Abstract
Recent studies show that the timeless (tim) gene is not an essential component of the circadian clock in some insects. In the present study, we have investigated whether the tim gene was originally involved in the insect clock or acquired as a clock component later during the course of evolution using an apterygote insect, Thermobia domestica. A cDNA of the clock gene tim (Td’tim) was cloned, and its structural analysis showed that Td’TIM includes 4 defined functional domains, that is, 2 regions for dimerization with PERIOD (PER-1, PER-2), nuclear localization signal (NLS), and cytoplasmic localization domain (CLD), like Drosophila TIM. Td’tim exhibited rhythmic expression in its mRNA levels with a peak during late day to early night in LD, and the rhythm persisted in DD. A single injection of double-stranded RNA (dsRNA) of Td’tim (dstim) into the abdomen of adult firebrats effectively knocked down mRNA levels of Td’tim and abolished its rhythmic expression. Most dsRNA-injected firebrats lost their circadian locomotor rhythm in DD up to 30 days after injection. DsRNA of cycle (cyc) and Clock genes also abolished the rhythmic expression of Td’tim mRNA by knocking down Td’tim mRNA to its basal level of intact firebrats, suggesting that the underlying molecular clock of firebrats resembles that of Drosophila. Interestingly, however, dstim also reduced cyc mRNA to its basal level of intact animals and eliminated its rhythmic expression, suggesting the involvement of Td’tim in the regulation of cyc expression. These results suggest that tim is an essential component of the circadian clock of the primitive insect T. domestica; thus, it might have been involved in the clock machinery from a very early stage of insect evolution, but its role might be different from that in Drosophila.
Circadian rhythms are about 24-hour oscillations observed in various physiological functions of a wide variety of organisms and are driven by an endogenous mechanism called the circadian clock. The molecular oscillatory mechanism of the circadian clock has been extensively studied in the fruit fly Drosophila melanogaster and is currently believed to consist of 3 transcriptional/translational loops (Hardin, 2005; Tomioka and Matsumoto, 2010). In the first loop, a heterodimer of CLOCK (CLK) and CYCLE (CYC) activates the transcription of negative elements, period (per) and timeless (tim) (Allada et al., 1998). The product proteins, PER and TIM, increase during the night, peaking at late night, and enter the nucleus by forming a heterodimer to repress their own transcription through inhibitory action on CLK-CYC. The repression of per and tim transcription results in a reduced level of PER and TIM, which eventually releases CLK-CYC from the inhibition to reactivate the transcription of per and tim. The CLK-CYC dimer also activates transcription of vrille (vri) and PAR domain protein 1ε (Pdp1ε) in the second loop (Cyran et al., 2003; Glossop et al., 2003), and VRI and PDP1ε in turn regulate the transcription of Clock (Clk) (Zheng and Sehgal, 2008). The third loop consists of cyclic expression of clockwork orange (cwo), which regulates the amplitude of the per-tim oscillation (Kadener et al., 2007; Lim et al., 2007; Matsumoto et al., 2007; Richier et al., 2008).
However, the circadian oscillatory mechanism seems to differ considerably among insect species. It has been shown that tim is not essential for the circadian clock mechanism in the cricket Gryllus bimaculatus (Danbara et al., 2010) and even does not exist in the genome of the honeybee, where mammalian-type CRYPTOCHROME (CRY2) is thought to form the first loop together with PER instead of TIM (Rubin et al., 2006). In the monarch butterfly Danaus plexippus, it has been suggested that the negative loop consists of PER, TIM, and CRY2, where CRY2 is shuttled into the nucleus, repressing CLK/CYC-mediated transcription, while TIM mostly stays in the cytoplasm and is mainly involved in light-dependent resetting of the clock (Zhu et al., 2008). These facts raise a question whether the tim gene was originally involved as an essential component in the insect clock or acquired later during the course of evolution.
As one of the ancestral insects, the firebrat appears to be suitable for searching the answer to this question. Our previous study showed that the firebrat Thermobia domestica has a circadian clock and possesses Clock (Td’Clk) and cycle (Td’cyc) genes (Kamae et al., 2010). Interestingly, the sequence and expression profile of these clock genes showed similarity to their mammalian orthologs (Kamae et al., 2010). Thus, we infer that primitive insects retain the common traits of Clk and cyc with vertebrates, but higher insects such as Drosophila later obtained new traits. In this study, we have cloned cDNA of the clock gene timeless (Td’tim) from the firebrat, examined its daily expression patterns using quantitative real-time RT-PCR (qPCR), and investigated the role of Td’tim in the rhythm generation with RNA interference (RNAi) technology. The results showed that Td’tim was structurally similar to its Drosophila homolog. Knockdown of Td’tim by RNAi disrupted circadian rhythms, suggesting that Td’tim plays an essential role in the circadian system in the firebrat like in Drosophila and that tim might have been involved in the insect circadian clock from an early stage of the insect’s evolution.
Materials and Methods
Animals
Adult firebrats, T. domestica, raised in the laboratory under a light cycle of 12-hour light and 12-hour darkness (LD12:12) at a constant temperature of 30 °C, were used for all experiments. They were fed laboratory chow (CA-1, Clea Japan, Tokyo, Japan).
Quantitative Real-Time RT-PCR
Quantitative real-time RT-PCR (qPCR) was used to measure mRNA levels. Both males and females were used for measurement because preliminary experiments revealed no significant gender differences in tim mRNA levels. Total RNA extraction from the entire bodies of adult firebrats was performed with TRIzol Reagent (Invitrogen, Carlsbad, CA), and the obtained RNA was treated with DNase I to remove contaminated DNA. About 500 ng of total RNA of each sample was reversely transcribed with random 6mers using PrimeScript RT reagent kit (Takara, Otsu, Japan). Real-time PCR was performed with Mx3000P Real-Time PCR System (Stratagene, La Jolla, CA) using Universal SYBR Green Master (Roche, Tokyo, Japan) including SYBER Green with primers 5′-TACAAGCCAGGTCCATCACA-3′ and 5′-TCAAGCGTCAATTCAGCATC-3′ for tim, 5′-ATCGCAAGGGTCTGGAAGTG-3′ and 5′-GGAAAACTCGCCAAGACAGG-3′ for Clk (GenBank/EMBL/DDBJ accession no. AB550828), 5′-CGTGTAATCTGTCGTGTTTGGTG-3′ and 5′-GAATCGTCCGCCTTTCCTC-3′ for cyc (GenBank/EMBL/DDBJ accession no. AB550829), and 5′-AGTCCGAAGGCGGTTTAAGG-3′ and 5′-TACAGCGTGTGCGATCTCTG-3′ for rp49 (GenBank/EMBL/DDBJ accession no. AB550830). The results were analyzed using the software associated with the instrument. The values were normalized with the values for rp49, a housekeeping gene, at each time point. Results of 3 or 4 independent experiments were pooled to calculate the mean ± SEM. Data were analyzed by a t test or ANOVA followed by the Tukey test.
