Abstract
Objective:
Given the global high incidence of colorectal cancer (CRC) and the need for subtype-specific molecular targets, this study aims to investigate the role and therapeutic potential of EIF4A1 in colon and rectal cancers.
Introduction:
The potential of Eukaryotic translation initiation factor 4A1(EIF4A1) to guide prognosis and inform immunotherapeutic strategies in colon and rectal cancers warrants further investigation.
Methods:
EIF4A1 expression levels were measured by quantitative real-time PCR (qRT-PCR), and EIF4A1 protein expression was evaluated via immunohistochemistry (IHC). Additionally, multi-dimensional database analyses were performed to characterize the biological functions of EIF4A1 and explore its heterogeneity in CRC.
Results:
EIF4A1 expression was upregulated in both colon and rectal cancers, with significant associations with poor survival outcomes. Functional enrichment analyses indicated that EIF4A1 was predominantly associated with the neutrophil extracellular trap (NET) network in colon cancer. Strong positive correlations between EIF4A1 expression and neutrophil infiltration were observed in both cancer subtypes. Notably, EIF4A1 exhibited divergent immune infiltration patterns: a strong positive correlation with CD8+ T cells in colon cancer, versus a negative correlation with CD4+ T cells in rectal cancer. EIF4A1 also showed strong positive correlations with immune checkpoint molecules (PD-1, PD-L1, CTLA4, and LAG-3) in colon cancer, whereas these correlations were weaker in rectal cancer.
Conclusion:
The heterogeneity of EIF4A1 and its differential roles in driving cancer progression between colon and rectal cancers highlight the need for subtype-specific immunotherapeutic strategies for these malignancies.
Introduction
Colorectal cancer (CRC) is recognized as one of the most common cancers and is characterized by one of the highest mortality rates. In China, its incidence has been steadily increasing, rendering it the third most prevalent cancer across the globe. 1 CRC can be further classified into colon and rectal cancers, with treatment principles often differing in clinical practice based on the disease’s location. The treatment options for both colon and rectal cancers includes surgery, chemotherapy, and radiotherapy; nonetheless, specific surgical techniques and treatment strategies vary according to the tumor’s location and stage. Surgery for rectal cancer can be notably complex due to its anatomical features. Focusing on the biological and clinical differences and similarities between rectal and colon cancers is critical, as these aspects significantly affect current cancer staging and treatment, and may shape the design of future targeted drug trials.2,3 Recent studies have revealed that within the realm of colorectal tumor therapy, multidimensional research provides significant perspectives for precision-oriented management. For instance, ensuring drug safety and individualized strategies are crucial: in response to the frequent occurrence of adverse reactions when treating tumors with drugs such as regorafenib, it is necessary to explore individualized dosing regimens based on patient tolerance, and to enhance treatment adherence and continuity through dynamic adjustment of dosing strategies. 4 In the epoch of immunotherapy, it is imperative to emphasize the early monitoring and multimodal management of immune-related adverse reactions (irAEs), and refine the treatment regimen by incorporating with inflammatory markers (e.g. NER, RMH scores), thereby safeguarding the antitumor efficacy and substantially enhancing the patients’ quality of life. 5 Moreover, integrating metabolism and inflammation regulation has become a cardinal direction: metabolic syndrome and chronic inflammation synergistically propel tumor progression, Clinicians need to incorporate blood glucose, lipid management and anti-inflammatory interventions into comprehensive treatment protocols and integrate surgery, radiotherapy and chemotherapy to devise individualized treatment regimens. 6 Regarding metabolic risk factors such as obesity and insulin resistance, lifestyle interventions including dietary adjustments and weight management should be integrated into the prevention and treatment system. Furthermore, it is essential to consider gender and age differences to facilitate the implementation of precise prevention and treatment strategies. Prognostic assessment tools exhibit substantial potential for clinical translation: the RMH score, which is based on albumin, 7 LDH level and the number of metastases, has emerged as a universal prognostic tool for colorectal and gastroesophageal cancers due to the consistency of cross-clinical studies and the real-world setting. For patients with high scores, interventions such as enhanced nutritional support, inflammation regulation and anti-angiogenic/metabolic regulation drugs are required. The neutrophil-eosinophil ratio (NER), as an emerging inflammation indicator, holds promise for assessing obesity-related and metabolic risk. 8 NER, as an emerging inflammatory indicator, can reflect the balance of the tumor microenvironment via simple detection. For patients with high NER values, combined interventions in immunotherapy should be prioritized to reverse immunosuppression. To this end, large-scale prospective studies are needed to clarify the standardized cutoff value and facilitate the integration of NER with molecular markers, thereby enabling the construction of Overall, tumor treatment strategies should transcend the single-model approach and establish a comprehensive precision strategy encompassing prevention, treatment, and monitoring. This strategy should integrate drug safety management, immune microenvironment regulation, metabolic inflammation intervention, and multi-dimensional prognosis assessment, ultimately aiming to achieve a dual improvement of therapeutic efficacy and quality of life. With societal progress, an increasing number of risk factors for colorectal cancer have been identified. Notably, individuals with colon or rectal cancer are being diagnosed at increasingly younger ages, often at advanced stages. This trend exerts a significant negative impact on their prognosis, treatment outcomes, and quality of life. Therefore, there is an urgent requirement for early diagnosis and prognostic assessment in clinical practice, alongside the development of novel diagnostic and prognostic markers to distinguish between colon and rectal cancers.9,10
Protein synthesis is frequently dysregulated in cancer; furthermore, the translation of mRNA into proteins is a highly complex and crucial process. 11 mRNA translation comprises the stages of initiation, elongation, and termination. Cells can rapidly reprogram their proteomes to dynamic environments through the regulation of these stages. 12 Accumulating evidence suggests that translational dysregulation of mRNAs associated with cancer development, and regulatory elements at the translational level are of critical importance.13–15 Dysregulated translation initiation is widely associated with cancer initiation and progression, and the eukaryotic translation initiation factor EIF4A1, as an important translation initiation regulatory element, is involved in the homeostatic regulation of translation initiation, which plays a crucial role in the translation initiation process by unraveling the secondary structure of the mRNAs upstream of the start codon through its own unraveling action, a mechanism that promotes the efficient recruitment of ribosomes to downstream gene sequences. This mechanism promotes the efficient recruitment of ribosomes to downstream gene sequences, thereby driving the dynamic process of protein synthesis.16–18 Dysregulation of mRNA translation is frequently dysregulated in various cancers, including colorectal cancer. Selective Targeting and inhibiting the translation of specific oncogenic mRNAs represents a promising anticancer therapeutic approach.
EIF4A1 is a member of the EIF4A family and plays a critical role in the initiation phase of translation by regulating the 5′ untranslated region (5′ UTR) of mRNA. As an RNA helicase, it catalyzes the unwinding of RNA secondary structures in an ATP-dependent manner.19,20 EIF4A1, serving as the enzymatic catalytic core of the EIF4F complex, is essential for translation initiation and plays a key role in reprogramming protein synthesis in cancer, thus making it a promising target for cancer therapy. 21 Accumulating evidence demonstrates that EIF4A1 expression correlates with epithelial-mesenchymal transition (EMT) in multiple cancers. EIF4A1 plays a crucial role in the EMT process of gastric cancer. In pancreatic cancer, genetic deletion of EIF4A1 compromises EMT and reduces the metastatic potential of pancreatic cancer cells. Furthermore, EIF4A1 functions as a biomarker for early tumor diagnosis, histological classification, and clinical staging, providing novel tools for precision and personalized cancer therapy.19,22–24
In this study, bioinformatics analysis was conducted using the TCGA database to characterize the expression disparity of EIF4A1 between colon and rectal cancers. Integrative analysis of EIF4A1 expression profiles and clinical datasets revealed consistent upregulation of EIF4A1in both colon and rectal cancers. This upregulation was statistically significant in colon cancer, implying stronger prognostic value for survival prediction and higher diagnostic specificity compared to rectal cancer. Functional enrichment analysis revealed that colon cancer is associated with DNA and protein interactions. Additionally, pathway enrichment analysis revealed significant connections with Neutrophil Extracellular Traps (NETs) and epigenetic modifications, such as methylation and acetylation. These findings suggest roles for EIF4A1 in cancer progression, immune evasion, and immune response mechanisms. Rectal cancer is primarily characterized by enrichment in pathways related to immune response recognition, immune and inflammatory responses, cellular stress, and energy metabolism. Notably, the Complement System and the MAPK signaling pathway play crucial roles in immune surveillance and immune response. Inhibition of these pathways may contribute to the development of cancer, highlighting their potential as therapeutic targets. Immunoassays revealed distinct immune infiltration patterns for EIF4A1 in colon and rectal cancers, with varying degrees of immune cell infiltration indicative of different immunotherapeutic responses. Correlation analysis between EIF4A1 and immune checkpoints indicated more complex interactions in colon cancer compared to rectal cancer, highlighting the potential for tailored immunotherapeutic strategies in colorectal cancers.
