Abstract
Enzyme Che plays an essential role in cholinergic and non-cholinergic functions. It is present in the fertilized/unfertilized eggs and sperm of different species. Inclusion criteria for data collection from electronic databases NCBI and Google Scholar are enzyme AChE/BChE, cholinergic therapy, genomic organization and gene transcription, enzyme structure, biogenesis, transport, processing and localization, molecular signaling and biological function, polymorphism and influencing factors. Enzyme Che acts as a signaling receptor during hematopoiesis, protein adhesion, amyloid fiber formation, neurite outgrowth, bone development, and maturation, explaining the activity out of synaptic neurotransmission. Polymorphism in the Che genes correlates to various diseases and diverse drug responses. In particular, change accompanies cancer, neurodegenerative, and cardiovascular disease. Literature knowledge indicates the importance of Che inhibitors that influence biochemical and molecular pathways in disease treatment, genomic organization, gene transcription, structure, biogenesis, transport, processing, and localization of Che enzyme. Enzyme Che polymorphism changes indicate the possibility of efficient and new inhibitor drug target mechanisms in diverse research areas.
Summary statement
Important information regarding enzyme AChE/BChE and cholinergic therapy literature is retrieved. The goal was to provide a description, conspectus, and critical evaluation of the literature through the logical development of arguments, ideas, and reference range of enzyme ChE.
Introduction
Genomic organization and gene transcription
Cholinergic neurons catalyze the hydrolysis of the cholinergic neurotransmitter Acetylcholine. 1 Acetylcholinesterase (AChE) and butyrylcholinesterase (BChE) classification depends on substrate specificity and inhibition. 1 AChE and BChE belong to the cholinesterase enzyme family with a critical role in cholinergic and non-cholinergic signaling.1,2 AChE is responsible for terminating neurotransmission and synaptic activity, whereas BChE handles detoxification and exhibits broader substrate specificity. Both enzymes share structural similarities.1,2 AChE and BChE differ based on the enzyme structure, active site, efficiency, substrate, biochemical roles, and tissue distributions.1–3 However, both share a common ancestor as a member of the cholinesterase family containing conserved structural domains essential for their catalytic activity, reflecting their evolutionary relationship.4,5 AChE and BChE genes arise from a duplication event. 6 Potential therapeutics arise with knowledge about similarities and differences between cholinesterase (ChE). 1


BChE has a role in drug and toxin metabolism, detoxifying organophosphates (used in pesticides and nerve agents), and metabolizing drugs in the liver. BChE is a physiological buffer of AChE hydrolysis. When AChE is inhibited and less efficient, it has clinical implications. Variations in BChE activity affect an individual’s response to certain drugs. 7
BChE is located in the liver and circulates in the blood plasma, while AChE is predominantly in the synaptic clefts, neuromuscular junction, and red blood cells.1,8 The gene for BChE localizes on chromosome 3 (q26.1), while the AChE gene localizes on chromosome seven (q22.1), as shown in Figure 1.9–11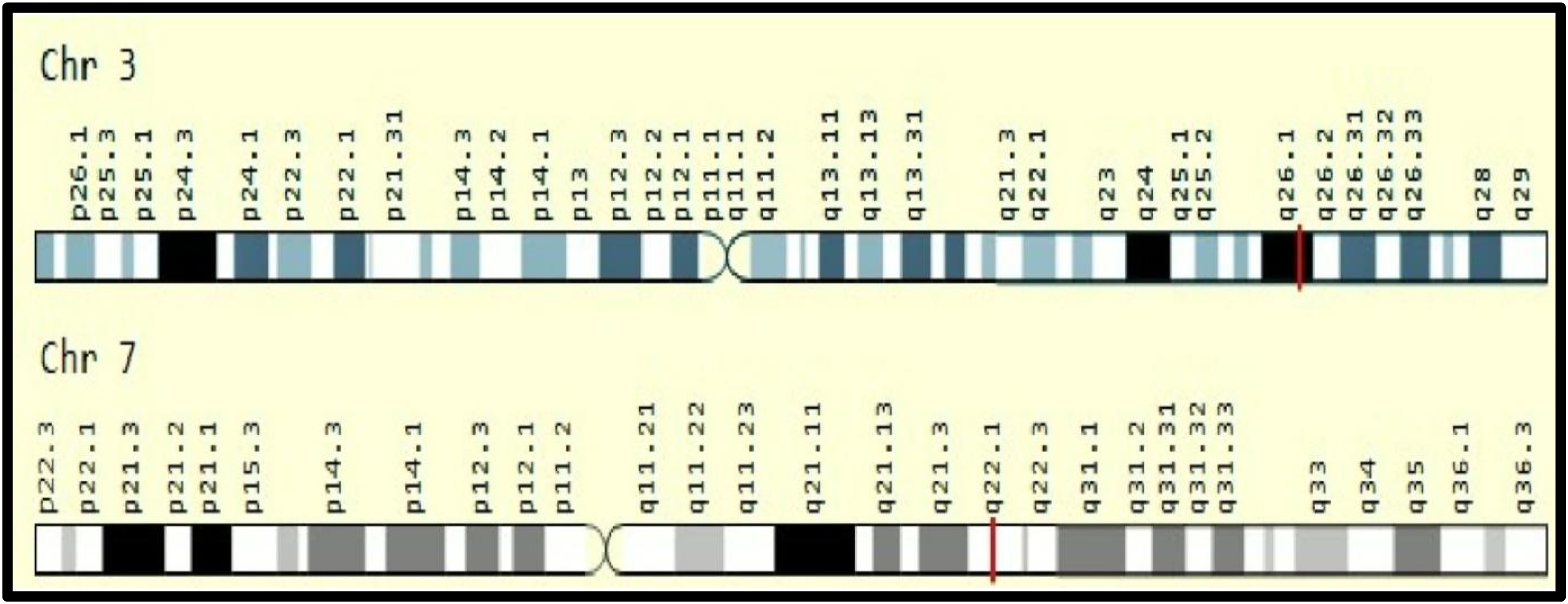
The genomic region of BChE spans around 70 kb and has four exons and three large introns, while the AChE gene spans seven kb and has six exons, Figure 2.12,13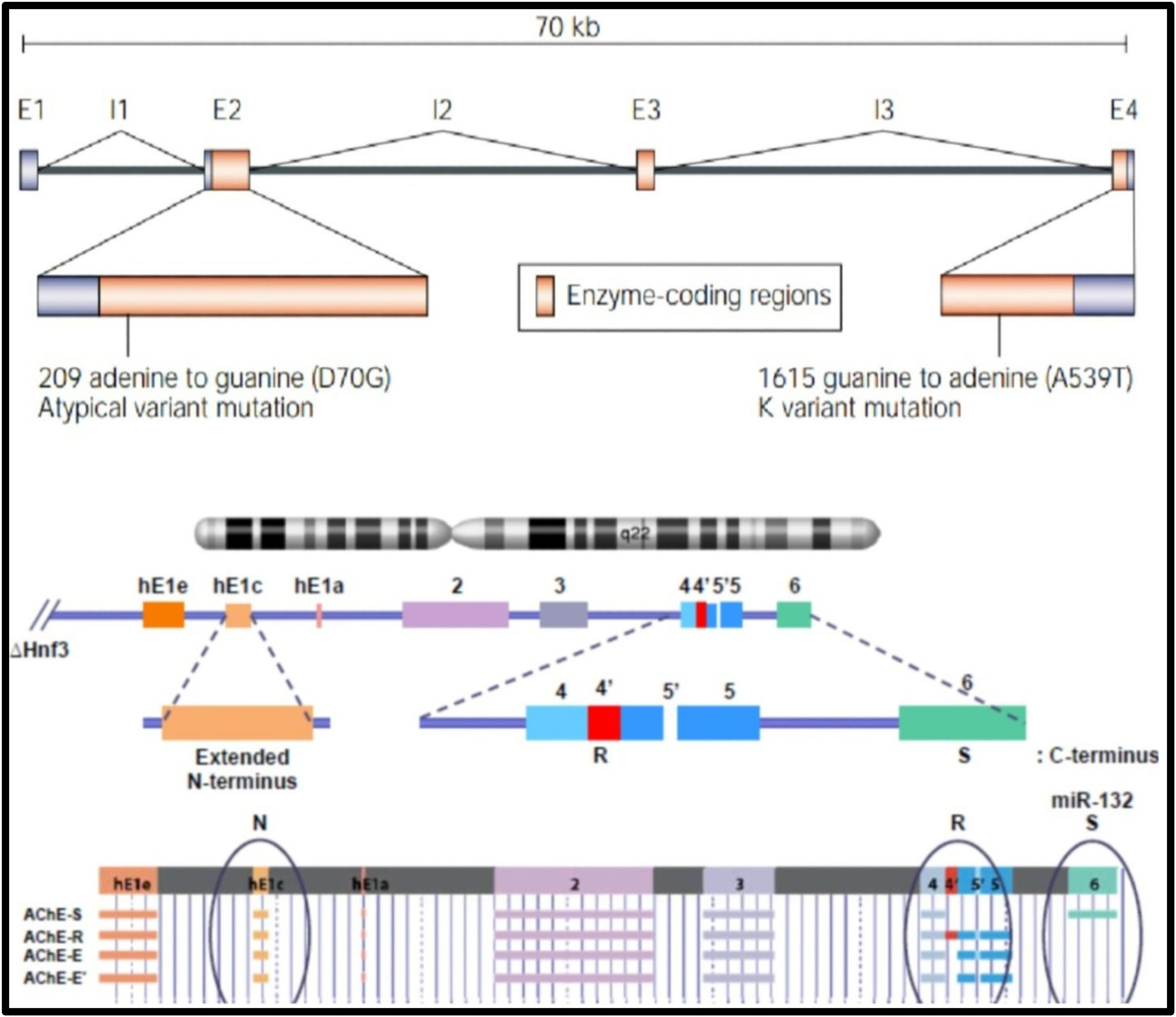
Generally speaking, more than 90% of transcripts undergo alternative RNA processing to generate about 100,000 different proteins from about 20,000 human genes. 14
BChE consists of different molecular forms called G1 (monomeric globular), G2 (dimeric), and G4 (tetrameric) with identical catalytic subunits that are symmetrical, hydrophilic, globular, and primary soluble forms. Globular BChE occurs in asymmetric, membrane-bound forms that are amphiphilic and consist of tetramers (G4) anchored to membranes by a protein, proline-rich membranes anchor (PRiMA).
