε locus (Suppl. Fig. S2C). In this context, we also examined potential off-target loci in CRISPR/Cas9-mediated targeting for Cry1, Cry2, CKIδ, and CKIε. As a result, no off-target mutations were detected in any of the analyzed clones (Suppl. Table S1). Collectively, we successfully generated a series of CKIδ/ε mutant ESC lines by using the CRISPR/Cas9 system. To examine the circadian rhythms of CKIδ/ε mutants, ESCs were differentiated in vitro and their PER2Luc bioluminescence rhythms were monitored. Interestingly, having a CKIεm/m mutation had little influence on the period of circadian rhythms (Fig. 2A,D). In contrast, having a CKIδm/m mutation would significantly lengthen PER2Luc rhythms (Fig. 2A,D). These results are compatible with the observation that mouse embryonic fibroblasts (MEFs) along with the SCN and liver explants from CKIε knockout mice show normal circadian period lengths, while those from CKIδ loss-of-function mutants exhibit longer periods (Etchegaray et al., 2009; Lee et al., 2009; Etchegaray et al., 2010; Lee et al., 2011). CKIδ+/m cells showed no drastic changes in period even if CKIε was mutated simultaneously, indicating that 1 allele of CKIδ is sufficient to maintain the circadian clock with a period change of less than 1 h (Fig. 2B,D). Intriguingly, a CKIε+/m mutation in CKIδm/m background resulted in a further elongation of the circadian period (~28 h) (Fig. 2C,D). Thus, CKIε should assume control over clock speed when CKIδ is disrupted. This result clearly demonstrates that CKIδ and CKIε have redundant roles in the regulation of the circadian clock.
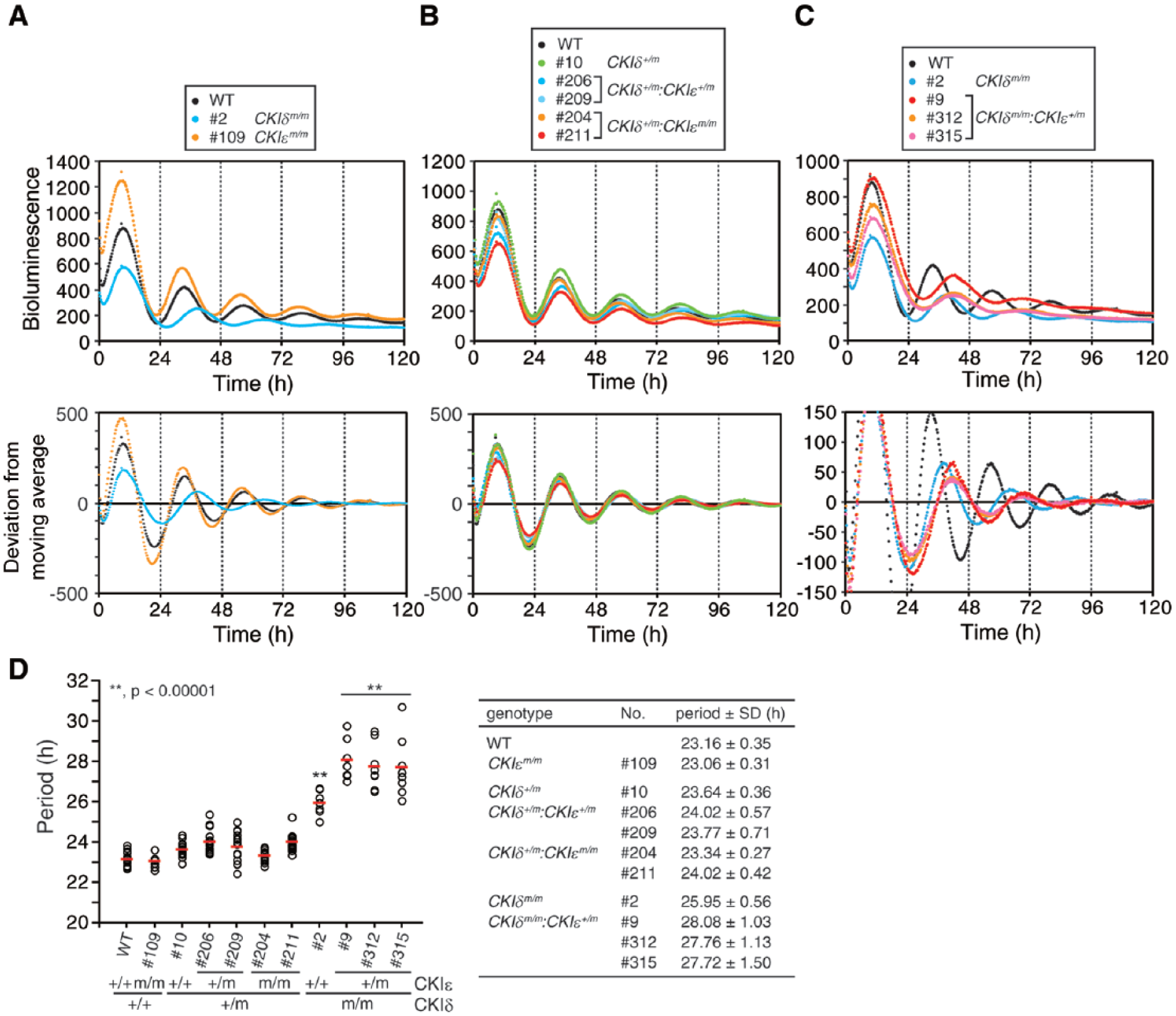