Result
Screening for differential prognostic genes
A total of 1549 differentially expressed genes (DEGs) in HCT116 were obtained from the RNA-binding protein (RBP) database cited in the literature. 25 A total of 2929 significant differential genes were identified by setting the parameters log2 fold change (|logFC|) > 1 and p.adj < 0.05 in the TCGA-COADREAD dataset. A list of colorectal cancer-related prognostic genes was derived through univariate Cox regression analysis of the TCGA-COADREAD dataset. A total of 631 prognostic genes were obtained by setting the parameters hazard ratios (HR) > 1 and p.adj < 0.05. The RBP library, TCGA-COADREAD differential genes, and prognostic genes were integrated to generate Venn diagram intersections. The results identified three overlapping genes (Figure 1(a)): TSFM, EIF4A1, and ARL6IP4. Visualization of these differential genes was conducted using volcano plot analysis in the TCGA-COADREAD database (Figure 1(b)). Co-expression analysis of the three DEGs with clinical Pathologic T stage, N stage, and M stage was performed using TCGA-COADREAD data. The results indicated that TSFM, EIF4A1, and ARL6IP4 exhibited higher expression in the Pathologic T3 stage, were predominantly expressed in the Pathologic N0 stage, and showed exclusive expression in Pathologic M0 stage (Figure 1(c)). Focusing on the eukaryotic initiation factor EIF4A1, which regulates translation initiation, we designated it as a key molecule for subsequent research.

Screening of RBP related differential prognostic proteins. (a) Cor1 is the RBP database, cor2 prognostic gene set, and cor3 differential gene set, and the intersection of the three sets is taken. (b) Volcano plot visualization of related differential genes. (c) Clinically relevant co expression maps of related genes.
Expression landscape of EIF4A1
EIF4A1 exhibits broad expression of the mRNA and protein levels across various organs and tissues (Figure 2(a)), Transcriptomic analysis indicated that EIF4A1 mRNA is preferentially expressed in the proximal digestive tract, kidney, urinary bladder, connective and soft tissue, as well as bone marrow and lymphoid tissues. Proteomic data from the The Human Protein Atlas (HPA) database showed that EIF4A1 protein was highly abundant in the stomach, colon, rectum, placenta, tonsil, and bone marrow (Figure 2(b) and (c)). IHC staining of EIF4A1 in normal colorectal tissues and colorectal cancer tissues from the HPA database (Figure 2(e)–(g)) revealed that normal colon tissues displayed stronger staining intensity than rectal tissues, with darker coloration indicative of higher expression. Notably, EIF4A1 expression was significantly upregulated in colorectal cancer tissues compared to normal tissues. Bioinformatics analysis via the UALCAN database demonstrated that EIF4A1 protein expression was markedly elevated in malignant tissues (Figure 2(d)). Subcellular localization analysis of EIF4A1 in CACO-2 cells, from the HPA database revealed that EIF4A1 is predominantly localized in the cytoplasm (Figure 2(h)).

EIF4A1 protein expression landscape and differences. (a) The expression landscape of EIF4A1 protein and mRNA in human tissues. (b) EIF4A1 expression data in human normal tissues. (c) EIF4A1 expression data in cancer. (d) EIF4A1 expression in CRC, p < 0.0001. (e) Representative IHC images of EIF4A1 expression in normal colon. (f) Representative IHC images of EIF4A1 expression in colon cancer. (g) Representative IHC images of EIF4A1 expression in rectal cancer. (h) Fluorescence staining images of EIF4A1 protein localization in colorectal cancer cell line CACO-2.
Pan-cancer analysis via the Timer 2.0 database (Figure 3(a)) indicated that EIF4A1 exhibited significantly elevated expression in multiple human malignancies, including CHOL, COAD, ESCA, GBM, HNSC, KIRC, KIRP, LIHC, LUAD, LUSC, PRAD, STAD, UCEC (each with p < 0.001), BRCA (p < 0.01), and READ (p < 0.05), when compared to normal tissues. Conversely, reduced expression was observed in KICH (p < 0.001) and PCPG (p < 0.05) compared to normal tissues. Pairwise tissue comparison using TCGA dataset (ID: 11123; Figure 3(b)) showed that EIF4A1 was significantly overexpressed in COAD (p < 0.001), CHOL (p < 0.05), KIRC (p < 0.001), and READ (p < 0.01), whereas under expressed in BRCA (p < 0.001) and THCA (p < 0.05). Based on the TCGA-COADREAD data of 698 samples, 51 cases of paraneoplastic tissues were analyzed for differences, and the results suggested (Figure 3(c)) that EIF4A1 showed high expression in COADREAD with significance p < 0.001, which was downloaded from the TCGA database (https://portal.gdc.cancer.gov) and organized RNAseq data from the STAR process of TCGA-COAD and TCGA-READ projects and extracted the data in TPM format, extracted the corresponding numbered paired paraneoplastic and cancer samples of TCGA-COADREAD dataset to perform the paired samples t-test, the results suggested that EIF4A1 was significantly higher in cancer tissues of COADREAD than in paraneoplastic tissues (Figure 3(c)), with significant p < 0.001. The TCGA-COAD, TCGA-READ datasets were downloaded and organized from the TCGA database, and the data were log2(value+1) processed, and cancer and paracancer difference analysis and paired analysis were performed in the same way, respectively. Both unpaired and paired analyses confirmed that EIF4A1 expression was significantly upregulated in COAD and READ tissues compared to paracancerous controls, with stronger statistical significance in COAD (p < 0.001) versus READ (p < 0.01). These findings imply a greater differential expression magnitude of EIF4A1 in colon versus rectal tissues (Figure 3(d) and (e)).

mRNA expression landscape and differences of EIF4A1. (a) The expression of EIF4A1 in various tumors. (b) Pairwise analysis of EIF4A1 in various tumors picture. (c) mRNA expression differences of EIF4A1 in COADREAD. (d) Differential expression of EIF4A1 in COAD. (e) Differential expression of EIF4A1 in read. (f) The expression of EIF4A1 in clinical tissue samples of colorectal cancer. *p < 0.05, **p < 0.01, ***p < 0.001
Clinical diagnosis and prognostic value evaluation of EIF4A1
Cox proportional hazards regression analysis was performed using the “survival” package to generate survival models. Results were visualized using the “survivminer” and “ggplot2” packages. Survival analyses included the COADREAD, COAD, and READ datasets, assessing three endpoints: Overall Survival (OS), Disease-Specific Survival (DSS), and Progress-Free Interval (PFI). Overall survival analysis revealed that low EIF4A1 expression was associated with improved survival outcomes in COADREAD (p = 0.045) and COAD (p = 0.007; Figure 4(b) and (c)), with statistical significance, whereas high EIF4A1 expression correlated with better prognosis in READ (p = 0.041; Figure 4(d)). DSS analysis indicated that high EIF4A1expression was linked to worse DSS outcomes in COADREAD (p = 0.024) and COAD (p = 0.012) had a poor survival prognosis (Figure 4(f) and (g)), while no significant association was observed in READ (p = 0.778; Figure 4(h)).

Analysis of prognostic and diagnostic value of EIF4A1 in colon and rectal cancer in TCGA database. (a, e, i) Correlation of EIF4A1 expression with OS, DSS, and PFI in colorectal cancer and colorectal cancer. (b–d) Correlation of EIF4A1 expression with OS and DSS and PFI in colorectal cancer. (f–h) Correlation of EIF4A1 expression with OS and DSS and PFI in colon cancer. (j–l) Correlation of EIF4A1 expression with OS and DSS and PFI in rectal cancer. (m–o) Clinical ROC analysis of colorectal cancer, colon cancer, and rectal cancer.
PFI analyses demonstrated that high EIF4A1 expression was significantly associated with worse PFI outcomes in COADREAD (p < 0.001) and COAD (p = 0.001; Figure 4(j)–(l)). Conversely, no significant differences were observed in READ (p = 0.256). To synthesize the prognostic significance of EIF4A1, Kaplan-Meier estimates for OS, DSS, and PFI were visualized using “ggplot2,” along with Cox regression results including HR and p values (Figure 4(a), (e), (i)). Using clinical datasets from the TCGA-COADREAD, TCGA-COAD, and TCGA-READ datasets, Receiver Operating Characteristic (ROC) analysis was performed using the “pROC” package, and ROC curves were visualized via “ggplot2.” ROC analysis demonstrated that EIF4A1 achieved Area Under the Curve (AUC) values of 0.827 in COADREAD, 0.836 in COAD, and 0.797 in READ (Figure 4(m)–(o)), There results indicate that EIF4A1 exhibits significant clinical diagnostic value across COADREAD, COAD, and READ, with subtype-specific diagnostic potential Notably, COAD displayed a higher diagnostic accuracy (AUC = 0.836) than READ (AUC = 0.797), underscoring EIF4A1’s superior diagnostic potential for colon cancer. To investigate the impact of EIF4A1 expression and clinicopathological parameters on survival outcomes, survival disparities between colon and rectal cancers were analyzed using univariate and multivariate Cox regression models. Univariate analysis in colon cancer identified that pathological T stage (particularly T4), N stage (particularly N1), advanced pathological stage, age, and EIF4A1 expression were significantly associated with increased mortality risk in colon cancer patients. In rectal cancer, elevated mortality risks were significantly associated with T4 stage, N2 stage, and advanced age (Tables 1 and 2, p < 0.05), Multivariate Cox regression analysis in colon cancer revealed that T4 stage, advanced pathological stages, and advanced age remained independent predictors of mortality after adjusting for confounding variables. In rectal cancer, N2 stage and advanced age were identified as independent risk factors for mortality following multivariate adjustment. In rectal cancer, missing data for certain variables and insufficient sample size when including pathological stage in multivariate analysis led to the exclusion of this variable. Hence the exclusion of this variable from the analysis. Age exceeding 65 years was consistently associated with an increased mortality risk in both univariate and multivariate analyses. Notable differences between the cancers included the significance of the T3 stage in colon cancer in univariate analysis, which did not persist in multivariate analysis. In contrast, the T3 stage in rectal cancer demonstrated no significant difference in either analysis. For colon cancer, the HR for the N1 stage was attenuated and lost statistical significance in the multivariate analysis, while the N2 stage remained significant. In rectal cancer, both N1 and N2 stages were significant predictors in univariate analysis; but only the N2 stage retained significance in the multivariate analysis. The M1 stage was a significant predictor in both univariate and multivariate analyses for colon cancer, but only in the univariate analysis for rectal cancer. In colon cancer elevated EIF4A1 expression was associated with an increased mortality risk in univariate analysis though this association was not maintained in multivariate analysis. Conversely, in rectal cancer, high EIF4A1 expression correlated with reduced mortality risk in univariate analysis, although this trend did not achieve statistical significance in multivariate analysis (Tables 1 and 2).