12
Asymmetric collagen-tailed BChE contains tetramers of catalytic subunits attached to membranes by a triple helical, non-catalytic collagen anchor called A4, where two tetramers make up the A8 form and three A12 forms.
12
Heteromeric multimers assemble with amino-terminal proline-rich attachment, and collagen Q. BChE structure forms are available in Figure 3.
12
Molecular forms of BChE. G1-monomer soluble globular form. G2-dimer soluble globular form. G4-tetrameric soluble globular anchored form. A12-asymmetric form with three tetrameric forms. Retrieved from ref. 12.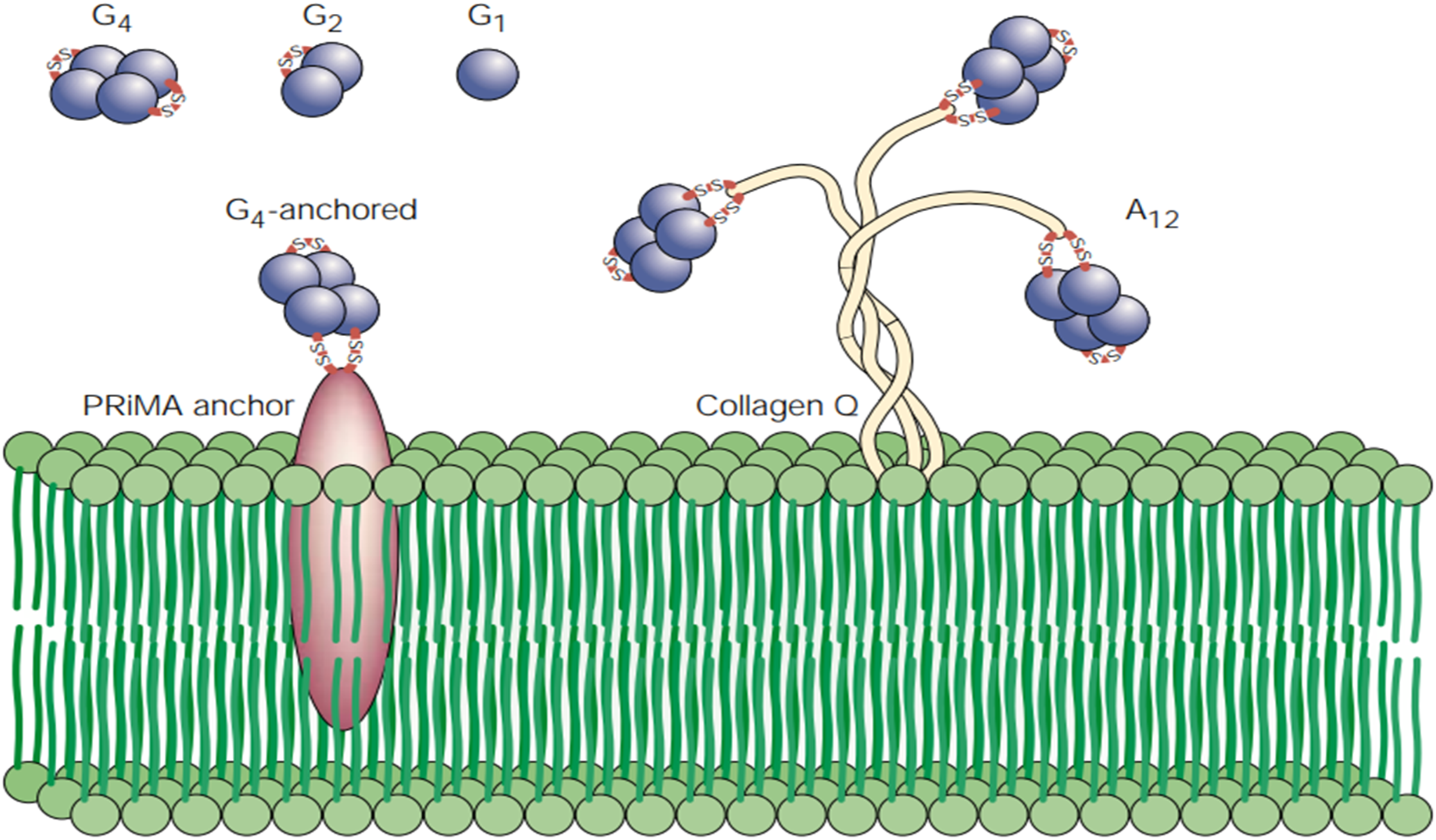
Acetylcholinesterase isoforms depend on alternative splicing and polyadenylation, Figure 4.
14
Molecular forms of AChE. AChET- tetrameric structure (red circles) that anchors to the PRiMA subunit (in brain) or trimeric CoLQ subunit (in muscle). AChEH isoform (green circles) - dimer that anchors to hematopoietic cell membrane through the GPI anchor. AChER isoform (gray circle) is rarely expressed and is a soluble monomer. PRiMA, proline-rich membrane anchor; CoLQ, collagen Q; MuSK, muscle-specific receptor tyrosine kinase; and GPI, glycophosphatidylinositol. Retrived from ref. 14.
The cap site at the 5’ end of the AChE gene allows the usage of promoters in different tissues. 15 The AChE gene has 6 exons. The exons include 3 invariant exons (exons 2, 3, and 4) encoding the catalytic domain nucleus and variable 5’ and 3’ regions, Figure 3.14,16 Exon 5 has two alternative splice sites (ss) 3’ ss: one at the boundary of intron 4. and exon 5a (proximal 3’ ss), and the other at the end of exon 5a and exon 5b (distal 3’ ss).14,16 The catalyst units produced differ based on the carboxyl end. 10 Alternative 3’ end splicing assembles three AChE (T, H, R) isoforms. 14 However, apart from this form, many others exist. The joining of exons 4 and 5a results in the creation of a hydrophobic dimeric isoform, AChE-H. Isoform is expressed in hematopoietic cells anchored to the membrane by glycophosphatidylinositol. 14 The fusion of exons 4 and 5b generates a tetrameric isoform, AChE-T, which is expressed in muscle, and brain, and anchored to a proline-rich membrane (PRIMA) or a trimeric collagen K tail. 14 The AChE-R monomeric isoform occurs when the transcript non-excretes after exon 4. The AChE-R isoform is rare and regulated in acute stress in the mouse brain. 14 Tissue-specific alternative polyadenylation site (PAS) is 21.2 kb downstream of canonical PAS in mouse muscles and brains. 17 Splicing factors SRSF1 (SF2 / ASF) and SRSF2 (SC-35) ubiquitous express and influence upstream and downstream expression regulation of AChE-T to AChE-R. 14
The soluble form is prevalent in plasma, while the membrane forms in muscles and the brain. 18 So far, more than 70 natural mutations are present in the BChE gene, which affects catalytic function and protein expression. 19 Leukaemias, megakaryocytopoiesis, breast cancer cells, and insects contain natural amplification of the cholinesterase gene. 20
SNPs in the non-coding 5′-untranslated region (5′-UTR) (rs1126680) and intron 2 (rs55781031) of the BCHE gene may play roles in the emergence and progression of AD and PD. 21 The most common genetic variant in the BChE gene is BChE-K substitution (c.1699G>A, p.A539T, rs1803274) associated with a lower level of BChE molecules in PL with decreased activity caused by the impaired quaternary organization of the tetramer. 22
Cholinesterase inhibitors are a significant focus for drug development, especially for conditions involving cognitive decline and memory impairment. 23 They offer a pathway for designing therapeutic agents to manage or potentially reverse symptoms. 23 Variations in Che genes (such as those coding for AChE and BChE) can influence individual responses to drugs and diseases. 24 Studying these polymorphisms aids in personalizing treatments and deepening our understanding of disease mechanisms. 24 The reason for choosing the Che enzyme is the role of Che, especially AChE, in AD, which makes it a critical target for research. Studying Che can lead to the development of better therapeutic strategies for managing cognitive disorders.25–27 Molecular mechanisms enable insight into neuronal signaling and pathology of physiological processes.25–27 Enzyme is a potential drug target for disease treatment and managing toxicity, making them relevant in pharmacology and medical chemistry research.25–27 Investigation of polymorphisms reveals how genetic differences impact drug efficacy, disease development, and therapy, making it significant for environmental and occupational health studies.25–27 Based on the abovementioned information, we can conclude that the Che enzyme is a noteworthy molecule in neurotransmission, physiological processes, disease, and treatment, with multifaceted functions impacting health outcomes. We chose Che for this study due to its well-established role in neurodegenerative diseases and its potential as a therapeutic target. By examining the ChE structure, function, and inhibition, this research aims to support the creation of new therapeutic approaches for diseases like Alzheimer’s, where cholinergic dysfunction is a fundamental aspect of the disease’s pathology. Additionally, natural compounds that inhibit Che offer a promising avenue for developing safer and more effective treatments.
Structure of enzyme ChEs
AChE classifies as a serine hydrolase/esterase with an ellipsoidal molecular structure featuring a central β-sheet and lateral α-helices.
17
In the narrow and deep depression, five regions involved in substrate and inhibitor binding (Torpedo Californica) have 5 different traits: (1) Catalytic triad residues: amino acids Serine 200, Histidine 440, and Glutamin 327, at the bottom of the gorge, which directly participates in the catalytic cycle.
28
(2) The oxyanion cavity: stabilizes a transient tetrahedral enzyme-substrate complex by negatively charged carbonyl oxygen acetylcholine (ACh).
28
(3) The anionic site, where Tryptophan 84 locates. All cholinesterases conserve Tryptophan 84 active in the orientation of the trimethylammonium group.
