Abstract
In assessing the effectiveness of DNA vaccines, it is important to monitor: (1) the kinetics of target gene expression in vivo; and (2) the movement of cells that become transfected with the plasmid DNA used in the immunization of a subject. In this study, we used, as a visual imaging marker, expression of the transfected human sodium/iodide symporter (hNIS) gene, which enhances intracellular radio-pertechnetate (TcO4–) accumulation. After intradermal (i.d.) and systemic injection of mice with pcDNA-hNIS and radioactive Technetium-99m (Tc-99m), respectively, whole-body images were obtained by nuclear scintigraphy. The migration of mice cells transfected with the hNIS gene was monitored over a 2-week period by gamma-radioactivity counting of isolated cell populations and was demonstrated in peripheral lymphoid tissues, especially in the draining lymph nodes (dLNs). Beginning at 24 h after DNA inoculation and continuing for the 2-week monitoring period, hNIS-expressing cells were observed specifically in the T-cell–rich zones of the paracortical area of the dLNs. Over the same time period, high levels of INF-γ–secreting CD8 T-cells were found in the dLNs of the pcDNA-hNIS immunized mice. Tumor growth was also significantly retarded in the mice that received hNIS DNA immunization followed by inoculation with CT26 colorectal adenocarcinoma cells that had been transfected with the rat NIS gene (rNIS), which is 93% homologous to the hNIS gene. In conclusion, mouse cells transfected with hNIS DNA after i.d. immunization were found to traffic to the dLNs, and hNIS gene expression in these cells continued for at least 2 weeks post immunization. Furthermore, sequential presentation of NIS DNA to T-cells by migratory antigen presenting cells could induce NIS DNA-specific Th1 immune responses and thus retard the growth of NIS-expressing tumors.
Introduction
In basic and clinical research approaches to cancer immunotherapy, a tumor vaccine is defined as one that increases specific immune responses to tumor antigens. 1 Early studies of DNA vaccines all emanate from a seminal study by Wolff et al. that characterized the in vivo effects of plasmid or naked DNA injection. 2 This study provides a strong basis for the belief that purified/recombinant nucleic acids (naked plasmid DNA) can be delivered by intramuscular inoculation and that these nucleic acids can direct protein expression in vivo. Given the relatively small amounts of protein likely to be synthesized as a result of immunization with DNA, probable explanations for the efficient induction of broad-based and sustained immune responses are likely related to the immune-enhancing properties of DNA (i.e. CpG motifs) and/or the type of antigen presenting cells (APCs) that are transfected with the injected DNA.3,4 Subsequent studies in mice have documented that both humoral and CD8+ cytotoxic T-lymphocyte (CTL) responses are elicited by DNA injection suggesting that these immune functions play a role in tumor cell abrogation.1–4 Specifically, tumor cell death is achieved by introducing a tumor specific DNA immunogen, which induces the production of Type 1 T helper cell (Th1) cytokines (such as tumor necrosis factor) and the resulting expansion of tumor-specific CTLs.1–4
The ability to monitor the in vivo kinetics of expression of the injected tumor genes is crucial for mechanistic studies of DNA-based vaccines, and several groups5–8 are currently working to achieve efficient in vivo DNA transfection using a number of different vectors. To date, the methods used to determine the efficacy of DNA vaccines have relied on evidence of T-cell−mediated immune responses.1–4 However, these assays provide only a snapshot in time and space, and do not reflect ongoing immune response plasticity. The development of a simple and reproducible method for detecting in vivo gene expression is thus critical for assessing the delivery of transgenes and the timeframe of their expression. To this end, whole-body non-invasive monitoring methods must be developed to track the spatial distribution of antigen-specific immune cells at any given time after immunization.
Reporter genes have been widely used to monitor transgene expression in various systems.9–11 One such reporter is the sodium/iodide symporter gene (NIS), which encodes an integral plasma membrane glycoprotein that is known to mediate the active transport of I− and pertechnate (TcO4−) into thyroid follicular cells, a crucial first step in thyroid hormone biosynthesis.12–14 The ability of thyroid cells to accumulate I− via the NIS gene product is the basis for diagnostic scintigraphic imaging of the thyroid using radio-iodide. 15 Moreover, newer methods use NIS as a reporter gene to detect the in vivo labeling of NIS-expressing cells with 124I, 125I, 131I, or Technetium-99m (99mTc); therefore, these techniques allow one to observe and quantify biochemical processes in the tissues of living subjects. 16 Use of the NIS reporter also provides a highly sensitive method for detecting even very low amounts of transgene expression.17,18 Furthermore, NIS has applications as an immunogen in autoimmune thyroiditis and is a potential molecular marker in residual thyroid cancer.19–21 On the basis of these observations, we conclude that the NIS gene transfer system is a useful tool for: (1) studying immune efficacy after DNA vaccine injection; and (2) in vivo visualization of target gene expression after systemic radioisotope administration.
