Abstract
The hematologic system performs a number of essential functions, including oxygen transport, the execution of the immune response against tumor cells and invading pathogens, and hemostasis (blood clotting). These roles are performed by erythrocytes (red blood cells), leukocytes (white blood cells), and thrombocytes (platelets), respectively. Critically, circadian rhythms are evident in the function of all 3 cell types. In this review, we describe these oscillations, explore their mechanistic bases, and highlight their key implications. Since erythrocytes are anucleate, circadian rhythms in these cells testify to the existence of a nontranscriptional circadian clock. From a clinical perspective, leukocyte rhythms could underlie daily variation in the severity of allergic reactions, the symptoms of chronic inflammatory diseases, and the body’s response to infection, while the rhythmic properties of thrombocytes may explain daily fluctuations in the incidence of heart attack and stroke. Consequently, the efficacy of treatments for these conditions is likely to depend on the timing of their administration. Last, we outline preliminary evidence that circadian disruption in the hematologic system could contribute to the deleterious effects of poor diet, shift work, and alcohol abuse on human health.
Circadian rhythms are endogenous 24-h oscillations in physiology and behavior that enable an organism to anticipate and adapt to the changing temporal demands of the environment (Johnson et al., 2004). The term circadian is derived from the Latin words circa, meaning “approximately,” and diem, meaning “day.” The sleep/wake cycle is perhaps the most obvious example of a circadian rhythm. Other examples include daily fluctuations in locomotor activity, core body temperature, heart rate, blood pressure, hepatic metabolism, and hormone secretion (Hastings et al., 2008). Circadian rhythms have been characterized well in the brain and various peripheral tissues, but until recently, relatively little attention had been paid to circadian rhythms in the hematologic system.
The body of an average adult contains approximately 4.5 liters of blood, which accounts for roughly 7% of total body mass (Cameron et al., 1999). Plasma constitutes 55% of whole blood, while blood cells account for the remaining 45% (Waugh and Grant, 2010). There are 3 types of blood cells, which serve distinct functions. Erythrocytes (red blood cells) transport oxygen from the lungs to body tissues, leukocytes (white blood cells) coordinate and execute the immune response against tumor cells and invading pathogens, and thrombocytes (platelets) are essential for hemostasis (blood clotting), which limits blood loss in the event of blood vessel damage (Waugh and Grant, 2010).
In this review, we describe the rhythmic properties of erythrocytes, leukocytes and thrombocytes, and explain how these properties could underlie daily rhythms in the severity of allergic reactions and the incidence of heart attack and stroke. In addition, we consider how changes in the rhythmic properties of blood cells could mediate the effects of poor diet, shift work, and alcohol abuse on human health.
Circadian Properties of Erythrocytes
Erythrocytes are the most common type of blood cell, accounting for approximately 25% of all cells in the human body (Waugh and Grant, 2010). They transport oxygen from the lungs to tissues, where it is used for aerobic respiration, generating energy in the form of adenosine triphosphate (ATP). To fulfill this role, the cytoplasm of erythrocytes contains hemoglobin, an iron-containing oxygen transport protein. To accommodate as much hemoglobin as possible, erythrocytes lack a nucleus and other organelles.
Until recently, it was widely believed that most (if not all) circadian rhythms were the product of autoregulatory transcriptional-translational feedback loops (TTFLs) (see Reppert and Weaver, 2002). In mammals, the core components of the primary TTFL are the genes Period (PER1 and PER2), Cryptochrome (CRY1 and CRY2), Brain and muscle ARNT-Like protein-1 (BMAL1), and Circadian locomotor output cycles kaput (CLOCK) (see Reppert and Weaver, 2002). Since erythrocytes are anucleate, they are incapable of transcription. Hence, a TTFL cannot exist in these cells.
The rhythmic properties of human erythrocytes first received attention when 24-h oscillations were observed in the enzymatic activity of hexokinase and adenonsine deaminase (Brok-Simoni et al., 1976; Cornelissen et al., 1985). In both studies, however, blood samples were collected at regular intervals from individuals who were not kept under constant conditions; therefore, these oscillations cannot be considered truly circadian (see Johnson et al., 2004). Instead, they could be driven by external cues.
In a more recent study by O’Neill and Reddy (2011), blood samples were taken from 3 healthy volunteers, and erythrocytes were extracted by centrifugation. Erythrocytes were cultured under constant conditions (37 °C; constant darkness), and over a period of 60 h, levels of overoxidized peroxiredoxins (PRX-SO2/3) were quantified at 4-h intervals by immunoblotting. Peroxiredoxins (PRXs) are highly conserved antioxidant enzymes that control intracellular levels of harmful reactive oxygen species (ROS), such as superoxide and hydrogen peroxide (Hall et al., 2009). In the process of reducing ROS, the catalytic cysteine of “typical” 2-Cys PRXs is oxidized to a sulfenic acid form (PRX-SO), which in turn can be overoxidized to a sulfinic acid form (PRX-SO2) (see O’Neill and Reddy, 2011). This renders the PRX catalytically inactive until it is reduced and reactivated by the ATP-dependent enzyme sulfiredoxin (Rhee et al., 2007; Jeong et al., 2012). Occasionally, PRX-SO2 can be hyperoxidized to a sulfonic acid (PRX-SO3) form, a change that is thought to be irreversible (see Fig. 1).