28
(4) An acyl pocket containing two phenylalanine residues at positions 288 and 290, which interact with a substrate acyl group
28
; (5) A peripheral anionic region: containing residues located at the perimeter of the gorge of the active site of the amino acids Tyrosine 70, Tyrosine 121, Tryptophan 279, and Asparagine 72.
28
The AChE enzyme is highly conserved because it possesses an 88% identical sequence (human and mouse) and a crystal structure (human, mouse, Torpedo Californica, and Drosophila) between different model systems.
29
The BChE belongs to the α/β-fold family of proteins. The enzyme catalyzes the hydrolysis of esters such as cocaine, acetylsalicylic acid, and heroin. It is significant in scavenging occurring (physostigmine) and synthetic (organophosphate) anticholinesterases.
12
Each human catalytic BChE subunit consists of 574 amino acid residues. The catalytic triad, set at the bottom of a 20 Å-deep gorge, includes serine 226, histidine 466, and glutamic acid 353, surrounded by six amino acids. In contrast, AChE features 14 amino acids in a similar region (Figure 1). Within the acyl pocket (A), the substrate’s acyl group reside, while tryptophan (W) creates an anionic site that facilitates interaction between the quaternary nitrogen and the anionic site. Substrates interact with aspartic acid (D) and tyrosine (Y) residues found at the lip of the active site gorge, Figure 5.
12
The active site of BChE is at the bottom of a 20 Å gorge. Serine (S), histidine (H), and glutamic acid (E), acyl pocket (A), tryptophan (W), aspartic acid (D), tyrosine (Y). Retrieved from ref. 12.
The anionic site interacts with the choline nitrogen through tryptophan. Positively charged substrates interact with aspartic acid 70 and tyrosine 332. During catalysis, the acyl pocket holds the acyl group of choline esters in a place where lysine 286 and valine 288 line this pocket. 12 BChE and AChE are glycoproteins where complex carbohydrates are covalently attached to human AChE at three amino acid residues, whereas BuChE at nine asparagines. 12 BChE has additional enzymatic activity, distinct from its esterase activity since it has an aryl acylamidase activity, catalyzing the hydrolysis of acyl amides of aromatic amines, and therefore is involved in functions other than cholinergic co-regulation. 12
The structure of Che changes when bound to a compound, differing it from its unbound form and impacting function and interactions.30,31 Differences like this enable an understanding of how inhibitors function and how they can be optimized for future therapeutic purposes.30,31 Changes in active site conformation depend on the Che structural difference.30,31 Upon binding a compound, the active site of Che undergoes induced fit conformational changes to accommodate the ligand shifting the key residues affecting the active site pocket, where compound binding affects enzyme conformation and interaction, enhancing the binding affinity of the compound.30,31 Binding can lead to the formation or disruption of hydrogen bonds between the enzyme and the compound. The bonds stabilize the compound within the binding site, changing overall conformation.30,31 The charge distribution in the active site can change upon binding, leading to altered electrostatic interactions.30,31 The compound may introduce new ionic interactions or modify existing ones, affecting enzyme stability and activity.30,31 Bounding compounds may cause enzyme flexibility of ligand interaction, changing enzyme structure.30,31 For example, in its free form, the active site of Che is more open and flexible.30,31 When an inhibitor binds, the enzyme reduces access to other molecules, enhancing the specificity of the inhibitor.30,31 Enzymes with multiple domains may exhibit shifts or rotations of domains upon compound binding. 32 These movements can change the relative positioning of different parts of the enzyme and influence its functional state. 32 Loops and helices near the binding site may adjust their positions to better interact with the compound, affecting form, dynamic, and complex stability. 32 In some cases, compound binding may induce allosteric changes in the enzyme, affecting regions distant from the active site and resulting in global conformational changes that impact the function. 32
Dynamic studies using molecular dynamics simulations can reveal how the structure of Che evolves when bound to a compound.33,34 These simulations can show how the conformation fluctuates and adapts during the binding process.33,34 Enzyme catalytic activity can be enhanced or inhibited by binding a compound to a catalytic triad that is significant for hydrolytic activity. 35 The structural changes upon compound binding can influence the binding affinity and specificity.36,37 Higher affinity binding often correlates with more extensive structural rearrangements and tighter interactions. Structural differences can also impact the selectivity of different compounds.36,37 Variations in binding modes and interactions can determine how well the enzyme recognizes and binds specific inhibitors or substrates.36,37 Interaction with the inhibitor is dependent on the structural difference between Che enzymes. AChE features a deep active site gorge where the compound binds.38,39 The structure of the gorge and the interactions with the compound can significantly affect enzyme activity.38,39 AChE has distinct esteratic and anionic sites.38,39 The binding of a compound can lead to changes in how these sites interact with the ligand, influencing the overall catalytic activity.38,39 BChE typically has an active site compared to AChE.40,41 The binding of compounds may involve different interactions due to this structural difference, affecting how the enzyme processes substrates and inhibitors.40,41 BChE also has a peripheral binding site that can influence the activity and interaction with compounds, leading to structural changes in the enzyme upon binding.40,41 The binding of a compound to Che can induce significant structural changes, including conformational shifts, alterations in side-chain positions, and modifications in binding site dynamics. These changes are crucial for understanding enzyme function, drug design, and the development of therapeutic agents targeting ChE, especially in neurodegenerative diseases.
Simulation of atomic and molecular physical movements over time, providing detailed insight into the dynamic behavior of macromolecules, is achieved by solving Newton’s equations of motion using the molecular dynamics (MD) computational method.42,43 The potential energy associated with the system depends on mathematical description and force field parameters like length, angles, torsions, and non-bounded interactions covering a time scale from femtoseconds (10^–15 s) to microseconds (10^–6 s). 44 Protein folding, conformational changes, and protein-ligand interactions are analyzed with MD using static data obtained from X-ray crystallography and NMR spectroscopy. 45 Advanced MD methods enable the exploration of rare events to overcome the energy barriers of conventional methodology. 45 These techniques enhance the exploration of conformational space and improve the accuracy of predictions. 45 MD simulations integrate with experimental data to validate computational models and refine structural predictions, enabling an understanding of biological processes and designing potential therapists.46,47 MD is a powerful tool for understanding molecular systems, which is significant and critical for drug discovery, structural biology, and biochemistry research.46,47
Biogenesis, transport, processing and localisation of ChE enzymes
All forms of AChE are glycoproteins that can secrete or membrane-bound undergo similar secretory pathways.
15
Secretory pathways include synthesis and molecule binding, transport and secretion, localization and binding, traffic, and degradation.
15
Enzyme AChE synthesizes on a granular endoplasmic reticulum and undergoes co-translational glycosylation, adding asparagine-linked oligosaccharides to a growing polypeptide chain.
15
Enzyme AChE passes bending, assembling, and quality control before leaving the endoplasmic reticulum.
15
The Golgi apparatus translocates for further processing.
15
In the case of a collagen-bound form, coupling with a non-catalytic subunit is performed.
15
The binding of the PRIMA (proline-rich binding site) subunit was associated with the sensitivity of PRIMA-bound G4 AChE to endoglycosidase H.
15
The oligomeric form of AChE assembles and relocates to the appropriate cellular domain.
15
Molecular chaperones increase the activity of AChE in the granulated endoplasmatic reticulum, while the clathrin transport mechanism enables packing in the Golgi apparatus.
15
The main steps of biogenesis, transport, processing, and localization of AChE enzymes are present in Figure 6. Biogenesis, transport, processing and localisation of AChE enzymes.
