Abstract
There is little evidence for the involvement of microRNAs (miRs) in the regulation of circadian rhythms, despite the potential relevance of these elements in the posttranscriptional regulation of the clock machinery. The present work aimed to identify miRs targeting circadian genes through a predictive analysis of conserved miRs in mammals. Besides 23 miRs previously associated with circadian rhythms, we found a number of interesting candidate genes, equally predicted by the 3 software programs used, including miR-9, miR-24, miR25, miR-26, miR-27, miR-29, miR-93, miR-211, miR-302, and miR-346. Moreover, several miRs are predicted to be regulated by circadian transcription factors, such as CLOCK/BMAL, DEC2, and REV-ERBalpha. Using real-time PCR we demonstrated that the selected candidate miR-27b showed a daily variation in human leukocytes. This study presents predicted feedback loops for mammalian molecular clock and the first description of an miR with in vivo daily variation in humans.
Approximately 5% to 10% of transcripts present a circadian oscillation (Pegoraro and Tauber, 2008). However, proteome analysis in mammals reveals that around 20% of proteins present cyclic expression, some of them with the corresponding mRNAs being constitutively expressed (Reddy et al., 2006). Moreover, cycling mature mRNAs are more abundant than cycling precursor RNAs, as recently reported in a genome scale analysis (Koike et al., 2012). These differences emphasize the importance of posttranscriptional and posttranslational mechanisms in the regulation of biological rhythms.
Recently, the participation of microRNAs (miRs) in the posttranscriptional control of circadian rhythms has been demonstrated (Alvarez-Saavedra et al., 2011; Balakrishnan et al., 2010; Cheng et al., 2007; Kojima et al., 2010; Luo and Sehgal, 2012; Tan et al., 2012; Yamakuchi, 2012). miRs are small noncoding transcripts responsible for the regulation of various genes (Filipowicz et al., 2008). It is estimated that in mammals, approximately 50% of coding genes are regulated by these elements (Krol et al., 2010). However, few miRs and target clock genes have been reported in the circadian literature.
The aim of this work was to perform an in silico analysis to identify candidate miRs potentially involved in mammalian circadian control. Based on the prediction of target genes and promoter analysis for clock transcription factors, we propose a series of posttranscriptional feedback loops that can be explored in subsequent studies. We also investigated the diurnal expression of miR-27b in human leukocytes.
Materials and Methods
Analysis of 3′UTR Region of Clock Genes
The bioinformatics tools Pictar (http://pictar.bio.nyu.edu), microrna.org (http://www.microrna.org), and TargetScan (http://www.targetscan.org) were used in this study. These programs use conserved elements for the prediction of targets (Lewis et al., 2003; Xia et al., 2009; Yoon and De Micheli, 2006).
We used the target mRNA search in microrna.org, with target sites of conserved miRs with good mirSVR scores. All miRs showed in the results were selected for our analysis. In TargetScanHuman and TargetScanMouse, we used the entrez gene symbol option. In the 3′UTRs of target genes, we selected those with conserved sites for miR families broadly conserved among vertebrates and conserved only among mammals. We chose the target predictions in PicTar for all human miRs based on conservation in mammals (human, chimp, mouse, rat, and dog). The gene ID was included in the search for all miRs predicted to the target gene.
The circadian genes and their isoforms Clock, Bma1l (Arntl), Per1-3, Cry1-2, Rerv-erbalpha (Nr1d1), Rerv-erbbeta (Nr1d2), Timeless, Dec1 (Bhlhe40), Dec2 (Bhlhe41), Rora (Ror-alpha), Rorb (Ror-beta), Rorc (Ror-gamma), Csnk1e (casein kinase 1, epsilon), Csnk1d (casein kinase 1, delta), and Nocturnin (Ccrn4L) were selected for analysis.
