Abstract
Objective:
To explore whether circulating plasma proteins have potential causal effects on gout risk by integrating genome-wide association study (GWAS) data with protein quantitative trait loci (pQTL) information, aiming to identify candidate proteins and pathways for therapeutic targeting.
Introduction:
Despite advances in gout pathophysiology, personalized therapies and validated drug targets remain limited. We conducted an exploratory Mendelian randomization (MR) analysis to assess whether circulating plasma proteins show evidence consistent with potential causal effects on gout risk, integrating GWAS with pQTL data to prioritize proteins and pathways for follow-up.
Methods:
We applied an integrated genetics framework combining proteome-wide MR, reverse MR, Bayesian colocalization, and sensitivity analyses. Plasma pQTL datasets were obtained from Zheng et al. and deCODE Genetics; gout GWAS were from UK Biobank and the FinnGen R12 study. Primary inference used inverse-variance weighting (IVW) with MR-Egger and complementary sensitivity tests. Colocalization used Bayesian methods; reverse MR evaluated potential reverse causation. Where available, findings were checked in external datasets. Analyses followed STROBE-MR guidance and were implemented in R with standard MR/colocalization packages.
Results:
Five circulating proteins (FN1, PLAU, CPQ, SPOCK2, and FAM213A) met the predefined discovery threshold. Reverse Mendelian randomization provided no evidence supporting reverse causality from gout to protein levels, while Steiger directionality tests supported the protein-to-gout causal orientation. Bayesian colocalization indicated moderate to strong evidence of shared genetic signals for PLAU, FAM213A, and FN1, whereas the evidence for CPQ and SPOCK2 was comparatively weak. In the independent replication analysis, genetically predicted higher levels of PLAU and CPQ were directionally associated with an increased risk of gout, while higher FN1 levels showed a consistent protective association. Nevertheless, effect estimates and precision varied across datasets, and the overall findings should be interpreted as exploratory rather than confirmatory.
Conclusion:
This exploratory, assumption-dependent MR work nominates five circulating proteins as hypothesis-generating candidates for gout risk and potential therapeutic targeting. Given method assumptions and cross-dataset heterogeneity, functional validation and independent replication are required to clarify mechanisms, confirm targetability, and assess translational relevance.
Introduction
Gout is a prevalent form of crystal-related arthritis characterized by the deposition of sodium urate in the joints, often associated with hyperuricemia. 1 This inflammatory joint disease predominantly affects men aged 40 years and older, manifesting as acute pain attacks and chronic joint conditions that can lead to significant morbidity and disability. 2 The global incidence and prevalence of gout are on the rise, with prevalence estimates ranging from 0.1% to 10% and annual incidence rates varying from 0.3 to 6 cases per 1000 individuals. 3 In the United Kingdom (UK), an analysis of health records from 1.2 million individuals indicated that the prevalence of gout in 2012 was approximately 2.49%. 4 In China, the cumulative incidence of gout among adults showed a slight increase from 0.9% to 1.1% between 2000 and 2005. 5 Concurrently, there is a rising trend in the annual incidence of hospitalization for gout among patients with type 2 diabetes, hypertension, coronary atherosclerotic heart disease, and chronic kidney disease.6–9 Despite advancements in understanding the pathophysiology of gout and the availability of effective urate-lowering treatments, the prevalence of gout continues to escalate. According to the 2019 Global Burden of Disease Study, the global incidence of gout has increased by 63.44% over the past two decades, resulting in a 51.12% increase in years lived with disability, which presents a significant public health challenge. 10 Gout is associated with a range of cardiometabolic, renal, and lifestyle-related factors, including hypertension, myocardial infarction, stroke, obesity, hyperlipidemia, type 2 diabetes, chronic kidney disease, and dietary factors such as high intake of purine-rich foods, red meat, seafood, and alcohol.11,12 Current pharmacological treatments are primarily categorized into two groups: the first includes nonsteroidal anti-inflammatory drugs and colchicine for managing acute attacks; the second consists of medications aimed at preventing gout recurrence, primarily by lowering uric acid levels, such as allopurinol and febuxostat.13,14 However, these treatment options can lead to significant side effects, including nausea, vomiting, and abdominal pain, as well as adversely impacting gastrointestinal function, which may reduce patient compliance with treatment.15–17
Circulating proteins are crucial components of plasma proteins, and their abnormal expression is closely linked to the onset and progression of various diseases. 18 These proteins include transport proteins such as albumin, tissue leakage proteins, and signaling molecules, which are essential for understanding pathological conditions. 19 Notably, the anti-inflammatory effect of Annexin A1 offers a novel strategy for treating gout by promoting macrophage recruitment and inducing neutrophil apoptosis.20,21 Previous studies have demonstrated that elevated plasma fibrinogen levels correlate with gout disease activity, 22 underscoring the significance of circulating proteins in the pathogenesis of gout. A proteomic study investigating the underlying mechanisms of gout revealed that hypoxanthine-guanine phosphoribosyltransferase activity is diminished in gout patients with Down syndrome, accompanied by abnormal protein expression. This suggests that renal dysfunction and abnormal lipid metabolism may be key factors in the development of gout. 23 Further research employing iTRAQ technology in conjunction with liquid chromatography-tandem mass spectrometry identified 369 differentially expressed proteins in the serum of gout patients. Additionally, a large sample enzyme-linked immunosorbent assay confirmed that tubulin beta-4A chain protein is differentially expressed between primary gout patients and controls, indicating that this protein may play a significant role in the pathogenesis of primary gout. 24 While this study underscores the potential value of circulating proteins, the specific clinical advantages of targeting these proteins remain unclear. Consequently, comparing the differential expression of proteins between gout patients and healthy individuals is crucial for a deeper understanding of the mechanisms underlying disease changes and progression.