RNA Interference
Double-stranded RNA (dsRNA) for Td’tim and DsRed2, derived from a coral species (Discosoma sp.), were synthesized using MEGAscript High Yield Transcription Kit (Ambion, Austin, TX). Primers fused with T7 promoter sequence were designed for the synthesis of dsRNA of Td’tim as follows: 5′-TAATACGACTCA-CTATAGGGTGCATTTGGTTGTGACTGCT-3′ and 5′-TAATACGACTCACTATAGGGAGAGGCGTGTGCCTTGTACT-3′. Standard PCR was performed using Td’s cDNA as a template. Resulting amplicons excluding the fused T7 promoters were 600 bp for Td’tim. DsRed2 was linearized from pDsRed2-N1 (Clontech, Mountain View, CA) and amplified with the forward and reverse primers. Then, RNAs were synthesized with T7 and SP6 RNA polymerases. The synthesized RNAs were extracted with phenol/chloroform and suspended in 50 µL of TE solution after isopropanol precipitation. The yield and quality of RNA were assessed by absorbance with a spectrophotometer (Genequant Pro, Amersham Biosciences, Piscataway, NJ), and the same amounts of sense and antisense RNA were mixed. The RNA was denatured for 5 minutes at 100 °C and annealed by a gradual cool down to room temperature. After ethanol precipitation, the obtained dsRNA was suspended in Ultra Pure Water (Invitrogen) and adjusted to a final concentration of 10 µM. The dsRNA solution was stored at −80 °C until use. A total of 70 nL (10 µM) dsRNA solution was injected with a nanoliter injector (WPI, Sarasota, FL) into the abdomen of adult firebrats anesthetized with CO2.
Recording of Locomotor Activity
To monitor locomotor activity, adult firebrats were individually housed in transparent acrylic rectangular tubes (6 × 6 × 70 mm). The tubes were plugged at one end with a piece of animal food and were sealed with plastic tape; at the other end, they were sealed with wet cotton connected to a water bottle. Movement of the firebrat was sensed by a photoelectrical detection system: A moving firebrat interrupted an infrared beam, and the number of interruptions during each 6-minute interval was recorded using a computerized system. The activity sensing system was placed in an incubator (MIR-153, Sanyo Biomedica, Osaka, Japan), in which the temperature was kept at 30 °C and lighting conditions were given by a cool white fluorescent lamp connected to an electric timer. The light intensity was 600 to 1000 lux at the animal’s level, varying with the proximity to the lamp. The raw data were displayed as conventional double-plotted actograms to judge activity patterns, and existence of the rhythmicity and free-running periods were analyzed by the χ2 periodogram written by Schmid et al. (2011). If peaks of the periodogram appeared above the 0.05 confidence level, the periods of the peaks were designated as statistically significant.
Results
Cloning and Sequencing of Clock Gene timeless
We cloned a tim homolog of T. domestica with primers constructed based on the conserved amino acid sequences of known insect tim genes and 5′ and 3′ RACE strategy (see supplementary material for method). We obtained a full length of 5490-bp cDNA of T. domestica timeless (Td’tim; GenBank/EMBL/DDBJ accession no. AB644410). Td’tim has 5′- and 3′-untranslated regions (UTR) of 128 bp and 1735 bp, respectively. Sequences were analyzed by Genetyx version 6 (Genetic Information Processing Software, Tokyo, Japan) and BioEdit version 7.0.9.0 (Biological Sequence Alignment Editor, Ibis Therapeutic, Carlsbad, CA). Amino acid sequence of TIMELESS (TIM) was analyzed with ClustalW (http://clustalw.ddbj.nig.ac.jp/top.html). The putative product protein consisted of 1209 amino acid residues, which is shown in Supplementary Figure S1. It contains PER-1 and PER-2 dimerization regions that are involved in protein-protein interaction and mediate TIM’s binding to its heterodimeric partner PER (Gekakis et al., 1995; Myers et al., 1995; Saez and Young, 1996), a cytoplasmic localization domain (CLD) (Saez and Young, 1996), and a nuclear localization signal (NLS) (Baylies et al., 1993; Vosshall et al., 1994). A BLAST database search indicated that the amino acid sequence has 43% to 57% identities along the entire length of the protein to those of known insect TIMs (Table 1). Among the putative functional motifs, NLS had the highest identity of 50% to 83% with other insect species, PER-1 and PER-2 had rather high identities of 34% to 74%, suggesting their conserved functional role, and CLD had a rather low identity of only 9% to 22% (Table 1). A phylogenetic tree based on the amino acid sequences of TIMs from known insects, sea urchin, and mice reveals that there are 3 clusters of tim and that Td’tim forms a cluster with the cricket (G. bimaculatus) and the louse (Pediculus humanus corporis) tim (Fig. 1).
Overall amino acid identity and similarity (%) of whole sequence and functional domains among insect TIM homologs.