Univariate and multivariate analyses of clinicopathological parameters in patients with COAD.

p < 0.05, and the results were statistically significant.
Univariate and multivariate analyses of clinicopathological parameters in patients with READ.
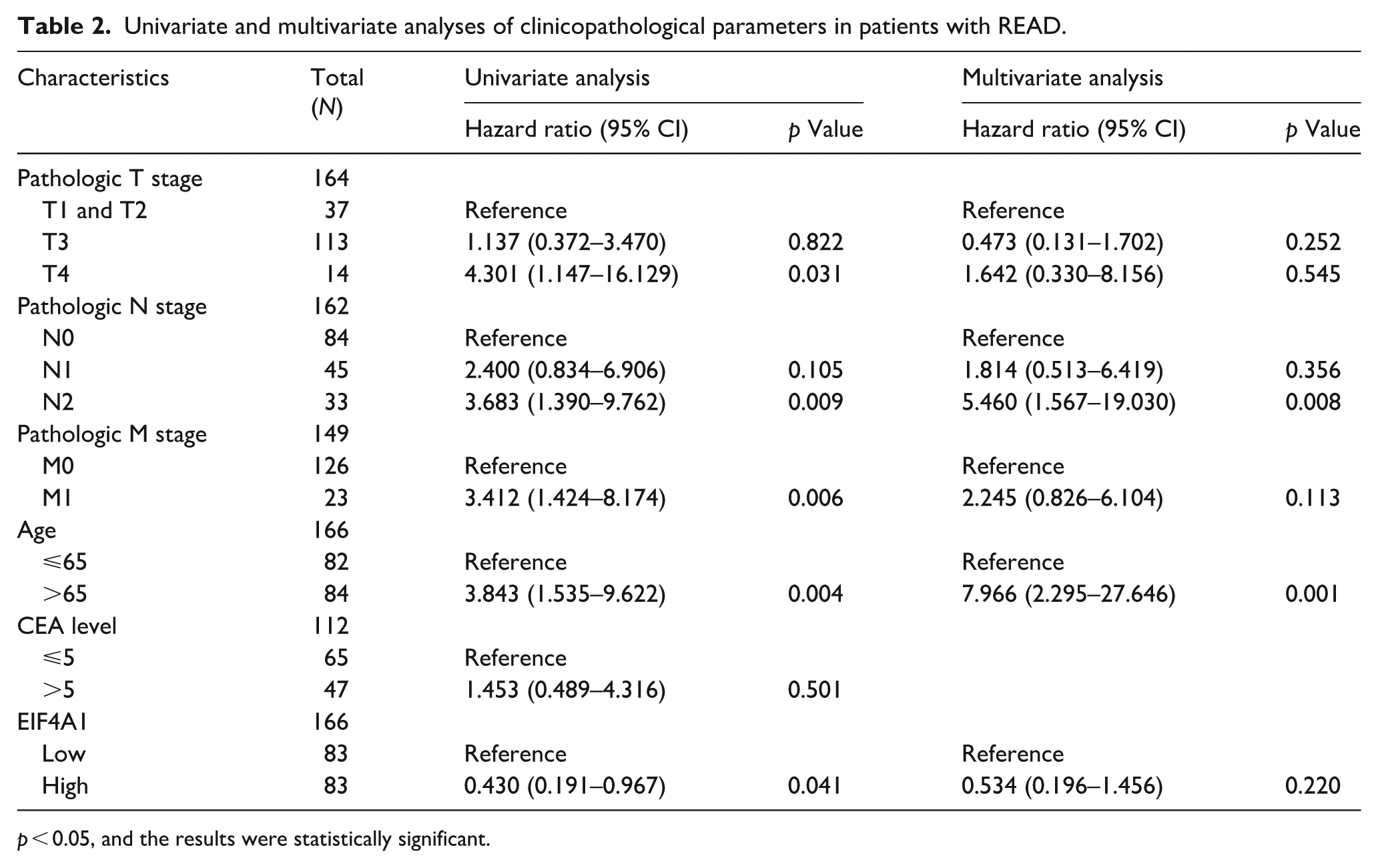
p < 0.05, and the results were statistically significant.
EIF4A1 transcription of expression and the clinical correlation analysis
We employed the UALCAN database to investigate the correlation between EIF4A1 mRNA and protein expression and clinical outcomes. The findings are presented below. Tumor Staging Analysis: EIF4A1 transcript levels were consistently elevated in COAD tissues relative to normal counterparts. Across all cancer stages, EIF4A1 mRNA expression exhibited significant upregulation (p < 1e-12–1.27e-08), underscoring statistically significant differences. In COAD, interstage comparisons revealed significant disparities in EIF4A1 transcript levels: between stage I and stage III (p < 4.858400e-2), stage I and stage IV (p < 3.00300e-3), and stage II and stage IV (p < 1.476810e-3), (Supplemental Figure 1(a)). When contrasting READ with COAD, EIF4A1 expression was significantly higher in stages I and IV, READ tissues. However, no significant differences were observed in stages II and III. Additionally, intrastage analysis in READ demonstrated significant difference in EIF4A1 expression of between stages I and II of READ (p < 1.47903e-03) and between stages I and III (p < 2.14800e-02; Supplemental Figure 1(b)). Overall, EIF4A1 mRNA expression in COAD predominantly accumulated in stage I-II, whereas in READ, it was primarily detected in stage I (Supplemental Figure 1(a) and (b)). Age analysis in COAD showed that EIF4A1 expression was significantly elevated across all age groups compared to the normal group, particularly in 41–60 (p < 5.55e-16), 61–80 (p < 1e-12), and 81–100 year olds (p < 8.89e-11; Supplemental Figure 1(c)). Conversely, in READ, EIF4A1 expression was significantly elevated in the 21–40 (p < 0.000956) and 41–60 year olds (p < 0.008163) compared to the normal group. Notably, expression in the 41–60 and 61–80 age groups was significantly higher than in the 81–100 year olds, all with statistical significance. However, no significant differences were observed in EIF4A1 expression between the 61–80 and 81–100 year old groups compared to the normal group (Supplemental Figure 1(d)). Compared to READ (Supplemental Figure 1(c) and (d)), elevated EIF4A1 expression in COAD mainly occurred in the 81–100 age group, whereas in READ, it was primarily observed in the 21–40 age range.
In COAD, EIF4A1 expression was significantly elevated in both adenocarcinoma and mucinous adenocarcinoma subtypes compared to normal tissues. Furthermore, EIF4A1 expression in mucinous adenocarcinoma was significantly higher than in adenocarcinoma (p = 0.04076), suggesting potential molecular differences between these subtypes (Supplemental Figure 1(e)). Conversely, in READ, EIF4A1 expression in mucinous was significantly higher than normal tissues, (p = 0.009957; Supplemental Figure 1(f)). In both COAD and READ, EIF4A1 expression was notably high in mucinous adenocarcinoma subtypes (Supplemental Figure 1(e) and (f)). Lymph node metastasis analysis revealed that EIF4A1 expression in COAD across different lymph node metastasis states(N0, N1, N2) was markedly higher than in normal tissues, demonstrating significant statistical differences (p values ranging from 1.62e-12 to 1.00e-9; Supplemental Figure 1(g)). No significant statistical correlation was found between EIF4A1 expression and lymph node metastasis states in READ (Supplemental Figure 1(h)). In COAD, the highest levels of EIF4A1 expression were predominantly observed in the N0 phase, whereas in READ, elevated EIF4A1 expression was primarily noted in the N1 phase (Supplemental Figure 1(g) and (h)). Gender analysis in COAD showed that EIF4A1 expression was significantly elevated in both male and female patients compared to normal tissues, with p-values of <1e-12 and 1.62e-12, respectively. However, the differences between genders were not statistically significant (p = 0.22666; Supplemental Figure 1(i)). In READ, EIF4A1 expression in male patients was notably higher than in normal tissues (p = 0.01266), whereas in female patients, this elevation did not reach statistical significance (p = 0.09802; Supplemental Figure 1(j)). In READ, the incidence of cancer among male patients may be higher than in female patients.