Randomized clinical trials (RCTs) represent the gold standard for assessing the efficacy of drug treatments. However, the high costs and inherent challenges associated with conducting RCTs have made it difficult to obtain large-scale, high-quality studies that investigate the potential effects of circulating plasma proteins on gout. This limitation may contribute to the scarcity of relevant research in this area. By employing Mendelian randomization (MR) techniques and utilizing genetic variation as an instrumental variable, this study elucidates the intricate causal relationships between specific circulating proteins and gout, thereby mitigating the biases often present in traditional observational studies.25,26 MR has become a widely used approach for elucidating causal mechanisms across diverse diseases, from cardiometabolic, cancer, and inflammatory disorders to emerging areas involving immunometabolic regulation and gut–microbiota interactions, thereby providing a powerful method for identifying and prioritizing therapeutic targets for drug development.27–31 Nevertheless, there have been few reports to date on MR studies that integrate genome-wide association studies (GWAS) with pQTL data related to gout. This study aims to leverage proteomic methodologies such as MR analysis, colocalization analysis, reverse MR, external validation, and replication meta-analysis to identify critical circulating plasma proteins and their associated biological pathways linked to gout, ultimately seeking novel drug targets.
Materials and methods
Study design
This was a two-sample Mendelian randomization study conducted between December 2024 and June 2025, based on publicly available GWAS and protein quantitative trait loci (pQTL) summary data. This investigation employed an integrated genetics methodology, as illustrated in Figure 1, which outlines the study’s workflow. Initially, a proteome-wide MR approach was used to explore potential genetic links between plasma proteins and gout susceptibility. Following this, we conducted reverse MR, Bayesian co-localization, and other supplementary analyses to evaluate the robustness of our findings. For external validation, plasma pQTL data from deCODE and the most recent release of the UK Biobank Whole-Genome Sequencing Consortium gout GWAS were examined to evaluate consistency of the associations in an independent dataset.
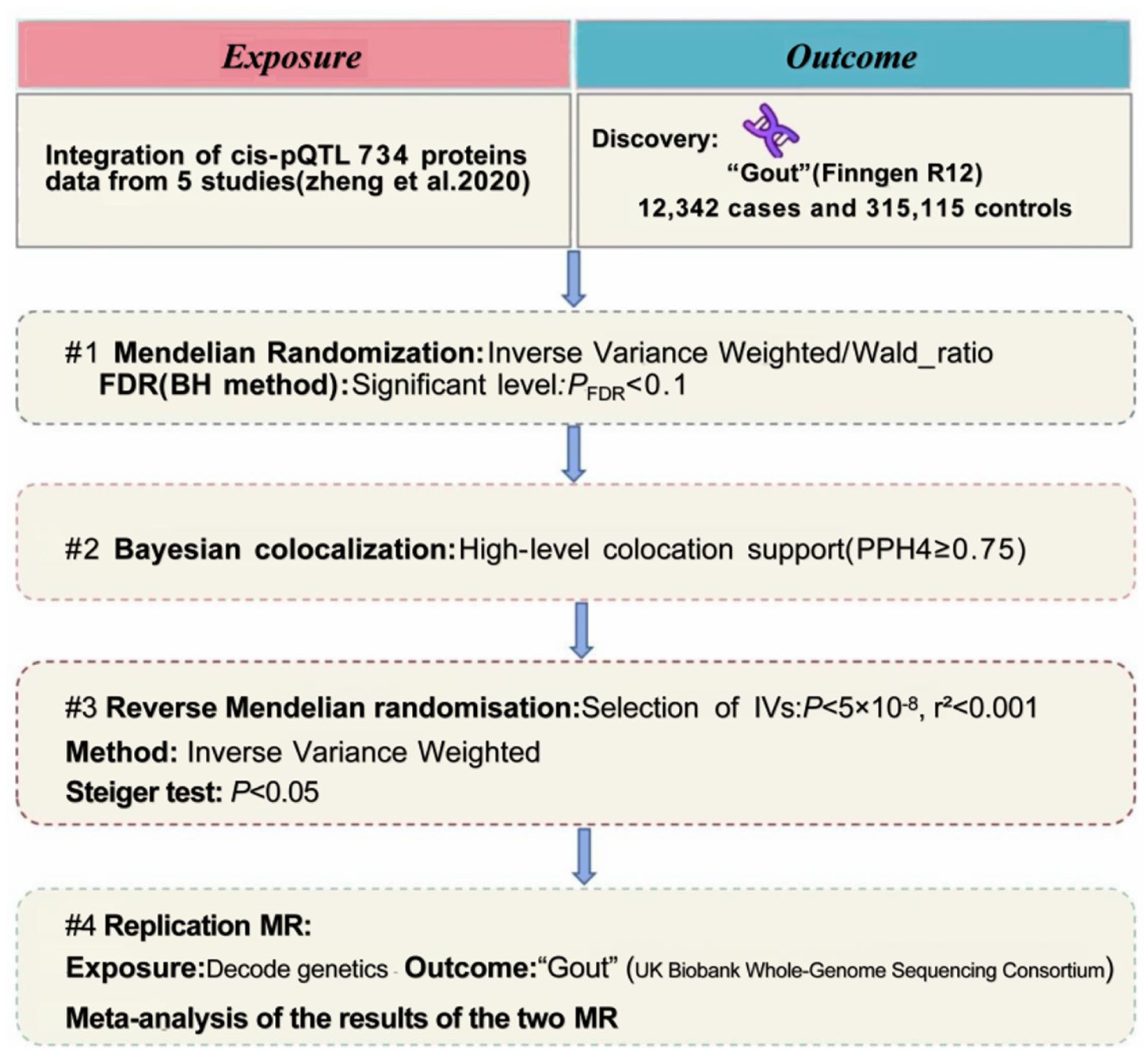
Flowchart of the study design.
This study involved the use of pre-existing data from studies involving human participants, with all necessary approvals and informed consent obtained according to the guidelines established by the ethics review boards. Moreover, this work adheres to the STROBE-MR checklist (Supplemental File 1). 32