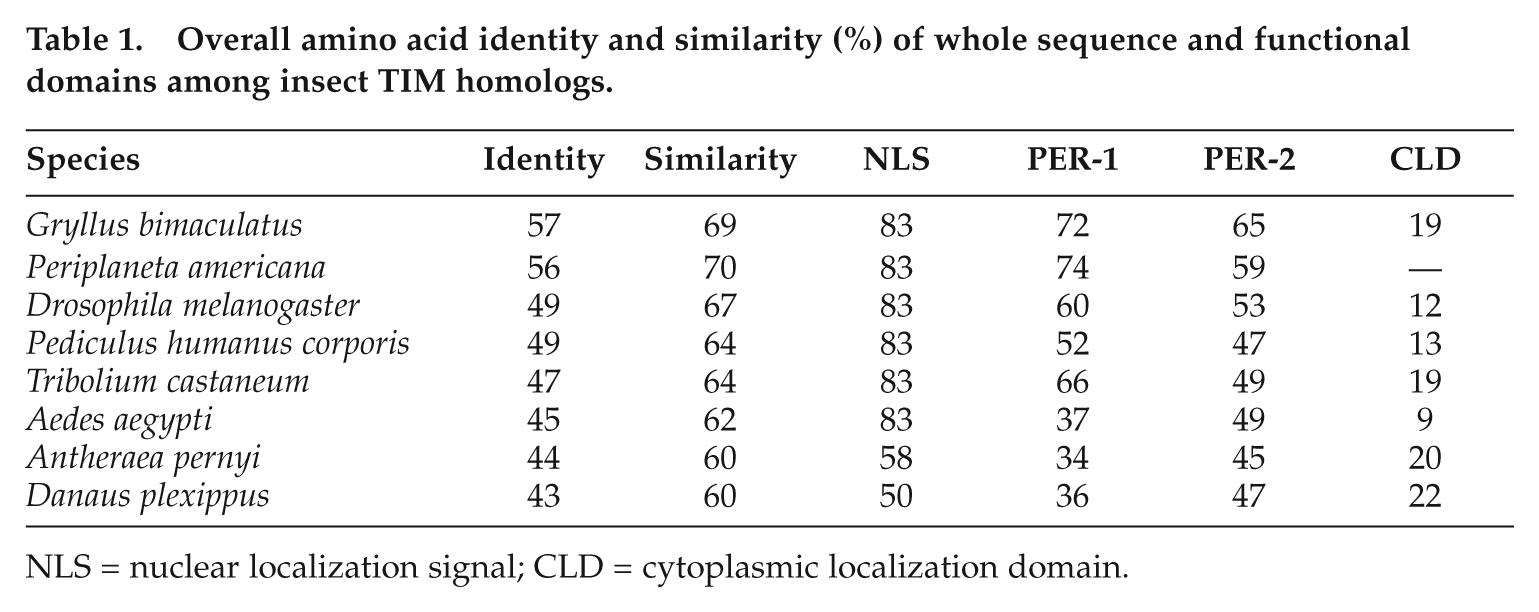
NLS = nuclear localization signal; CLD = cytoplasmic localization domain.
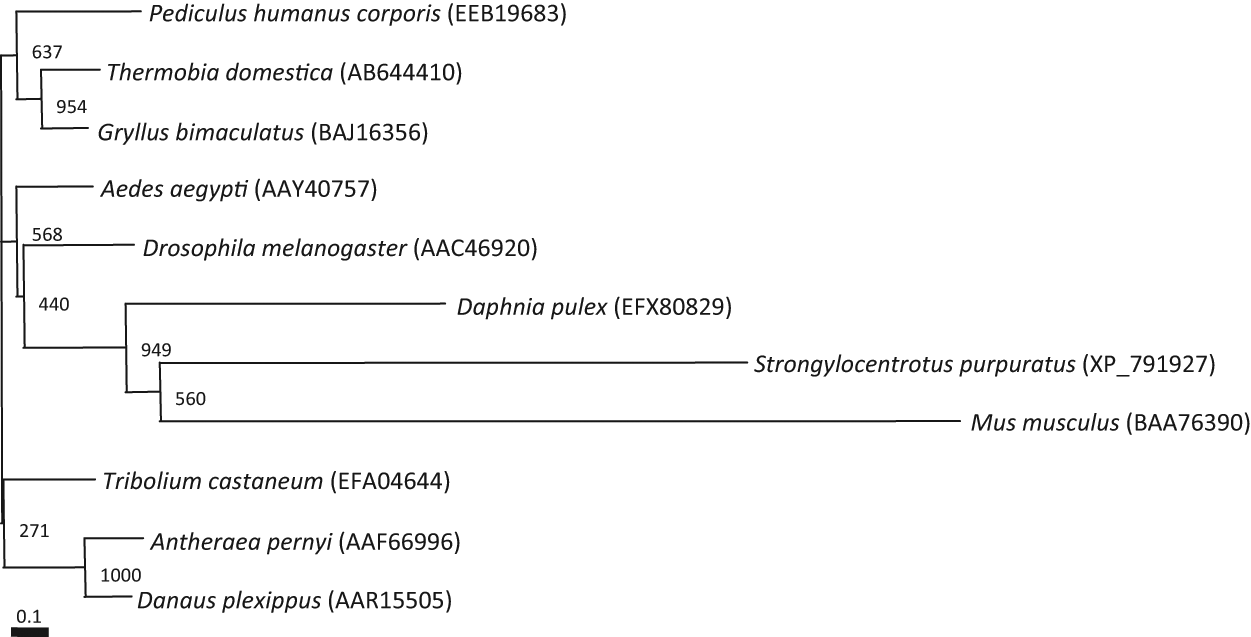
Phylogenetic neighbor-joining tree of known insect, sea urchin, and mouse TIM proteins. Neighbor-joining trees were inferred with ClustalW. Sequences of known insect, sea urchin, and mouse TIM were obtained from GenBank. A reference bar indicates distance as the number of amino acid substitutions per site, and the numbers indicate bootstrap values. Numbers in parentheses indicate GenBank or RefSeq accession number.
Temporal Expression Pattern of Td’tim mRNA
To determine whether transcripts of Td’tim oscillated in a circadian manner, we measured levels of Td’tim mRNA in T. domestica under LD12:12 and on the second day of DD by qPCR (Fig. 2). Sample collections were performed every 4 hours starting at zeitgeber time (ZT) 2 (ZT0 corresponds to lights-on and ZT12 to lights-off) or circadian time (CT) 2 (CT0 corresponds to projected lights-on and CT12 to projected lights-off in DD). In LD, Td’tim mRNA showed a rhythmic expression with a peak at late day to early night (ANOVA, p < 0.01); the peak value at ZT10 (2 hours before lights-off) was about 2 times higher than the trough level, and the difference was statistically significant (Tukey test, p < 0.05). The rhythmic Td’tim mRNA expression persisted in DD with a peak at late subjective day (CT10; ANOVA, p < 0.01), and the peak was about 3-fold of the trough level.