Regarding weight in COAD, EIF4A1 expression was significantly up-regulated across various weight categories: Normal Weight (p = 3.64e-12), Extreme Weight (p = 1.62e-12), Obese (p = 3.56e-10), and Extremely Obese (p = 0.007; Supplemental Figure 1(k)). In READ, EIF4A1 expression was up-regulated in Normal Weight, Extreme Weight, and Obese categories, with only Normal Weight reaching statistical significance (p = 0.00918). Additionally, EIF4A1 expression in Normal Weight was significantly higher than in Extreme Weight (p < 0.032; Supplemental Figure 1(l)), Notably, in both COAD and READ, the highest EIF4A1 expression occurred in the Normal Weight category, with expression generally decreasing as body weight increased (Supplemental Figure 1(k) and (l)).
The correlation between EIF4A1 protein expression and clinical analysis
Given that the protein database only includes clinical data for Colon cancer, our analysis focused on evaluating between EIF4A1 protein expression levels and various clinical parameters in Colon cancer, such as cancer stages, patient age, tumor history, chromatin modifier status, patient gender, and weight.
In Colon cancer, EIF4A1 protein expression exhibited significant variability across pathological stages. Compared to normal tissues, EIF4A1 expression was substantially elevated in all cancer stages, particularly peaking in stages II (p = 8.77e-12) and III (p = 1.99e-12), with these findings reaching significant statistical levels (Supplemental Figure 2(a)). However, there was no significant difference in EIF4A1 expression between cancer stages. This suggests that once colon cancer develops, EIF4A1 expression remains consistently high across advanced stages, indicating minimal correlation with cancer progression stages.
Regarding patient age, EIF4A1 expression was significantly up-regulated across all age groups compared to the normal tissues, with the most striking elevation in 61–80 year-old group (p = 9.92e-16; Supplemental Figure 2(b)). These findings indicate that variations in EIF4A1 protein expression are potentially age-related during cancer progression, particularly among middle-aged and elderly individuals.
Tumor History Analysis: EIF4A1 protein expression was significantly elevated in both mucinous and non-mucinous tumors compared to normal samples (Supplemental Figure 2(c)). Notably, the increase was more pronounced in non-mucinous tumors (p = 2.19e-21), although the differences between mucinous and non-mucinous tumors were not statistically significant (p = 0.527). This suggests that elevated EIF4A1 expression is a characteristic common among cancer patients, particularly in non-mucinous tumors.
Chromatin Modifier Status Analysis: EIF4A1 protein expression was significantly up-regulated in samples with modified chromatin (Supplemental Figure 2(d)), with a statistically significant increase compared to normal samples (p = 2.82e-22), suggesting a significant correlation between elevated EIF4A1 expression and chromatin modification status, which may promote EIF4A1 protein expression upregulation.
Patient Gender: Correlation analysis revealed that EIF4A1 protein expression was significantly higher in both males (p = 6.50e-13) and females (p = 1.75e-15) compared to the normal group (Supplemental Figure 2(e)). However, there was no significant difference in EIF4A1 expression between males and females (p = 0.36).
Patient Weight Analysis: Compared to the normal group, EIF4A1 protein expression was significantly elevated in cancer patients across all weight categories: Normal Weight (p = 2.55e-11), Overweight (p = 3.32e-07), Obese (p = 4.30e-09), and Extremely Obese (p = 0.027; Supplemental Figure 2(f)). Specifically, EIF4A1 protein expression in Normal Weight was markedly higher than in other weight categories (Supplemental Figure 2(f)). Compared to Overweight, EIF4A1 protein expression was significantly higher in the Normal Weight category (p = 0.0148), demonstrating statistical significance. Body weight appears to be a contributing factor to variations in EIF4A1 protein expression levels.
EIF4A1 interaction analysis
The EIF4A1 interaction network was constructed using the GeneMANIA database (Figure 5(a)). Eleven genes closely associated with EIF4A1 were identified via the STRING database, and the protein-protein interaction (PPI) network was constructed at a specified threshold, subsequently visualized via Cytoscape software (Figure 5(b)). Key genes in this network include EIF4G1, EIF4A1, EIF4E, EIF4B, EIF3B, EIF1, EIF4G2, EIF4H, EIF4G3, PABPC1, Notably, PDCD4. We intersected the molecular datasets from GeneMANIA and STRING to identify the most relevant gene (Figure 5(c)). Intersected interaction molecules were visualized and analyzed for expression in COADREAD using the ComplexHeatmap package (Figure 5(d)). Intriguingly, PDCD4 exhibited significantly elevated expression in paracancerous tissues.
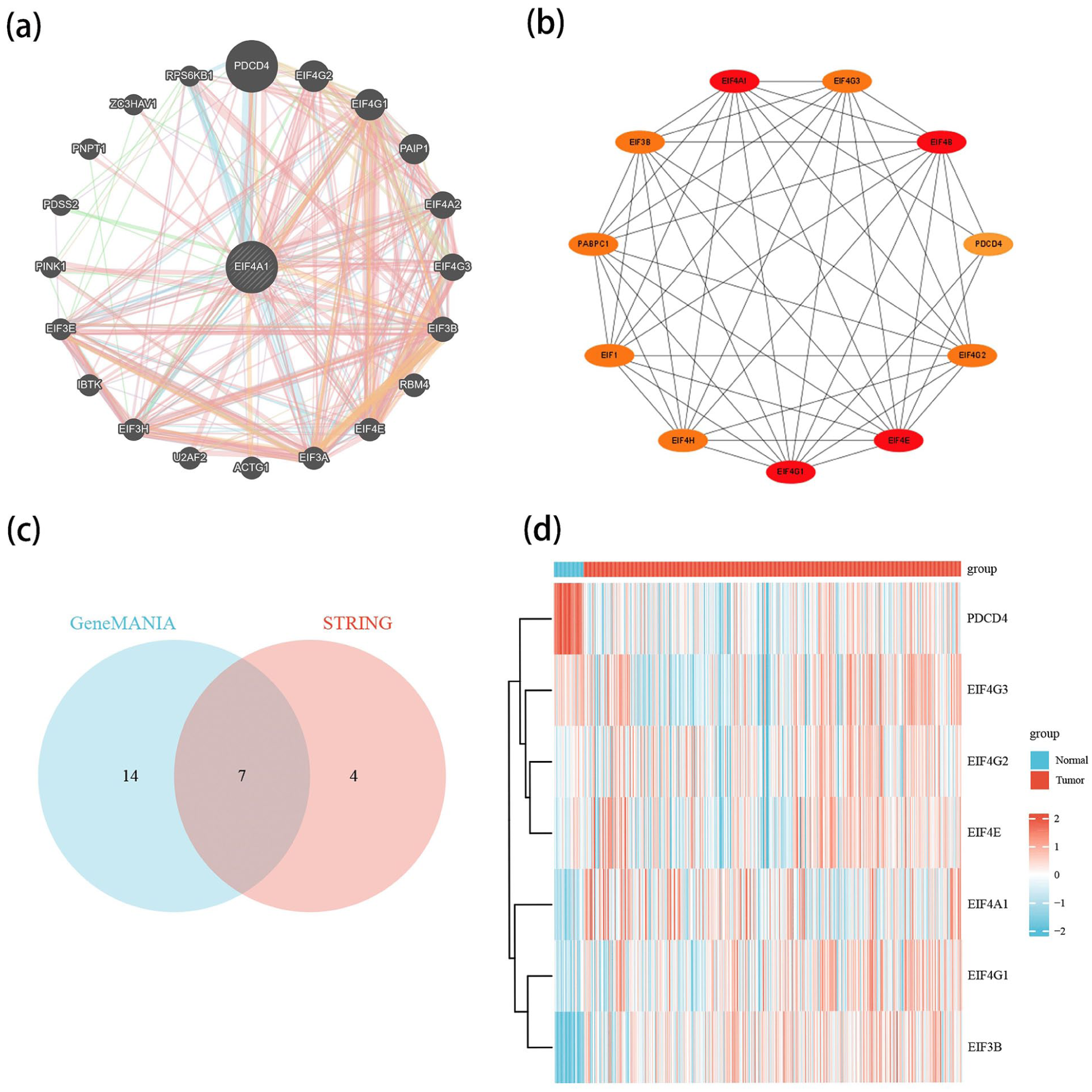
PPI network and related gene expression analysis of EIF4A1. (a) The interaction network of EIF4A1 in GeneMANIA. (b) String interaction network of EIF4A1. (c) Wayne diagram of the intersection of genemania and string datasets. (d) Co expression analysis of genemania and string intersection genes in colorectal cancer.