We also included Ccne1 (Cyclin e1), Ccnd1 (Cyclin d1), Scp2, Rgn, Khk, Hspd1, Hao1, Grp58 (Pdia3), Gnpat, Eno1, Cat, Calr, Atp5b, Atp5a1, Apoa4, Aldo2 (AldoB), Aldh1a1, and Acaa2, which present a circadian pattern of protein expression with mRNAs expressed constitutively (Balakrishnan et al., 2010; Reddy et al., 2006), and Adcy6 (adenylyl cyclase VI), targeted by the circadian miR-96/miR-182 in mice retina (Xu et al., 2007). Only miRs conserved in mammals and targeting the mentioned genes in humans and mice were selected.
Identification of Putative Target Sites for Circadian Transcription Factors in the Promoter of Candidate miRs
The candidate miRs that presented targets equally predicted in 3 bioinformatics software programs were selected for promoter analysis. The promoter sequences of miRs were obtained in Eukaryotic Promoter Database (EPD). The predictive analyses of putative target sites for circadian transcription factors were performed with the MatInspector program (Cartharius et al., 2005), using an interval of −499 to 100 bp relative to the transcription starting site. ENSEMBL coordinates for promoter sequences and sequence motifs used are available in Supplementary Tables S1 and S2, respectively.
Quantitative Real-Time PCR
For the analysis of daily variation of miR-27b in human leukocytes, we selected 5 young men (mean age 22.8 ± 3.42 years), named herein as S1-S5, of the same ethnicity and intermediate chronotype according to the Horne-Ostberg Morningness-Eveningness Questionnaire (Horne and Ostberg, 1976). These individuals did not experience shift-work or jet lag, were not smokers, and did not regularly use medications. They also presented normal hematological findings. The subjects were told that during the week that preceded the study they should perform their daily activities under a regular routine of sleeping and eating, should avoid excessive light at night, should not ingest foods with caffeine or energetics, and should not perform extensive physical activities.
On the day of blood sampling, food was available from 0700 to 0800 h (breakfast), 0930 to 1030 h (snack), 1200 to 1300 h (lunch), 1530 to 1630 h (snack), and 1900 to 2030 h (dinner), with water available ad libitum. Blood was collected in EDTA tubes every 4 h along a circadian cycle. The study was approved by the Committee on Ethics of the Federal University of Alagoas (#007138/2011-41).
Red blood cells were lysed with Buffer EL (Qiagen, Hilden, Germany), and RNA purification of the remaining cells was performed with miRNeasy Mini Kit (Qiagen). The samples were then stored at −80 °C for further analysis.
The cDNA was synthesized from 1 µg of total RNA using the miScript Reverse Transcription Kit (Qiagen) according to the manufacturer’s protocol. As suggested, we added 4 µL of miScript RT Buffer (includes Mg2+, dNTPs, and primers) and 1 µL of miScript Reverse Transcriptase. The reactions were adjusted to a final volume of 20 µL with RNase-free water. All cDNA samples were diluted (1:10) in RNase-free water for analysis.
The qRT-PCR reactions were performed using StepOne Plus (Applied Biosystems, Foster City, CA, USA) with 2 µL of miScript Primer Assays (Qiagen) for miR-27b and 10 µL of 2x QuantiTect SYBR Green PCR Master Mix, 2 µL of 10x miScript Universal primer, and 4 µL of RNase-free water, which compose the miScript SYBR Green PCR Kit (Qiagen). The cycling conditions were 15 min/95 °C for initial activation of HotStarTaq DNA Polymerase, 15 sec/94 °C for denaturation, 30 sec/55 °C for annealing, and 30 sec/70 °C for extension in a total of 40 cycles. All reactions were performed in duplicates with negative controls, and efficiency tests with melting curve analysis were performed for each primer.
For normalization of miR-27b expression data, we selected SCARNA-17 (Small Cajal body-specific RNA 17) and SNORA-73A (small nucleolar RNA, C/D box 73A) (miScript PCR Controls, Qiagen) as reference genes. The stability analysis of SCARNA-17 and SNORA-73 in the geNorm program (Vandesompele et al., 2002) showed that both genes are stable across our samples (subjects and time points). However, SNORA-73 had a better stability score (0.582) than SCARNA-17 (0.613). Given these results, we decided to use SNORA-73 to normalize the expression data.