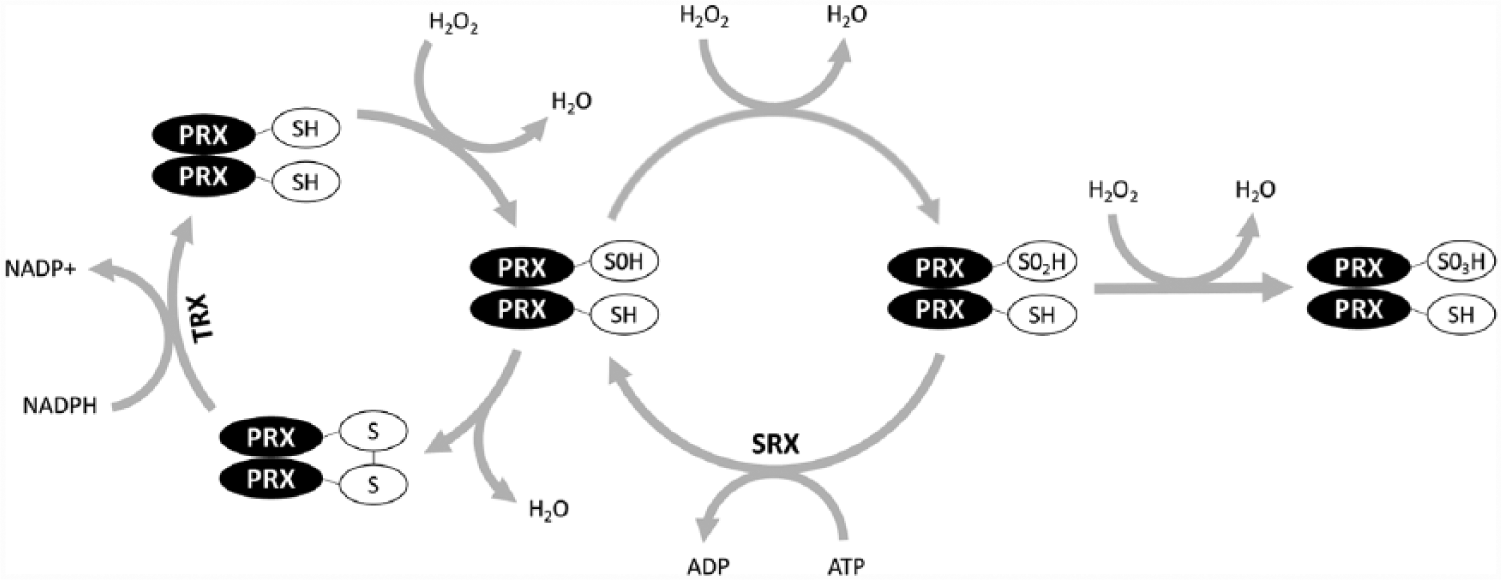
The “typical” 2-Cys peroxiredoxin system. The catalytic cysteine of peroxiredoxin (PRX) can be oxidized to a sulfenic acid form (PRX-SO), overoxidized to a sulfinic acid form (PRX-SO2), and hyperoxidized to a sulfonic acid form (PRX-SO3). The latter step is likely irreversible, whereas PRX-SO and PRX-SO2 can be recycled with the aid of thioredoxin (TRX) and sulfiredoxin (SRX), respectively. ADP = adenosine diphosphate; ATP = adenosine triphosphate; H2O = water; H2O2 = hydrogen peroxide; NADP+ = nicotinamide adenine dinucleotide phosphate; NADPH = reduced NADP+; SH = sulfhydryl group. Figure adapted from Reddy and Rey (2014).
Significantly, robust oscillations in PRX-SO2/3 levels were observed in the cultured human erythrocytes. More important, these oscillations met the 3 criteria of circadian rhythmicity (see Johnson et al., 2004): they persisted under constant conditions with an endogenous period length of approximately 24 h, they could be synchronized to environmental cues (in this case, 24-h temperature rhythms), and they were temperature compensated (period length did not differ between erythrocytes cultured at 32 °C and 37 °C). The observed PRX-SO2/3 rhythms were not the result of sample contamination with nucleated cells, since they persisted in the presence of transcriptional and translational inhibitors. Besides PRX-SO2/3 rhythms, circadian oscillations were observed in several other parameters, including Peroxiredoxin-2 (PRX2) expression, hemoglobin oxidation, ATP levels, and the abundance of the reducing agents NADH and NADPH (O’Neill and Reddy, 2011). Taken together, these data clearly demonstrate that circadian rhythms can exist in the absence of transcription, which provides strong evidence for the existence of a nontranscriptional circadian clock. Future studies should determine whether other antioxidant enzymes display circadian characteristics in erythrocytes. The activities of glutathione peroxidase, catalase, and superoxide dismutase all show daily variation in human erythrocytes (Baskar et al., 2004), although cells were not maintained under constant conditions in these experiments.
There are 6 isoforms of PRX in mammals, but only PRX1, PRX2, and PRX6 are expressed in erythrocytes (see Cho et al., 2014). Using an antibody specific for PRX6-SO2/3 and another specific for PRX1/2-SO2/3, Rhee and colleagues observed rhythmic oxidation of PRX1/2, not PRX6, in wild-type mouse erythrocytes (Cho et al., 2014). Crucially, these rhythms were also present in erythrocytes harvested from PRX1–/– mice, but not in erythrocytes taken from PRX2–/– mice. These data indicate that PRX-SO2/3 rhythms in mammalian erythrocytes reflect the modification of PRX2 rather than PRX1 or PRX6. In addition, Rhee and colleagues demonstrated that PRX-SO2/3 rhythms could be abolished by exposing wild-type mouse erythrocytes to carbon monoxide (CO) (Cho et al., 2014). Since hemoglobin has a higher affinity for CO than oxygen, CO prevents the auto-oxidation of hemoglobin and the resultant production of superoxide and hydrogen peroxide. These data imply that the rising phase of PRX-SO2/3 oscillations results from rhythmic ROS generation, which in turn is a consequence of rhythmic hemoglobin oxidation (O’Neill and Reddy, 2011; Cho et al., 2014).