In this study, we describe a novel, simple, and reproducible method for monitoring gene expression in vivo and its trafficking to secondary lymphoid organs after intradermal (i.d.) DNA injection. With real-time in vivo imaging and gamma-counting, we show that, after DNA inoculation, target gene-expressing cells were found in the paracortical areas of lymph nodes (LNs), in intracytoplasmic interferon-γ (IFN-γ)–secreting T-cells, and in greater amounts in the draining LNs (dLNs). Finally, the in vivo efficacy of this DNA vaccine was evaluated by challenge with a rat NIS (rNIS)–expressing tumor (CT-26/rNIS), and our results showed significant tumor growth retardation.
Materials and methods
Plasmid DNA vectors
The human NIS (hNIS)–expressing vector, pcDNA3.1-FL-hNIS (pcDNA-hNIS), was kindly provided by Dr. S. Jhiang (Ohio State University, OH, USA) and also β-galactosidase expressing pCMVβ vector was purchased from (Clontech, CA, USA). hNIS gene expression was controlled using cytomegalovirus (CMV) promoter, and expression of the neomycin resistance gene expression by Simian Virus 40 (SV40) promoter. Plasmid DNAs were amplified in E. coli DH5α bacteria and the large-scale plasmid preparations were purified using endotoxin-free Giga Prep columns (QIAGEN, CA, USA).
Immunization
Specific pathogen-free 6-week-old female BALB/c mice were obtained from SLC (Japan) and handled under specific pathogen-free conditions according to the guidelines issued by the Seoul National University Animal Research Committee. To monitor in vivo plasmid vector trafficking, mice given i.d. injections (in the thigh using a 30 G insulin syringe [Becton Dickinson, NJ, USA]) of 100 μg of pcDNA-hNIS (100 μg) suspended in endotoxin-free Tris-ethylenediaminetetraacetic acid buffer (50 μL; TE; QIAGEN). To identify tumor-protective or antigen (Ag)-specific cellular immune responses, mice were immunized three times at 2-week intervals. The mice were anesthetized by injecting intraperitoneally (i.p.) 0.3 mL of a 1:1:9 solution of Rompun (Parke Davis, Germany): Ketamine (Bayer, Germany): saline (RKS).
Whole-body imaging of immunized mice with nuclear scintigraphy
At the designated times (2, 16, 24 h and 2, 3, 11, 18, 25 days) after injection with pDNA-hNIS, mice were administered 300 μCi of Tc-99m (11.1 MBq) i.p. and anesthetized with 0.3 mL of the RKS solution. Thirty minutes after the Tc-99m injection, mice were placed in a prostrate position and scanned with a gamma-camera (ON 410; Ohio Nuclear, Solon, OH, USA) equipped with a pinhole collimator. Relative radioactivity accumulation was assessed in the entire body over a period of 5 min, and dynamic frames were obtained. 22 One hour later, LNs, spleen, liver, and skin near the injection site were taken from each mouse and weighed to determine the organ distribution patterns of the injected DNA; blood was also collected. The tissues were stored at −70°C for 16 h and then Tc-99m uptake was measured using a gamma-counter. The uptake of Tc-99m by tissues via the hNIS gene product was calculated using % of relative radioactivity: (cpm/pcDNA-hNIS– injected tissue weight (g) ÷ injection dose) × 100} ÷ (cpm/pcDNA3.1 (parent plasmid)–injected tissue weight (g) ÷ injection dose) × 100} × 100.
hNIS reverse transcription-polymerase chain reaction (RT-PCR)
Various organs were taken from immunized mice and lysed using a homogenizer. Total RNAs were extracted from lysates in the presence of RNase inhibitors according to the TRIzol reagent protocol (Molecular Research Center, OH, USA). The isolated RNAs were dissolved in diethyl pyrocarbonate-treated water (DEPC; Sigma-Aldrich, MO, USA). cDNA was generated from an mRNA template by incubating the mRNA with a 15-mer poly dT oligonucleotide (Invitrogen, CA, USA) and the Superscript reverse transcriptase enzyme (GIBCO-BRL, CA, USA) at 37°C for 1 h using the Superscript Preamplification System protocol. hNIS gene expression was detected in various organs (dLNs, non-dLNs, and spleens) by RT-PCR using the following gene-specific primers: 5’-GGCTCCTCGGTGACTCTAGGATGC-3’ (forward) and 5’-CATGAATTCTGGGCTCAATTTTCTTGTCC-3’ (reverse). To control for DNA integrity, the mouse β-actin gene (codons 135-223) was amplified using the following gene-specific primers: 5’-GGCTCCTCGGTGACTCTAGGATGC-3’ (forward) and 5’-CATGAATTCTGGGCTCAATTTTCTTGTCC-3’ (reverse). For amplification of the β-actin gene, the following PCR conditions were used: 34 cycles of 60 s at 94°C, 60 s at 55°C, and 60 s at 72°C.