The expression of the clock gene Per3 was used as a positive control for diurnal variation. The reactions were also performed with miScript SYBR Green PCR Kit reagents (Qiagen) using the specific forward primer (5′-GCAGAGGAAATTGGCGGACA-3′) and reverse primer (5′-AGAGTGTTGGCTTATTGCGTC-3′). The primer sequences of the noncoding RNAs used were not supplied by the company (Qiagen). Data are presented as relative expression (2–ΔΔCt), and the nadir (time point with the lowest expression level) was used as the calibrator sample.
Statistical Analysis
The grouped time-series data were analyzed by repeated-measures ANOVA and Bonferroni multiple comparison posttest. Considering interindividual differences, we grouped subjects with the same phase according to their peak of expression (based on fold change). The values for each time point are presented as means and SEM. In addition, we analyzed the expression individually with the cosinor test with periods adjusted to 24 h. We also calculated the mean, amplitude, acrophase, and fitted cosine wave from relative expression data. In all analyses, p < 0.05 was considered significant.
The calculations were performed using GraphPad Prism (GraphPad Software, San Diego, CA) for ANOVA and Acro software (http://www.circadian.org/softwar.html) for calculation of the mean, amplitude,acrophase, and fitted cosine wave.
Results
We found 237 predicted miRs in humans and 227 in mice using microrna.org, 202 in humans and 159 in mice using TargetScan, and 114 miRs in both species using PicTar for analysis (data not shown). Sixty-nine miRs were equally predicted using the 3 programs in humans and/or mice (Suppl. Table S3). In Table 1 we present a selected list of these candidate miRs based on literature search for validated targets previously associated with circadian rhythms.
Selected candidate miRs for circadian regulation based on bioinformatic screening and literature search for validated targets or pathway genes.
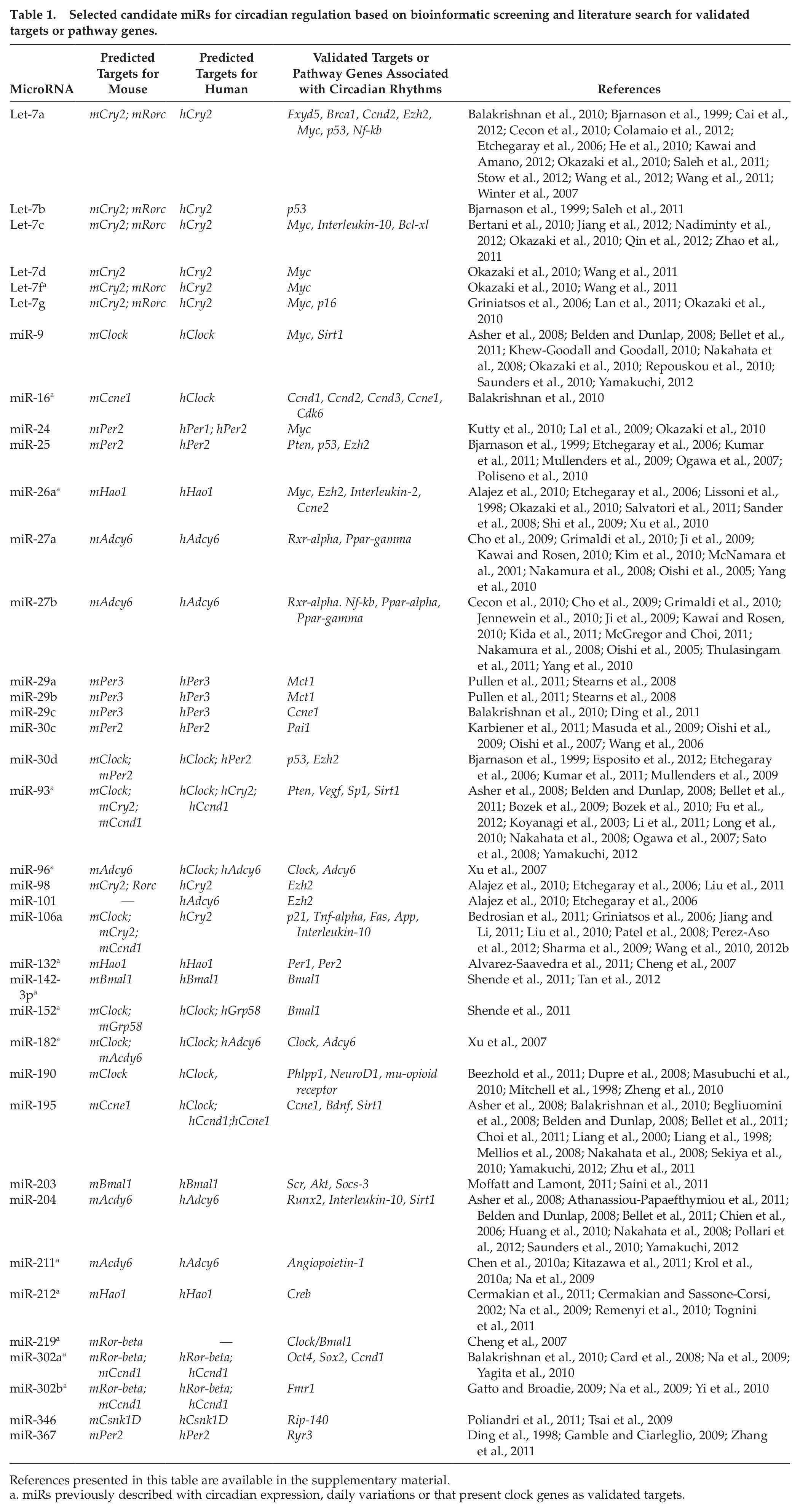
References presented in this table are available in the supplementary material.
miRs previously described with circadian expression, daily variations or that present clock genes as validated targets.
We used MatInspector to investigate whether the 69 miRs have themselves conserved sites for circadian transcription factors in their promoters. Table 2 shows the 33 miRs that meet this criterion. Several hypothetical feedback loops potentially mediated by some of these miRs can be predicted from these data and are represented in Supplementary Figure S1.
Candidate miRs presenting putative sites for circadian transcription factors in their promoters.