What causes the decay phase of PRX-SO2/3 oscillations? Sulfiredoxin was a potential candidate, since it is important for recycling PRX proteins back to their reduced states. However, sulfiredoxin is not responsible to any appreciable degree, given that PRX-SO2/3 rhythms are largely unaltered in erythrocytes derived from Sulfiredoxin knockout (SRX–/–) mice (Cho et al., 2014). This is perhaps unsurprising given that SRX levels are relatively low in erythrocytes (Cho et al., 2014), and given that PRX-SO2/3 oscillations have been reported in eukaryotic organisms that do not express SRX (see Stangherlin and Reddy, 2013). By contrast, the application of the proteasome inhibitors MG132 and lactacystin eliminates the decay phase of the PRX-SO2/3 rhythm in wild-type mouse erythrocytes (Cho et al., 2014). This finding suggests that PRX-SO2/3 oscillations are dependent on the proteasomal degradation of PRX-SO2/3. Consistent with this, the coincubation of the 20S proteasome complex with PRX2 and PRX2-SO2/3 results in the selective degradation of PRX2-SO2/3 (Cho et al., 2014).
Besides human and mouse erythrocytes, PRX-SO2/3 rhythms have been observed in mouse liver, adrenal cortex and suprachiasmatic nucleus, mouse embryonic fibroblasts (MEFs), heads of Drosophila melanogaster, seedlings from the plant Arabidopsis thaliana, mycelia of the fungus Neurospora crassa, and the unicellular alga Ostreococcus tauri (O’Neill and Reddy, 2011; O’Neill et al., 2011; Edgar et al., 2012; Kil et al., 2012). PRX-SO2/3 rhythms have also been reported in 2 prokaryotic species: the archaeon Halobacterium salinarum and the bacterium Synechococcus elongatus (Edgar et al., 2012). Given that PRX-SO2/3 rhythms are present in both eukaryotic and prokaryotic organisms, and given that the proteins themselves are highly conserved across species, PRX-SO2/3 rhythms must be evolutionarily ancient. Indeed, they likely predate the emergence of circadian TTFLs, which are believed to have evolved independently in separate species (Edgar et al., 2012). In the future, it will be important to establish whether PRX-SO2/3 rhythms are a core component of the nontranscriptional clockwork, or a downstream marker of redox activity. Moreover, it will be interesting to see if there is any crosstalk between PRX-SO2/3 rhythms and the TTFL in eukaryotic organisms. Preliminary evidence suggests that there is a reciprocal relationship between the two. In one direction, the period length of PRX-SO2/3 rhythms is extended in embryonic fibroblasts derived from CRY1/2–/– mice (O’Neill and Reddy, 2011), while in the other, the RNAi knockdown of specific PRX isoforms affects the period length and amplitude of clock gene rhythms in human U2OS cells (Zhang et al., 2009; O’Neill and Reddy, 2011).
Last, it should be noted that it is not only subcellular processes within erythrocytes that undergo 24-h oscillations. The number of erythrocytes in the blood also varies rhythmically (e.g., Oishi et al., 2006a). This parameter appears to be under clock gene control, since the rhythm is suppressed and phase-advanced in CLOCK mutant mice (Oishi et al., 2006a).
Circadian Properties of Leukocytes
Leukocytes account for about 1% of total blood volume and are involved in the immune response against infection and cancer (Alberts et al., 2002). They can be divided into 5 categories—lymphocytes, neutrophils, monocytes, eosinophils, and basophils—which differ in their abundance, structure, and function (see Table 1). In turn, lymphocytes can be subdivided into natural killer (NK) cells, B cells, and T cells. Like erythrocytes, the number of leukocytes in the blood displays 24-h oscillations (e.g., Oishi et al., 2006a). Again, this parameter seems to be under clock gene control, as the rhythm is damped and phase-delayed in CLOCK mutant mice (Oishi et al., 2006a). Unlike erythrocytes, leukocytes are nucleated. Rhythmic clock gene expression has been reported in a wide range of mammalian leukocytes (see Labrecque and Cermakian, 2015), and, more important, these rhythms persist in leukocytes cultured under constant conditions, which implies that they are genuinely circadian (e.g., Ebisawa et al., 2010; Perez-Aso et al., 2013).
Rhythmic Properties of Leukocytes.