EIF4A1 enrichment of function analysis
The DESeq2 package was utilized to analyze differential gene expression TCGA-COAD, TCGA-READ datasets, and the original counts matrix. EIF4A1 expression in COAD and READ was categorized into high and low for differential expression analysis. Data analysis was conducted using ggplot2 (version 3.3.6), with set thresholds of logFC: 1 and p-value: <0.05. Results indicated that in COAD, 1038 genes were up-regulated and 13 were down-regulated; in READ, 117 genes were up-regulated and 150 were down-regulated. These findings were visualized in a volcano plot (Figure 6(a) and (b)). The differentially up-regulated genes from COAD and READ were sorted by logFC, and the top 100 were selected for GO/KEGG enrichment analysis. GO analysis of the EIF4A1-associated up-regulated genes in COAD revealed associations mainly with DNA, protein, and nucleosome processes (Figure 6(c)). KEGG analysis identified links primarily to NET (neurotrophil extracellular trap) formation, atherosclerosis, and systemic lupus erythematosus. GO analysis of differential gene sets in READ indicated relations to intermediary metabolism, receptor ligands, and hormonal activity (Figure 6(d)). KEGG enrichment analysis highlighted its primary association with the MAPK signaling pathway. Differential gene sets for EIF4A1 high and low expression groups in COAD and READ were identified using the DESeq2 package with a threshold of p < 0.05. These gene sets were subsequently analyzed for functional enrichment using GSEA. Parameters were set as follows based on the analysis results: p < 0.05, q-value < 0.25. Five pathways with the highest positive and negative NES correlations were selected for further analysis. GSEA analysis of COAD tissues revealed significant enrichment of differential genes in DNA replication, signal transduction, gene expression regulation, epigenetic regulation, cell cycle, chromosome dynamics, and gene silencing (Figure 6(e) and (f)). Notably, modifications such as methylation and acetylation were closely linked to cancer progression. Conversely, the top five pathways negatively correlated with NES predominantly involved energy metabolism, viral host responses, and mitochondrial function. According to GSEA analysis of READ tissues (Figure 6(g) and (h)), pathways exhibiting positive NES correlations were predominantly enriched in functions pertaining to immune response recognition, neural signal transmission, and sensory perception. Conversely, pathways with negative NES correlations were chiefly enriched in areas involving energy metabolism, immune and inflammatory responses, and cellular stress. The complement system and complement cascade are critical for immune surveillance and response, and their inhibition may contribute to the development and progression of cancer.

Relevant functional enrichment analysis of EIF4A1. (a–b) Volcano plot of EIF4A1 related differential genes in colon and rectal cancer. (c) KEGG and go analysis of EIF4A1 in colon cancer. (d) KEGG and go analysis of EIF4A1 in rectal cancer. (e–f) Top 5 pathway with positive and negative correlation in colon cancer. (g–h) Top 5 pathway with positive and negative correlation in rectal cancer.
Immune infiltration analysis
In COAD, EIF4A1 expression was significantly inversely correlated with tumor purity (correlation coefficient:cor = −0.163, p = 9.79e-04), indicating an increase in EIF4A1 expression as tumor purity decreased, which suggests enhanced immune cell infiltration. Furthermore, EIF4A1 expression in COAD demonstrated a strong positive correlation with the infiltration levels of CD8+ T cells (partial correlation:cor = 0.327, p = 1.34e-11), neutrophils (partial correlation:cor = 0.269, p = 4.38e-08), and dendritic cells (partial correlation:cor = 0.282, p = 8.80e-09; Figure 7(a) and (c)). In READ, a similar inverse correlation was observed between EIF4A1 expression and tumor purity (correlation coefficient:cor = −0.249, p = 3.02e-03). Contrary to COAD, in READ, EIF4A1 expression was negatively correlated with CD4+ T cell infiltration (partial correlation:cor = −0.21, p = 1.31e-02) and positively correlated with neutrophil infiltration (partial correlation:cor = 0.181, p = 3.39e-02), highlighting the diverse regulatory roles of different immune cells in the colorectal cancer microenvironment (Figure 7(b) and (c)).

Correlation between EIF4A1 expression and immune infiltration in colon and rectal cancer. (a) In the TIMER database, the infiltration scores for six accessible immune cell types (B Cells, CD4+ T Cells, CD8+ T Cells, Neutrophils, Macrophages, and Dendritic Cells) and the relationship between EIF4A1 expression and the degree of immune cell infiltration in colon and rectal cancer. *p < 0.05, **p < 0.01, ***p < 0.001. (b) Scatter plot of the correlation between EIF4A1 expression and CD8+ T Cell, Neutrophil, and DendriticCell in colon cancer. (c) Scatter plot of the correlation between EIF4A1 expression and CD4+ T Cell and Neutrophil in rectal cancer.
Correlation of EIF4A1 expression with immune checkpoint regulation and its implications for immunotherapy efficacy
Given the critical link between the expression of immune checkpoint genes and the efficacy of immunotherapy, we initially investigated the association between EIF4A1 expression and genes identified as immune response-related checkpoints via the TIMER 2.0 database. 26 In COAD, EIF4A1 demonstrates a strong correlation with most immune checkpoints, whereas in READ, the correlation with EIF4A1 is comparatively weaker (Figure 8(a)). Key immunosuppressive genes, including PD-L1 (CD274) and CTLA-4 (CTLA4), exhibit significant positive correlations with EIF4A1 across colon, colorectal, and rectal cancers (p < 0.001; Figure 8(a)). We identified correlations between EIF4A1 and several immune checkpoint blocker genes, including PD-1, PD-L1, CTLA4, and LAG-3. In both READ and COAD, EIF4A1 demonstrated significant positive correlations with CTLA4 and PD-L1, with these associations being more pronounced in colon cancer. Furthermore, in colon cancer, EIF4A1 expression along with PD-1, PD-L1, CTLA4, and LAG-3 was up-regulated, showing significant positive correlations. Conversely, in rectal cancer, the correlations between EIF4A1 and LAG-3, PD-1 were not statistically significant (Figure 8(b)–(e)).

Correlation of EIF4A1 expression on immune checkpoints and immunotherapy. (a) The correlation of EIF4A1 with more than 40 immune checkpoint genes in pan-cancer (*p < 0.05, **p < 0.01, ***p < 0.001). (b–e) The correlation of EIF4A1 with PD-1, PD-L1, CTLA-4, and LAG-3 in the TIMER 2.0 database. (f–g) In colorectal cancer, as well as in colon and rectal cancer, the analysis of the correlation between the expression of EIF4A1 and microsatellite instability (MSI) and tumor mutational burden (TMB).
Recent research has demonstrated that high microsatellite instability (MSI-H) and tumor mutation burden (TMB) are promising predictive biomarkers for the efficacy of immunotherapy, with MSI identified in a diverse range of cancers.27,28 The presence of MSI signifies an unstable intracellular genomic state with the potential to precipitate cancer development. Specifically, tumors with high microsatellite instability (MSI-H) exhibit extensive genomic instability, associated with genetic disorders like Lynch syndrome, a hereditary form of colorectal cancer. We investigated the correlation between Tumor Mutation Burden (TMB) and microsatellite instability (MSI) of EIF4A1 across COADREAD, COAD, and READ. The findings revealed that EIF4A1 expression in COADREAD, COAD, and READ positively correlates with both TMB and MSI. Notably, this correlation was more pronounced in COAD than in READ (Figure 8(f) and (g)).
IHC and Hehematoxylin and Eosin staining (HE)
Comparison with paracancerous tissues shows that in colorectal cancer sections, cellular anomalies are remarkable, manifested by cell size and shape irregularities, increased nucleoplasmic ratio, uneven cytoplasmic staining; cellular nuclei have prominent anomalies, with irregular sizes and shapes, and some of them are deeply stained and enlarged, and the genetic material may be altered. The glandular structure is severely damaged, with irregular morphology, a large number of fusion or disappearance, forming irregular cell clusters; the tissue structure is disordered, the cells are arranged in a chaotic manner, the boundary is unclear, the cancer cells are densely packed in solid clusters, and the interstitium is few and loosely connected, which is highly invasive and highly malignant (Figure 9(a)–(h)). Comparative analysis revealed distinct immunohistochemical profiles between colorectal carcinoma subtypes. In paracancerous tissues of colon adenocarcinoma patients, significantly weaker immunoreactivity was observed compared to matched normal tissues from rectal carcinoma cases (Figure 10(a), (c), (e), and (g)). Paracancerous areas exhibited uniformly distributed faint staining with minimal regional heterogeneity. Malignant tissues from both cancer types demonstrated marked intratumoral heterogeneity, characterized by alternating foci of intense brownish-black deposits (indicative of high protein density) and pale-brown zones (Figure 10(b), (d), (f), and (h)). Histopathological examination revealed disorganized tissue architecture with loss of normal glandular structure. Overall, EIF4A1 expression in colorectal cancer specimens showed a non-uniform distribution pattern, displaying strong cytoplasmic staining associated with high levels of EIF4A1 protein expression. In contrast, neighboring paracancerous tissues showed predominantly negative to weak immunoreactivity with weak staining (Figure 10(a)–(h)).

HE staining images of cancer and adjacent tissues. (a–d) HE staining images of colon cancer and adjacent tissues. (e–h) HE staining images of rectal cancer and adjacent tissues.

Immunohistochemical staining images of colorectal cancer tissues and adjacent non-tumor tissues. Panels (a–d) show the staining patterns in colorectal cancer tissues and adjacent non-tumor tissues from colon cancer patients. Panels (e–h) illustrate the staining patterns in colorectal cancer tissues and adjacent non-tumor tissues from rectal cancer patients.
Discussion
The proliferation of molecular diagnostics and immune-targeted therapies has accelerated the global adoption of precision medicine, particularly, these advancements have significantly enhanced therapeutic outcomes in colorectal cancer, with the development of immune checkpoint inhibitors demonstrating notable clinical efficacy.29–31 Clinically, the diagnosis of colorectal cancer predominantly depends on methods such as colorectal colonoscopy and CT scans, which are invasive. The distinct anatomical locations of the colon and rectum suggest that treatment approaches may vary. Additionally, the site of the tumor within the colorectal region is correlated with survival outcomes. 32 Therefore, identifying target molecules specific for molecular diagnosis and treatment in colon and rectal cancers is crucial, promising to significantly enhance precision medicine.