Immunofluorescent staining
dLNs were collected from the pcDNA-hNIS–immunized mice X min post-injection. Tissues were fixed in 10% formalin and embedded in paraffin. Tissue cross-sections (4 μm) were deparaffinized in xylene and rehydrated through two graded alcohol solutions (100% or 95%). Sections were boiled in target retrieval solution (citrate buffer) at pH 6.0 (DakoCytomation, Denmark) and washed using a mixture of 50 mM Tris and 0.15 M sodium chloride, pH 7.5 (Tris-buffered saline). Endogenous Fc receptor and biotin binding sites were blocked by incubation in 3% H2O2 in methanol, and non-specific binding was blocked by incubation in 3% bovine serum albumin (BSA) in phosphate-buffered saline (PBS) (both for 30 min at room temperature [RT]). The sections were then incubated overnight at 4°C with rabbit polyclonal anti-hNIS antibodies (at 20 μg/mL; Alpha Diagnostic International, TX, USA), followed by incubation with anti-rabbit Cy-3 (Caltag, CA, USA) for 30 min at RT. All antibodies were diluted in 10% fetal bovine serum (FBS)-PBS. Biotin-labeled anti-CD3 antibody (BD Pharmingen, CA, USA) and streptavidin-FITC (BD Pharmingen) were used to identify the T-cell zones in the tissue cross-sections. All slides were counterstained with 4′,6′-diamidino-2-phenylindole (DAPI; Sigma-Aldrich) and then mounted with Vectashield (Vector Laboratories, UK) to preserve fluorescence. Images were acquired using an OLYMPUS BX51 (Optical Elements Corporation, VA, USA) fluorescent microscope and Viewfinder Lite v.1.0 (Pixera Corporation, CA, USA) software. Separate green fluorescein isothiocyanate (FITC) and far red Cy-3 images were collected for each section analyzed. Final image processing was performed using Photoshop 6.0 software (Adobe, CA, USA).
Intracellular cytokine flow cytometry analysis
Ten days after the third immunization, splenocytes and lymph node cells were isolated from either pcDNA3.1 or pcDNA-hNIS inoculated mice, and these cell populations (5 × 106 cells/mL) were restimulated in vitro by incubation for 36 h with irradiated (50 Gy) CT26 or CT26/rNIS cells (1 × 106/mL) prior to intracellular cytokine staining. Anti-mouse CD3 antibody (3 μg/mL; BD Pharmingen) and anti-mouse CD28 antibody (3 μg/mL; BD Pharmingen) were added to the culture media to optimize restimulation.23,24 The restimulated cells were washed in PBS and fixed in 4% paraformaldehyde (PFA)–PBS for 5 min at 37°C after CD4 (BD Pharmingen) or CD8 (BD Pharmingen) staining to define the T-cell type. Cells (1 × 106) were then resuspended in 1% Saponin/5% skim-milk-PBS (S/M-PBS) and incubated with fluorescence-conjugated IL-4 (BD Pharmingen) or IFN-γ (BD Pharmingen) primary antibodies for 30 min at RT. Isotype-matched antibody was used as a control. Cells were then washed twice with S/M-PBS and resuspended in 0.1% BSA-PBS. Staining was analyzed with a Coulter EPICS XL Flow Cytometry (BD Coulter, FL, USA).
Tumor challenge
Two weeks after the final Hnis DNA injection, mice were challenged by subcutaneous (s.c.) injection of 5×104 or 5×105 CT-26/rNIS cells suspended in 100 μL of 10% FBS in Dulbecco’s modified eagle medium (DMEM). Tumor dimensions were measured twice a week, and tumor sizes (mm3) were calculated using the following equation: horizontal (mm) × vertical (mm) × depth (mm).
Results
Expression of the hNIS gene in mouse tissues after DNA inoculation
After i.d. injection of mice with hNIS DNA, expression of the hNIS gene in mouse skin biopsies was evaluated by RT-PCR, which was carried out at the designated times over a period of 18 days post injection (Figure 1a). As expected, hNIS gene expression was detected in the lingual skin, including at the injection site, at both 11 and 18 days after DNA injection. To trace the path of the injected DNA beyond the injection site, we measured radioactive Tc-99m uptake in skin and soft tissue biopsies at 11 and 18 days after hNIS DNA injection. At both time points, γ-radioactivity was detected in soft tissue, including the LN-free lymphatic systems, but was not detectable in skin, even at the injection site (Figure 1b). As a control, we also immunized mice with bacterial LacZ DNA (which encodes the β-galactosidase enzyme) and showed that β-galactosidase activity also was detected far from the injection site in dorsal tissues (data not shown).

hNIS expression after i.d. DNA injection. (a) hNIS RT-PCR (n = 5 in each group). Total RNA was extracted from lingual skin, including the injection site, and hNIS mRNA was amplified by RT-PCR. β-actin primers were used as a control. h, hours post injection; D, days post injection; hNIS, RT-PCR using hNIS primers; β-actin, RT-PCR carried out with primers specific for the β-actin gene. (b) γ-radioactivity in mice injected (i.d.) with pcDNA-hNIS (n = 5 in each group). At the designated times post-injection, mice were injected i.p. with 300 μCi of Tc-99m (11.1 MBq). Thirty minutes later, the soft tissue that encompassed the lingual lymphatic system (without the LNs) was isolated from the mice, and uptake of radioactive Tc-99m was measured with a γ-counter. In order to exclude non-specific radioactivity, the Tc-99m uptake level was calculated as % of relative radioactivity as follows: {[cpm/pcDNA-hNIS–injected tissue weight (g) ÷ injection dose] × 100} ÷ {[cpm/pcDNA3.1–injected tissue weight (g) ÷ injection dose] × 100} × 100.