Plasma pQTL data source
For the preliminary analysis, plasma pQTL data were derived from Zheng et al.’s 33 study, which merged results from five earlier GWAS studies34–38 spanning aptamer-based SomaLogic SOMAscan and antibody-based Olink PEA platforms. Ethical approval for this study was waived because this study utilizes published data from prior research involving human participants. All relevant data were obtained with formal written approval and informed consent from the institutional ethics review board. Such as, all participants gave informed consent before joining the study and the National Research Ethics Service approved this study (11/EE/0538). The contributing studies primarily involved participants of European ancestry, typically including ~3000–3500 individuals each and quantifying from ~80 to >3000 plasma proteins depending on the assay panel. Inclusion criteria for plasma pQTLs were: (i) genome-wide significance (p < 5 × 10−8); (ii) outside the major histocompatibility complex region (chromosome 6, 26–34 Mb); (iii) independent associations, with linkage disequilibrium (LD) r2 < 0.001; and (iv) confirmed cis-acting pQTLs. In total, 738 cis-SNPs related to 734 proteins were included, with missing or unsuitable data excluded.
For external validation, data from the deCODE Genetics database, profiling approximately 35,000 Icelandic participants of European ancestry using SomaScan v4 (~4700 plasma protein aptamers), were used, applying the same pQTL selection criteria (https://www.decode.com/). 39 Data were cross-referenced with original sources for accuracy.
Gout GWAS data source
Gout-related GWAS data were obtained from FinnGen release R12 (endpoint M13_GOUT). This dataset comprises Finnish-ancestry participants (European population subset), including 12,342 gout cases and 315,115 controls. Because we also evaluated potential reverse causation (gout→proteins), we prespecified a bidirectional MR framework, in which gout served as the exposure and circulating proteins as outcomes for the reverse direction. Accordingly, instrumental variants (IVs) for gout were selected at genome-wide significance (p < 5 × 10−8) and LD clumped at r2 < 0.001 (European LD reference), with standard harmonization (effect-allele alignment and exclusion of ambiguous palindromic SNPs at intermediate allele frequency).
As an external validation cohort, we analyzed gout summary statistics from the UK Biobank Whole-Genome Sequencing (WGS) Consortium, restricted to non-Finnish European ancestry (19,142 cases and 439,298 controls), ensuring ancestry consistency with the primary datasets to minimize population stratification bias and enhance comparability across analyses.