EIF4A1, a member of the eukaryotic initiation factor family, is a core component of the EIF4F complex and is crucial for the initiation of translation. Given EIF4A1 central role in regulating translation and the link between mRNA translation dysregulation and cancer progression, it has garnered significant attention. EIF4A1 is a potent and effective RNA helicase. Leveraging this property, researchers have developed inhibitors targeting the eIF4 family, achieving promising preclinical results in human solid tumors. 33 The EIF4F complex, crucial for translation initiation, serves as a nexus for cancer-promoting signaling pathways. Its inhibition reduces the expression of key oncoproteins and induces apoptosis, with EIF4A1 functioning as a critical enzymatic active center within this complex. Research indicates that targeting EIF4A1 and the EIF4F complex offers promising avenues for developing and exploring tumor suppressive strategies.21,34
Through a literature review, we identified the RBPs associated with EMT progression and EIF4A1, a molecule critical for prognostic stratification. 25 Given its crucial role in translation regulation, EIF4A1 became the focal point of our study. Employing bioinformatics, we examined EIF4A1 expression in colon and rectal cancers and its correlation with molecular mechanisms, biological functions, immune infiltration, and immune checkpoints. We mapped the protein expression landscape of EIF4A1 across human organs, which revealed its expression in normal colorectal tissues and colorectal cancer pathological samples, Utilizing the TCGA database, we analyzed its mRNA expression levels in pan-cancer, colorectal, colon, and rectal cancers, and characterized its predominant cellular localization in the cytoplasm.
The findings demonstrated that EIF4A1 mRNA exhibited upregulation across most cancer types, with both mRNA and protein levels elevated in colon cancer. EIF4A1 mRNA levels were significantly higher in COAD versus rectal cancer. Immunofluorescence analysis revealed that EIF4A1 was predominantly localized to the cytoplasm and enriched in cancer cells. These observations suggest that the up-regulation of EIF4A1 is linked to tumor progression. Furthermore, we assessed the correlation between EIF4A1 mRNA and protein expression and clinical features. Analysis indicated that EIF4A1 mRNA expression was primarily observed in stages I and II of colon cancer, and stage I of rectal cancer, with comparatively lower levels in stage II. In both colon and rectal cancers, EIF4A1 mRNA levels were notably higher in younger patients, potentially linking younger onset of cancer with increased expression. In colon cancer, the highest levels of EIF4A1 mRNA expression were observed in N0 staging, while in rectal cancer, although peak levels were noted in N1 staging, these were not statistically significant.
Contrary to expectations, we observed that EIF4A1 mRNA and protein expression levels were higher in individuals of normal weight and tended to decrease slightly with increasing body weight in both COAD and READ cases. Analysis revealed that EIF4A1 protein expression was associated with tumor stage, age, histological type, body weight, and gender. EIF4A1 protein expression was predominantly observed in stages II and III, with peak expression in mid-to-late disease stages. This feature allows for the detection and assessment of the patient’s condition. High EIF4A1 expression predominantly occurs in individuals aged 61–80 years, indicative of a tendency toward middle-aged and elderly patients and associated with a poorer survival prognosis. EIF4A1 protein expression in normal-weight individuals was significantly higher compared to their overweight counterparts. Subsequently, we assessed EIF4A1 diagnostic. We observed that high expression of EIF4A1 may be associated with poorer prognosis in colon and rectal cancers. Clinical ROC curve analysis showed that EIF4A1 in colorectal cancer, colon cancer and rectal cancer were AUC: 0.827 AUC: 0.836, AUC: 0.797. The findings suggest that increased EIF4A1 expression significantly contributes to the onset and progression of colorectal cancer and may serve as a novel clinical diagnostic marker. Notably, EIF4A1 diagnostic utility exhibits greater diagnostic efficacy in colon cancer versus rectal cancer. Following comprehensive univariate and multivariate Cox regression analyses, it is clear that although colon and rectal cancers share critical prognostic factors such as the significant roles of pathological staging and age, they differ markedly in T, N, M stages, and biomarkers like CEA levels and EIF4A1 expression. These variations likely arise from the distinct biological behaviors and treatment responses inherent to each cancer type. Consequently, in clinical decision-making, it is crucial for physicians to carefully evaluate the specific risk factors and prognostic indicators to develop more personalized and precise treatment strategies. Survival and ROC curve analyses of EIF4A1 indicated that its expression correlates with survival outcomes and diagnostic accuracy, suggesting its potential as a prognostic biomarker and diagnostic indicator.
The PPI interaction network of EIF4A1 was constructed through the database download data and online data, and the expression relationship of the genes whose interaction intersection was analyzed in colorectal cancer. Analysis revealed that PDCD4 was negatively correlated with the expression of other related genes, and PDCD4 was significantly expressed in normal tissues. PDCD4 is one of the genes related to programmed cell death, which encodes a tumor suppressor protein that inhibits tumor cell growth and proliferation through multiple mechanisms. There was a high correlation between elevated EIF4A1 expression and reduced PDCD4 expression. 35 Through conducting a functional enrichment analysis of differential genes related to EIF4A1, we identified that in COAD, key pathways are primarily involved in DNA replication, signal transduction, gene expression regulation, epigenetic regulation, cell cycle, chromosome dynamics, gene silencing, developmental regulation, and energy metabolism and synthesis. Notably, processes such as neutrophil extracellular trap formation, and modifications like methylation and acetylation, are critically associated with cancer development, immune escape, and response. In rectal cancer, the primary enriched pathways include immune recognition, immune and inflammatory responses, cell stress, and energy metabolism. The complement system and the MAPK signaling cascade are fundamental to immune surveillance and response, and are deeply intertwined with the progression and development of cancer. Inhibiting these pathways could potentially influence the development and progression of cancer.
Finally, an investigation was conducted into the relationship between EIF4A1 and immunity. By analyzing immune infiltration, it was observed that colon cancer showed a strong positive correlation with neutrophils, CD8+ T cells, and dendritic cells, whereas rectal cancer was positively correlated with neutrophils and negatively correlated with CD4+ T cells. Interestingly, consistent with earlier KEGG analyses in colon cancer, EIF4A1 demonstrated a significant positive correlation with neutrophil infiltration. A crucial pathogen defense mechanism—the hijacking of neutrophils for the formation of neutrophil extracellular traps (NETs)—has been implicated in cancer progression. Studies have shown that NETs promote CRC metastasis through potent immunosuppression. 36 Furthermore, specific components of NETs, including histones and elastases, have been shown to induce angiogenesis, thus supplying nutrients and oxygen to tumors. This leads to vascular dysfunction and thrombosis, thereby contributing to increased mortality risk in cancer patients.37,38 Elevated expression and infiltration of neutrophils have been implicated in promoting cancer progression. In certain scenarios, neutrophils may transform into cells with immunosuppressive capacities, notably myeloid-derived suppressor cells (MDSCs). Increasing evidence supports the role of neutrophils as key mediators in creating an immunosuppressive environment favorable for tumor progression across multiple cancer types, and elevated EIF4A1 expression may facilitate this aggregation of immunosuppressive cells, thereby aiding tumor evasion of immune surveillance.39,40
CD8 cells, which can directly recognize and kill tumor cells, mediate anti-tumor immunity through multiple mechanisms, including inhibition of tumor growth and metastasis, and release of cytolytic agents such as perforin, granzyme, and major histocompatibility complex I (MHC I)-presented antigens to eliminate target cells. Additionally, EIF4A1 expression is positively correlated with CD8 T cells; this correlation may promote the inflammatory environment in cancer and reduce the efficacy of CD8+ T cells by inducing overexpression of PD-L1. Although infiltration of CD8+ T cells increases, their activity may be suppressed. This suppression likely reflects an adaptive immune resistance mechanism within the complex tumor microenvironment.41–44 Dendritic cells are pivotal for antigen presentation, primarily offering Major Histocompatibility Complex I (MHC I) molecules to present antigens to CD8+ T cells, thus activating these cells. 45 EIF4A1 is positively correlated with dendritic cells and CD8+ T cells. While this gene’s expression might theoretically promote the immune response. Within the complex tumor immune microenvironment, EIF4A1 expression, which promotes cancer progression, may enable tumor cells to regulate antigen expression and alter antigen processing and presentation mechanisms, thus evading immune recognition. Despite high expression and infiltration of CD8+ T cells and dendritic cells, elevated EIF4A1 expression may be associated with a complex tumor microenvironment. Factors such as impaired function or maturity of dendritic cells may contribute to the progression of cancer.46,47
In rectal cancer, EIF4A1 is predominantly negatively correlated with CD4+ T cells. Elevated EIF4A1 expression is associated with a complex tumor microenvironment.48–50 Based on the current research landscape, while definitive evidence directly elucidating the complete mechanisms by which EIF4A1 interacts with CD4+ T cells, CD8+ T cells, and related immune cells within the tumor immune microenvironment (TIME) remains lacking, its established role in regulating tumor cell biological behavior suggests that EIF4A1 likely interacts with immune cells in the TIME through multiple avenues. These may include influencing the intrinsic antigenic properties of tumor cells, modulating the secretion of signaling molecules within the immune microenvironment, and interfering with key signaling pathways in immune cells, thereby playing a significant role in the formation of the immunosuppressive microenvironment. Future research urgently needs to delve deeper into the specific molecular mechanisms of EIF4A1 within the TIME. For instance, employing gene editing technologies to precisely modulate EIF4A1 expression could reveal its direct impact on immune cell function. Alternatively, utilizing cutting-edge techniques such as single-cell sequencing could dissect the relationship between EIF4A1 expression and the transcriptomic and proteomic profiles of immune cells in the tumor microenvironment.