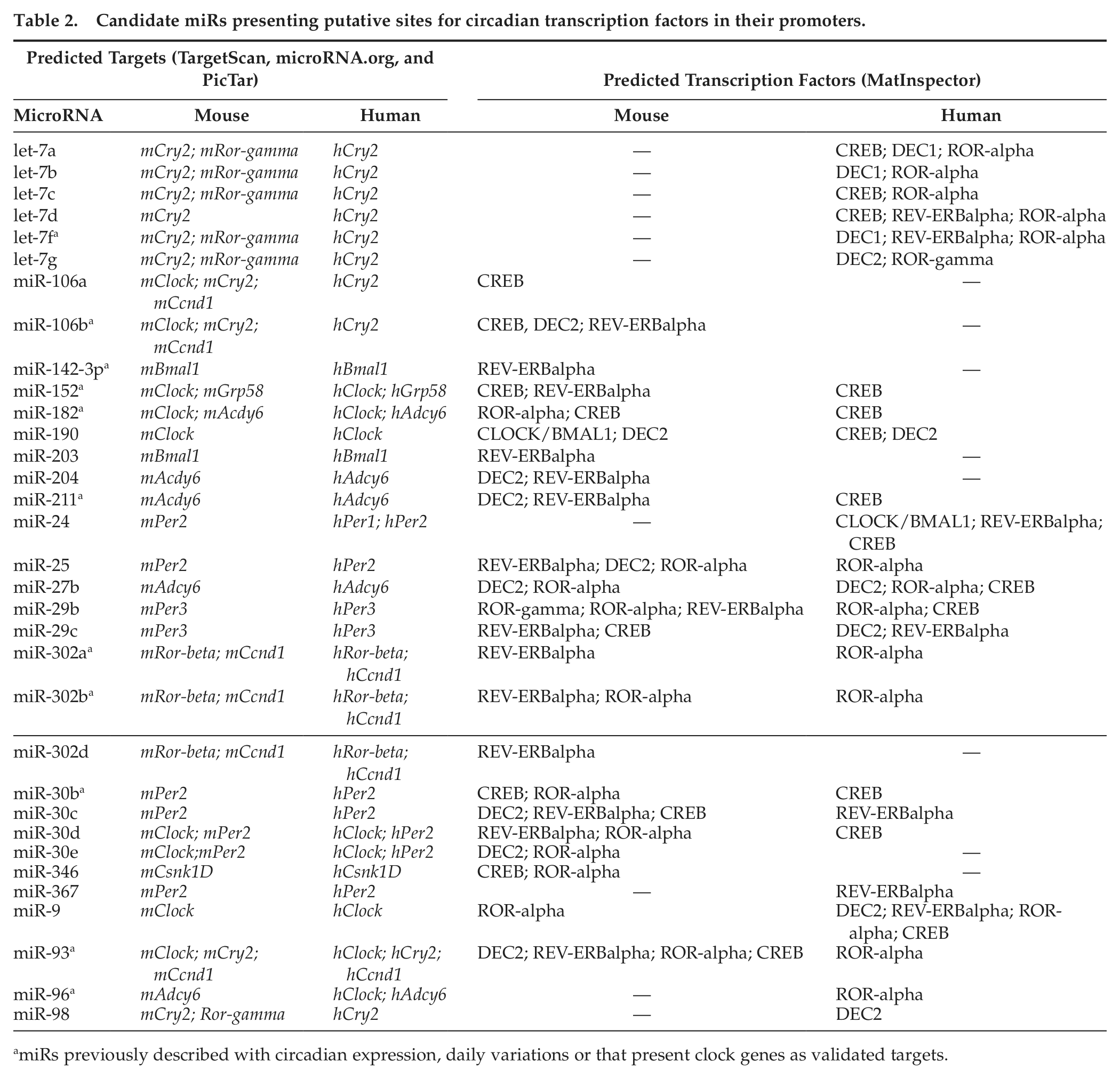
miRs previously described with circadian expression, daily variations or that present clock genes as validated targets.
As an initial approach for experimental evaluation of the potential contribution of the identified miRs to circadian process, we collected blood samples from healthy donors over a period of 24 h and evaluated the expression of miR-27b by qRT-PCR. miR-27b was selected for expression analysis in human leukocytes based on its potential involvement in hematopoiesis and immune response.
The individual analysis showed that miR-27b expression data from S3 and S4 present a significant difference in cosinor test. In the analysis of Per3 expression, only S4 presents p < 0.05 (Fig. 1). ANOVA showed a significant temporal variation in miR-27b when S1 and S5, with peak expression at 00 h, were combined (p = 0.041), and when S3, S4, and S5, with peak expression at 12 h (S3) and 8 h (S4 and S5), were combined for Per3 analysis (p = 0.040) (Fig. 1). Table 3 shows the descriptive analysis of miR-27b and Per3 expression. The average fold change in relative expression between nadir and acrophase (sampled time point with the highest expression level) was 2.01 (±0.11 SEM) for miR-27b and 2.31 (±0.10 SEM) for Per3.