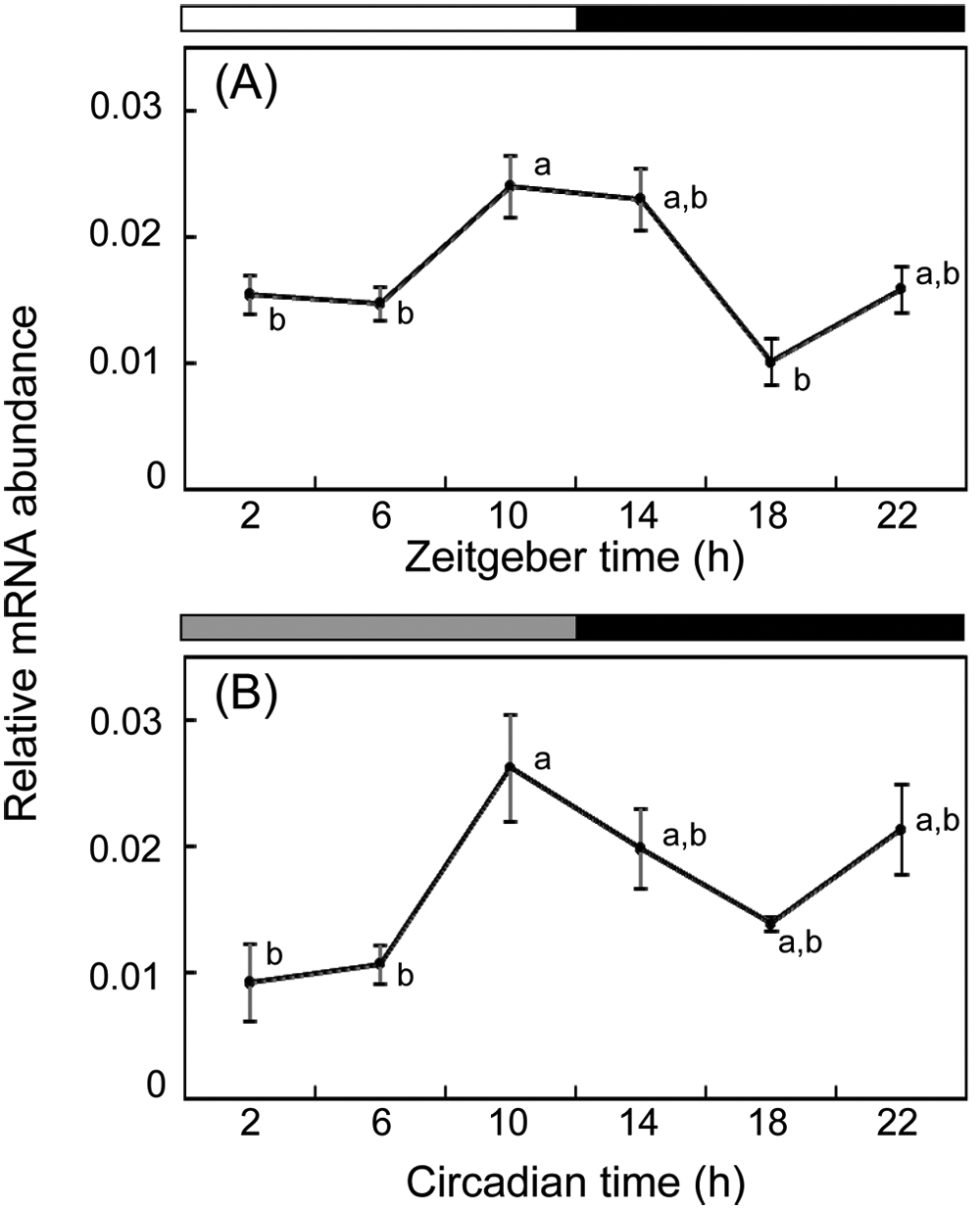
Daily and circadian expression patterns of Td’tim mRNA in LD12:12 (A) and DD (B). The abundance of Td’tim mRNA was measured by quantitative real-time RT-PCR. Total RNA was extracted from firebrats collected at 4-hour intervals starting at 2 hours after lights-on (ZT2) or projected lights-on (CT2). White, black, and gray bars indicate light phase, night/subjective night, and subjective day, respectively. The data collected from 4 independent experiments were averaged and plotted as mean ± SEM values relative to the value of rp49 mRNA used as reference. Values with different letters significantly differ from each other (p < 0.05, ANOVA with Tukey test).
Td’tim dsRNA Suppressed Td’tim mRNA Levels
To examine whether RNAi of the tim gene worked effectively in T. domestica, we measured Td’tim mRNA levels by qPCR in adult firebrats injected with Td’tim dsRNA (dstim) into the abdomen. The firebrats were sampled 7 days after the injection. The Td’tim mRNA expression was found to be arrhythmic with significantly lower levels than those of intact firebrats (ANOVA, p > 0.9) (Fig. 3A), suggesting that dstim suppressed Td’tim mRNA levels and abolished its rhythmic expression through RNAi.