The expression of immune checkpoint genes is associated with tumor cells evasion of immune surveillance. Several immune checkpoint inhibitors have been incorporated into therapeutic research and exhibited substantial efficacy.51,52 Immune checkpoint genes, including PD-1 (PDCD1), PD-L1 (CD274), CTLA-4 (CD152), and LAG-3, are involved in tumor cells’ evasion of immune surveillance. 53 Subsequent analysis was conducted to examine the correlation between EIF4A1 expression and various immune checkpoint-related genes. The analysis revealed that EIF4A1 was strongly correlated with several key immune checkpoints in colon cancer, notably PD-1 (PDCD1), PD-L1 (CD274), CTLA4 (CD152), and LAG-3. The correlation between EIF4A1 expression and these immune checkpoints was examined using the correlation between EIF4A1 expression and these immune checkpoints. The expression levels of PD-1 (PDCD1), PD-L1 (CD274), CTLA4 (CD152), and LAG-3 were significantly up-regulated in colon cancer. Further, these checkpoint expressions, which were significantly correlated with CD8+ T cell infiltration in colorectal cancer, suggest that elevated EIF4A1 levels may contribute to T cell activation inhibition, impairing T cell functionality and potentially fostering cancer progression. In rectal cancer, however, only the up-regulation of PD-L1 and CTLA4 was statistically significant, whereas PD-1 up-regulation was not significant. Additionally, the up-regulation of PD-L1 and CTLA4 may also relate to the activation and infiltration of CD4+ T cells, potentially inhibiting T cell activation and thus promoting cancer. 52 In treating colon and rectal cancers, treatment strategies should be tailored to the expression of immune checkpoint inhibitors such as PD-1, PD-L1, LAG-3, and CTLA4. Further exploring the relationship between EIF4A1 and these immune checkpoint genes in both cancer types will enable us to better personalize treatment strategies and minimize immune tolerance and resistance.
This study only focused on tissue samples from pathological conditions such as colon cancer and rectal cancer, and was unable to comprehensively explore the impact of the disease on the morphological and molecular expression continuity at the junction of the sigmoid colon and rectum. In the future, emphasis will be placed on the characteristic changes in the forms at the junctions. This study highlights EIF4A1 potential as a diagnostic and prognostic biomarker for colorectal cancer. However, limitations exist, such as the relatively small sample size, which necessitates further validation using larger independent datasets. While correlations between EIF4A1 expression and various clinical/molecular features have been identified, the underlying mechanisms warrant further mechanistic investigation. The clinical data used in this study were obtained from the TCGA database, which may have inherent limitations in terms of data completeness and accuracy. Some information related to postoperative survival as well as detailed treatment histories may be missing or incomplete, which may affect survival analysis and assessment of the correlation between EIF4A1 expression and prognosis. Second, our study was mainly based on bioinformatics analysis and retrospective data analysis, and these methods have limitations in establishing a direct causal relationship between EIF4A1 expression and prognosis in colon and rectal cancer. Although we observed a significant correlation, further experimental and clinical studies are needed to confirm this relationship and explore the underlying mechanisms. Finally, the treatment modalities of the patients included in this study may be varied and complex. Different chemotherapy regimens and other therapeutic factors may have a significant impact on prognosis. Looking ahead, it is anticipated that significant advancements in this field. In the treatment of gastrointestinal tumors, adverse drug reactions should be closely monitored, exploring the dose individualization of regorafenib and other drugs, minimizing selection bias in clinical trials and establishing an early monitoring mechanism 4 ; immunotherapy should strengthen the monitoring and management of irAEs in order to optimize the regimen. 5 Prospective studies should focus on special populations and the efficacy and safety of combination therapy. Clinical treatment should integrate metabolic modulation and anti-inflammatory interventions, incorporate lifestyle interventions into the prevention and treatment system, and take into account gender and age differences. 6 The Royal Marsden Hospital score can stratify patient prognosis, provide a universal prognostic tool for a wide range of tumors, and can be combined with molecular markers to optimize precision therapy. 7 The neutrophil-eosinophil ratio could be a new prognostic indicator for tumors, with potential for use in immunotherapy and gastric oncology practice, and a standardized cut-off value should be defined to facilitate the translation of precision therapeutic strategies. Over the next 5 years, we expect more in-depth studies on EIF4A1’s role in cancer biology and its therapeutic potential. With technological and methodological advancements, we aim to bridge knowledge gaps and translate these findings into more effective clinical applications for colorectal cancer patients.
Conclusion
Through bioinformatics analysis, we examined the differences in EIF4A1 expression, clinical parameters, and immune infiltration between colon and rectal cancers. We discovered that EIF4A1 is upregulated in both cancers, suggesting its potential as a diagnostic marker. Notably, EIF4A1’s association with survival prognosis and clinical outcomes is more statistically significant in colon cancer than in rectal cancer. Regarding immunity, EIF4A1 exhibits distinct infiltration patterns in colon and rectal cancers, positioning it as a viable target for immunotherapy. However, the immunological differences between colon and rectal cancers necessitate tailored therapeutic strategies for each.
Materials and methods
Screening for intersecting genes
RNA-seq data from the TCGA database were analyzed, and DESeq2 was utilized to identify differentially expressed genes in the TCGA-COAD and TCGA-READ datasets. 54 With a threshold of |logFC| > 1 and P.adj < 0.05, we discovered 2929 genes with significant expression differences. Additionally, univariate Cox regression analysis was performed using the R packages survival and survminer, 55 and 631 prognostic genes associated with colorectal cancer were identified based on the criteria of HR (Hazard Ratio) > 1 and P.adj < 0.05. The overlap between these gene sets and the RBP library 25 was graphically visualized using ggplot2 and VennDiagram in R. 25
Analysis of differentially expressed mRNA (DEmRNA)
Using the TCGA-COADREAD dataset, differential mRNAs were identified through R-based analysis with parameters setting the log2 fold change (|logFC|) > 1 and P.adj < 0.05.The results of the differential analysis were visualized using the R package ggplot2.
Gene co-expression analysis
RNAseq data from TCGA database were downloaded and processed using the STAR pipeline of TCGA-COAD and TCGA-READ projects and extracted in TPM (Transcripts Per Kilobase Million) format, data processing method: log2(value+1), using R language to visualize the clinical Pathologic T stage, Pathologic N stage, Pathologic M stage with gene expression analysis and heatmap visualization by R package: ComplexHeatmap. 56
Analysis of EIF4A1 expression
HPA database (https://www.proteinatlas.org) was used to investigate the mRNA and protein expression levels of EIF4A1 in normal human tissues. The expression levels of the EIF4A1 gene in a variety of cancer tissues were retrieved through the “Gene_DE” module in TIMER 2.0 (http://timer.cistrome.org/). 57 RNA sequencing data from the TCGA-COAD and TCGA-READ cohorts within the TCGA database were rigorously curated and normalized to Transcripts Per Million (TPM) values. Paired samples of tumor and adjacent non-tumor (paracancerous) tissues were specifically extracted and validated for numerical correspondence.. The expression profiles of the EIF4A1 gene in colorectal adenocarcinoma (COAD) and rectal adenocarcinoma (READ) tissues were compared with their paracancerous counterparts to identify differential expressed patterns. A paired-sample t-test was employed to statistically evaluate the expression levels of EIF4A1 within these paired samples. The resultant data were visualized using the R software environment, with graphical representation facilitated by the ggplot2 and stats packages for graphical representation.
Survival analysis
RNA-seq data from the TCGA database were downloaded and organized for the STAR process of the TCGA-COAD and TCGA-READ projects. The data were extracted in TPM format, along with the corresponding clinical data. Survival analysis of EIF4A1 in COADREAD, COAD, and READ was conducted using Cox regression with the R survival package to test the proportional risk hypothesis and fit survival regression models to determine p-values. Comparisons of EIF4A1 gene expression were made between Overall Survival (OS), Disease-Specific Survival (DSS), and Progression-Free Interval (PFI). The results were visualized using the R packages survminer and ggplot2. 55 Forest plots displaying hazard ratios (HR), 95% confidence intervals, and P-values of survival curves were generated and visualized using the R packages survminer and ggplot2.
ROC analysis of EIF4A1
ROC (Receiver Operating Characteristic) curves were employed to assess the diagnostic value of EIF4A1 in COADREAD, COAD, and READ. Data for constructing ROC curves were derived from the mRNA expression of EIF4A1 in cancer and corresponding normal tissues in the TCGA database. ROC curves were calculated using the pROC (v1.17.0.1) package in R software and plotted with the ggplot2 package. The area under the curve (AUC), cutoff value, sensitivity, specificity, positive predictive value, negative predictive value, and Youden index (YI) were calculated. AUC values closer to 1 indicate better diagnostic performance; an AUC of 0.5–0.7 indicates low precision, 0.7–0.9 indicates better accuracy, and above 0.9 indicates high accuracy.