CKIδ and CKIε have an overlapping role in regulation of circadian period. (A) CKIδ but not CKIε mutant cells exhibit rhythms with a longer period. The average of the raw (upper) and detrended (bottom) data is shown (n = 8). (B) One allele of intact CKIδ is sufficient to maintain a normal circadian period. The average of raw (upper) and detrended (bottom) data is shown (n = 16). (C) A heterozygous mutation of CKIε results in a longer period in CKIδm/m background. The average of raw (upper) and detrended (bottom) data is shown (n = 8). (D, E) The period length of the mutant cells is analyzed in A to C. Each open circle indicates the value calculated from the data of 1 embryoid body, and red bars indicate the mean (**p < 0.00001).
CKIδ:CKIε:Cry2 Triple Gene-Targeted Cells Show Circadian Rhythms with a Yet Longer Period
To generate ESCs that developed circadian clocks with even longer periods, we targeted Cry2 in a CKIδm/m:CKIε+/m background. Consequently, we obtained CKIδm/m:CKIε+/m:Cry2m/m triple gene-targeted ESCs (Fig. 3A). Upon in vitro differentiation, the cells developed bioluminescence rhythms with much longer periods (~31 h) than any other single or double gene-targeted mutant cells (Fig. 3B).
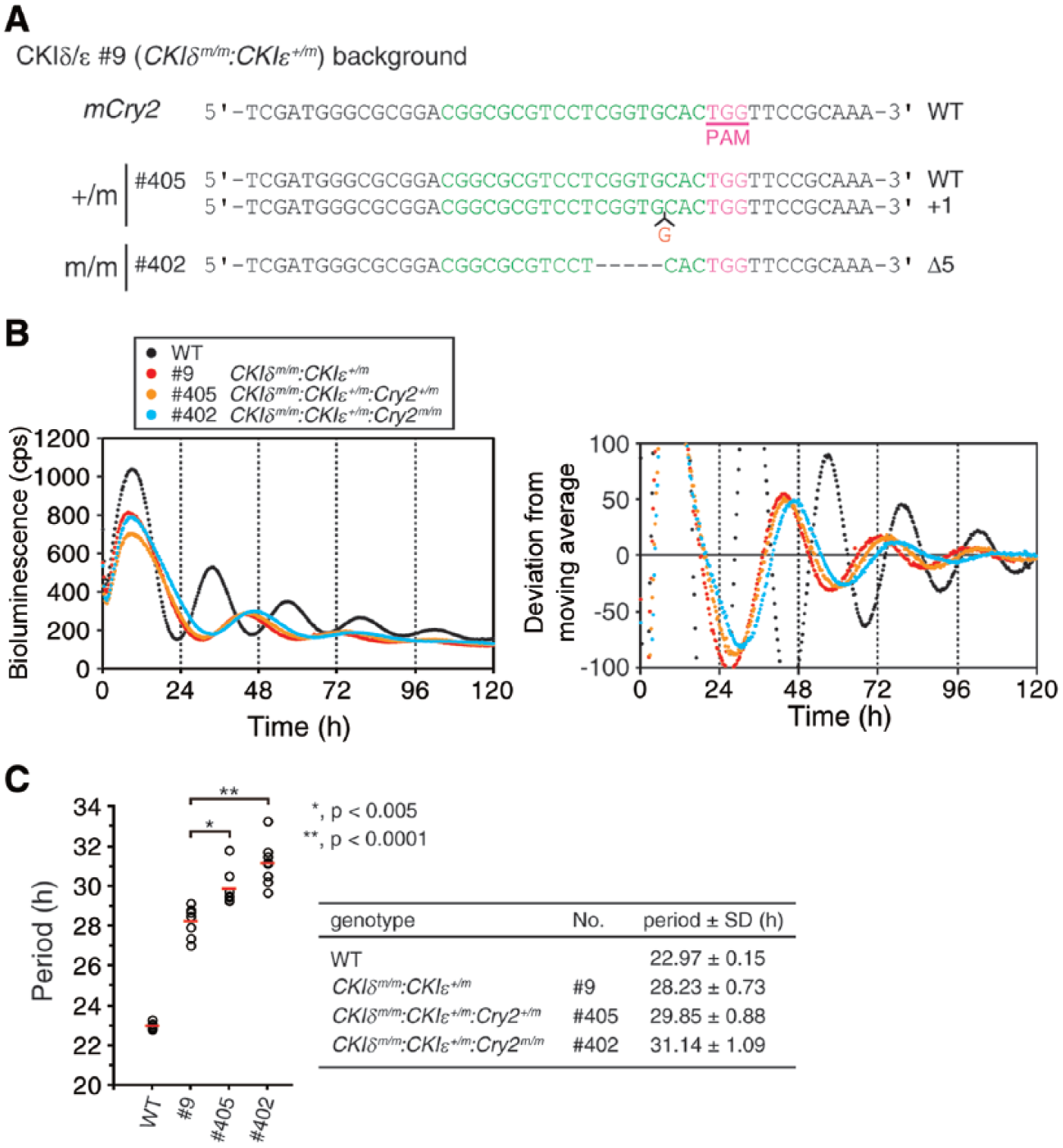
CKIδ/CKIε/Cry2 multiplexed mutant cells show circadian rhythms with an extraordinarily long period. (A) The schematic of Cry2 target region and the sequences of the targeted clones. (B) CKIδm/m:CKIε+/m:Cry2m/m cells show a longer period than CKIδm/m:CKIε+/m cells. The average of the raw (left) and detrended (right) data is shown (n = 8). (C) The period length of the mutant cells is analyzed in B. Each open circle indicates the value calculated from the data of 1 embryoid body, and red bars indicate the mean (*p < 0.005; **p < 0.0001).
Temperature Compensation of Circadian Rhythms in Long Period Mutants