13
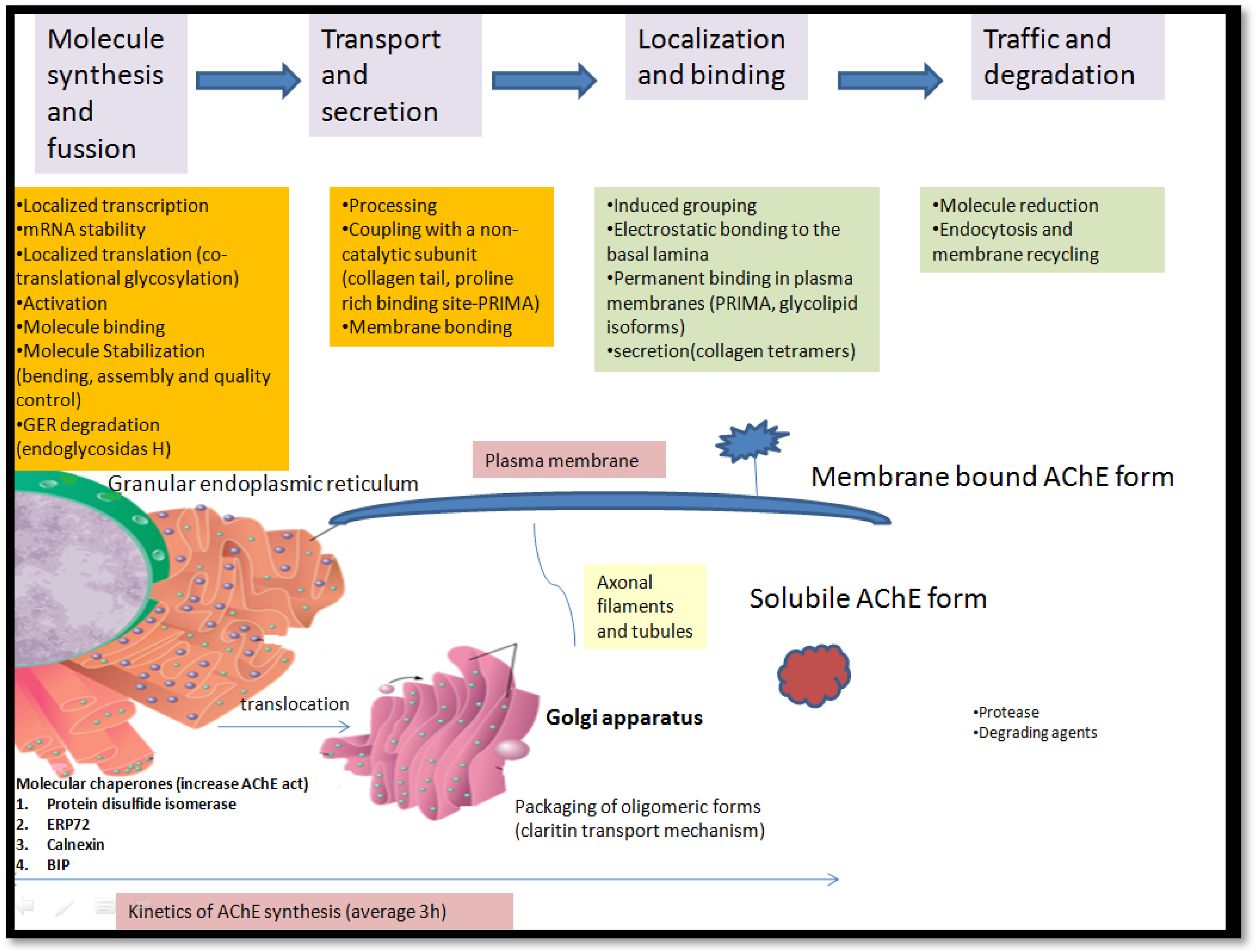
The definition of the physiological role of the BChE is missing since it doesn’t have a single physiological substrate. Substrates interact with various endogenous substrates and xenobiotics like ACh, serving as a backup in ACh hydrolysis when inhibiting AChE. 48
Molecular signalization and biological function
Cellular processes are affected by a complex network of biochemical and molecular reactions influencing gene expression through signaling pathways, receptors, signal transduction, effector molecules, and secondary messenger. 49 AChE and BChE affect gene expression with various molecules within these particular signaling pathways. AChE has diverse roles and applications. For example, AChE has catalytic and non-catalytic functions leading to synaptic and nonsynaptic hydrolysis, terminating synaptic transmission, and expressing developmental and regenerative roles.50,51 AChE is a potential disease biomarker, with increased expression during physiological stress, and is involved in cellular processes activating pathways like PI3K, MAPK, WNT/β-catenin, NF-κB, and many others involved in cell proliferation, differentiation, survival, and immune response.29,52–58
MAPK/ERK signaling pathway is activated by receptor tyrosine kinases (RTKs) and G-protein coupled receptors (GPCRs), leading to sequential phosphorylation of MAP kinase kinases. 29 The WNT/β-catenin pathway enables binding to Frizzled receptors and LRP5/6 co-receptors, while in the PI3K/Akt/mTOR pathway, PI3K activation leads to the generation of phosphatidylinositol-3,4,5-triphosphate (PIP3), recruits Akt to the cell membrane and activating mTOR.29,52 NF-κB Signaling Pathway releases NF-κB dimers (p50/p65) from IκB inhibitors and causes translocation into the nucleus. 57 These signaling pathways activate specific gene expression, leading to cell response.29,52–58 The communication between pathways exists with negative and positive feedback, leading to the coordination of multiple signals. AChE interacts with diverse molecules within biochemical and molecular interactions like microRNA (miRNA), erythropoietin, and miRNA-132, leading to inflammatory response and gene expression.29,52–58
AChE inactivates mitogen-activated protein kinase (MAPK) and PI3K/Akt pathways and activates GSK3ß in hepatocellular carcinoma (HCC) cells. 29 Moreover, AChE can activate the JNK pathway and contribute to the activation of Apaf1, cytochrome c release, and Caspase 9. 29 There are also indirect interactions with MAPK/PI3K pathways and potential involvement with the WNT/β-catenin signaling pathway. 52
AChE downstream regulates microRNA-124 and modulates macrophage activation by inhibiting the STAT3 signaling pathway in patients with bowel failure. 53 Downstream regulation of erythropoietin regulates the expression of the AChE dimeric form during erythroblast differentiation, affecting protein function through the interaction of AChE with the erythropoietic receptor. 54 Erythropoietin regulates the expression of the dimeric form of AChE during erythroblast differentiation. 55 Based on analyses using online tools and a luciferase reporter assay, miR-132 protects Ach from degradation by ACHE, where miR-132 exerts favorable effects on CNS neurons via brain-derived neurotrophic factors (BDNF). 56 Regarding inflammation and Gene Expression contributing to cholinergic function and disease is inhibition of AChE, which leads to the attenuation of retinal inflammation via NF-κB suppression, and upstream regulation leads to Early Growth Response 1 (ERG1) affecting transcription.57,58
BChE has a non-specific function and influences nervous system development due to structural similarities between AChE and BChE and cell adhesion molecules with the possible non-catalytic role. The enzyme BChE is a serine hydrolase. It catalyzes the hydrolysis of choline esters like butyrylcholine, succinylcholine, and acetylcholine. Hydrolysis of acetylcholine is less efficient than AChE.
12
BChE influences PKC/ERK pathways in R28 cancer cells, immune response pathways, transforming growth factor-β (TGF-β) signaling, and cancer immunotherapy via PD1 blockade pathways.59,60 BChE expression positively correlates with CD28, ADORA2A, BTNL2, and TNFRSF18.
60
BChE expression is under regulation by iron through Iron Response Elements (IREs) in the 3′-UTR, where under low iron conditions, IRP1 and IRP2 bind to the IREs, stabilizing BChE mRNA, preventing degradation, and increasing BChE levels.
61
A positive correlation exists between BChE activity and red blood cell (RBC) count, Figure 7.
61
Erythropoietin (EPO) synthesis increases through iron-regulated transcriptional activation.
61
When iron levels rise, HIF-2α mediates EPO gene transcription, indirectly regulated by IRP interactions with IREs in BChE mRNA.
61
The lifetime of RBCs is about 100–120 days. The aging RBCs undergo eryptosis and programmed death.
61
Much of the breakdown products recirculate by the spleen and the liver. The iron is released into the plasma to be recirculated by protein transferrin (Tf).
61
Thus, the recirculated Tf-bound iron (TBI) is proportional to the RBC count. Transferrin receptor protein 1 regulates serum BChE biosynthesis by IRPs in the liver.
61
EPO synthesis is triggered by the transcriptional activation induced by iron, leading to an upregulation of erythropoiesis.
61
EPO gene transcriptional activation mediates by binding HIF-2α to the hypoxia response element.
61
As iron levels rise, IRP1 cannot bind to the IRE sequence at the 5′-UTR of HIF-2α. This results in an increase in EPO due to the extended translation of HIF-2α. Based on this information, levels of BChE and RBC represent the cellular iron concentration or TBI.
61
Proposed model of BChE expression regulation by iron in the activated glial cells. Retrived from ref. 61.
The putative IRE occurs in the 3’-UTR of BChE and the 5’-UTR. 38 The consensus IRP binding sequence resides in the 5’-UTR region. 61 Under high iron cell concentrations, the IRPs cannot bind the IREs. IRPs bind to IRE at the 5’-UTR of mRNA and inhibit translation inhibition by preventing ribosome binding to the mRNA. 61 Binding IRPs to IREs at the 3’-UTR prevents mRNA degradation and endonucleolytic cleavage by decreasing mRNA levels. 61 Based on this information, BChE expression is positively regulated by iron in the 5’-UTR IRE element and 3-UTR by one stem loop with additional regulator factors as a negative feedback mechanism. More testing will be needed to validate this hypothesis. 61
Disease mechanism research depends on a choice model system (in vivo, in vitro) to validate disease signaling biomarkers. Bioinformatics and Systems Biology enable modeling and simulation of signaling networks, analyze high-throughput data, and predict pathway interactions and dynamics in a cost-effective, fast, and precise way. Clinical application of AChE and BChE inhibitors arise from a multidisciplinary scientific approach, combining knowledge.