Statistical analysis
MR methodology
To ensure the accuracy of the MR results, three main assumptions were made 40 : (i) the correlation assumption, which ensures that IVs are strongly associated with the exposure; (ii) the independence assumption, which asserts that IVs are not influenced by confounders; and (iii) the exclusivity assumption, stating that IVs affect the outcome only through the exposure, without horizontal pleiotropy. For the MR analysis, when multiple IVs were available, the inverse variance weighting (IVW) method was primarily used. If there was no significant heterogeneity, a fixed-effects IVW model was applied, and if heterogeneity was detected, a random-effects IVW model was employed. 41 To assess horizontal pleiotropy and test the robustness of the results, MR-Egger regression was applied. This method tests for pleiotropy and generates corrected estimates. 42 Additionally, it includes an intrinsic distortion test, which assesses the differences in causal estimates before and after the removal of outliers. 43 In this study, we conducted a proteome-wide MR analysis utilizing cis-pQTL as the genetic IV for circulating proteins to investigate the association between circulating plasma proteins and the risk of gout. 44 In the case of a single pQTL for a protein, the Wald ratio was used to calculate the odds ratio (OR) for gout risk, expressed as a 10-fold standard deviation increase in plasma protein levels. For multiple hypothesis testing, a false discovery rate (FDR) correction was applied, with P(FDR) < 0.1 considered statistically significant. This threshold was selected based on previously published MR studies, where an FDR cutoff of 0.1 is commonly adopted to balance the discovery of biologically meaningful associations against the risk of false positives.45,46 To validate our preliminary findings, we employed the same analytical approach.
Finally, the F statistic was calculated to assess the power of each SNP, using the formula: F = R2(N−K−1)/K, where R2 represents the variance explained by genetic variation, K is the number of SNPs, and N is the sample size. SNPs were selected if their F statistic exceeded 10 to avoid weak instrumental variable bias. 47
Colocalization analysis
We tested whether plasma protein and gout signals at the same locus reflect a shared causal variant using the coloc (coloc.abf) framework on ancestry-matched (European) summary statistics. Analyses used ±500 kb windows around the protein sentinel or gout index SNP (harmonized alleles; ambiguous palindromes removed). Priors followed common practice. Evidence for colocalization was summarized by PP.H4; PP.H4 > 0.70 was interpreted as strong support for a shared causal signal, whereas high PP.H3 indicated nearby but distinct signals. Because coloc assumes one causal variant per trait, we report sensitivity to window size and priors
External verification
For validation, we repeated the MR analyses using pQTL data from the deCODE Genetics database and gout GWAS data from the UK Biobank Whole-Genome Sequencing (WGS) Consortium, applying identical instrument-selection and harmonization procedures to confirm the robustness of the causal estimates.
Reverse causality detection
To detect reverse causality, 22 gout IVs from the Finnish R12 GWAS database were analyzed alongside plasma circulating proteins, using the MR IVW method. Steiger filtering was applied to verify the direction of the association between circulating proteins and gout. 48 Results were considered significant if p < 0.05.
Data availability
We analyzed publicly available, de-identified GWAS summary datasets—plasma pQTLs for 734 proteins and gout GWAS from UK Biobank and FinnGen (European ancestry). All datasets are openly accessible to promote data sharing and research transparency. Statistical analyses were performed in R (v4.2.0) using the TwoSampleMR (v0.5.6) and coloc (v5.2.3) packages.43,49–52
Results
Screening gout-related proteins in the proteome
After FDR correction (PFDR < 0.1), this MR analysis identified five circulating proteins with associations consistent with potential effects on gout risk (Table 1, Figure 2, and Supplemental Table 1): Fibronectin 1 (rs1250258), Plasminogen Activator Urokinase (PLAU, rs2227551), SPARC/Osteonectin, Cwcv and Kazal-Like Domains Proteoglycan 2 (SPOCK2, rs1245540), Family with Sequence Similarity 213 Member A (FAM213A, rs10887868), and Carboxypeptidase Q (CPQ, rs145746079). Specifically, genetically predicted higher levels of PLAU (OR = 1.25; 95% CI, 1.11–1.39; p = 1.29 × 10−4), FAM213A (OR = 1.36; 95% CI, 1.18–1.57; p = 1.67 × 10−5), and CPQ (OR = 1.22; 95% CI, 1.09–1.37; p = 7.53 × 10−4) were associated with increased gout risk, whereas higher levels of FN1 (OR = 0.93; 95% CI, 0.88–0.97; p = 7.72 × 10−4) and SPOCK2 (OR = 0.81; 95% CI, 0.71–0.91; p = 6.45 × 10−4) were associated with reduced risk.