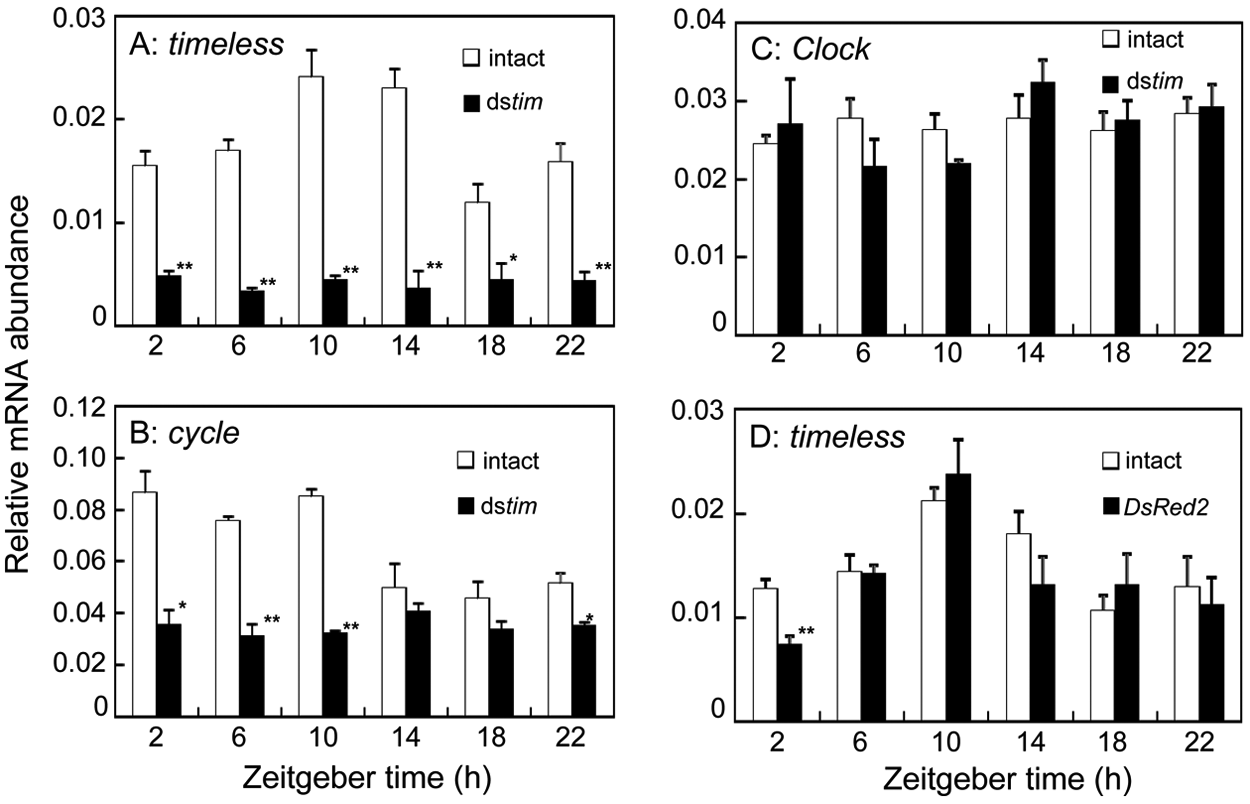
Effects of tim dsRNA (dstim) on mRNA levels of tim (A), cyc (B), and Clk (C) and effects of dsDsRed2 on tim mRNA levels (D) in Thermobia domestica. (A) dstim effectively knocked down tim mRNA levels and prevented its rhythmic expression. It also knocked down cyc mRNA to abolish its daily rhythm (B) but had no significant effect on Clk mRNA (C). dsDsRed2 had no significant effect on the tim mRNA levels except at ZT2, where a significant reduction was observed (D). The abundance of mRNAs was measured by quantitative real-time RT-PCR. Total RNA was extracted from firebrats collected 7 days after the dsRNA injection at 4-hour intervals starting at 2 hours after lights-on (ZT2). The data collected from 3 independent experiments were averaged and plotted as mean ± SEM values relative to those of rp49 mRNA used as reference. *p < 0.05, t test. **p < 0.01.
To examine the effects of tim RNAi on the expression of other clock genes, cycle (Td’cyc) and Clock (Td’Clk), we measured levels of those mRNAs in firebrats treated with dstim. Td’cyc mRNA levels were significantly reduced at most of the examined time points (ZT2-ZT10, ZT22; t test, p < 0.05) (Fig. 3B), with the greatest suppression of 41% at ZT2. The rhythm that was evident in intact firebrats was abolished (ANOVA, p > 0.5) (Fig. 3B). In contrast, Td’Clk mRNA showed both expression levels and patterns similar to those of intact firebrats, and no significant difference was observed at any time point (t test, p > 0.05) (Fig. 3C).
As a control, we measured mRNA levels in dsDsRed2-treated firebrats (Fig. 3D). Although a significant reduction was observed at ZT2, Td’tim mRNA levels showed a clear rhythm similar to that of intact firebrats with a peak at ZT10 (ANOVA, p < 0.05). The peak to trough ratio was about 3.
Effects of dsClk and dscyc on Td’tim
In Drosophila, transcription of tim is hypothesized to be activated by transcription factors CLK and CYC. To test this possibility in the firebrat, we also measured levels of Td’tim mRNA in firebrats treated with dscyc and dsClk. The knockdown of Td’cyc and Td’Clk by dscyc and dsClk, respectively, has been confirmed previously (Kamae et al., 2010). In both treatments, Td’tim mRNA levels stayed at near basal levels of intact animals throughout the day, and no significant rhythm was observed (ANOVA, p > 0.7) (Fig. 4): reductions at ZT6 to ZT14 and at ZT10 and ZT14 were statistically significant for dsClk and dscyc, respectively (t test, p < 0.01 for dscyc at ZT10; p < 0.05 for the remaining).
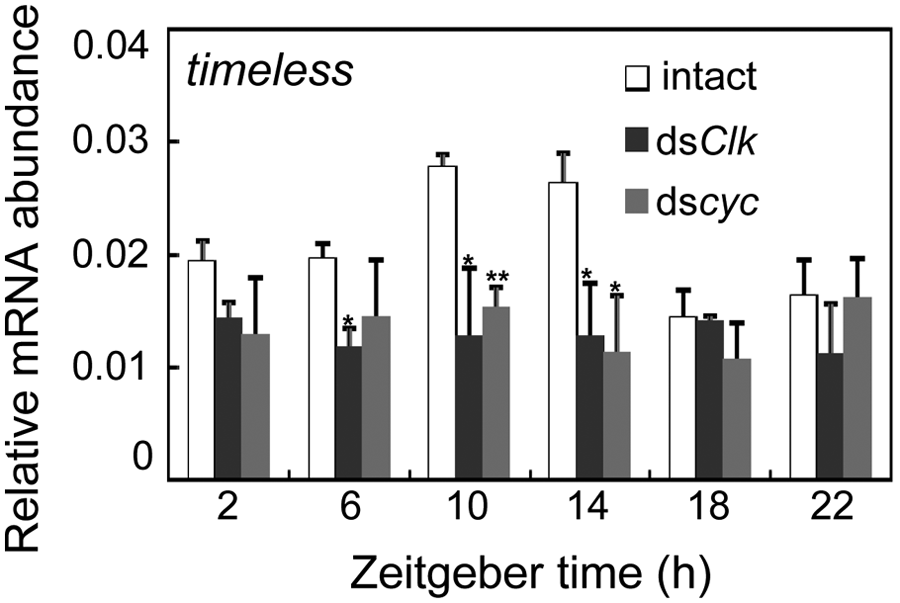
dscyc and dsClk knocked down tim mRNA levels to its basal level of intact animals in Thermobia domestica. Total RNA was extracted from firebrats collected 7 days after dsRNA injection at 4-hour intervals starting at 2 hours after lights-on (ZT2). The data collected from 3 independent experiments were averaged and plotted as mean ± SEM values relative to those of rp49 mRNA used as an internal reference. *p < 0.05, t test. **p < 0.01.