Polymorphisms
AChE enzyme polymorphism is present in the eggs and sperm of different species.62,63 AChE acts as a signaling receptor during hematopoiesis, protein adhesion, amyloid fiber formation, neurite outgrowth, bone development, and maturation.62,63 Roles from the above explain the nonsynaptic hydrolysis of the AChE enzyme. So far, to present knowledge, 35 gene variations of clinical significance (pathogenic, benign, and unknown) exist. The list provided by the National Library of Medicine and the National Center for Biotechnology Information is available in Supplemental File 1. By 2005, an accurate estimate of the genetic polymorphism (Single Nucleotide Polymorphism_SNP) of the AChE gene in the healthy population was called into question. 64 There was a limitation of ethnic groups, a small number of respondents, and incomplete coverage of the AChE coding region. 64 There were 18 SNPs on the list, of which 13 lacked biological validation of population frequency. 64 Some SNPs are harmful due to amino acid substitution. 64 Clinically significant polymorphisms include the distal portion of the AChE promoter and the His 322 substitution. 64 Polymorphism of the distal part of the AChE promoter (substitution T> A) interrupts or interferes with the glucocorticoid response. 64 It is associated with acute susceptibility to anticholinesterase agents (pesticides). 64 Essential for the development of Gulf War syndrome. 64 Substitution of the amino acid His 322 with Asp is responsible for the YT-2 phenotype of the blood group, which is a significant factor for matching donor and recipient blood groups. 64 CDNA change: c. 1057 C>A in protein (Histidine 353 with Asparafine) does not affect function. 64 In 2005, researchers identified 13 SNPs in the AChE gene within various ethnic groups and explored their possible effects on the structure and function of AChE. 64 Moreover, amino acid substitutions exist (Arginine 34Glicine, Glycine57Arginin, Glutamine 344Glycine, Histidin353Asparagin, Proline 592Arginine). 64 The basis for the functional and pharmacogenetics of natural AChE protein variants has been established. 64 The detected SNPs assess the involvement of occurring AChE polymorphisms during the disease mechanism and different drug responses. 64 AChE HN353N protein polymorphism is associated with a low prognosis of chronic Chaga disease. 65 The intron point mutation AChE rs2571598 has C / T substitution. 63 It affects enzyme activity and therapeutic response in patients with Alzheimer’s disease (AD) and increases AChE activity in patients with multiple sclerosis. 63 A variant of the rs17228602 enzyme AChE is associated with drug dependence, such as hashish and heroin. 66 Polymorphism rs17228616 in the AChE gene enables overcoming the post-traumatic stress symptom correlated with the co-intensified emotional response of the amygdala and ventromedial prefrontal cortex. 67 This SNP disrupts the suppression of AChE miRNA-608, increases AChE in the brain, and decreases cortisol and a miRNA-608 targeted GABAergic modulator CDC42, which is associated with stress in soldiers. 67 A correlation exists between miRNA-608, CD44, CDC42, and interleukin 6 expression levels in the human amygdala. 67 The role of synaptic AChE in β-amyloid deposition and aggregation in individuals with AD has been demonstrated. 68 Prolonged exposure to electromagnetic waves (Wi-Fi) changes gene expression, behavioral patterns, and neurodegenerative diseases. 69 Primor-specific recognition of polymorphism of the primate-specific miRNA-608 element in the 3’ untranslated region of the AChE transcript increases the level of AChE in the brain during anxious and inflammatory reactions accompanied by disturbances in the regulatory network of non-coding RNA (CholinomiRs-608). 70 The genetic polymorphism of the nicotinic cholinergic receptor α7 (CHRNA7) and CHRNA7 (CHRFAM7A) genes is a possible susceptibility trait for dementia. 71
The enzyme AChE is a target for chemical warfare. 72 The enzyme AChE applies organophosphate, carbamate, food poisoning, disease diagnosis, therapy, and treatment.72,73 The enzyme AChE is a marker of the aging process. 74
Mutations in BChE produce a range of enzyme catalytic activity. The wild-type BuChE (BuChE-WT) is sensitive to dibucaine inhibition. Atypical BChE variant with a mutation at codon 70 causes the substitution of aspartic acid for a glycine residue (D70G) which leads to insensitive inhibition by dibucaine, where some show apnoea after administration of succinylcholine or ingestion of the cholinesterase inhibitor pyridostigmine including severe depression, insomnia and profound weight loss, pointing to effects of the mutation in both the central and the peripheral nervous systems. 12 K variant mutation has a mutation at codon 539 that leads to the replacement of an alanine by a threonine residue (A539T)76 and has about 33% lower enzymatic activity relative to BuChEWT. J mutation variant results from a mutation at codon 497, which changes a glutamic acid residue to valine (E497V)78, leading to 66% lower enzymatic activity about BuChE-WT. H variant has a valine residue at codon 142 replaced by a methionine (V142M), resulting in a protein with 90% lower activity than the wild-type enzyme. 12 Fluoride inhibits BuChE-WT, and two fluoride-resistant forms exist. At first, threonine at codon 243 changes to methionine (T243M). Secondly, glycine at codon 390 shifts by valine (G390V). Finally, there is the Silent variant, where there are at least 12 additional mutant forms of BuChE with no known activity, and they can’t catalyze the hydrolysis of choline esters.
Allelic frequencies of the W, K, and atypical forms of BuChE reports being about 85%, 10%, and 13%, with some people carrying double mutations (K and A), whereas other BuChE mutations are present in much lower frequencies. The reliability of these frequency estimates has to be confirmed, as differences in allelic frequencies are present within various ethnic populations. 12 On the other hand, the allelic frequency of the atypical variant is higher in populations from areas where plants of the Solanaceae family are Indigenous, favoring the natural selection of people resistant to poisoning through diet. The functional consequences of most of these variants are not well known. Neurodegenerative disease relates to the K variant. 12
Cholinergic therapy with enzyme ChE inhibitors
Brief overview of the natural origin and the compounds selected for enzyme ChE inhibitors.

Organizes the information regarding natural compounds with ChE inhibitory properties, their sources, and effects/uses relevant to neuroprotection and cognitive enhancement.
Natural compounds in Alzheimer’s disease treatment.

Mechanisms of action summary.

This mechanism enables natural sources to reduce oxidative damage and neuroinflammation, protecting neurons from apoptosis, excitotoxicity, and other forms of damage, thereby preserving cognitive function.85,86
Clinical trials and applications.

Tables 1–4 captures the natural products, their sources, properties, mechanisms of action, and some clinical trials or applications related to AD treatment.
Enzyme AChE is a target biological molecule for the diagnostics, prognosis, and treatment of various diseases. 29 Enzyme AChE is a visible candidate gene for pharmacogenetic studies. 29 Potential therapeutics for Alzheimer’s disease(AD) and Myasthenia gravis (MG) are AChE inhibitors, which can be natural or synthetic, leading to the enhancement of cholinergic transmission.1,87 Physostigmine (Eserine) is a natural reversible AChE inhibitor derived from the Calabar bean, used to treat glaucoma and AD due to its ability to increase acetylcholine (ACh) levels in the brain. 88 Synthethic inhibitor Tacrine is eradicated in AD therapy due to its hepatotoxicity. 88 A widely used synthetic, reversible, non-competitive AChE inhibitor, known for its effectiveness in improving cognition and global function in AD, is Donepezil and Galantamine, while for treatment MG is Pyridostigmine affecting APP processing and Aβ levels. 88 Specific reversible inhibitors like BW284c51, GUK-987, and GDK-510 are often used in basic research studies of the cholinergic system.89,90 Pseudo pseudo-irreversible inhibitor targeting AChE and BChE is Rivastigmine. 88 Reversible and irreversible inhibitors increase the level of ACh and improve neurotransmission, cognitive function, and synaptic plasticity associated with mutations in Amyloid Precursor Protein (APP) and Presenilins 1 and 2 (PSEN1 and PSEN2 ) genes.88,91 These inhibitors serve as a research tool for studying the cholinergic system, potentially investing in ChE inhibition, APP processing, and Aβ production.87–91 Depending on the site of action, they may be reversible inhibitors (therapeutics) or irreversible inhibitors (toxins). 87 Reversible inhibitors can be competitive or non-competitive based on their binding characteristics. 92 The competitive inhibitor binds to the enzyme and blocks substrate binding. 92 The non-competitive inhibitor binds to another site on the enzyme, impairing the further normal reaction of the enzyme, although the substrate binds to its active site. 92 Over 59 plant species contain natural inhibitors of AChE activity. 93 The most commonly used synthetic AChE inhibitors in the literature are pyridostigmine and BW284C51.87,94
Inhibitor Eserin (synonym: physostigmine, physostol) is a reversible AChE inhibitor that applies to eliminate the side effects of anticholinesterase drugs (bromazepam, barbiturate, antihistamine) and poisoning (plants Tatula-Datura stramonium and Velebilje-Atropa belladonna).1,95 It belongs to the drugs for nervous system disorders.1,95 Some of the completed clinical studies of physostigmine inhibitors include phase 4 with the treatment of adolescent antimuscarinic Toxidrome (CTID: NCT03090620), phase 3 with opioid analgesia (CTID: NCT01394445), and phase 2 as an analgesic treatment after hysterectomy (EudraCT: 2006-006839-51)].96–99 A virtual search of the Eserin database indicated new variants of inhibitors that could improve the development of new AChE inhibitors. 100 The available literature confirms that Eserin interacts with the anionic/eserine portions of the AChE active site. 89 Eserin may function as a competitive inhibitor when in solution or as a non-competitive inhibitor when bound to membranes, depending on its concentration. 94 Competitive inhibitors bind to the active site and are non-competitive to the allosteric site. 100
BW284C51 (1,5-bis (4-allyldimethylaminoniumphenyl)) pentane-3-one dibromide inhibitor, eserine analog or 4- (5- {4- [Dimethyl (prop-2-enyl) ammonio] phenyl} -3-oxopentyl) - N, N-dimethyl-N-prop-2-enylbenzenaminium) is a reversible AChE inhibitor, a nicotinic receptor antagonist. 89 Dysfunction of the nicotinic receptor is associated with neurodegenerative diseases like Alzheimer’s and Parkinson’s. 100 The BW284C51 inhibitor applies in the phase of pre-clinical tests at room temperature.101,102 The available literature indicates that BW284C51 is a selective and non-competitive AChE inhibitor.101,102
The GUK-987 inhibitor is a reversible BChE inhibitor used to treat cancer and neurodegenerative diseases. 90 GUK-guanylate kinase genes convert drugs for the treatment of cancer and viral infections. Drugs transform into pharmacologically active metabolites (acyclovir) and are bound to the analog 6-mercapto purine. 103 Preliminary pre-clinical data on cell lines related to the GUK-987 inhibitor exist. 101 No information is available in the literature on whether the GUK-987 inhibitor is competitive or non-competitive.
Information on the GDK-510 inhibitor is not publicly available. The literature on whether the GDK-510 inhibitor is competitive or non-competitive is lacking. The most used AChE inhibitors are tacrine, donepezil, galantamine, and rivastigmine.
104
The reversible inhibitor binds to the anionic or peripheral anionic site of the AChE enzyme.
28
So-called dimeric (double) inhibitors bind simultaneously, combining two biological activities in one molecular entity.
28
The correlation of pharmacological properties with the 3D description of drug molecules and identification of drug characteristics contribute to its potency and represent a problem in drug design.
28
Independent molecule overlap analysis (3D Q-SAR-Quantitative Structure-Activity Relationship) on 110 structurally distinct dual AChE inhibitors showed that the minimum energy conformation, the co-crystallizing ligand conformation, and the anchored conformation have good results and predictability.
28
The model revealed the significance of the protonated pyridine nitrogen of the tacrine part of anti-AChE activity and recognized the interaction of donors and hydrogen acceptors as extremely important for potency.
28
A molecular interaction field involving the AChE enzyme active site gorge was defined using probes that mimic structural elements of a ligand in a data set.
28
The molecular interaction in the active site and predictive power are present in Figure 8. Schematic representations of calculated molecular interaction sites for the active site of the AChE enzyme using probes: (a) probe that mimics protonated pyridinium, (b) dry probe that mimics aromatic moieties of tarin inhibitors and similar inhibitors, (c) probe that mimics amino group linkers, (d) probe that resembles an alkoxy substituent of the benzene ring compound 68–79, 97–110. Retrieved from ref. 28.