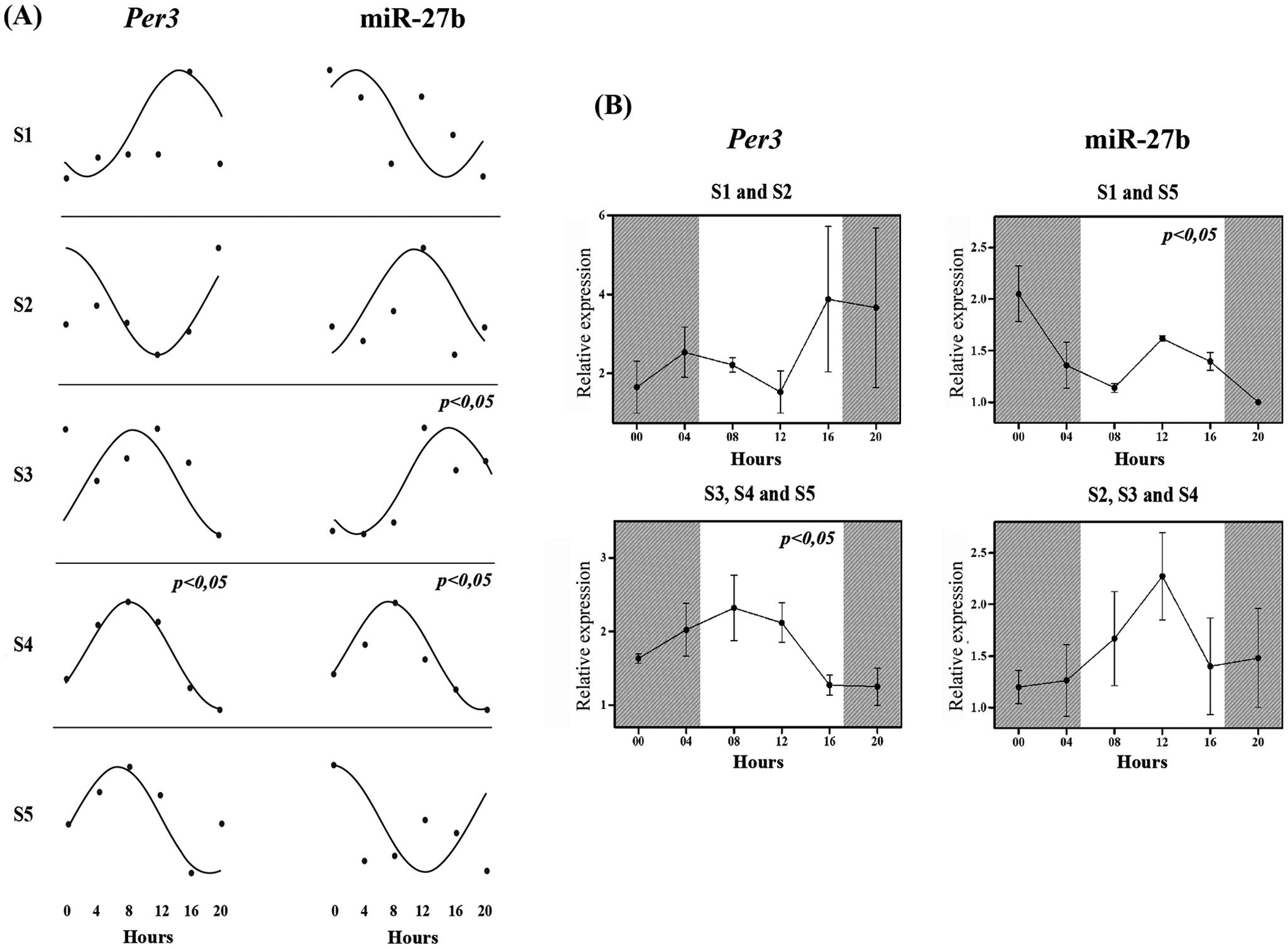
Daily variation of miR-27b and Per3 expression in human leukocytes measured by qRT-PCR. (A) Cosinor analysis of each subject. The lines show the fitted cosine wave and the dots correspond to the relative expression data (fold change). (B) Analysis by repeated-measures ANOVA of different groups. Above are clustered subjects with peak of expression in dark phase and below in light phase. The data are presented as mean and SEM. Shaded area marks the dark phase of the day. Nadir (time point with the lowest expression level) was used as calibrator sample in fold change equation (2−ΔΔCT) and SNORA-73A as endogenous reference gene. p < 0.05 was considered significant for both statistical tests.
Descriptive analysis of miR-27b and Per3 expression in leukocytes of 5 subjects.