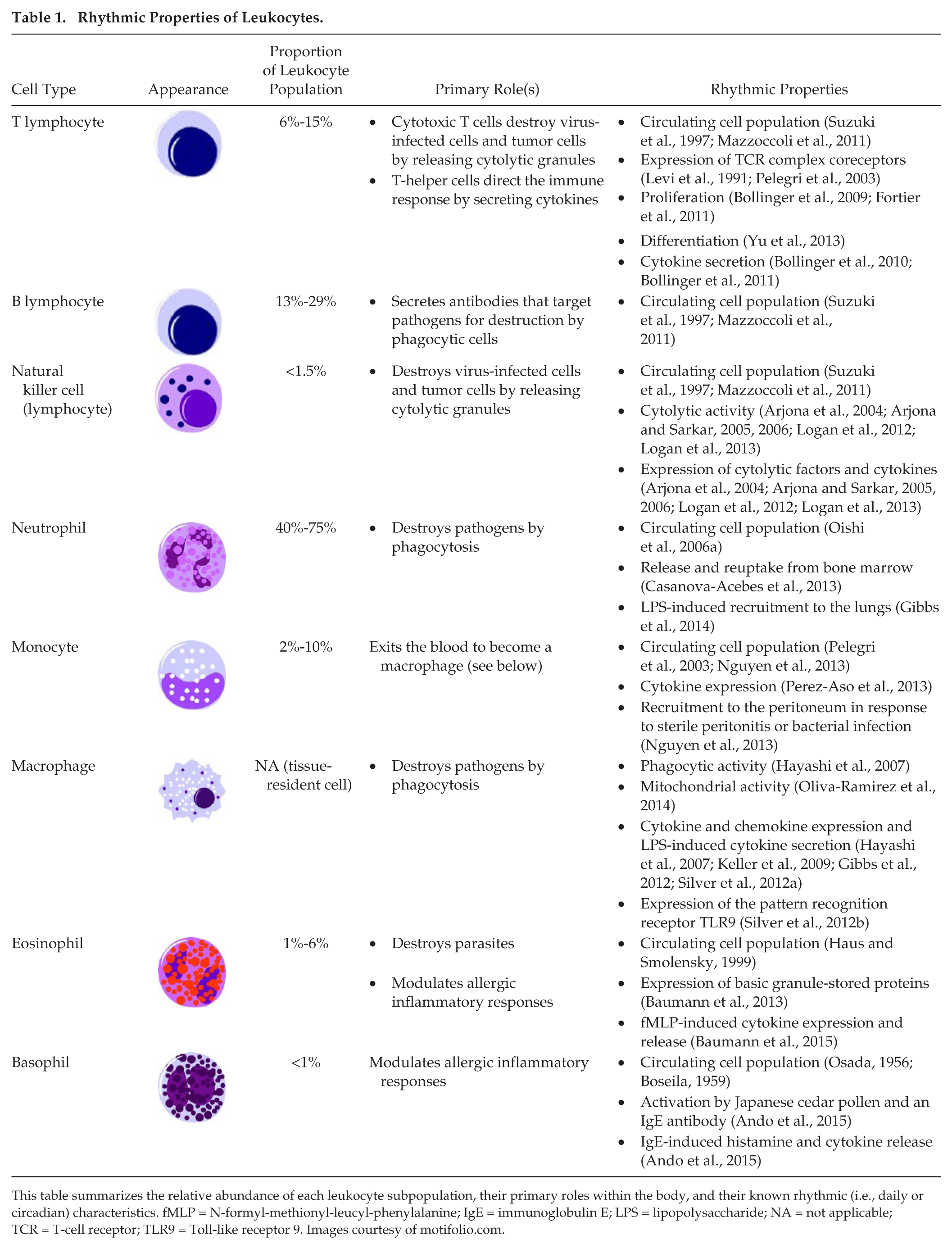
This table summarizes the relative abundance of each leukocyte subpopulation, their primary roles within the body, and their known rhythmic (i.e., daily or circadian) characteristics. fMLP = N-formyl-methionyl-leucyl-phenylalanine; IgE = immunoglobulin E; LPS = lipopolysaccharide; NA = not applicable; TCR = T-cell receptor; TLR9 = Toll-like receptor 9. Images courtesy of motifolio.com.
Since it is relatively easy to quantify clock gene expression in leukocytes, and because the phase of the observed rhythms often correlates with the individual’s physiological and behavioral rhythms (e.g., the sleep/wake cycle), leukocyte clock gene rhythms could be used as a biomarker during the diagnosis and treatment of sleep and circadian rhythm disorders (Boivin et al., 2003; Takimoto et al., 2005; Archer et al., 2008; Kusanagi et al., 2008). In fact, they provide a straightforward and noninvasive means to evaluate circadian system function in any disease (Fukuya et al., 2007). However, it is important to note that clock gene rhythms in leukocytes are by no means identical to those in the suprachiasmatic nucleus (SCN), the “master” circadian pacemaker in the brain (Teboul et al., 2005; Weigl et al., 2013). Moreover, leukocyte and SCN rhythms can become uncoupled under certain circumstances (e.g., Haimovich et al., 2010).
Interestingly, there are daily rhythms in the severity of allergic conditions such as allergic rhinitis (e.g., Smolensky et al., 2007); the symptoms of chronic inflammatory diseases such as asthma, chronic obstructive pulmonary disease, and rheumatoid arthritis (e.g., Kessler et al., 2011; Gibbs and Ray, 2013; Durrington et al., 2014); and the body’s response to infection (e.g., Pollmacher et al., 1996; Alamili et al., 2014). As the major cellular effectors of the immune system, leukocytes are likely responsible for these rhythms. At this point, the reader is referred to a recent review by Labrecque and Cermakian (2015), which outlines the rhythmic properties of lymphocytes, neutrophils and monocytes, and explains how these properties might contribute to rhythmic immune responsiveness. In the paragraphs that follow, we describe the rhythmic properties of eosinophils and basophils. Table 1 summarizes the key discoveries in this field.
Eosinophils
Eosinophils comprise 1% to 6% of the total leukocyte population (Nigam and Ayyagari, 2007), and are the body’s primary defense against external parasites such as tapeworms and hookworms (Alberts et al., 2002). They also modulate allergic inflammatory responses through the release of cytokines, chemokines, growth factors, lipid mediators, and basic granule-stored proteins, including major basic protein (MBP), eosinophil peroxidase (EPO), eosinophilic cationic protein (ECP), and eosinophil protein X/eosinophil-derived neurotoxin (EPX/EDN) (Kariyawasam and Robinson, 2006; Hogan et al., 2008; Stone et al., 2010). Like other blood cells, the number of eosinophils in the blood exhibits daily variation in healthy adults (see Haus and Smolensky, 1999), although the causative mechanism is not yet known.
It is well established that the symptoms of allergic conditions, such as allergic rhinitis, are worst between midnight and early morning (Smolensky et al., 2007). It is possible that eosinophils contribute to this daily variation, since cultured human eosinophils exhibit circadian oscillations in ECP and EPX/EDN expression (Baumann et al., 2013). Consistent with this finding, a study of 7 healthy children revealed 24-h oscillations in serum ECP and EPX/EDN concentration, with peak levels of both proteins occurring at 0600 h (Wolthers and Heuck, 2003). Daily rhythms have also been observed in nasal ECP levels and in sputum and urine EPX/EDN levels (Aoyagi et al., 1999; Storm van’s Gravesande et al., 1999; Panzer et al., 2003). Significantly, the immune responsivity of eosinophils varies in a circadian manner. In a recent experiment, cultured human eosinophils were activated with the chemotactic tripeptide N-formyl-methionyl-leucyl-phenylalanine (fMLP) at 4-h intervals over a 24-h period (Baumann et al., 2015). The fMLP-induced expression and release of the proinflammatory cytokines IL-8/CXCL8 and CCL2/MCP-1 displayed circadian rhythmicity, as did fMLP-induced phosphorylation of the signaling molecule ERK1/2, which has been linked to cytokine and chemokine expression in eosinophils (Shamri et al., 2013).