To reveal the relationship between the period length and temperature compensation of circadian rhythms, we examined whether ambient temperature would affect the period of the circadian rhythms that developed in the mutant cells that we generated. We measured the bioluminescence rhythms of in vitro differentiated cells from various genotypes under different temperature conditions. In control PER2Luc cells, the period length was well compensated for temperatures in a range of 33.5 to 36.0 °C as we previously reported (Fig. 4 and Suppl. Fig. S3) (Umemura et al., 2013). Although higher temperatures (over 38.5 °C) led to longer periods (data not shown), they were likely to be associated with impaired cellular physiology. The temperature coefficient Q10 value was 1.00, which indicates the temperature-insensitive property of the rhythms and is comparable to previously reported values for circadian rhythms (Barrett and Takahashi, 1995; Ruby et al., 1999; Izumo et al., 2003; Tsuchiya et al., 2003). Significantly, in the period mutants with single or combined mutations in the CKIδ, CKIε, Cry1, and Cry2 loci, the period length was largely as temperature insensitive as the wild-type (WT) cells (Fig. 4 and Suppl. Fig. S3). Furthermore, even the longest period mutant (CKIδm/m:CKIε+/m:Cry2m/m) kept temperature-compensated rhythms with a Q10 value of 1.09, although the Q10 values were slightly affected by mutations of CKIδ/ε (Fig. 4; compare the Q10 values of CKIδm/m, CKIδm/m:CKIε+/m, and CKIδm/m:CKIε+/m:Cry2m/m mutants with that of WT). These results demonstrate that the genetically determined period ranging from 22 to 31 h is temperature compensated in mammalian circadian rhythms and furthermore suggest that CKIδ/ε may be involved in the maintenance of temperature compensation.