Association of EIF4A1 transcript and protein expression with clinicopathological characteristics in cancer
The associations between transcript levels, protein levels, and various clinicopathological parameters (cancer stage, lymph node metastasis, age, overweight, gender, etc.) of EIF4A1 in colorectal cancer were analyzed using the UALCAN (https://ualcan.path.uab.edu/) web resource with data from the TCGA database. 58
Protein-protein interaction (PPI) network analyses of EIF4A1
The GeneMANIA database (https://www.genemania.org) was employed to reconstruct a protein-protein interaction (PPI) network for EIF4A1, 59 while the STRING online database was utilized to generate an additional network incorporating EIF4A1. 60 The proteins obtained from both sources were taken as intersections and visualized using ggplot2 [3.3.6] and the VennDiagram package [1.7.3] for Venn diagram plotting. The most relevant genes identified at the intersections were visualized using the R package ComplexHeatmap [2.13.1] to depict the expression patterns of EIF4A1-related genes in the TCGA-COADREAD dataset. 56
Functional enrichment analysis of EIF4A1-related differential genes
To elucidate the function of EIF4A1, EIF4A1 associated differential expressed genes (DEGs) were s filtered and visualized using the ggplot2 package. The top 100 up-regulated genes from the volcano plot were subjected to Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses in TCGA-COAD and READ. Gene Set Enrichment Analysis (GSEA) was performed using the full set of DEGs identified from the volcano plot. The clusterProfiler package was used to automate the GO and KEGG enrichment workflows, 61 and the results were visualized as bubble plots and mountain plots using “ggplot2.”
Immunoassay of EIF4A1 in COAD and READ
Correlation data were downloaded from the TIMER 2.0 (http://timer.cistrome.org/) database, cleaned, and visualized using the ggplot2 package for heatmap representation. Correlation scatter plots of EIF4A1 with related immune cells were visualized via the TIMER database (https://cistrome.shinyapps.io/timer/). Expression and correlation data between EIF4A1 and related checkpoint genes were organized from TIMER 2.0 and visualized as heatmaps using the ggplot2 package. Data on EIF4A1 and four immune checkpoints obtained from TIMER 2.0 were organized and visualized as dot plot using the ggplot2 package. The harmonized and standardized pan-cancer dataset TCGA Pan-Cancer (PANCAN.N = 10535, G = 60499) was downloaded from the UCSC (https://xenabrowser.net) database. Expression data of the EIF4A1 gene in COAD and READ were extracted, and samples derived from Primary Blood Derived Cancer and Peripheral Blood were screened. Additionally, samples processed by MuTect2 software were downloaded from GDC (https://portal.gdc.cancer.gov/), providing a Simple Nucleotide Variation dataset of all TCGA samples at level 4. The Tumor Mutation Burden (TMB) of each tumor was calculated using the R package maftools (version 2.8.05). TMB and gene expression data were integrated, followed by log2(x+0.001) transformation for each expression value. The data were then cleaned, organized, and visualized as radar plots using ggplot2. 62
Immunohistochemistry (IHC) and Hematoxylin and Eosin (H&E) staining
This study was approved by the Ethics Committee of the First Affiliated Hospital of Hebei North University (Approval No. K2024147). Cancer tissues and adjacent para-cancerous tissues were collected from colorectal adenocarcinoma patients in the Department of Vascular and Glandular Surgery at the institution. Following fixation in 4% paraformaldehyde, specimens were paraffin-embedded and sectioned into 4-μm slices. After standard dewaxing, rehydration, and antigen retrieval procedures, tissue sections were incubated with primary antibodies (catalog #250072, Zhengneng Biotechnology, Chengdu, China; dilution 1:5000) followed by chromogenic development using a 3,3′-diaminobenzidine (DAB) detection kit.
Statistical analysis
Student’s t-test was used to compare the differences between the two groups, and data are reported as mean values ± SD. The p-value < 0.05. Statistical analyses were performed using GraphPad Prism version 9.
Supplemental Material
sj-docx-1-iji-10.1177_03946320251379175 – Supplemental material for Translation element EIF4A1 is a potential divergent immune biomarker between colon cancer and rectal cancer
Supplemental material, sj-docx-1-iji-10.1177_03946320251379175 for Translation element EIF4A1 is a potential divergent immune biomarker between colon cancer and rectal cancer by Zhenpeng Zhu, Peng Wang, Chenyang Hou, Jiajia Xiao, Fengxu Yan, Shan Liu, Xiran Wang, Xuejun Zhi, Jun Xue, Dandan Xu, Fei Guo and Weizheng Liang in International Journal of Immunopathology and Pharmacology
Supplemental Material
sj-docx-2-iji-10.1177_03946320251379175 – Supplemental material for Translation element EIF4A1 is a potential divergent immune biomarker between colon cancer and rectal cancer
Supplemental material, sj-docx-2-iji-10.1177_03946320251379175 for Translation element EIF4A1 is a potential divergent immune biomarker between colon cancer and rectal cancer by Zhenpeng Zhu, Peng Wang, Chenyang Hou, Jiajia Xiao, Fengxu Yan, Shan Liu, Xiran Wang, Xuejun Zhi, Jun Xue, Dandan Xu, Fei Guo and Weizheng Liang in International Journal of Immunopathology and Pharmacology
Supplemental Material
sj-jpeg-3-iji-10.1177_03946320251379175 – Supplemental material for Translation element EIF4A1 is a potential divergent immune biomarker between colon cancer and rectal cancer
Supplemental material, sj-jpeg-3-iji-10.1177_03946320251379175 for Translation element EIF4A1 is a potential divergent immune biomarker between colon cancer and rectal cancer by Zhenpeng Zhu, Peng Wang, Chenyang Hou, Jiajia Xiao, Fengxu Yan, Shan Liu, Xiran Wang, Xuejun Zhi, Jun Xue, Dandan Xu, Fei Guo and Weizheng Liang in International Journal of Immunopathology and Pharmacology
Supplemental Material
sj-png-4-iji-10.1177_03946320251379175 – Supplemental material for Translation element EIF4A1 is a potential divergent immune biomarker between colon cancer and rectal cancer
Supplemental material, sj-png-4-iji-10.1177_03946320251379175 for Translation element EIF4A1 is a potential divergent immune biomarker between colon cancer and rectal cancer by Zhenpeng Zhu, Peng Wang, Chenyang Hou, Jiajia Xiao, Fengxu Yan, Shan Liu, Xiran Wang, Xuejun Zhi, Jun Xue, Dandan Xu, Fei Guo and Weizheng Liang in International Journal of Immunopathology and Pharmacology
Footnotes
Acknowledgements
We thank The First Affiliated Hospital of Hebei North University for their approval of the study. We thank Bioimaging Core of Shenzhen Bay Laboratory for providing imaging support. We also would like to acknowledge Bioimaging Core engineers Zhao Huijuan and Ren Xiaodie for assistance with the samples preparation. This study was supported by the Zhangjiakou City Key R&D Plan Project (No.2322088D, No.2421118D, and No.2311038D), Medical Science Research Subject Plan Project of Hebei Provincial Health Commission (No.20240805), Natural Science Project of Hebei North University (No.XJ2024034 and No.XJ2024035), 2025 Government-funded Training Project for Outstanding Clinical Medicine Talents (ZF2025264), Hebei Health Commission Scientific Research Foundation Project (No.20240240), Hebei Provincial Administration of Traditional Chinese Medicine Research Project (No.2024062), Hebei Provincial Administration of Traditional Chinese Medicine Project (No.2022147 and No.2023331), Central Government Guides Local Science and Technology Development Fund Projects (No.246Z7768G), Hebei Provincial Administration of Traditional Chinese Medicine Project (No.2025392 and No.2025395), Hebei North University 2024 school-level cultivation research project (XJPY2024026), Medical Science Research Subject Plan Project of Hebei Provincial Health Commission (No.20240782), Hebei Province Innovation Capacity Enhancement Program Project (No.24457711K), The Provincial Funding Project for Cultivating the Innovation Ability of Postgraduate Students Reading in 2025 (CXZZSS2025120), 2025 Hebei Province Graduate Education Project (CX202410), Research Project of Medical Innovation for Chinese Youth (Project Leader: Jun Xue). Research Project of Medical Innovation for Chinese Youth (Project Leader: Xuejun Zhi).
Author contributions
Weizheng Liang: Conceptualization, funding acquisition. Jun Xue, Fei Guo and Dandan Xu: Supervision, funding acquisition. Zhenpeng Zhu and Shan Liu: Data curation, Writing-Original draft preparation. Peng Wang and Xiran Wang: Methodology, Software. Chenyang Hou and Fengxu Yan: Writing—Reviewing and Editing. Jiajia Xiao and Xuejun Zhi: Software, Validation, funding acquisition. Fei Guo and Dandan Xu: Supervision. All authors have reviewed the results and approved the final version of the manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by Medical Science Research Subjects of Hebei Provincial Health and Wellness Commission (No.20250144).
Ethics approval and consent to participate
Ethical approval for this study was obtained from the Ethics Committee of the First Affiliated Hospital of Hebei North University (APPROVAL NUMBER/K2024147).
Informed consent
Written informed consent was obtained from all subjects prior to the study.
Data availability statement
The datasets presented in this study can be found in the online repositories, including The Human Protein Atlas (HPA) database (https://www.proteinatlas.org), TIMER 2.0 (http://timer.cistrome.org/), UALCAN (https://ualcan.path.uab.edu/), STRING (https://cn.string-db.org/), and TIMER database (https://cistrome.shinyapps.io/timer/), GeneCards (https://www.genecards.org), GeneMANIA database (http://genemania.org/), UCSC (![]() ).
).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