In vivo hNIS expression kinetics and organ distribution of γ-radioactivity after DNA injection
γ-radioactivity was monitored to trace hNIS-expressing cells in the bodies of living animals after pcDNA-hNIS injection. At 2 h post-injection, Tc-99m uptake was clearly detectable at the injection site in the pcDNA-hNIS–inoculated mice. In addition, γ-radioactivity was sustained at the injection site over a 2-week period following pcDNA-hNIS injection To quantify the intensity of in vivo radioactivity, regions of interest (ROIs) in the scintigraphic images were analyzed by gamma-counter at various times after immunization. Relative radioactivity intensity adjacent to the injection site was significantly higher in mice immunized with the hNIS plasmid than in control mice (Figure 2). Furthermore, the relative radioactivity intensity in the ROI around the injection site peaked at day 11 and declined to near baseline by day 18 (Figure 2).

In vivo hNIS expression kinetics following i.d. injection of hNIS DNA. Imaging of live mice injected with hNIS DNA. pcDNA3.1 or pcDNA-hNIS (100 μg) was administrated by injection into a single thigh of each BALB/c mouse (6–8-week-old female mice, n = 5 in each group). (a) At the designated times post injection, Tc-99m (300 μCi; 11.1 MBq) was administered to mice under general anesthesia by i.p. injection of 0.3 mL of a 1:1:9 RKS solution. Thirty minutes after Tc99m administration, Tc99m-injected mice were placed in a prostrate position and scanned with a gamma-camera (ON 410; Ohio Nuclear, Solon, OH, USA) equipped with a pinhole collimator. The accumulation of radioactivity was assessed during these 5-min whole-body scans, and dynamic frames were obtained subsequently. White circles indicate the area of γ-radioactivity accumulation in pcDNA-hNIS–immunized mice. h, hours post injection; D, days post injection. (b) Intensity of Tc99m radioactivity in a ROI from pcDNA-hNIS–immunized mice (n = 5 in each group). The ROI was determined from the scintigraphic image using an image analysis program (Photoshop 6.0 [Adobe, CA, USA]). The scintigraphic image was first scanned as a whole, and then differences between pictures of the hNIS-immunized mice and those of the pcDNA controls were analyzed, focusing on the designated area of the skin in and around the injection site (white circle in Figure 2a). % Relative radioactivity in ROI, (radioactivity in ROI of pcDNA-hNIS–injected mice ÷ radioactivity in ROI of pcDNA3.1–injected mice) × 100. These experiments were done three times independently.
Organ radioactivity distributions were determined at various times after immunization (Figure 3a) using the whole-body images. Relative radioactive emissions were significantly higher in the dLNs and non-dLNs of pcDNA-hNIS–inoculated mice relative to controls, and these emissions peaked at day 11 post immunization. In spleens of pcDNA-hNIS–inoculated mice, radioactivity began to rise around day 18 and was increased twofold by day 25 post immunization. The patterns of organ-specific radioactivity in the LNs of pcDNA-hNIS–inoculated mice were similar to the patterns observed in the whole-body, real-time scintigraphic images (Figure 2b) and to the patterns of LacZ expression after LacZ DNA inoculation (data not shown).

LN-specific trafficking of hNIS DNA. (a) Organ distribution of γ–radioactivity in mouse tissues at various times after immunization with pcDNA3.1 (control) or pcDNA-hNIS (n = 5 in each group). At the designated times after DNA injection, Tc-99m (300 μCi; 11.1 MBq) was injected i.p. into mice that had been inoculated with pcDNA3.1 or pcDNA-hNIS. One hour later, heart, liver, kidneys, spleen, non-dLNs, and dLNs were extracted from the mice and weighed. Tissues were kept at −70°C for 16 h, and Tc-99m uptake levels were measured using a gamma-counter. To exclude contributions from non-specific radioactivity, the Tc-99m uptake level was calculated as % relative radioactivity using the following formula: {[cpm/pcDNA-hNIS–injected tissue weight (g) ÷ injection dose] × 100} ÷ {[cpm/pcDNA3.1–injected tissue weight (g) ÷ injection dose] × 100} × 100. (b) hNIS RT-PCR. At the designated times after injection of mice with pcDNA3.1 or pcDNA-hNIS, total RNA was extracted from spleens, non-dLNs, and dLNs. RT-PCR products were generated from the cDNA template with two pairs of primers, one that was specific for hNIS and one that was specific for β-actin (control). h; hours post injection, D; days post injection, hNIS; RT-PCR product obtained using the hNIS primers, β-actin; RT-PCR product obtained using the β-actin primers. Lane 1, dLN; lane 2, non-dLN; lane 3, spleen. These experiments were done three times independently.