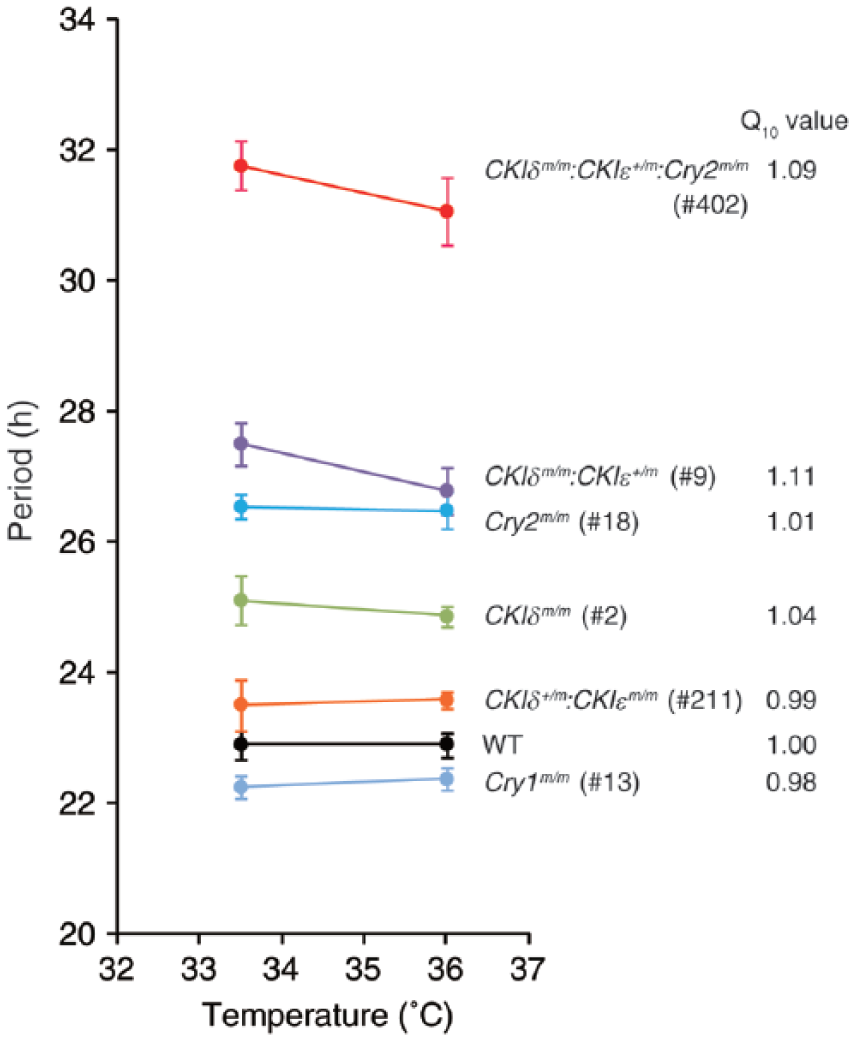
Temperature compensation of circadian period in Cry1/2 and CKIδ/ε mutant cells. PER2Luc bioluminescence rhythms were measured at 33.5 °C and 36.0 °C. The clone numbers of each mutant are indicated in parentheses. The values are the means ± SEM (n = 3 or 4). The Q10 values are also indicated.