Basophils
Basophils are the least common type of leukocyte, accounting for less than 1% of the total population (Nigam and Ayyagari, 2007; Marone et al., 2014). The number of circulating basophils displays daily variation in healthy adults (Osada, 1956; Boseila, 1959). Together with tissue-resident mast cells, basophils release the chemical histamine (Alberts et al., 2002; Marone et al., 2014), which is central in allergic inflammatory responses, and exhibits daily variation in humans (Friedman et al., 1989). They also secrete the anticoagulant heparin, eosinophil- and neutrophil-attracting chemokines (e.g., IL-8/CXCL8), and cytokines that promote T-helper cell differentiation (e.g., IL-4) (Stone et al., 2010; Marone et al., 2014). In a recent study, cultured human basophils were activated by Japanese cedar pollen or an anti–immunoglobulin E (IgE) antibody (Ando et al., 2015). Both interventions activated a significantly greater number of basophils (as determined with the activation marker CD203c) at 0700 h relative to 1900 h. To determine whether basophil-mediated immune responses are under clock gene control, the IgE-induced secretion of histamine and the cytokine IL-4 was quantified at 2 circadian time points, 12 h apart, in cultured basophils derived from wild-type and CLOCK mutant mice (Ando et al., 2015). Time-of-day–dependent variation in IgE-induced histamine and IL-4 release was seen in wild-type basophils but not in CLOCK mutant basophils (Ando et al., 2015). This implies that the rhythmic immune responsivity of basophils is clock gene-regulated, and that it contributes to rhythmicity in the symptoms of allergic conditions.
Circadian Properties of Thrombocytes
Thrombocytes are fragments of cytoplasm derived from bone marrow–resident cells called megakaryocytes (Machlus et al., 2014). Like erythrocytes, they are anucleate like erthrocytes, but possess some organelles such as mitochondria. In a similar fashion to red blood cells, they display circadian characteristics despite lacking a nucleus. For example, levels of the antioxidant enzyme glutathione oscillate with 24-h periodicity in human platelets cultured in constant light or constant dark conditions (Radha et al., 1985).
Together with the fibrous protein fibrin, platelets form blood clots to limit blood loss in the event of blood vessel damage (“hemostasis”). Although this is part of the healthy healing process, blood clots occasionally obstruct the flow of blood within a blood vessel (“thrombosis”), which can lead to myocardial infarction (heart attack) or ischemic stroke. Interestingly, the occurrence of these events demonstrates daily variation; they take place most frequently in the morning, between 0600 and 1200 h (Muller et al., 1985; Muller et al., 1987; Casetta et al., 2002). This may reflect 24-h rhythms in platelet function (see Hartley, 2012). The first evidence of this was provided in 1987, with the demonstration of daily rhythms of platelet aggregation in healthy adult males (Tofler et al., 1987). More recently, the same topic was investigated using a forced desynchrony protocol, in which healthy volunteers spent 13 days in constant dim light while they were subjected to an enforced 20-h rest/activity cycle (Scheer et al., 2011). Circadian phase was determined from measurements of core body temperature. Regular blood sampling revealed robust circadian oscillations in markers of platelet aggregation, platelet-leukocyte adhesion, and platelet-endothelial cell adhesion. Crucially, each of these rhythms peaked at circadian time points corresponding to 0800 to 0900 h (Scheer et al., 2011).
Platelet aggregation is under clock gene control, since it demonstrates daily variation in wild-type mice but not CLOCK mutant mice (Ohkura et al., 2009). There is also evidence that CLOCK regulates the rhythmic expression of Thrombopoietin (TPO), which in turn governs the generation of platelets from megakaryocytes (Tracey et al., 2012). This mechanism may contribute to daily rhythmicity in the number of circulating platelets, which peaks in the late afternoon in healthy humans (Bremner et al., 2000; Ohkura et al., 2009; Scheer et al., 2011; Hartley, 2012). The clock genes BMAL1 and PER2 are also implicated in platelet function. BMAL1–/– mice exhibit a prothrombotic phenotype characterized by more rapid hemostasis and wound healing, enhanced platelet aggregation and adhesion, and increased expression of the prothrombotic genes von Willebrand factor (vWF), Fibrinogen, and Plasminogen activator inhibitor-1 (PAI-1) (Somanath et al., 2011). vWF is rhythmically expressed, and its expression is directly regulated by BMAL1 (Somanath et al., 2011). In contrast to BMAL1–/– mice, PER2–/– mice demonstrate slower hemostasis, a 50% reduction in blood platelet content, and significant impairments in platelet formation, apoptosis, and aggregation (Zhao et al., 2011).
Plasminogen Activator Inhibitor-1 (Pai-1) Rhythms
The prothrombotic protein PAI-1 is secreted by endothelial cells, and inhibits tissue plasminogen activator (tPA) and urokinase (uPA), which promote fibrinolysis or the breakdown of blood clots (see Fig. 2A). Plasma levels of PAI-1 oscillate with 24-h periodicity, with levels peaking at the start of the active phase in both humans and rodents (Andreotti and Kluft, 1991; Oishi, 2009) (see Fig. 2B). This may explain why fibrinolysis is lowest at this time of day, and by extension, why the incidence of myocardial infarction and ischemic stroke both peak at this time (Andreotti and Kluft, 1991; Oishi, 2009). A polymorphism in the PAI-1 promoter is associated with mildly elevated plasma PAI-1 levels (see Chong et al., 2006) and markedly increased rhythmicity in PAI-1 expression (Hoekstra et al., 2002; van der Bom et al., 2003), which may confer a greater susceptibility to cardiovascular events in carriers.