Effects of dsRNA on Locomotor Rhythm
We have shown that intact firebrats exhibit a nocturnal rhythm synchronizing to LD and free running in DD with a period slightly longer than 24 hours (Kamae et al., 2010). To examine the effect of dstim on the circadian locomotor rhythm, we recorded locomotor activity of 23 adult firebrats injected with dstim. As a control, we recorded the locomotor activity of adults injected with dsDsRed2. The results are summarized in Table 2. Under LD, the majority (93%) of dsDsRed2-injected controls showed a nocturnal locomotor rhythm, which was interrupted by an inactive phase associated with molting (Fig. 5C and Table 2). The rhythm persisted in the ensuing DD in 84% of them, with a period slightly longer than 24 hours (Table 2). The ratio of rhythmic animals and free-running periods were very close to those of intact animals (Table 2), which are reanalyzed with the χ2 periodogram (Schmid et al., 2011), suggesting that injection of dsDsRed2 has no significant effect on the rhythm.
Effects of dstim and dsDsRed2 on the locomotor rhythm of the firebrat Thermobia domestica.
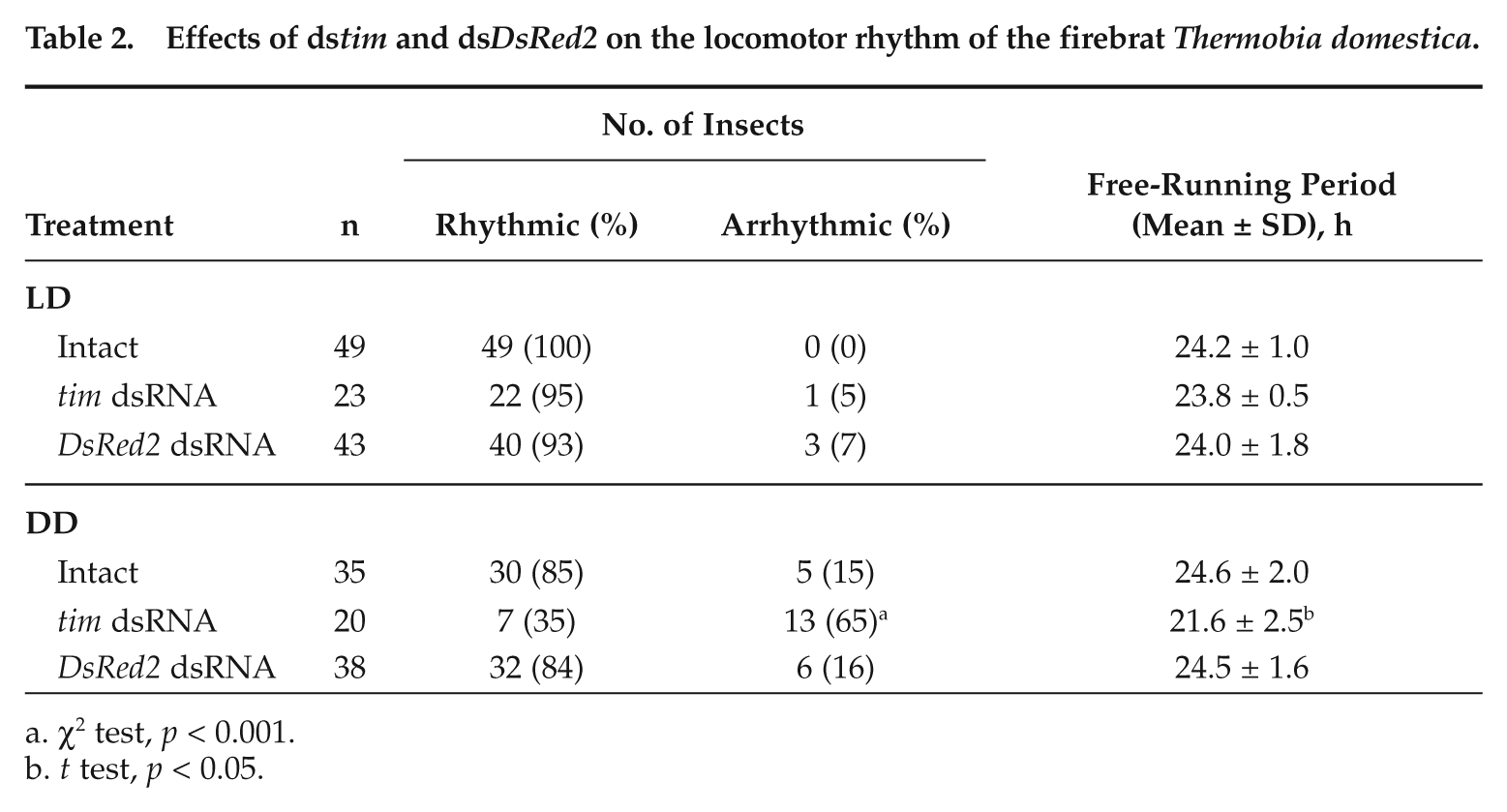
χ2 test, p < 0.001.
t test, p < 0.05.