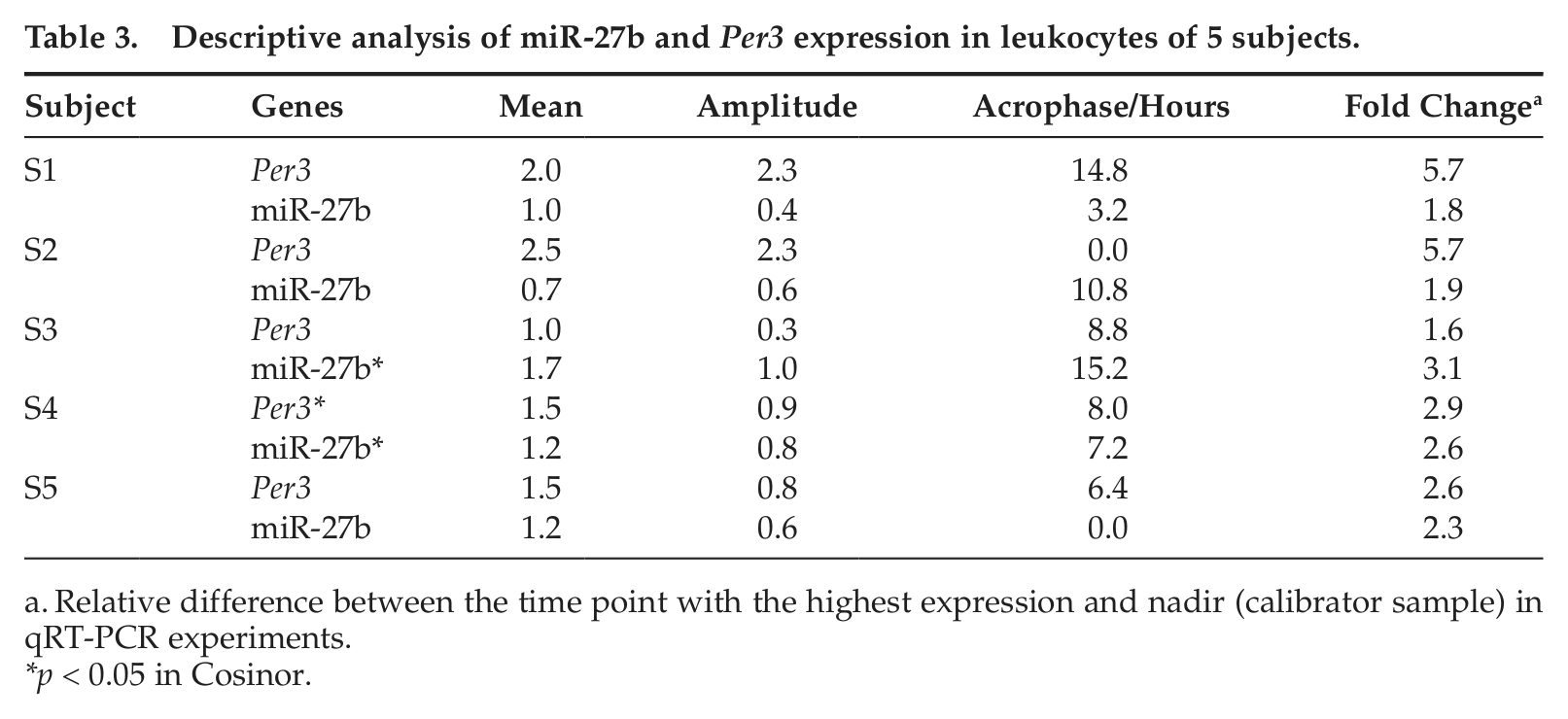

Relative difference between the time point with the highest expression and nadir (calibrator sample) in qRT-PCR experiments.
p < 0.05 in Cosinor.
Discussion
Despite the pivotal role of posttranscriptional mechanisms in the regulation of circadian rhythms and the involvement of miRs in this process, there are few examples in the literature linking these genetic elements to the control of circadian clock. Using an in silico approach for identifying miR target sites in clock genes or genes with circadian expression at a protein level only, coupled with the investigation of putative sites for clock transcription factors in the promoter of these miRs, we present a list of candidate genes that might be important for the circadian molecular machinery. Since this strategy is independent of the biological sample examined, it has the potential to reveal miRs that show circadian expression in any particular tissue.