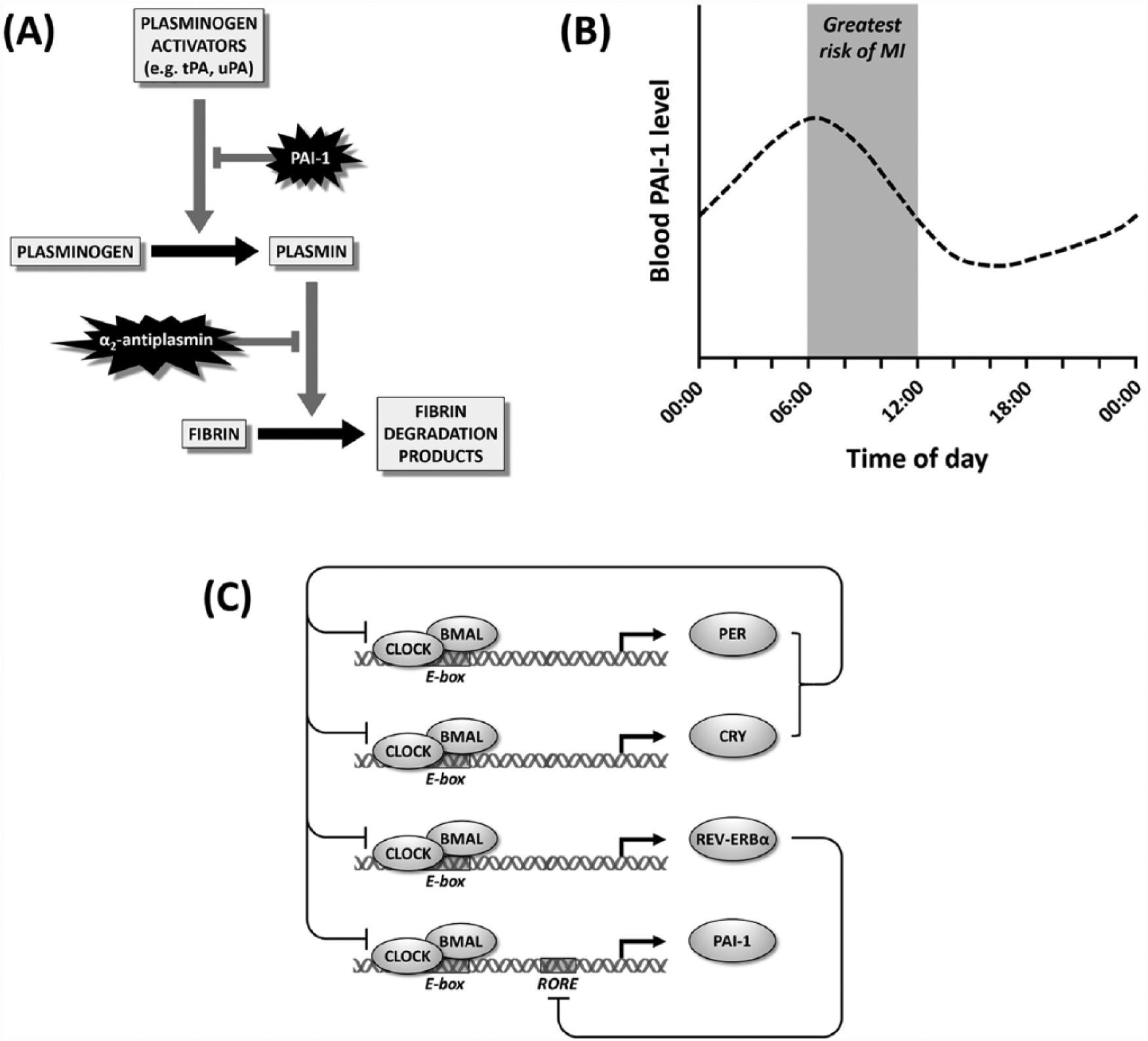
(A) The fibrinolytic pathway. The plasminogen activators tissue plasminogen activator (tPA) and urokinase (uPA) convert plasminogen into plasmin, which breaks down fibrin clots. Plasminogen activator inhibitor-1 (PAI-1) inhibits tPA and uPA, while α2-antiplasmin inhibits plasmin. (B) Blood PAI-1 level varies rhythmically, peaking at approximately 0630 h in humans. This coincides with the peak incidence of myocardial infarction (MI). (C) PAI-1 expression is under clock gene control. CLOCK:BMAL heterodimers drive transcription of PER, CRY, REV-ERBα, and PAI-1 by binding to E-boxes in their promoters. PER and CRY heterodimerize and repress CLOCK:BMAL-mediated transcription. REV-ERBα inhibits PAI-1 transcription by binding to retinoic acid receptor–related orphan receptor response elements (ROREs) in the PAI-1 promoter. Figure 2B adapted from Scheer and Shea (2014).