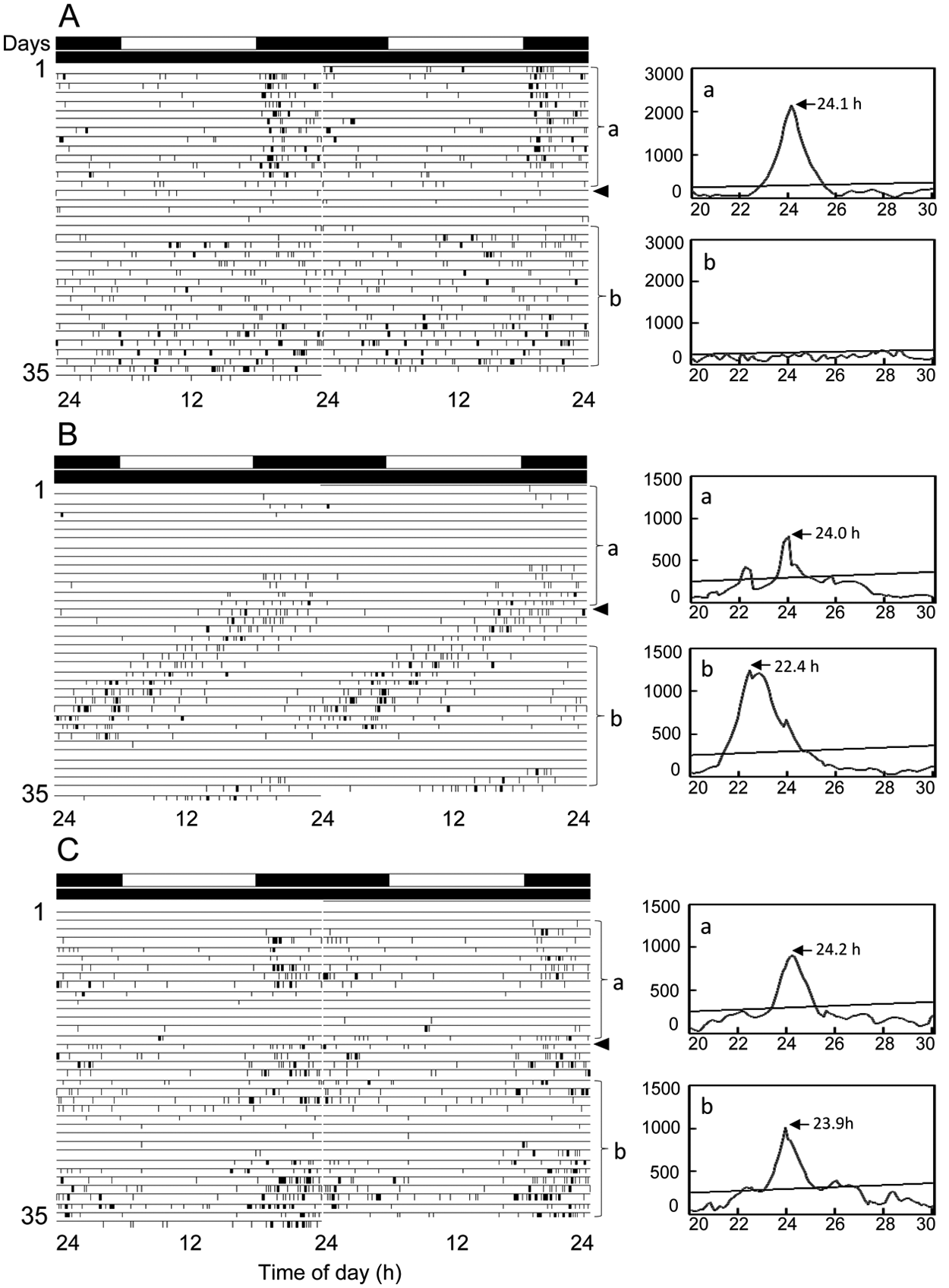
Double-plotted actograms (left) and χ2 periodograms (right) of locomotor rhythms of firebrats Thermobia domestica injected with Td’tim dsRNA (A, B) or DsRed2 dsRNA (C) under LD12:12 and DD at a constant temperature of 30 °C. Arrowheads indicate the day when the firebrats were transferred from LD to DD. a and b indicated in the periodogram correspond to the analyzed time span, a and b, indicated in the actogram, respectively. An oblique line in the periodogram indicates the significance level of p < 0.05; peak value above the line was designated as significant. The firebrat in A showed a clear rhythm in LD, which disappeared on transfer to DD, but the insect in B showed a rhythm both in LD and DD. The control firebrat injected with DsRed2 dsRNA showed a clear rhythm throughout the recording period (C). For further explanations, see text.
In the firebrat injected with dstim, 22 animals (95%) exhibited a nocturnal activity rhythm under LD with a peak in the early night (Fig. 5A and 5B). The remaining animal was arrhythmic. In the ensuing DD, 13 (65%) animals almost immediately lost the nocturnal rhythm and became arrhythmic (Fig. 5A), suggesting that the rhythm observed in LD was a masking effect of light. Only 7 (35%) of the injected firebrats showed the nocturnal peak free running with an average period of 21.6 ± 2.5 h (Fig. 5B). The free-running period was significantly shorter than that of intact and dsDsRed2-injected firebrats (t test, p < 0.05) (Table 2).
Discussion
timeless Gene in the Firebrat
In the present study, we obtained the timeless (Td’tim) gene from the firebrat T. domestica. This is the first tim homolog in Thysanura insects. Phylogenetic analyses with sequence data from other species revealed that the insect’s tim consists of 3 clusters. The firebrat lineage belongs to a rather primitive group, including the cricket and the louse, compared with 2 other clusters including dipteran, coleopteran, and lepidopteran species (Fig. 1). Td’TIM includes 4 defined functional domains, that is, PER-1, PER-2, NLS, and CLD like Drosophila TIM (Dm’TIM) (Suppl. Fig. S1). The CLD has a very low similarity to other insects’ TIM, suggesting its structural and/or functional uniqueness in the firebrat. A comparison between TIMs of the firebrat and fruit fly, D. melanogaster, however, revealed that PER-1, PER-2, and NLS domains have high identity (Table 1), suggesting that Td’TIM retains a functional ability similar to that of Dm’TIM. In the fruit fly, tim is one of the major genes constituting the negative feedback loop together with per to regulate its own rhythmic expression (Tomioka and Matsumoto, 2010). In fact, our qPCR measurement revealed that Td’tim mRNA showed circadian oscillation in both LD and DD conditions with a peak at late day to early night like in Drosophila (Fig. 2). The fact that Td’tim dsRNA abolished the locomotor rhythm in most of the treated firebrats suggests tim’s involvement in the rhythm generation (Figs. 3 and 5). The significant reduction and arrhythmic profile of mRNA expression in the tim RNAi also indicate tim’s importance in the clock machinery of firebrats. Together with the structural similarity to its Drosophila homolog, these results suggest that tim had been involved as an essential component in the ancestral insect clock.