The bioinformatics tools TargetScan, PicTar, and microrna.org have been successfully used to identify candidate genes further validated experimentally, including miRs previously mentioned in circadian studies, such as miR-182, miR-132, miR-219, miR-142-3p, and miR-16 (Alvarez-Saavedra et al., 2011; Balakrishnan et al., 2010; Cheng et al., 2007; Saus et al., 2010; Tan et al., 2012). The intersection of the results using different prediction algorithms increases the confidence of the findings. In fact, one-third of the 69 miRs were previously described with circadian expression or daily variation in different tissues (Alvarez-Saavedra et al., 2011; Balakrishnan et al., 2010; Cheng et al., 2007; Na et al., 2009; Nagel et al., 2009; Saus et al., 2010; Shende et al., 2011; Tan et al., 2012; Xu et al., 2007). Some of them modulate circadian rhythms targeting clock genes (Alvarez-Saavedra et al., 2011; Cheng et al., 2007; Nagel et al., 2009; Shende et al., 2011; Tan et al., 2012).
We selected miR-27b to investigate its expression in human leukocytes collected over a period of 24 h as an experimental model of biological rhythmicity. Studies have demonstrated that clock genes oscillate in human leukocytes and that changes in their expression are associated with different pathologies (Ando et al., 2009; Boivin et al., 2003; Cai et al., 2010; Gouin et al., 2010; Haimovich et al., 2010; Kusanagi et al., 2004; Taniguchi et al., 2009; Watanabe et al., 2012; Yang et al., 2011).
We selected miR-27b for this experimental study based on its potentially relevant functional participation in leukocyte biology. miR-27b expression is altered in T-lymphocytes from patients with multiple sclerosis (Guerau-de-Arellano et al., 2011). Moreover, there is evidence for the possible involvement of this miR in hematopoiesis and immune response through the modulation of retinoid X receptor α (RXR-alpha), peroxisome proliferator–activated receptors (PPARs), and nuclear factor-κB (NF-κB), which were previously associated with circadian rhythms (Cecon et al., 2010; Cho et al., 2009; Gerondakis et al., 2012; Grimaldi et al., 2010; Jennewein et al., 2010; Ji et al., 2009; Karbiener et al., 2011; Kawai and Rosen, 2010; Kida et al., 2011; Kim et al., 2010; McGregor and Choi, 2011; McNamara et al., 2001; Oishi et al., 2005; Rasooly et al., 2005; Stephensen et al., 2007; Thulasingam et al., 2011; Yang et al., 2010). Thus, the study of miR-27b may add to our understanding of its contribution to the normal and altered physiology of leukocytes.
Herein, miR-27b is consistently predicted to target Adcy6, which presents circadian expression in the pineal gland (Han et al., 2005) and daily variation in the retina (Xu et al., 2007). ROR-alpha and DEC2 are predicted to be potential regulators of miR-27b in human and mouse and CREB only in human (Table 2).
We showed that miR-27b presented daily variation in some subjects when they were studied individually with cosinor analysis or when they were grouped in ANOVA according to peak phase of expression (Fig. 1). The divergences in the acrophase observed between individuals may result from differences in behavior, environmental conditions, or genetic background and may have contributed to the lack of statistical significance found in some grouped data. In fact, interindividual variations have been reported in diurnal expression of clock genes in leukocytes (Balmforth et al., 2007; Teboul et al., 2005) and may be an interfering factor for ANOVA, since grouping subjects with different phases could mask temporal variation.
The amount of data across time series (interval of sampling) might not have been ideal for cosinor testing in our samples, which may have resulted in nonsignificant values, even for Per3 expression in some individuals. These, together with the total number of subjects evaluated, are limitations of our study. However, we believe that the statistical significance found in the analysis of miR-27b in different individuals, using cosinor or ANOVA, reinforces the hypothesis of a circadian variation of this miR in human leukocytes and its possible contribution to the circadian control of RXR-alpha, PPAR-gamma, NF-κB, and ACDY6 activity.
To our knowledge, this is the first study that demonstrates an in vivo diurnal variation of an miR in human tissue. Although the gene list corresponds to predicted candidates based on bioinformatic strategies, which is the main limitation of our study, the data presented in the literature and herein suggest the potential involvement of the identified miRs with the posttranscriptional regulation of biological rhythms and highlight them as potential candidates for circadian genes. Further validation studies could confirm these hypotheses.
Footnotes
Acknowledgements
This research was supported by CAPES and CNPq. The authors thank Bruna P. dos Santos for collecting blood samples.
Conflict of Interest Statement
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