MR results for plasma proteins significantly associated with gout.

PVE denotes the proportion of variance explained. Only cis-acting SNPs were utilized.

Mendelian randomization (MR) estimates of the causal effects of circulating plasma proteins on gout risk.
Sensitivity analysis of the gout pathogenic proteins
Based on the preliminary MR findings, five circulating proteins—FN1, PLAU, CPQ, SPOCK2, and FAM213A—emerged as hypothesis-generating candidates for potential therapeutic targeting in gout. First, the reverse MR analysis did not indicate any causal effect of gout on the levels of these five plasma proteins (Table 2), and the results from Steiger filtering further corroborated the consistency of these findings. Secondly, Bayesian co-localization analysis strongly suggests a relationship between gout and the following proteins: PLAU (PP.H4 = 0.88), FAM213A (PP.H4 = 0.92), and FN1 (PP.H4 = 0.76). These three proteins exhibit a significant association with gout and share the same genetic variation. In contrast, CPQ (PP.H4 = 0.01) and SPOCK2 (PPH4 = 0.04) demonstrated a weak co-localization relationship with gout (Table 2 and Figure 3, Supplemental Table 1).
Overview of reverse causality detection and Bayesian co-localization analysis for five candidate causal proteins.

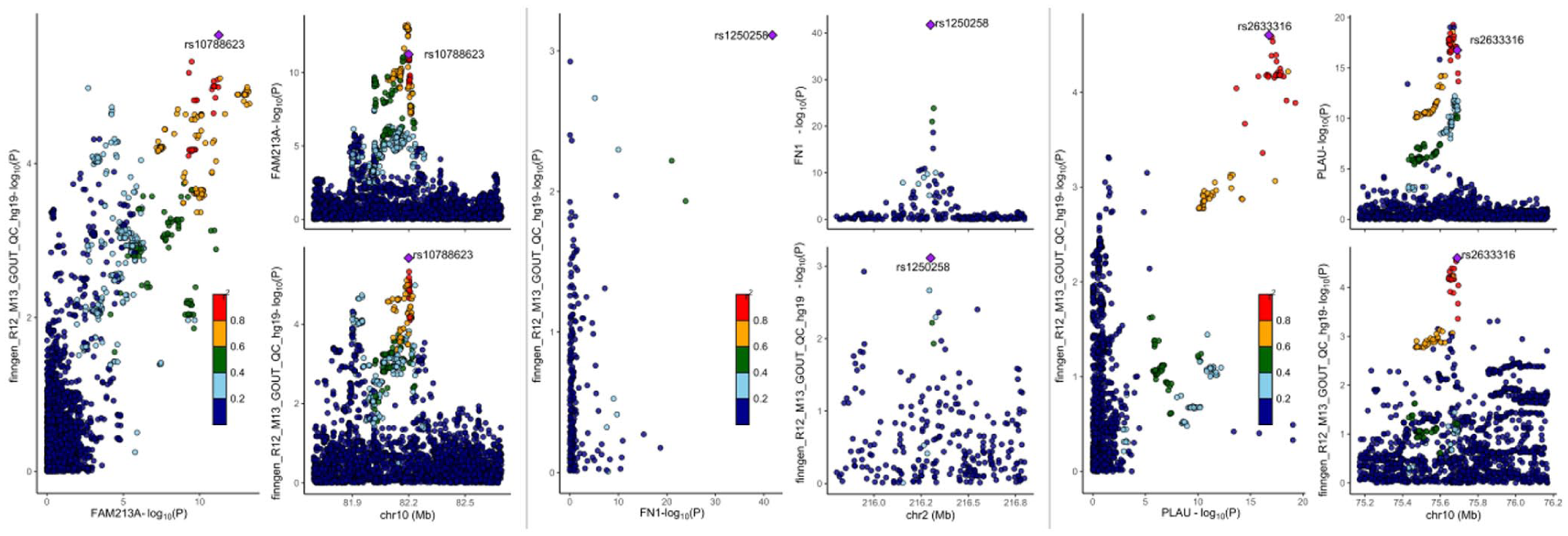
It shows the sensitivity analysis of three plasma proteins (FN1, PLAU, and FAM213A) as potential drug targets for gout.
External validation and replication meta-analysis
We performed external validation in the independent UK Biobank Whole-Genome Sequencing Consortium dataset for five circulating plasma proteins (FN1, PLAU, CPQ, SPOCK2, and FAM213A), applying harmonized instrument-selection criteria. Per a pre-specified concordance criterion, FAM213A was excluded from meta-analytic interpretation due to discordant effect directions across analyses. Fixed-effects meta-analysis supported positive associations of CPQ (OR = 1.09; 95% CI, 1.03–1.15; p = 0.003) and PLAU (OR = 1.18; 95% CI, 1.08–1.29; p < 0.001) with gout risk, whereas FN1 showed a protective effect (OR = 0.95; 95% CI, 0.93–0.98; p < 0.001). SPOCK2 exhibited a concordant direction across datasets but did not reach statistical significance (OR = 0.97; 95% CI, 0.92–1.01; p = 0.138; Figure 4).