Numerous patients treated with these or similar drugs have side effects, limiting the maximum administered dose. Inhibitors containing nitrogen and sulfur atoms are employed across various industries to inhibit AChE and BChE. 105 Compounds containing nitrogen and sulfur atoms are applicable for AChE and BChE inhibition. 106 For homeostatic cholinergic system functioning, immune cells like T and B lymphocytes, macrophages, and dendritic cells are necessary. 107 Knowledge about the cholinergic system and immune cells applies to drug design and disease therapy. 107 Such an analysis would enable the association of the natural AChE variant with a predisposition to the disease. Based on the above notions, cholinesterase inhibitors increase and prolong the availability of acetylcholine, improving cognitive processes. 12
It is significant in inhibiting BChE in neurodegenerative disease treatment. 12 BChE inhibitors are usually specific to BChE, like tetrahydroaminoacridine (THA), an early AD stage nonselective and late AD stage selective cholinesterase inhibitor like Rivastigmine, Cymserine, and analog with action on AChE and BChE. 12
In clinical practice for MG and AD therapy, nonselective inhibitors (Tacrine, Donepezil, Rivastigmine), selective irreversible inhibitors (Ethyl 4-nitrophenyl ethyl phosphonate, Haloxon), and reversible inhibitors (Neostigmine Bromide) are retrieved due to its neurotoxicity. 12 However, many others (Carbonic Anhydrase, Bentamapimod, and Turofexorate Isopropyl target affecting) apply in clinical practice with neuroprotective effects targeting neuroinflammation, oxidative stress, and metabolic regulation preventing the accumulation of Aβ plaque due to APP, Aβ42 peptide due to nonfunctional γ-secretase complex, which processes APP caused by mutation in PSEN genes and Apolipoprotein E (APOE) ε4 mutations affecting Aβ aggregation and clearance in lipid metabolism. 12
Numerous BChE inhibitors exhibit activity against AChE, reporting the existence of 100 selective BChE compounds alongside 25 compounds that inhibit both AChE and BChE. 108 Donepezil, rivastigmine, and tacrine are known as nonselective cholinesterase inhibitors. Other dual inhibitors are ethyl 4-nitrophenyl ethyl phosphonate, haloxon, and neostigmine bromide. Novel nonselective cholinesterase inhibitors are LX-7101 and Orlistat. LX-7101 applies to treat ocular hypertension and associated glaucoma, while Orlistat is for obesity. 108
Docking analysis applies to identify the differences in the active site between AChE and BChE molecules. 108 All compounds are docked well within the binding pocket of BChE. 108 In the AChE active site, bentamapimod and turofexorate isopropyl compounds do not interact. BChE has a more variety of small molecules that interact with the enzyme than AChE. 108
Molecular docking explores molecular interactions between proteins and ligands. Molecular docking methods suggested binding modes with active sites for BChE. As in Figure 9, an example can be compound 1 (carbon to magenta) fitted well in the active site gorge of BChE and tetrahydroisoquinoline fragment where catalytic site spatial overlap with tacrine and the amino side-chain was oriented towards the peripheral anionic site of BChE on figure A, while on figure B is shown the interactions between compound 1 and the residues of BChE, including π-π and cation-π interactions of tetrahydroisoquinoline fragment with W82 and π-π interaction of benzene fragment with Y332.29,109 Since the interactions with W82 and Y332 are significant in BChE inhibition, the two key segments remain before performing chemical modifications.
109
Schematic representation of binding molecules to enzyme BChE. (a) The predicted orientation of 1 (carbon in magenta) in the active pocket of BChE (PDB: 48DS) with tacrine as a ligand in green colour; (b) The key interaction in the BChE active site. π-π stacking and cation. π interaction is displayed in green and red dashed lines, respectively. Retrieved from ref. 109.
Combining inhibitors leads to the development of efficient therapies affecting molecular pathways, reducing neuroinflammation, and providing neuroprotection by providing a more comprehensive approach to disease management.
Factors that influence enzyme AChE
The quality of life depends on factors affecting health that are under/out of the control of the individual. Disease in the population is cured and eradicated by discovering factors leading to the illness and applying suppression methods. Detection of factors includes consideration of the current situation in the population, diagnosis, understanding of the impact on disease onset and death, monitoring of health status, detection of the causes of disease and death, application of measures to prevent and control the outcome and assessment measures for disease control and eradication. Numerous environmental factors influence the prognosis and incidence of the disease in both developed and developing countries. The most significant factors are biological characteristics of the organism and genetics (hereditary diseases), public policies and regulations (vaccination campaigns), health (financing, diagnosis, and treatment of diseases), life habits (smoking, physical activity, hydration of the organism), social and environmental factors (crime, environmental pollution).110,111 Today it is known that the onset and course of the disease require the combined action of various environmental factors. Studies on ChE levels generally examine genetic variability, health conditions, and exposure to specific chemicals, which can influence enzyme levels in different populations. For example, cholinesterase tests assess exposure to organophosphates and diagnose certain liver conditions, and results can vary widely based on individual health and genetic background. 112 Muslims suffer more from contagious jaundice and spotted fever and less often from trichinosis because they do not eat pork. 113 There is no statistically significant relation between alcohol consumption habits and the intra-racial difference in BChE activity between the Black and non-black population in Salvador, Bahia. There are statistically significant differences between genders and age groups within the Salvador, Bahia population. However, other communities show interracial differences between white and black people. 114 Disease diversity depends on biological characteristics, environment, and habits. 115 Enzyme AChE activity decreases due to chemical exposure for tea cultivation and agriculture.110,111 Moreover, the enzymes AChE and BChE change depending on the profession.112,116
Residents who smoke or consume drugs due to religious or religious customs, such as Indians, have a higher risk of cancer. 115 Cigarette consumption affects AChE and not BChE activity.117,118 Smoking leads to blood and urine biomarker changes.119,120 Drug addicts show the involvement of AChE and BChE depending on the substrate.66,121 Alcohol consumption affects the activity of the enzyme AChE and its concentration. 122 This information addresses different aspects of ChE activity, focusing on demographic differences and examining the impact of smoking, providing valuable insight into factors influencing ChE activity.
In situations with life-threatening cocaine-induced cardiovascular failure, enzyme BChE is an effective and fast therapy. 123 There was no relationship between alcohol use and lower BChE activity (p = .725, Mann–Whitney). Women using hormonal contraceptives had an activity median of 9.2% lower than the non-users. 114
Food consumption influences the immune system of an organism. 124 Insufficient vitamins, minerals, proteins, fats, and carbohydrates can lead to disease. 124 Dairy foods such as potatoes, tomatoes, peppers, strawberries, broccoli, and carrots contain natural cholinesterase inhibitors. 125
Drugs have different effects on biomarker activity. 126 For example, AChE inhibitors apply to quit smoking.127–129 Sulfonamide derivatives are good candidates for novel AChE inhibitors. Sulfonamides have low cytotoxicity and tumor selectivity in oral squamous cell carcinoma. 130 AChE enzyme inhibitors kill disease vectors such as mosquitoes. 131 Air pollution reduces AChE activity. 132 Water pollution influences the AChE range. 133 Oil spillage can lead to the inhibition of the AChE enzyme. 134
During World War II, the Germans weaponized nerve agents utilizing phosphonofluoridate-based nerve toxins like sarin, soman, and tabun dispersed through the air, inhibiting the AChE. 72
Modeling and simulation of molecular dynamics
In computational chemistry, biochemistry, and biophysics, simulation of the interaction movement and interaction of atoms and molecules during time is done with GROMACS (GROningen MAchine for Chemical Simulations), ensuring the validity and reliability of the model simulation. 134
For studying ChE inhibitors and cholinergic therapy, Linux applies with GROMACS, making it reliable and robust support. 135 Molecular Docking Studies in simulations can predict and validate the binding modes of ChE inhibitors within the active site of AChE or BChE.135,136 The molecular basis of inhibition makes sense by investigating hydrogen bonds, van der Waals interactions, and electrostatic interactions between inhibitors and enzymes. 135 In this way, molecular interactions between ChE inhibitors, AChE, and BChE are investigated, along with conformational changes and flexibility induced by inhibitors.135,136
Prediction and validation simulations of ChE inhibitor interaction with AChE and BChE arise from molecular docking studies where the primary goal is to grasp the molecular basis of interaction. 137 Determining the specific orientation and position of the inhibitor relies on the binding node and docking software that generates multiple conformations within the enzyme’s active site. 137 Each pose is scored based on how accurately it aligns with factors such as the complementarity between the inhibitor and the enzyme’s active site. 137 Investigating diverse bounds ranging from hydrogen, Van der Waals, and Electrostatic interaction reveals information regarding the reinforcement of the inhibitor complex, increasing the binding affinity and stability.138,139 Docking simulations examine how these interactions help the inhibitor to fit snugly into the active site.138,139
Electrostatic forces between charged regions of the inhibitor and the enzyme also play a significant role in binding. The docking process evaluates these interactions to understand how they contribute to the inhibitory effect. Enzyme conformational changes may arise during inhibitor binding, dependent on enzyme flexibility and conformational changes. 138 Docking studies validation applies through experimental methods such as X-ray crystallography and NMR spectroscopy. 138 This process generates a hypothesis of molecular inhibition and supports the rational design of potential therapeutic targets, incorporating a scoring function. 138 The scoring functions include empirical, force field-based, and knowledge-based methods. Diverse algorithms function as rigid bodies and flexible docking studies. 138 This study’s initial screening identifies flexibility, making results more computationally intensive, accurate, and stable in time. 138 Proper enzyme preparations like determining protonation states, optimizing geometries, and defining rotatable bounds with software tools like AutoDockTools or MOE (Molecular Operating Environment) enable accurate docking information. 138 A grid map was applied around the enzyme’s active site for docking study. 138 This grid helps evaluate the interactions between the enzyme and the inhibitor by evaluating spatial arrangements and energy terms. 138