We next performed RT-PCR to determine whether the hNIS gene was present in all tissues that displayed detectable γ-radioactivity. As shown Figure 3b, hNIS transcripts were detected in the LNs of pcDNA-hNIS–inoculated mice at all designated times, while small amounts of hNIS transcripts were observed in spleens from these mice beginning at 11 days post injection. No hNIS transcripts were observed in any other organs from pcDNA-hNIS–inoculated mice (data not shown).
hNIS expression in the T-cell–rich zone of dLNs
To determine whether hNIS-expressing cells localize in the dLNs after immunization of mice with pcDNA-hNIS, we used an immunofluorescent co-staining method that involved the application of monoclonal antibodies (mAbs) to the hNIS protein and to the mouse CD3 protein (a marker for T-cells) to sectioned tissue from the dLNs (Figure 4). At 16 h after pcDNA-hNIS injection, small numbers of hNIS-expressing cells (red fluorescence staining) were observed in the subcapsular sinuses of the dLNs. At later time points, red fluorescence was also detected in the areas between B-cell–rich follicles within the dLNs and was increased in number. By co-immunostaining with the anti–hNIS and anti–mouse CD3 mAbs, we found that red fluorescent staining was strongest in the T-cell–rich zones of the dLNs, although some staining was observed in regions adjacent to the T-cells.

Localization of hNIS-expressing cells in dLNs from pcDNA-hNIS–immunized mice. At the indicated times after pcDNA-hNIS immunization, dLNs were collected and fixed with paraformaldehyde (n = 5 in each group). The fixed tissues were sectioned (4 μm) and stained with an anti-hNIS mAb (red) to detect hNIS-expressing cells and an anti-CD3 mAb for T-cells (green). The two types of fluorescence were merged and analyzed with Photoshop 6.0. (a) pcDNA3.1 injection (control), (b) 16 h after, (c) 1 day after, (d) 2 days after, (e) 3 days after, and (f) 11 days after hNIS DNA injection (×100). B, B-cell follicle; white arrows, area encompassing hNIS-expressing cells.
Immune responses induced after i.d. injection of hNIS DNA
To determine whether hNIS DNA injection induces an Ag-specific antibody response, we determined the IL-4 and IFN-γ profiles in cells isolated from dLNs, non-dLNs, MLNs (mesenteric LNs), and spleens. To assess hNIS-specific T-cell responses following DNA injection, splenocytes and LN cells were harvested 10 days after the final injection. The isolated cells were transferred to culture dishes and then restimulated with irradiated CT-26/rNIS cells, after which we performed intracellular cytokine analysis to detect IFN-γ– or IL-4–producing CD4+ T-cells. hNIS-specific IFN-γ−producing CD4+ T-cells were more numerous in the splenocytes and lymph node cells from the dLNs DNA−immunized mice than in those from animals immunized with the control vector (Figure 5a). In particular, these T-cells were found in 2.3-fold greater numbers in dLNs from the hNIS DNA−immunized mice, relative to control mice. In contrast, no significant IL-4 staining was observed in dLN cells from the hNIS-treated mice, although some was noted in the corresponding spleen cells (Figure 5b).

hNIS-specific CD4+ T-cell responses. To characterize the hNIS-specific CD4+ T-cell response in mice immunized with pcDNA-hNIS or pcDNA3.1, splenocytes and LN cells were harvested 10 days after the third DNA injection (n = 5 in each group). Cells were stained first for IFN-γ or IL-4 expression and then for CD4 expression after in vitro restimulation of the isolated cells by incubation with CT-26/rNIS cells for 36 h. (a) IFN-γ−producing CD4+ T-cells, (b) IL-4−producing CD4+ T-cells. The numbers without parentheses indicate the numbers of IFN-γ− or IL4−secreting CD4+ T-cells per 30,000 immune cells tested. The numbers within parentheses represent mean fluorescence. This experiment was done three times independently. (c, d) Statistics.
The same splenocytes and LN cells that we tested for CD4+ IFN-γ or IL-4 responses were also assessed for CD8+ T-cell IFN-γ and IL-4 responses. Significantly, IFN-γ−producing CD8+ T-cells were found in greater numbers than were IL-4−producing T-cells in tissues from mice immunized with hNIS DNA (Figure 6a and b); furthermore, the numbers of hNIS-specific, IFN-γ−secreting CD8+ T-cells were highest in the LNs, especially the dLNs (Figure 6a). These results demonstrate that hNIS-specific immune responses, especially IFN-γ−secreting CD8+ T-cell responses, are induced by hNIS DNA immunization.