It presents the meta-analysis of the causal associations between five serum proteins (FN1, PLAU, CPQ, SPOCK2, and FAM213A) and gout. The results are expressed as ORs with corresponding confidence intervals.
Summary of high-confidence protein targets
By integrating evidence from the discovery, sensitivity, and external validation stages, PLAU, CPQ, and FN1 were identified as high-confidence protein candidates with consistent genetic associations with gout. Elevated genetically predicted levels of PLAU and CPQ were robustly associated with increased gout risk across multiple datasets and analytical frameworks, supported by strong colocalization and meta-analytic concordance. In contrast, higher levels of FN1 were consistently linked to a reduced risk of gout, suggesting a potential protective or reparative role in disease pathophysiology. Collectively, these results highlight PLAU and CPQ as risk-enhancing therapeutic targets, whereas FN1 may serve as a protective target, offering genetically grounded insights for the development of protein-based interventions in gout.
Discussion
In this study, we used a two-sample MR framework to evaluate whether genetically proxied levels of circulating plasma proteins are associated with the risk of gout. By integrating large-scale pQTL datasets from European-ancestry cohorts with genome-wide association data for gout, and by applying complementary approaches including reverse MR, Steiger directionality tests, Bayesian colocalization, and external validation in an independent cohort, we aimed to genetically prioritize proteins and pathways that may contribute to gout pathogenesis and could ultimately inform therapeutic development. Our intention is to highlight protein candidates that are supported by converging genetic evidence and that warrant mechanistic follow-up.
While the primary screen implicated five proteins, not all signals generalized across cohorts. Using FinnGen R12 and UK Biobank WGS, we now classify proteins by replication: PLAU, CPQ, and FN1 showed concordant directions and retained statistical support in meta-analysis; SPOCK2 showed the same protective direction but did not reach significance in UKB and is treated as exploratory; FAM213A displayed a discordant effect direction and was excluded from meta-analytic interpretation. These discrepancies may reflect differences in case definitions, allele frequencies, LD structure, sample size, or proteomic platforms. Accordingly, proteins that fail replication or show opposing directions are interpreted as hypothesis-generating rather than confirmed causal mediators. We also use colocalization to prioritize signals with convergent genetic support (e.g. higher PP.H4 for PLAU and FN1).
Gout, a commonly encountered inflammatory arthritis, is characterized by persistently elevated serum uric acid levels exceeding 408 μmol/L, leading to the deposition of monosodium urate crystals in joints and soft tissues. 1 Cytokines such as IL-1β, TNF-α, and IL-6 are key mediators in its pathogenesis, primarily through activation of the NLRP3 inflammasome.53,54 Interestingly, our results also showed that IL1RN, which encodes the interleukin-1 receptor antagonist, was included in the MR analysis but did not reach the statistical significance threshold (P(FDR) > 0.1), and was therefore not prioritized. Despite its relatively clear pathophysiology, gout is often underestimated and inadequately treated in clinical practice. 55 With the rising prevalence of hypertension, obesity, metabolic syndrome, type 2 diabetes, coronary artery disease, and chronic kidney disease, the global incidence of gout has escalated in recent decades.3,56–58 Acute gout attacks are typically associated with intense pain, and if not promptly addressed, they may restrict joint mobility and cause further tissue damage, significantly impairing the patient’s quality of life. 59 While the pathogenesis of gout is well understood and medications such as colchicine, allopurinol, and urate oxidase have been utilized in clinical practice for many years, traditional therapies have notable limitations. They do not directly target the site of inflammation and are prone to rapid inactivation within the body, which diminishes their efficacy and heightens the risk of systemic side effects.60,61 Consequently, there is an urgent need to identify new drug targets and enhance the treatment of gout.