Evaluation of Binding Modes by Pose Ranking generates multiple poses of the inhibitor after binding. 138 These poses are ranked based on their docking scores, which reflect their predicted binding affinity. 138 The highest-ranked poses apply for further analysis. Detailed analysis of the interactions within the top poses includes examining hydrogen bonds, hydrophobic interactions, and electrostatic interactions. 138 Conformational flexibility is significant in determining accurate conformational change to accommodate inhibition in addition to the simulation of molecular dynamics after docking, helping confirm the accuracy of the predicted binding modes and interactions. 138 The docking models improve accuracy based on experimental findings. This iterative process helps optimize the inhibitor design and enhance its binding affinity. 138
Molecular docking studies have applications in Drug Design regarding lead optimization and virtual screening. 138 Molecular docking is essential in lead optimization, where initial hits improve their efficacy and selectivity. 138 By analyzing docking results, chemists can design analogs with better binding properties. Docking is used in virtual screening to identify potential inhibitors from large chemical libraries. 138 This process aids in prioritizing compounds for additional experimental evaluation. 138 Despite the positive impact of molecular docking studies, potential limitations and challenges exist regarding the accuracy of enzyme and inhibitor structures. These issues can depend on the algorithm and scoring function used, which can affect the reliability of the results. 138 Advanced Docking Techniques include docking with solvent and protein-ligand binding kinetics. Some docking studies incorporate solvent effects by using implicit or explicit solvent models. Implicit models treat solvent effects as a continuous medium, while explicit models include actual solvent molecules, facilitating a more precise simulation of the binding environment. 138 Inhibitor association and dissociation through advanced docking techniques analyze the kinetics of protein-ligand binding. 140 The Free Energy Perturbation method computes the changes in free energy related to ligand binding, providing a more accurate estimate of binding affinity. 140 Although it demands substantial computational resources, it delivers valuable quantitative data. 141 The MM-PBSA/MM-GBSA methods, combined with Poisson-Boltzmann or Generalized Born solvent models, facilitate accurate estimation of free energy in molecular mechanics. 141 The indirect action of enzymes in protein-protein interactions depends on allosteric sites in molecular docking studies. 140 By integrating molecular docking studies with quantitative structure-activity relationships and pharmacophore modeling, researchers can acquire valuable insights into biological activity data and design therapeutic compounds. 142 Software tools such as PyMOL, Chimera, and VMD enable the visualization of these results. 143 Challenges in Docking Studies represent the flexibility of enzymes and scoring functional limitations.144–146 Enzymes are often flexible, and their active sites may undergo conformational changes upon ligand binding.144–146 Addressing this flexibility remains challenging and requires advanced methods like flexible docking or molecular dynamics simulations.144–146 While scoring functions provide binding affinity estimates, they may not accurately reflect experimental results.144–146 Continuous improvement and validation of scoring functions are necessary for better prediction accuracy.144–146
Docking studies have a broader application, expanding their domain in biomolecular research, environmental science, and agriculture by investigating interactions within molecular pathways and designing inhibitors for specific biological targets by utilizing machine learning algorithms and artificial intelligence (AI) that enhance the accuracy of scoring functions.147–149 Deep learning techniques apply to dock large-scale datasets and identify patterns in protein-ligand interactions that might not be apparent through traditional methods.147–149 Enhanced Sampling Techniques include metadynamics and adaptive sampling.147–149 Metadynamics overcomes the limitations of conventional docking by exploring multiple conformational states of the enzyme and ligand.147–149 It enhances the sampling of conformational space and provides a more comprehensive view of binding interactions.147–149 Adaptive biasing and umbrella sampling techniques are employed to improve the study of ligand binding pathways and sampling events.147–149
Virtual screening of large libraries to identify potential inhibitors through high throughput docking narrow compounds to the highest likelihood, while automation in docking workflow processes large datasets, which is significant in drug discovery and development.150–152 Combining and integrating molecular docking studies with Cryo-Electron Microscopy (Cryo-EM) and NMR spectroscopy improves binding predictions by validating interaction and conformational changes.150–152 The cross-docking approach involves docking a ligand to multiple protein targets to identify off-target interactions and potential side effects and understanding the specificity of inhibitors.150–152 Docking studies compare enzyme isoforms or homologous proteins to identify binding features.150–152 Mode analysis complements docking by providing insights into how receptor flexibility affects protein binding.150–152 Colective motion of protein domains and their effect on ligand binding are studied with elastic network models (ENM), while drug quality of potential inhibitors includes prediction of Absorption, Distribution, Metabolism, Excretion, and Toxicity (ADMET) properties.150–152 Adverse effects of drugs are assessed with toxicity studies to optimize the safety of drug candidates. Binding interactions and the electronic environment of the active site combine quantum mechanical calculations with quantum mechanics and molecular mechanics methods.150–152 This approach enables studying enzyme catalysis and the effects of specific interactions on binding affinity.150–152 Ensemble docking involves multiple enzyme ligand docking generated from molecular dynamics simulations.150–152 This approach accounts for the dynamic nature of proteins and provides a more accurate picture of potential binding modes and affinities.150–152
Investigation of the allosteric sites focuses on the evolution of allosteric modulators that provide alternative personalized therapeutic strategies in pharmacologics.153–156 Combining chemical libraries to identify novel inhibitors with fragment-based drug design and scaffold hopping with simulations allows better exploration of dynamic change during time by incorporating experimental data, enhancing the reliability of predictions.153–156 Implementing software like AutoDock, Dock, and FlexXX represents a powerful tool in docking studies by integrating large-scale molecular databases and repositories, enabling efficient access.153–156 Docking studies increasingly use pharmacophore models to define features required for binding. Cross docking and consensus confidence enhance the reliability of the results.153–156 Ensemble docking involves ligand docking to multiple enzymes generated through molecular dynamics or other conformational sampling methods.153–156 This approach accounts for the inherent flexibility of proteins and provides a more comprehensive assessment of binding potential.153–156
Integrating docking with pharmacophore modeling helps identify the essential features required for binding.157–159 Pharmacophore models represent spatial interactions and can guide the design of new inhibitors by highlighting crucial interaction points.157–159 Advanced docking approaches incorporate flexibility in the ligand, allowing for the exploration of multiple conformations of the inhibitor for accurately predicting binding modes, especially for flexible ligands with multiple rotatable bonds.157–159
Utilizing cloud computing resources can significantly speed up docking simulations, particularly for high-throughput or complex docking studies with computational workload, reducing time and enabling more detailed and extensive analysis.160–162 Structural bioinformatics includes protein structure databases and prediction tools.160–162 Integrating docking results with systems pharmacology approaches helps inhibitors affect entire biological systems, considering both molecular interactions and physiological effects.160–162
Docking studies can explore how mutations in the enzyme may affect binding affinity and contribute to drug resistance to designing inhibitors that can overcome resistance in pathogenic targets.163–165
Utilizing interactive visualization tools allows for dynamic exploration of docking results.163–165 Features such as molecular dynamics playback, interactive analysis of binding interactions, and real-time manipulation of structures enhance the interpretation of results.163–165 The advancement of user-friendly commercial and open-source docking platforms makes sophisticated docking techniques more accessible to researchers.163–165 These platforms often come with intuitive interfaces and integrated tools for analysis and visualization.163–165
Protein flexibility and binding affinity reveal the dynamic behavior of ChE enzymes upon inhibitor binding, including induced-fit mechanisms and conformational changes that may affect inhibitor efficacy, calculating binding free energies and affinity constants to quantify the strength of inhibitor and predict potential improvements or modifications.136,166–169 Investigating mutations in the ChE enzyme associated with inhibitor resistance unravels residues influencing inhibitor binding and efficacy.136,166–169 Novel compounds arise from exploring modification of inhibitor structure, activity, potency, selectivity, and pharmacokinetics, accelerating drug discovery in neurodegenerative diseases like AD affecting catalytic and non-catalytic activity.124–139
GROMACS runs on Linux through a few steps, starting from installation, command line interface following system requirements, parallel execution, simulation setup and execution, analysis, and visualization. GROMACS is installed using package managers (Ubuntu/Debian, ‘yum’ on CentOS/RHEL) or compiling from source, starting bash.