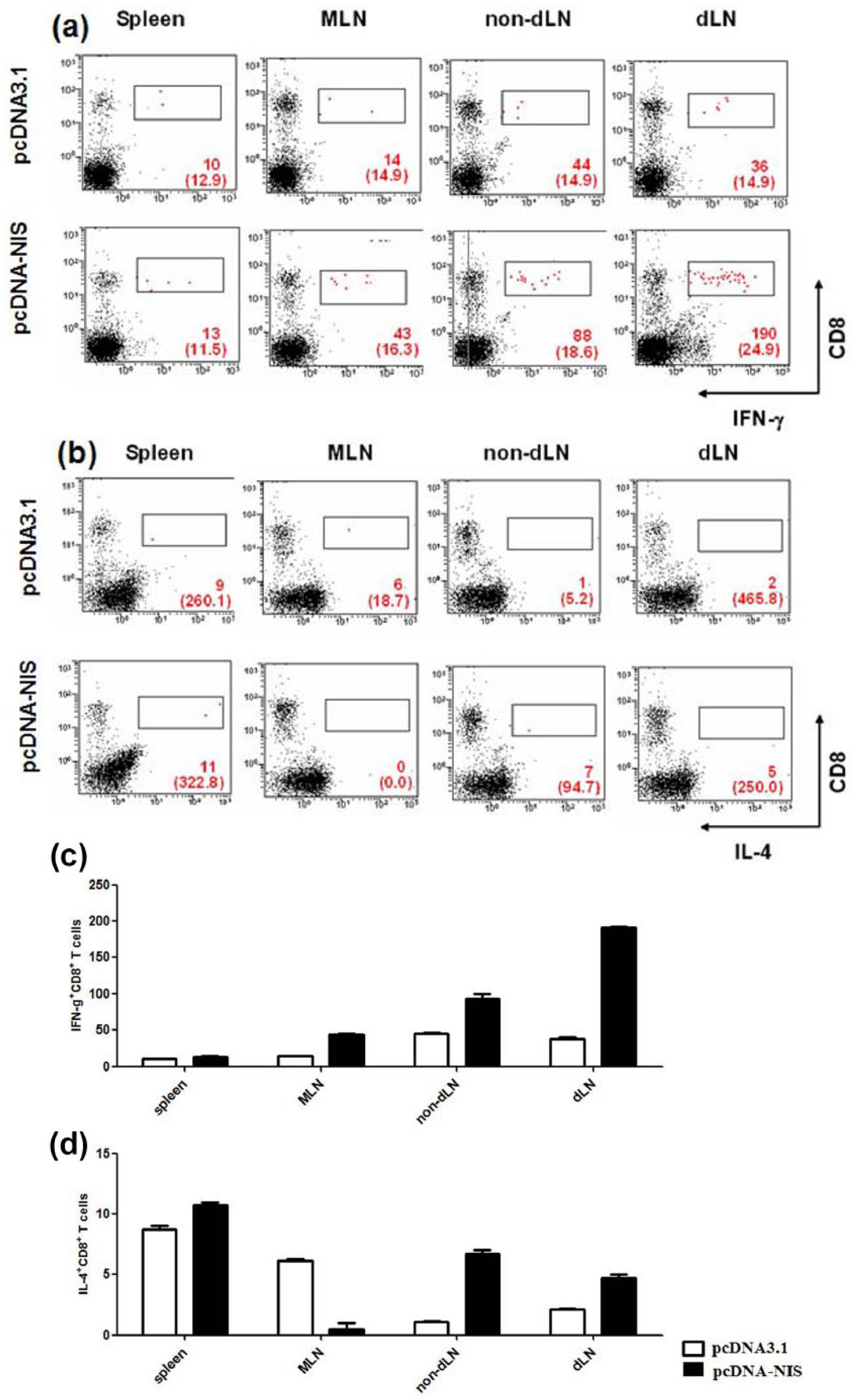
hNIS-specific CD8+ T-cell responses. To characterize the hNIS-specific CD8
Protective tumor immunity induced by DNA immunization
On the basis of the results described above, we investigated whether immunization of mice with hNIS DNA induced protection against an rNIS-expressing tumor challenge. Mice were inoculated s.c. with CT-26/rNIS tumor cells 2 weeks after the final hNIS DNA injection (Figure 7a). The mice immunized with hNIS DNA showed significant retardation of tumor growth compared with mice injected with pcDNA3.0 (Figure 7b). Tumor mass calculations were confirmed with Tc99m uptake measurements (data not shown).

Tumor protection following immunization of mice with hNIS DNA. Two weeks after the third injection of either pcDNA3.1 (n = 6) or pcDNA-hNIS (n = 6), mice were injected subcutaneously with 5×104 or 5×105 CT-26/rNIS tumor cells suspended in 100 μL of 10% FBS-DMEM. Tumor size was monitored twice per week and tumor volume (mm3) was calculated as follows: horizontal (mm) × vertical (mm) × depth (mm). (a) A schematic showing the injection protocol. wk, weeks. (b) Tumor growth. Tumor volume was presented by mean value. The numbers in parentheses indicate the number of tumor cells injected into the immunized mice.