Discussion
In this study, we demonstrated that CRISPR/Cas9-mediated gene targeting could recapitulate the circadian phenotypes of well-known clock genes in knockout mice and knockout mice cells. Taking advantage of the CRISPR/Cas9 system that enables multiple gene targeting easily and quickly, we could also generate CKIδm/m:CKIε+/m:Cry2m/m triple gene mutant cells, which turned out to exhibit extra-long circadian periods. The low rate of off-target mutations indicates the specificity of the CRISPR/Cas9-mediated gene targeting in ESCs. Thus, the ESC-based analysis of the mammalian circadian clock combined with the CRISPR/Cas9-system is a useful tool to investigate the molecular mechanisms underlying the 24-h periodicity and the robustness of the circadian clock. Interestingly, CKIδm/m:CKIεm/m ESCs could not obtained in this study. We speculate that it may due to the function of CKIδ/ε on cellular proliferation and losing the population during the selection to pick up the mutant clones.
Our data demonstrate that the period length of circadian rhythms is temperature compensated, even in clock gene mutant cells that exhibit an extra-long period. This means that temperature compensation is maintained even in a circadian oscillator lacking multiple clock components that are required for keeping a normal period length. Also, in Neurospora and Drosophila, several clock gene mutants that exhibit abnormal period lengths have been found to keep temperature-compensated rhythms (Hong et al., 2007). Thus, we speculate that temperature compensation is not solely achieved by a delicate balance of period-increasing and period-decreasing processes of which rate constants are temperature dependent. Rather, as proposed previously (Hong et al., 2007; Francois et al., 2012), there may be a limited number of processes that determine the temperature-insensitive property of the circadian period. Given that there are mutations found to cause both an altered period and compromised temperature compensation in Neurospora and Drosophila (Gardner and Feldman, 1981; Loros and Feldman, 1986; Huang et al., 1995; Rutila et al., 1996; Liu et al., 1997; Sawyer et al., 1997; Matsumoto et al., 1999; Ruoff et al., 2005; Hong et al., 2007), it seems that some processes would be critical for regulating both the period and its temperature sensitivity. It has been reported that in cultured retina of CKIεtau mutant hamsters, temperature compensation of melatonin secretion rhythms is altered (Tosini and Menaker, 1998). In addition, a previous study has reported that phosphorylation of PER2-derived peptides by CKIδ/ε is a temperature-insensitive reaction (Isojima et al., 2009). As the Goodwin model has predicted that the degradation rate of clock proteins is responsible for setting the period (Ruoff, 1992; Ruoff et al., 1996), it is possible that CKIδ/ε-dependent phosphorylation and degradation of PER proteins contribute to the temperature-insensitive property of the period length. In this context, slight changes in the Q10 value of CKIδ/ε disruption may imply the involvement of CKIδ/ε in the maintenance of temperature compensation, and residual CKIε might be sufficient to keep rhythms in CKIδm/m:CKIε+/m mutants temperature compensated.
As a recent study has shown that in cyanobacteria, the structurally determined slow ATPase activity sets the circadian periodicity (Abe et al., 2015), it may be also possible that some intramolecular mechanisms define the circadian period and its temperature compensation in mammals. For a complete understanding of the principle of the temperature compensation property in mammalian circadian clocks, further study is necessary. CRISPR/Cas9-mediated gene targeting and the differentiation-dependent emergence of circadian rhythms in ESCs would be a useful assay system for introducing the desired mutations into clock genes as well as analyzing the genetically determined properties of the mammalian circadian clock.
Footnotes
Acknowledgements
We thank Drs. Munehiro Ohashi and Tasuku Hara (Kyoto Prefectural University of Medicine) for their valuable discussions and technical support. The authors were supported in part by a Japan Science and Technology Agency Precursory Research for Embryonic Science and Technology program (to K.Y.), grants-in-aid for scientific research from the Japan Society for the Promotion of Science (to K.Y., Y.T., and Y.U.), and the Takeda Science Foundation (to K.Y.).
Author Contributions
Y.T. and K.Y. designed research; Y.T., Y.U., T.H., and M.H. performed experiments; Y.T., Y.U., H.I., Y.M., N. K., H.I., and K.Y. analyzed data; H.I. and K.Y. contributed analytic tools; and Y.T. and K.Y. wrote the paper.
Conflict of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.