Critically, the PAI-1 rhythm persists in humans subjected to a “forced desynchrony” protocol, which implies that it is truly circadian (Scheer and Shea, 2014). Indeed, there is plentiful evidence that PAI-1 expression is under clock gene control. Its expression is reduced but still rhythmic in CLOCK RNAi mice and PER2-overexpressing mice (Oishi et al., 2009a; Cheng et al., 2011), reduced and nonrhythmic in CLOCK mutant mice (Minami et al., 2002; Ohkura et al., 2006), and increased and nonrhythmic in CRY1/CRY2–/– mice and BMAL1–/– mice (Ohkura et al., 2006; Masuda et al., 2009; Somanath et al., 2011). Accordingly, fibrinolysis is increased but still rhythmic in CLOCK RNAi mice (Cheng et al., 2011), increased and nonrhythmic in CLOCK mutant mice (Ohkura et al., 2006), and reduced and nonrhythmic in CRY1/CRY2–/– mice (Ohkura et al., 2006). Rhythmic PAI-1 expression is also absent in mice subjected to SCN lesions (Kudo et al., 2004). In addition, PAI-1 is implicated in the entrainment of the “master clock” in the SCN, since the application of PAI-1 blocks glutamate-induced phase shifts in cultured mouse SCN slices (Mou et al., 2009).
CLOCK:BMAL1/2 heterodimers drive PAI-1 transcription by binding to E-box sequences in the PAI-1 promoter, and this process is synergistically enhanced by the binding of peroxisome proliferator–activated receptor-α (PPARα) to the PAI-1 promoter (Chong et al., 2006; Oishi et al., 2007; Singletary et al., 2008; Oishi et al., 2009a; Hayashida et al., 2010). CLOCK/BMAL2 heterodimers are more potent activators of PAI-1 transcription than CLOCK/BMAL1 heterodimers (Schoenhard et al., 2003; Chong et al., 2006), while the basic helix-loop-helix (bHLH) protein cycle-like factor (CLIF) can also heterodimerize with CLOCK to drive PAI-1 transcription (Maemura et al., 2000; Maemura et al., 2001). CLOCK/BMAL and CLOCK/CLIF-mediated PAI-1 transcription are both inhibited by Period and Cryptochrome proteins (Maemura et al., 2000; Maemura et al., 2001; Chong et al., 2006; Oishi et al., 2007; Oishi et al., 2009a; Hayashida et al., 2010). In addition, the clock accessory protein REV-ERBα represses PAI-1 transcription by binding to retinoic acid receptor–related orphan receptor response elements (ROREs) in the PAI-1 promoter (Wang et al., 2006) (see Fig. 2C). Mutations of these binding sites increase PAI-1 expression, as does the RNAi knockdown of REV-ERBα (Wang et al., 2006). Under physiological conditions, daily fluctuations in plasma PAI-1 levels may also be regulated by glucocorticoids (Singletary et al., 2008) and the peptide hormone angiotensin (Tsujino et al., 2005).
Clock Genes, Cancer, and Hematologic Rhythms
Clock Gene Expression in Lymphocytic Cancer Cell Lines
Clock genes are also relevant to B-cell cancers, such as lymphocytic leukemia, Burkitt lymphoma, and diffuse large B-cell lymphoma (DLBCL) (Taniguchi et al., 2009). BMAL1 is transcriptionally silenced by promoter CpG island hypermethylation in lymphocytic leukemia and Burkitt lymphoma cell lines (Taniguchi et al., 2009). Consistent with this, the RNAi knockdown of BMAL1 enhances tumor growth in cells unmethylated at the BMAL1 CpG island, while the reintroduction of BMAL1 slows tumor growth in methylated lymphoma and leukemia cells (Taniguchi et al., 2009). BMAL1 is not the only clock gene implicated in aberrant B-cell proliferation; CRY1 is also transcriptionally silenced by promoter CpG island hypermethylation in lymphocytic leukemia cells (Hanoun et al., 2012), while PER2 mRNA is significantly reduced in DLBCL cells (Thoennissen et al., 2012). In addition, polymorphisms in the clock genes CRY2 and NPAS2 modify the risk of B-cell cancers (Zhu et al., 2007; Zhu and Zheng, 2008; Hoffman et al., 2009).
Perturbed Hematologic Rhythms in Cancer Patients
Rhythmic clock gene expression is significantly perturbed in the leukocytes of myeloid leukemia patients (Yang et al., 2011). Altered clock gene expression has also been reported in peripheral blood samples from lymphocytic leukemia patients and in the leukocytes of patients with head and neck squamous cell carcinoma (HNSCC), although both studies included just a single time point (Hsu et al., 2014; Rana et al., 2014). In another study, 24-h rhythms in blood lymphocyte counts were lacking in 13 of 18 colorectal cancer patients (Mormont et al., 2002). Last, daily oscillations in the activity of antioxidant enzymes are significantly perturbed in the erythrocytes of oral cancer patients (Baskar et al., 2004; Manoharan et al., 2005) and Wistar rats with mammary tumors (Kolanjiappan and Manoharan, 2005). Hence, there is plentiful evidence that the rhythmic properties of blood cells are disrupted in organisms with cancer. However, based on these results alone, it is impossible to determine whether these abnormalities are a cause or a consequence of the disease process.
Hematologic Rhythms and Cancer Treatment
Hematologic rhythms are also relevant to the treatment of cancer, as the immunosuppressive effects of anticancer drugs vary markedly depending on the timing of drug administration (Peleg et al., 1989; Kobayashi et al., 2002; Gorbacheva et al., 2005; Levi et al., 2007; Van Dycke et al., 2015). The chemotherapeutic agent cyclophosphamide (CY), for example, inhibits the production of lymphocytes, and has a greater impact on lymphocyte (and in particular B-cell) numbers when administered at ZT 2 than at ZT 14 (Gorbacheva et al., 2005). The impact of interferon (IFN)–α treatment on leukocyte numbers also depends on when it is administered (Shinohara et al., 2008). Although the mechanistic basis of these effects is unknown, they have clear therapeutic implications. By administering these compounds at their least toxic time points, larger doses could be safely employed, thereby improving the efficacy of cancer treatment (Levi et al., 1991; Antoch et al., 2005; Levi et al., 2007). These optimal time points may vary between individuals, depending on their circadian clock genotype. Indeed, BMAL1–/– and CLOCK mutant mice lack the rhythmic sensitivity to CY seen in wild-type mice (Gorbacheva et al., 2005).