Discussion
Since Wolff et al. 2 first demonstrated that the direct transfection of plasmid DNA could lead to the expression of exogenous genes in vivo; many studies have reported success in expressing exogenous genes using local/regional plasmid DNA administration.5–7 Dendritic cells have been demonstrated to be efficient carriers of transgenes and/or their products, and these were found to migrate to LNs from the site of administration.25–27 However, the kinetics of exogenous DNA protein expression, the physical pathway of this DNA post injection, and its mechanism of eliciting an antigen-specific immune response are unclear. In the present study, we investigated the kinetics and in vivo trafficking of DNA following i.d. injection. To develop the sensitivity of experimental system and to exclude the bystander effects by proteins secreted after transfect to cells, the hNIS gene was chosen that its products are not secreted.
At first, hNIS gene expression was examined at the protein level in transiently transfected CT-26 cells by using radioactive 125I uptake assays for pcDNA-hNIS. The result showed us pcDNA-hNIS vector was transfected and could be used to follow in vivo DNA trafficking post injection. Figure 1a, RT-PCR data showed in vivo hNIS gene transfection was also successful.
We noted that β-gal activity was detectable far from the injection site after LacZ DNA injection and assumed these movements were caused by trafficking of cells taken-up DNA. To define the route of them, mice were i.d. injected with blue dyes to visualize DNA inoculation sites and these were shown to be linked with soft tissue, particularly with the lymphatic system. 30 Radioactive Tc-99m uptake activity was also examined on LN free lymphatic systems 11 and 18 days post injection of LacZ DNA at which β-gal activity was shown far from injection site. Radioactivity was significant in lymphatic systems from hNIS DNA immunized mice compared with control vector (Figure 1b). In regards, sensitized cells appeared in the dermal lymphatics 31 and within LNs after challenge with contact sensitizers.32,33
On the other hand, Wolff et al. 28 and Liu et al. 29 demonstrated that plasmid DNA injected into a mouse via a tail vein is degraded quickly due to the presence of numerous endonucleases in both the blood stream and in lymph plasma. Even within 10 min of a DNA injection, no intact DNA remained in the liver, suggesting that the injected plasmid DNA was degraded soon after injection. Therefore, possibility of the movement of DNA by from the injection site could be excluded in the present study. Taken together, our results suggest that transfected hNIS DNA containing cells migrate from the injected site via the lymphatics.
To assess the injected hNIS DNA distribution in whole body, radioactive Tc99m (232,700 ± 35,300 cpm/g) was administrated systemically 1 h post hNIS DNA. In all experiments, γ-radio-images of animal were taken prior to the introduction of Tc99m, and no change in background radioactivity was detected in animals injected with plasmid vectors. Also, after Tc-99m uptake, no difference in radioactive intensity was observed in blood acquired from mice injected with mock pcDNA alone or with pcDNA-hNIS (Figure 2a).
Targeted protein was found to be highest in LNs 11 days after injection (Figure 3a) and this result concurs with radioactivity patterns obtained by real-time whole-body imaging (Figure 2b). These results were confirmed by RT-PCR analysis (Figure 3b). The observed alterations in expressional strengths and patterns over time are in agreement with observations made by Hengge et al. 34 in pig skins, which showed a decline of expression after 3 weeks and a gradual reduction in size of the transfected area. This decline in expression could be considered to be an immunological process in muscle, due to an inflammatory reaction in response to the injection of naked DNA. 35 In addition, PCR analysis was used to confirm hNIS gene localization and detected hNIS DNA in LNs by day 18, but not in liver, kidneys, or heart. Also, small amounts of hNIS DNA were also detected in the spleen 3–18 days after injection (data not shown). Therefore, it is possible that non-secretary protein corresponding to the injected DNA could be expressed over a 2-week period after homing to LNs.
It has been well documented that intradermal injections generate an immune response via skin DCs acquiring antigens and presenting these antigens to T-cells via the conventional class I pathway.36–38 In the present study, we experimented to expand upon these findings by evaluating both the kinetics of inoculated DNA expression and to determine the length of time necessary to generate a transgene-specific immune response. We observed by immunohistochemical analysis hNIS expressing cells at the capsular and sinus regions of LNs as early as 6 h after hNIS DNA injection (data not shown). Kamath et al. 39 recently demonstrated that dermal DCs appear in lymph nodes as early as 24 h after skin painting, whereas Langerhans cells require 3–4 days. Thus, we presume hNIS-expressing LN sinus cells are dermal DCs migrated from the injection site.
Furthermore, we investigated the movement of them in dLN following DNA injection by immunofluorescence analysis. hNIS-expressing cells were observed in the subcellular sinus of dLNs 16 h after DNA injection, but gradually in T-cell–rich zones after that (Figure 4). However, our data do not differentiate between the possibilities as to whether hNIS-expressing cells in the sinus moved to T-cell zones or whether other cells were also involved in this movement. Studies by Shortman et al. 39 and Ruedl et al. 40 demonstrated that Langerhans cells and dermal DCs home to LNs, and that they persisted and presented antigens for over 2 weeks, thereby prolonging the T-cell response. Therefore, we conclude that dermal DCs that have acquired hNIS DNA home to LNs and then to the T-cell zone. This might also be important for the induction of T-cell response in an antigen specific manner.