However, some tim RNAi firebrats (35%) maintained a locomotor rhythm in DD, although their free-running periods were significantly shorter than those of intact animals. Persistence of the rhythm after dstim treat-ment was reported for the cricket G. bimaculatus, where tim RNAi resulted in only shortening of the free-running period and never abolished the rhythm (Danbara et al., 2010). The role of the cricket’s tim is thus probably only modi-ficational and not essential in the clock mechanism. In the firebrat, however, the fact that the majority of dstim-treated animals lost the rhythm suggests that the persistence of the rhythm is rather exceptional and probably caused by an insufficient knockdown of tim mRNA. The insufficient knockdown may associate with the systemic RNAi strategy because uptake of injected dsRNA may be in a rather passive manner (Huvenne and Smagghe, 2010). The shortening of the free-running periods in the rhythmic tim RNAi firebrats may be caused by faster degradation of the TIM protein due to a less amount of TIM resulting from reduced tim mRNA levels. This possibility has also been suggested for the cricket G. bimaculatus (Danbara et al., 2010) but needs to be experimentally addressed in future studies.
Possible Role of tim in the Firebrat Circadian Clock
The RNAi experiment revealed that knockdown of Td’tim had no significant effect on Td’Clk mRNA levels but reduced the mRNA levels of a clock gene Td’cyc and eliminated its rhythmic expression (Fig. 3B and 3C). Together with the previous finding that Td’Clk is constitutively expressed and Td’cyc shows rhythmic expression (Kamae et al., 2010), the fact contradicts what is expected from the Drosophila’s clock hypothesis. In Drosophila, the transcription of Clk is known to be rhythmically controlled by VRI and PDP1ε, of which transcriptions are activated by a CLK-CYC heterodimer, which is repressed by a PER-TIM heterodimer (Cyran et al., 2003). According to this Drosophila hypothesis, a knockdown of Td’tim would affect Td’Clk mRNA levels because of a loss of repression by the PER-TIM complex. However, there was no apparent effect of Td’tim RNAi on Td’Clk mRNA levels. Regarding the Td’cyc gene, there seems to be some regulatory mechanism includ-ing Td’tim for its rhythmic expression unlike in Drosophila, where cyc is thought to be expressed constitutively. In vertebrates, a clock gene Bmal1, a structural counterpart of cyc, is expressed rhythmically by a loop including transcription factors ROR and REV-ERB, which are under the regulation of CLK-BMAL1 (Ko and Takahashi, 2006). In the firebrat, the knockdown of Td’tim might affect Td’cyc mRNA levels through the yet unknown regulatory pathway for Td’cyc by a reduction of inhibitory effects on Td’CLK and Td’CYC.
Regulation of tim Expression
The present study revealed that Td’tim mRNA is rhythmically expressed in both LD and DD to peak late day to early night (Fig. 2) like Drosophila tim (So and Rosbash, 1997). In Drosophila, the transcription of tim is activated by a CLK and CYC heterodimer, which binds to a promoter region of tim, called E-box (Hardin, 2006). This Drosophila’s hypothesis may account for the present results of Td’Clk and Td’cyc RNAi experiments that a knockdown of these transcription factors resulted in a reduced Td’tim mRNA level and eliminated its cyclic expression. We have previously shown that Td’cyc has a BMAL C-terminal region (BCTR) domain with a highly conserved sequence to that of mouse BMAL1 with a transactivational function (Kamae et al., 2010). It is thus likely that in the firebrat, Td’CLK and Td’CYC form a heterodimer and activate the transcription of Td’tim like in Drosophila and that the rhythmic Td’tim expression may be due to the rhythmic expression of Td’cyc (Kamae et al., 2010).
The Clock Machinery in the Firebrat
Most interesting is that the firebrat circadian clock possesses both Drosophila-type and mammalian-type operation parts. The molecular structure and functional role of Td’tim in the clock mechanism appear to be similar to those in the Drosophila clock (Table 1 and Fig. 3). The rhythmic expression of Td’tim is most likely regulated by a negative feedback loop of Td’tim, of which transcription is activated by Td’CLK/Td’CYC. period gene probably participates in the loop because we have obtained its cDNA fragment (Kamae and Tomioka, unpublished results) and because a Western blot analysis has revealed a cycling of PER-like antigen (Závodská et al., 2003, 2005). However, the expression profiles of Td’cyc and Td’Clk seem to resemble those in vertebrate clocks rather than the Drosophila clock (Kamae et al., 2010). The mechanism of rhythmic expression of Td’cyc is currently unknown. Investigation of the mechanism will reveal whether vrille and Pdp1 participate in it like in Drosophila or the orthologs of ror and rev-erb are involved like in vertebrates. Investigation of the role of per in the firebrat clock mechanism also needs to be carried out. Elucidation of the firebrat clock machinery will contribute to the understanding of the origin of the insect circadian clocks and how it has been diversified among insects because the firebrat is, to our knowledge, the most primitive insect species, of which the circadian clock has been investigated at the molecular level.
In conclusion, it is likely that tim is a gene that characterizes the insect’s clock machinery because a member of the most primitive insect group incorporates it as a major component of the clock. Those possessing a clock functioning without tim, such as honeybees and crickets (Rubin et al., 2006; Danbara et al., 2010), may have lost tim or its function during the evolutionary process.
Footnotes
Acknowledgements
The authors thank Drs. Akira Matsumoto and Taishi Yoshii for critically reading the article. This study was supported in part by Grants-in-Aid for Scientific Research from the Japan Society for the Promotion of Science to K.T. Y.K. is a JSPS research fellow.
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