sudo apt-get install gromacs
GROMACS is operated through a command-line interface (CLI) to run simulations and perform analysis using commands in the terminal. 140 Linux system needs to meet the hardware requirements for GROMACS for computational performance (multi-core CPUs, GPU acceleration). 140 GROMACS takes advantage of parallel processing using Message Passing Interface (MPI) and OpenMP, which are well-supported on Linux systems, improving simulation speed and efficiency. 141 For simulation setup, an Execution file is prepared using gmx grompp to compile the input file into a simulation run file (‘.tpr’). 140 Simulations execute using ‘gmx mdrun’. 142 Post-simulation analysis and visualization tools (e.g., ‘gmx analyze’, VMD, PyMOL) is compatible with Linux, allowing comprehensive study of simulation results. 142 Using GROMACS on Linux has multiple advantages like performance, stability, community, and support, leveraging its robustness and scalability for advanced molecular dynamics and providing foundation.124–142
Molecular Dynamics Simulation Workflow with GROMACS includes system preparation (system setup and file formats), force field selection, parametrization, system energy minimization, equilibration, production of molecular dynamics, analysis and visualization (trajectory analysis, visualization, optimizing parameter, parallelization, and advanced techniques).140–142 A wanted molecular system is prepared by obtaining or building a structure and adding necessary solvents to mimic physiological conditions and factors contributing to change from experimental data (protein data bank, X-ray, NMR spectroscopy) or reliable computational model (homology modeling). 143 Before the simulation model and parameters are validated against known values, reliable experimental data, or well-established computational models using visualization tools (PyMOL, VMD) to inspect the overall structure and any other anomalies.140,142 Appropriate file format input is significant for GROMACS like PDB, GRO, and TOP.140,142 Atomic interaction is governed by a force field of choice like GROMOS, AMBER, CHARMM, and OPLS, where nonstandardized molecules are parameterized with energy minimization to relax atomic position and remove steric clashes using algorithms like steepest descent in bash by making protocols to conjugate gradient until convergence criteria making it compatible with molecular system and validated, reaching stable value equilibrium.140,142
gmx grompp -f minim.mdp -c input.gro -p topol.top -o em.tpr
gmx mdrun -v -deffnm em
Physiological conditions are mimicked with a thermostat (Berendsen, Nose-Hoover) heating the system to the desired temperature, allowing solvent and ions to equilibrate the biomolecular structure.140,142
gmx grompp -f nvt.mdp -c em.gro -p topol.top -o nvt.tpr
gmx mdrun -v -deffnm nvt
and pressure coupling (Parrinello-Rahman, Berendsen), maintaining constant pressure during equilibration and system stabilization.140,142
gmx grompp -f nvt.mdp -c em.gro -p topol.top -o nvt.tpr
gmx mdrun -v -deffnm nvt
gmx grompp -f npt.mdp -c nvt.gro -p topol.top -o npt.tpr
gmx mdrun -v -deffnm npt
Production molecular dynamics is run over time to observe the system with integrators (Verlet, leap-frog) and time step (typically 2. fs), enabling settings for electrostatic calculation, temperature, and pressure coupling.140,142
gmx grompp -f md.mdp -c npt.gro -p topol.top -o md_0_1.tpr
gmx mdrun -v -deffnm md_0_1
Trajectory files (XTC, TRR) are analyzed to extract thermodynamic properties (temperature, pressure), structural information (RMSD, RMSF), and dynamic behaviors (hydrogen bonds, protein-ligand interactions) to asses system stability, dynamics, and thermodynamic properties and visualized with VMD, PyMOL, or GROMACS built-in tools to visualize molecular structures, trajectories, and interaction networks.140,142 Root Mean square Deviation (RMSD) and Square Fluctuation (RMSF) monitor structural integrity and fluctuations along with protein-ligand interactions, validating post-stimulation with experimental data or literature and dynamic behavior.140,142 The results should be consistent across all replicates and divergent in structural dynamics during time to assess reproducibility and consistency of results.140,142
To make effective stimulation parameter optimizes with fine tune simulation (cutoff distances, integration time step) for accuracy and efficiency, GROMACS parallel capabilities (MPI, OpenMP) for faster simulations on multi-core CPUs or GPU-accelerated systems and exploring advanced simulation techniques like enhanced sampling (umbrella sampling, metadynamics) to study rare events and complex dynamics.140,142
GROMACS enables powerful tools for researching biomolecular systems at the atomic level. Following this systematic workflow, the reader can simulate and explore diverse biological processes in drug discovery, enzyme mechanisms, and protein folding, contributing to molecular biology and biophysics. Applying this workflow in model validation enables the best practice and validation of simulation protocols in drug discovery and biomolecule engineering. In this way, computational methodology brings a gap between molecular insight and clinical application, offering a comprehensive insight into ChE inhibitors from molecular interaction to therapeutic application, enabling future data refinement of drug design. During reporting and result interpretation, discussing limitations and recommendations is essential to offer insights into further experimental validation and computer method refinement support in decision-making in drug discovery. Following this step-by-step guide will enable systematic integration of modeling validation into molecular dynamics using GROMACS, pushing boundaries further in biological research of molecular simulation that will help patients in a real-time struggle with the disease.124–142 The detailed script is present in the Supplemental file 2.
An important consideration and future perspectives
Analytical techniques detect and quantify AChE inhibitors and their metabolites in blood, urine, and cerebrospinal fluid samples.143,144 It is one decade of techniques in the research practice, enabling the detection of simultaneous changes in the organism during homeostatic and pathological organism states. Changes in enzyme AChE concentration and structure are specific. Enable the development of potential drug target molecules. A series of experiments in vitro human blood samples can show that enzyme AChE activity and inhibitor inhibition efficacy change in plasma, erythrocyte, and whole blood fraction, depending on pre-analytical factors like time of freezing, the temperature of freezing, sample frequency, physical activity, and water intake. Investigating the effect of pre-analytical factors is significant for proper result interpretation. Enzyme AChE is not a generic marker of extracellular vesicles, meaning that AChE activity and inhibition in blood fractions are reliable blood biomarkers. 145 Blood fractions enriched with AChE participate in inflammatory processes during health and disease, membrane integrity, aging, gender differences, neurotoxicity, and pesticide poisoning. 146
The primary task of developing countries is to improve health outcomes and disease eradication by providing high-quality health care using available drugs. Increasing the effectiveness of drugs and connecting and understanding the physiological changes of the organism due to environmental factors will improve the quality of the market. Creating a new clinical and scientific chain system that uses ingredients, permitted doses, and is safe to work with humans improves drug access and increases safety.
The plan for further scientific analysis in this area includes increasing the research value by conducting a systematic analysis with guiding steps. An up-to-date literature review of existing experimental and computational findings will enable the revealing of potential inconsistencies and gaps with further validation and research development. The literature review encompasses experimental protocols and statistical data analysis to validate the information from the materials and methods section and deepen the understanding of current research. Collaboration with interdisciplinary experts will enable rigor and robust methodology validation of the experimental result designs. To further understand the action mechanism for BChE/AChE inhibition series of compounds, the kinetic assay and molecular docking studies determine the optimal composite. The inhibition assay should evaluate a range of bioactive compounds and natural products to assess their potential to inhibit AChE and BChE enzymes, thereby facilitating their clinical application in disease treatment. Before clinical application in vivo, models provide pharmacokinetic properties and investigate clinical performance. A combination of biochemical, molecular biological, genetic, and molecular docking studies will allow the identification of crucial compounds and specific genomic and biochemical locations for disease treatment. Providing extensive data will help develop models capable of predicting the effects and clinical efficacy of AChE and BChE inhibitors.
Structural analysis with X-ray crystallography and NMR spectroscopy will validate protein-protein interactions and detect structural changes to validate existing findings. The objective would be to increase the robustness and reliability of data by designing and executing an experimental approach with computational methodology. However, future experimental validation should include patients with AD and MG from patients. In silico studies of potential biomarkers in neurodegenerative diseases and neurotoxicity are currently under review. Potential computational methodology should investigate and confirm the findings from the literature and experiments. Accuracy and reliability of computational models with approaches like molecular dynamics simulations by extending simulation times (from nanoseconds to microseconds) improve sampling stability and validate simulation results by comparing root mean square deviations (RMSD) of enzyme structures against experimental results or known structures, quantum mechanics calculations to refine the understanding of reaction mechanisms and binding energies of inhibitor-enzyme complexes and to study enzymatic reaction mechanisms involving cholinesterase inhibitors, and force field optimization for fine-tune force field parameters (partial charges, van der Waals parameters) based on experimental validation or community standard, and comparing force field with an experimental observation like binding affinities and structural stability with GROMACS. The computational analysis should validate and refine computational models and simulations comparing predictions with experimental results.
Conclusion
This study highlights the diversity of ChE roles ranging from cholinergic transmission to various biological processes. Revealing the ChE polymorphisms and their correlation in diseases and treatment could provide valuable insights into personalized medicine and improve therapeutic outcomes. The findings of this study greatly enhance scientific understanding of ChE function and its involvement in diseases, highlighting the potential of ChE inhibitors as novel therapeutic targets by elucidating their roles and mechanisms that influence various biochemical and molecular pathways. Future studies should based on uncovering specific polymorphisms associated with particular disease phenotypes, exploring the role of ChE inhibitor development, and investigating the non-cholinergic function by revealing new pathways and mechanisms in personalized medicine. The study explicitly contributes to the literature and suggests clear directions for further research, emphasizing the importance of ChE polymorphisms and inhibitors in disease treatment and drug development. Multitarget therapeutic strategies are recommended as a part of natural compounds in AD treatment as part of natural sources, providing a wide range of bioactives varying from antioxidants with anti-inflammatory and neuroprotective roles addressing the complex pathophysiology of the disease. Future research on these natural compounds will lead to a complete understanding of the implementation of standard disease treatment protocols. Molecular docking studies enable understanding the complete picture of the interaction between ChE inhibitors alongside ChE at the molecular level, revealing information regarding inhibitor affinity, stability, and efficacy and improving therapeutic outcomes. Despite its potential limitations, it remains valuable to drug discovery and development. A comprehensive understanding of the enzyme inhibitor and accuracy of prediction with molecular docking studies result in effective disease therapy, protein engineering, and system biology. These advancements enhance the precision of docking studies and expand their applicability across various research areas. By incorporating these advanced considerations and technologies, researchers can further refine molecular docking studies, enhance the accuracy of predictions, and gain deeper insights into enzyme-inhibitor interactions. This comprehensive approach contributes to more effective drug discovery and development processes.
Blood enzyme AChE widely applies in clinical and scientific research. AChE represents an excellent basis for understanding the origin of human diseases. This review contributes to a better understanding of the enzyme structure, biogenesis, and function while indicating significant considerations and future perspectives related to genomic organization, polymorphism, cholinergic therapy, and influencing factors. Following the step-by-step guide, you can systematically integrate modeling validation into your molecular dynamics simulations using GROMACS. This approach ensures that your simulation results are robust, reliable, and scientifically sound, contributing to advancements in understanding biomolecular interactions and supporting drug discovery efforts. Continual refinement and validation of computational models are essential for pushing the boundaries of molecular simulation in biological research. Understanding disease biochemistry, genetics, and molecular biology is rising with the understanding of AChE and BChE molecular signalization, function, and clinically significant inhibitors. The current problems should be more thoroughly explored and investigated by scientists. Understanding the physiology of the ChE enzymes, their interaction and functionality, and the mechanism that contributes to disease development enables full clinical significance and applicability. There has been substantial progress in our understanding of the biochemistry and genetics of BChE.
Supplemental Material
Supplemental Material - Enzyme ChE, cholinergic therapy and molecular docking: Significant considerations and future perspectives
Supplemental Material for Enzyme ChE, cholinergic therapy and molecular docking: Significant considerations and future perspectives by Snežana M Jovičić in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplemental Material - Enzyme ChE, cholinergic therapy and molecular docking: Significant considerations and future perspectives
Supplemental Material for Enzyme ChE, cholinergic therapy and molecular docking: Significant considerations and future perspectives by Snežana M Jovičić in International Journal of Immunopathology and Pharmacology
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
List of abbreviations
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