Environmental Factors, Hematologic Rhythms, and Disease
Poor diet, shift work, and alcohol abuse confer increased susceptibility to a wide range of diseases. As explained in the paragraphs that follow, these effects may be at least partially mediated by the disruption of hematologic rhythms.
Poor Diet
The rhythmic expression of PAI-1 is significantly altered in obese leptin-deficient mice (Oishi et al., 2006b), and in rodents fed high-fat, high-cholesterol, or high-salt diets (Mohri et al., 2003; Kudo et al., 2004; Oishi et al., 2009b; Hayashida et al., 2010; Oishi et al., 2010). Similar changes could therefore contribute to the increased incidence of cardiovascular events in individuals with poor dietary habits. The rhythmic expression of PAI-1 is also perturbed in hypertensive (Naito et al., 2003) and diabetic rodent models (Oishi et al., 2004; Oishi et al., 2005; Oishi et al., 2006b), while 24-h oscillations in the lymph node T-cell population are perturbed in calorie-restricted rats (Esquifino et al., 2004). In humans, there is preliminary evidence that rhythmic clock gene expression is altered in the peripheral blood cells of obese individuals (Tahira et al., 2011), while the amplitude of rhythmic clock gene expression is reduced in the leukocytes of diabetes patients (Ando et al., 2009; Pappa et al., 2013).
Shift Work
Shift work causes significant circadian disruption, and is associated with an increased incidence of stroke, heart disease, and cancer (see Castanon-Cervantes et al., 2010; Logan et al., 2012). This may in part be explained by the negative impact of circadian dysregulation on the hematologic system. For example, chronic exposure to 6-h phase shifts (one a week for 4 weeks) leads to an increase in plasma PAI-1 levels in mice (Oishi and Ohkura, 2013). If a similar process occurs in humans, this could contribute to the increased risk of cardiovascular disease among shift workers. When rodents are subjected to a chronic shift-lag paradigm (ten 6-h phase advances administered at 2-day intervals), clock gene expression in NK cells and macrophages is severely perturbed, as is the rhythmic expression of cytolytic factors and cytokines in NK cells (Castanon-Cervantes et al., 2010; Logan et al., 2012). In the same animals, NK cell cytolytic activity is suppressed, lung tumor growth is enhanced, and the mortality rate is higher after lipopolysaccharide exposure (Castanon-Cervantes et al., 2010; Logan et al., 2012). In hamsters, exposure to an abnormal light/dark schedule (a nocturnal light pulse followed by a phase delay) damps rhythmic variation in the number of circulating leukocytes (Prendergast et al., 2013). Similarly, in zebrafish, exposure to alternately shortened and lengthened days increases the number of aged erythrocytes in the blood, via a reduction in the rate of erythrocyte apoptosis (Egg et al., 2014). The oxygen transport capacity of aged erythrocytes is reduced, and their tendency to aggregate is increased. Again, this may contribute to the increased incidence of cardiovascular events in shift workers.
Alcohol Abuse
Chronic alcohol abuse impairs the immune system and confers increased susceptibility to infections and cancer (Garro et al., 1992; Watson et al., 1994). In rats, chronic ethanol administration suppresses the cytolytic activity of splenic NK cells by perturbing the rhythmic expression of the cytolytic factors Granzyme B and Perforin, and that of the cytokine IFN-γ (Arjona et al., 2004). It also disrupts daily rhythms in the total number and mitogenic responses of B cells and T cells in the thymus and spleen (Jimenez et al., 2005). Last, evidence from the clinic indicates that clock gene expression is reduced in the peripheral blood cells of male alcoholics (Huang et al., 2010).
Conclusions and Future Directions
Over the past decade, evidence has emerged for circadian variation in the function of erythrocytes, leukocytes, and thrombocytes. The discovery of circadian rhythms in erythrocytes was particularly striking, since it challenged the prevailing view that all circadian rhythms originate from TTFLs. There is also increasing evidence of 24-h rhythmicity in the function of leukocytes and thrombocytes. Not all of these rhythms have been demonstrated under constant conditions, so it is unclear whether they are all genuinely circadian. It is also unclear how clock gene rhythms in individual blood cells are synchronized with each other and with the SCN under physiological conditions, although there is some evidence that glucocorticoids mediate this process (Oishi et al., 1998; Burioka et al., 2005; Nebzydoski et al., 2010; Ohmori et al., 2013; Cuesta et al., 2015), while endogenous temperature cycles could also play a significant role (Buhr et al., 2010; O’Neill and Reddy, 2011). Whatever the mechanism(s) responsible, it is highly likely that the rhythmic function of leukocytes and thrombocytes underlies rhythmicity in the symptoms of various diseases, including asthma, chronic obstructive pulmonary disease, rheumatoid arthritis, allergic rhinitis, myocardial infarction, and ischemic stroke. Consequently, the efficacy of pharmacological treatments for these disorders is likely to vary according to the timing of drug administration. Moreover, the optimal time for drug delivery may vary between individuals with different circadian clock genotypes. Although these ideas are far from novel (see Levi et al., 1991), a concerted research effort is required before they are translated into clinical practice. Finally, the deleterious effects of poor diet, alcohol abuse, and shift work on human health may be at least partially mediated by the disruption of hematologic rhythms.
Footnotes
Acknowledgements
ABR is a Wellcome Trust Senior Clinical Fellow. He receives funding from the Wellcome Trust (Grant No. 100333/Z/12/Z), the European Research Council (ERC Starting Grant No. 281348, MetaCLOCK), the European Molecular Biology Organization (EMBO) Young Investigators Programme, and the Lister Institute of Preventative Medicine. DP is supported by the Wellcome Trust.
Conflicts of Interest Statement
The author(s) declare no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