PLAU is a gene that encodes a secreted serine protease, primarily responsible for converting plasminogen into plasmin, which functions to degrade fibrin in the blood. 62 Additionally, PLAU plays a pivotal role in the remodeling of the extracellular matrix (ECM) and the activation of growth factors, both of which are essential for biological processes such as cell migration, wound healing, and tumor development.63,64 Proteolytic joint destruction in inflammatory arthropathy is mediated, at least in part, by the plasminogen activators(PA)/fibrinolytic system, with plasmin serving as the central enzyme. Plasmin is a broad-spectrum serine protease involved in fibrinolysis and thrombolysis, and it is also necessary for cell migration and the tissue remodeling associated with ECM degradation. 65 Previous studies have suggested a potential pathophysiological role for the PA/fibrinolytic system in destructive joint diseases.66–68 Furthermore, research indicates that the PA/fibrinolytic system is significantly involved in the pathophysiological processes of gouty arthritis. Notably, the activity of this system in gouty arthritis is greater than that observed in osteoarthritis; during acute gout attacks, the synovium produces increased levels of PA, which accumulate in the synovial fluid, potentially promoting further degradation of articular cartilage and bone through the proteolytic activity mediated by the PA/fibrinolytic system. 69 Fibrin deposition, cell migration, and tissue remodeling are core processes in inflammatory joint diseases, including gouty arthritis. Studies have demonstrated that elevated levels of Urokinase-type Plasminogen Activator (uPA), Soluble Urokinase-type Plasminogen Activator Receptor, and Plasminogen Activator Inhibitor-1 in patients experiencing acute gout attacks correlate positively with the activity of the PA/fibrinolytic system, underscoring the importance of this system in the inflammatory response. 70 Other studies have demonstrated that miRNA-181b inhibits the Protein kinase B-Nuclear Factor kappa-light-chain-enhancer of activated B cells (AKT/NF-κB) pathway and reduces inflammation by targeting PLAU. Conversely, increased PLAU expression or inhibition of miRNA-181b enhances the inflammatory response. 71 The results of our study indicate that PLAU and Gout exhibit strong co-localization evidence. We speculate that PLAU is likely to influence Gout through these mechanisms and may have the potential to serve as a therapeutic target for Gout in the future.
FN1 showed an inverse association with gout risk, suggesting a potentially protective or reparative role. Fibronectin is a key structural and signaling component of the extracellular matrix, involved in cell adhesion, wound repair, matrix assembly, and tissue remodeling.72–74 In the context of crystal-driven synovitis, features of the joint matrix microenvironment are closely linked to tissue damage and remodeling; notably, monosodium urate crystals can nucleate on damaged cartilage collagen fibrils, implicating extracellular matrix organization in gout pathogenesis.75,76 Accordingly, adequate matrix organization might limit erosive damage or modulate immune-cell trafficking. 77 Early rheumatologic studies showed that fibronectin associates with complement-fixing/IgG immune complexes in synovial fluid, including in gout, suggesting a link between fibronectin and immune complex–mediated inflammation. 78 Moreover, activated polymorphonuclear leukocytes in inflammatory synovitis, including gout, have been shown to synthesize and release fibronectin de novo, indicating that inflammatory cells themselves may contribute to the fibronectin pool within the joint microenvironment. 79 Our findings are consistent with the hypothesis that higher circulating or tissue-available FN1 may help maintain joint integrity or dampen destructive remodeling during gout flares. Importantly, FN1 also demonstrated supportive colocalization evidence in our analyses, which increases confidence that fibronectin biology—rather than a nearby correlated gene—might be relevant to gout pathogenesis. This raises the possibility that pathways enhancing fibronectin-dependent stabilization of the joint microenvironment could have protective effects.
By contrast, three proteins require more caution in interpretation. SPOCK2, which is involved in extracellular matrix turnover and has been studied in cartilage biology and tumor microenvironment remodeling,80,81 showed a directionally protective MR association in the discovery dataset and the same direction in the replication cohort, but without statistically significant replication. Colocalization support for SPOCK2 was also weak. These findings make SPOCK2 an interesting but lower-confidence candidate: it may participate in matrix remodeling and immune infiltration in inflamed synovium, but current genetic evidence is insufficient to prioritize it for therapeutic exploration. Similarly, FAM213A (also known as PAMM), which has been linked to redox regulation, macrophage activation, and osteoclast differentiation,82,83 showed a strong primary MR association and robust colocalization support, but the direction and magnitude of effect did not consistently reproduce in the independent gout GWAS. Additionally, CPQ emerged as another risk-associated protein. Although CPQ is less well characterized in gout biology than PLAU, carboxypeptidases can participate in peptide processing, extracellular proteolysis, and inflammatory signaling cascades. 84 Our MR and meta-analysis suggested that higher genetically proxied CPQ levels are associated with an increased risk of gout. Colocalization support for CPQ was weaker, indicating that further work is needed to determine whether CPQ itself is the mediator or whether CPQ levels are correlated with a nearby causal signal. This heterogeneity could reflect differences in gout case definition, statistical power, allele frequency, or local LD structure between Finnish and non-Finnish European samples. In light of these inconsistencies, we refrain from proposing FAM213A as a confirmed causal mediator of gout risk and instead consider it an exploratory signal that merits follow-up only after independent replication.