To trace the organ preference and the progress of T-cell priming, 10 days after final injection, dLNs, non-dLNs, MLNs, and spleens were extracted from DNA immunized mice. We used rat NIS gene transfected CT-26 murine colon carcinoma cell line (CT-26/rNIS) as in vitro restimulator. Until now, the immunity against NIS has not yet been reported such as NIS is immunogenic and also there are specific T-cell repertoires. So, we considered naturally expressed NIS as a stimulator for in vitro restimulation to detect T-cell immunity and a capture for ELISA to detect humoral immunity. In this paper, we used Balb/C syngeneic CT-26/rNIS; rNIS transfected colon cancer cell line provided by Dr. Jung (Seoul National University, Republic of Korea). Smanik et al. 13 identified a cDNA encoding hNIS and it exhibits 84% identity and 93% similarity to rNIS. hNIS differs from rNIS mostly on account of a 5-amino acid insertion between the final two hydrophobic domains and a 20-amino acid insertion in the COOH terminus. Also, Dohan et al. 15 demonstrated a secondary structure of NIS contains 13 putative transmembrane segments, the NH2 faces extracellular milieu, and the COOH terminus faces the cytosol. They also showed that the NH2 terminus and hydrophobic loop face the external milieu are important to NIS function not COOH terminus face cytosol. According to these reports, we regarded CT-26 cells expressing rNIS as a suitable material being able to hNIS-specific immunity induced by DNA vaccination.
Intracellular cytokine staining also showed that hNIS-specific IFN-γ producing T-cells, especially CD8+ T-cells in dLNs, were highly detected whereas IL-4 producing CD4+ T-cells were merely detected in spleen after restimulation in vitro with irradiated CT-26/rNIS (Figures 5 and 6). MAbs to CD28 and CD3 were used to optimize IL-4 and IFN-γ-producing cell detection, but no non-specific cytokine production was observed in cells incubated only with these Abs. Also, there were no significant changes in the batch pulsed with PMA and ionomycin (Supplementary Figure 1). In addition, cells from immunized mice were explored for surface markers, i.e. CD3, CD4, CD8, and CD22, without restimulation in vitro. We found that the number of T-cells, especially CD8+ T-cells, was significantly increased only in the dLNs of hNIS DNA immunized mice (data not shown). It means increased IFN-γ producing CD8+ T-cells are not generated in vitro by restimulation but reactivated from memory or resting hNIS specific T-cells.
Tumor retardation was also found to be associated with hNIS DNA immunization (Figure 7). This Th1 immunity activating pattern and tumor intimated that IFN-γ secreting CD8+ T-cells may play an important role in retarding tumor growth. As IFN-γ can skew immune responses by acting on APC, which may inhibit Th2 response, 24 IL-4 would be expected to have a detrimental effect if Th1 cells and CTLs are the main mediators of tumor rejection. Therefore, our results suggest that skin DCs acquired hNIS DNA, and present them to T-cells following i.d. injection, and that dLN is the main secondary immune organ to induce hNIS-specific anti-tumoral immune response, particularly IFN-γ secreting CD 8+ T-cells response.
There are at least three mechanisms by which antigen encoded by plasmid DNA is processed and presented to elicit an immune response: (1) direct priming by somatic cells (myocytes, keratinocytes, or any MHC class II-negative cells); (2) direct transfection by professional APCs (i.e. DCs); and (3) cross-priming in which plasmid DNA transfect somatic cells and/or professional APCs and secreted protein is taken up by other professional APCs and presented to T-cells. 41 Our results suggest that the direct transfection of professional APCs is a main process after the i.d. injection of non-secretary protein expressing DNA. Plasmid-containing cells homed to LNs and showed long-lasting corresponding gene function following immunization. In addition, foreign gene expressing cells moved toward T-cell zones in dLNs and induced an antigen-specific Th1 immune response and tumor growth retardation.
In conclusion, this study describes the kinetics of gene expression following i.d. DNA injection and analyzes the traffic of injected DNA in vivo without sacrificing test animals. The traffic patterns and kinetics of this transgene could be useful in various DNA vaccine strategies. These issues clearly warrant further studies to develop simple tool being able to evaluate adjuvant effects using estimated γ-radioactivity of LN homing cells after vaccination or to develop hNIS DNA cancer vaccine to clear thyroid cancer cells.
Footnotes
Acknowledgements
The authors would like to thank Professor Sang-Hoon Choi (Department of Aquatic Life Medicine, Kunsan National University, Republic of Korea) and Professor Vasso Apostolopoulos (The Austin Research Institute, Melbourne University, Australia) for critically reviewing this paper.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by grants from the Korea Science & Engineering Foundation (KOSEF) through the Tumor Immunity Medical Research Center at Seoul National University College of Medicine, Korea. This work was also supported by the National Research Foundation of Korea (NRF) grant for the Global Core Research Center (GCRC) funded by the Korea government (MSIP) (No. 2016005276). H-Y Son and Y-H Jeon were supported by the BK21 Project for Medicine, Dentistry, and Pharmacy in Korea.