This study employed MR, a method that utilizes genetic variation present at conception to reduce bias from reverse causation, thereby ensuring that the findings are not influenced by disease status. This approach mitigates the impact of confounding factors by associating specific genetic variants with disease, as these variants are typically inherited independently of environmental influences. The robustness of our results is further reinforced through co-localization analysis, reverse causality analysis, external validation, and replication meta-analysis. However, this study does have certain limitations. First, MR analysis is constrained to a specific subset of proteins characterized by cis-indexed pQTL signals. Second, since the study sample predominantly comprises individuals of European ancestry, the generalizability of the results may be restricted for other ethnic groups. Third, the effect sizes identified through MR analyses were modest, reflecting lifetime genetic exposures rather than short-term clinical effects. Moreover, our MR-based findings do not define absolute plasma concentration thresholds or simulate the effects of therapeutic protein inactivation, and pQTL comparisons are further limited by tissue specificity and post-transcriptional regulation. Lastly, there is a deficiency of real-world research and basic experimental support; future studies should incorporate these elements to provide a more comprehensive understanding of our findings.
Conclusion
This exploratory and assumption-dependent Mendelian randomization study identifies five circulating proteins as hypothesis-generating candidates for gout risk and potential therapeutic targeting. Considering the inherent assumptions of MR and the heterogeneity across datasets, further functional validation and independent replication are essential to clarify their biological mechanisms, confirm targetability, and evaluate their translational potential.
Supplemental Material
sj-docx-1-iji-10.1177_03946320261428859 – Supplemental material for Mendelian randomization implicates circulating plasma proteins in gout risk and identifies candidate therapeutic targets
Supplemental material, sj-docx-1-iji-10.1177_03946320261428859 for Mendelian randomization implicates circulating plasma proteins in gout risk and identifies candidate therapeutic targets by Han Zhang, Zixian Dang, Pixue Yu, GuanHong Chen, Yingze Zhang, Tianrui Wang, Tengbo Yu, Yongtao Zhang and Zhen Shang in International Journal of Immunopathology and Pharmacology
Supplemental Material
sj-docx-2-iji-10.1177_03946320261428859 – Supplemental material for Mendelian randomization implicates circulating plasma proteins in gout risk and identifies candidate therapeutic targets
Supplemental material, sj-docx-2-iji-10.1177_03946320261428859 for Mendelian randomization implicates circulating plasma proteins in gout risk and identifies candidate therapeutic targets by Han Zhang, Zixian Dang, Pixue Yu, GuanHong Chen, Yingze Zhang, Tianrui Wang, Tengbo Yu, Yongtao Zhang and Zhen Shang in International Journal of Immunopathology and Pharmacology
Supplemental Material
sj-xlsx-3-iji-10.1177_03946320261428859 – Supplemental material for Mendelian randomization implicates circulating plasma proteins in gout risk and identifies candidate therapeutic targets
Supplemental material, sj-xlsx-3-iji-10.1177_03946320261428859 for Mendelian randomization implicates circulating plasma proteins in gout risk and identifies candidate therapeutic targets by Han Zhang, Zixian Dang, Pixue Yu, GuanHong Chen, Yingze Zhang, Tianrui Wang, Tengbo Yu, Yongtao Zhang and Zhen Shang in International Journal of Immunopathology and Pharmacology
Footnotes
Acknowledgements
This study was supported by the Shandong Provincial Natural Science Foundation, China [grant number ZR2021MH090, Yongtao Zhang] and the Qingdao Outstanding Health Professional Development Fund [Yongtao Zhang]. We gratefully acknowledge their financial support.
Abbreviations
Author contributions
Z.H.: Writing–original draft, review & editing, Conceptualization, Methodology and Project administration; D.Z.X. and Y.P.X.: Writing–review & editing, Conceptualization and Methodology; S.Z., Z.Y.Z. and C.G.H.: Writing–original draft, Conceptualization and Methodology; W.T.R. and Y.T.B.: Writing–review & editing and Formal Analysis; Z.Y.T. and S.Z.: Writing–review & editing.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding for this study was provided by the Shandong Provincial Natural Science Foundation, China [grant number ZR2021MH090, Yongtao Zhang]. Supported by Qingdao Outstanding Health Professional Development Fund [Yongtao Zhang].
Ethics approval and consent to participate
This study utilizes published data from prior research involving human participants. All relevant data were obtained with formal written approval and informed consent from the institutional ethics review board.
Consent for publication
All authors agree to publish in your journal and to abide by all your journal’s regulations without dispute and this data is all based on public databases.
Availability of data and materials
The original contributions presented in the study are included in the article and supplementary material, further inquiries can be directed to the corresponding authors.*
Supplemental material
Supplemental material for this article is available online.
