Abstract
Sepsis, critical condition marked by severe organ dysfunction from uncontrolled infection, involves the endothelium significantly. Macrophages, through paracrine actions, play a vital role in sepsis, but their mechanisms in sepsis pathogenesis remain elusive. Objective: We aimed to explore how macrophage-derived exosomes with low miR-141 expression promote pyroptosis in endothelial cells (ECs). Exosomes from THP-1 cell supernatant were isolated and characterized. The effects of miR-141 mimic/inhibitor on apoptosis, proliferation, and invasion of Human Umbilical Vein Endothelial Cells (HUVECs) were assessed using flow cytometry, CCK-8, and transwell assays. Key pyroptosis-related proteins, including caspase-1, IL-18, IL-1β, NLR Family Pyrin Domain Containing 3 (NLRP3), ASC, and cleaved-GSDMD, were analyzed via Western blot. The interaction between miR-141 and NLRP3 was studied using RNAhybrid v2.2 and dual-Luciferase reporter assays. The mRNA and protein level of NLRP3 after exosomal miR-141 inhibitor treatment was detected by qPCR and Western blot, respectively. Exosomes were successfully isolated. miR-141 mimic reduced cell death and pyroptosis-related protein expression in HUVECs, while the inhibitor had opposite effects, increasing cell death, and enhancing pyroptosis protein expression. Additionally, macrophage-derived exosomal miR-141 inhibitor increased cell death and pyroptosis-related proteins in HUVECs. miR-141 inhibits NLRP3 transcription. Macrophages facilitate sepsis progression by secreting miR-141 decreased exosomes to activate NLRP3-mediated pyroptosis in ECs, which could be a potentially valuable target of sepsis therapy.
Introduction
Sepsis is considered a life-threatening condition marked by organ dysfunction, arising from an excessively activated systemic immune reactions in the host. It affects approximately 31.5 million cases and accounts for 5.3 million deaths occurring annually.1,2 During sepsis, the immune system releases various signaling molecules, precipitating widespread tissue damage, and culminating in the failure of multiple organ systems.3,4 Notwithstanding significant strides in elucidating the pathophysiological intricacies of sepsis, progress in developing effective treatments has been slow. Therefore, there is an urgent need to uncover the mechanisms underlying sepsis to identify new therapeutic targets for its treatment.
As the primary permeability barrier between vascular tissue and blood, vascular endothelial cells (ECs) play an indispensable role in the development of sepsis.5,6 During sepsis, ECs undergo a transformation towards a state characterized by augmented apoptosis, inflammation, adhesiveness, and coagulation.7,8 Additionally, the dysregulation of vascular tone precipitates disturbances in microcirculatory blood flow, culminating in organ damage and the potential for critical organ failure. 6 Pyroptosis, a caspase-1-dependent programmed cell death pathway, is induced by various infectious and non-infectious triggers. 9 Emerging research has highlighted the involvement of ECs pyroptosis in sepsis-related organ injuries.10,11 ECs possess remarkable plasticity and can swiftly respond to cytokine stimulus secreted by macrophages or other cells, adapting their phenotype accordingly. 12
Macrophages are pivotal to the immune response and represent the vanguard in defense against pathogenic incursions.13–17 They exhibit a high degree of plasticity, allowing them to adapt their phenotype along a spectrum ranging from M1 pro-inflammatory to M2 anti-inflammatory states, contingent upon environmental cues. 18 The promotion of the phenotypic transition from M1 to M2 macrophages plays a crucial role in attenuating inflammatory responses, augmenting tissue repairprocesses, and enhancing regenerative capabilities. 19 Numerous studies have indicated that uncontrolled macrophages activation is a key mechanism driving sepsis.20,21 This is due to their widespread presence in various organs and their multifaceted roles, including antigen presentation, phagocytosis, and production of inflammatory cytokines.22,23 Upon the initiation of sepsis, macrophages respond to inflammatory stimuli with heightened activity and morph into a pro-inflammatory phenotype. This transformation results in the release of an abundance of cytokines, such as tumor necrosis factor-alpha (TNF-α), interleukin-1-beta (IL-1β), IL-12, IL-18, and IL-23.24,25 These cytokines, in turn, amplify the inflammatory response and disrupt immune function, thereby playing a significant role in the pathophysiology of sepsis. Given their crucial role in both infection and inflammatory responses, macrophages stand as an indispensable cellular model in the realm of sepsis research. However, the specific effects of macrophages on the process of pyroptosis in ECs have not been extensively studied or reported.
Recent studies have proposed that macrophages exert their effects through the paracrine activity of their secreted trophic factors, anti-inflammatory proteins, and exosomes. 26 Exosomes, delineated as nanoscale extracellular vesicles with dimensions ranging from 30 to 100 nm, encompass an array of constituents, including nucleic acids, lipids, proteins, mRNA, miRNA, and other non-coding RNAs. 27 Upon fusion with the cellular membrane, these components of exosomes are liberated into the extracellular environment. 28 Therefore, exosomes play a crucial role in transmitting signals and transporting materials between different cells and tissues. 29 Emerging evidence suggests that macrophages transfer exosomal miRNAs, which carry epigenetic information, to recipient cells, thereby influencing gene expression and diseases development.30–32 miR-141, a specific microRNA belonging to the miR-200 family, plays a regulatory role in a myriad of cellular processes, including apoptosis, proliferation, differentiation, and cell adhesion. 33 In various human tumors, miR-141 is downregulated and has been considered a tumor repressor. It has also been found to be downregulated in the nasopharyngeal carcinoma (NPC) biopsy samples.34–36 Additionally, miR-141-3p plays a role in promoting tumor progression in ovarian cancer by contributing to M2-like macrophage polarization. 37 Exosomal miR-141 derived from bone marrow-mesenchymal stem cells has been shown to mitigate myocardial injury in septic mice by targeting PTEN and stimulating β-catenin. 38 However, the expression and mechanisms of miR-141 in the context of pyroptosis remain elusive.
This research aims to investigate the impact and underlying mechanism of macrophages on the occurrence of pyroptosis in ECs, which will improve our understanding of mechanisms involved in the development of sepsis, thus may pave the way for the discovery of innovative therapeutic strategies aimed at preventing or ameliorating sepsis.
Materials and methods
Culture of HUVECs and macrophages
HUVECs (passage 2–10, no. PCS-100-010, ATCC) and human macrophages THP-1 cells (passage 4-15, no. TIB-202, ATCC) were cultured in DMEM (no. 10313039, Gibco), supplemented with 10% FBS (no.16140071, Gibco, pH = 7.0) and 1% penicillin/streptomycin (no.15140122, Gibco), within a humidified atmosphere containing 5% CO2 at a temperature of 37°C.
Isolation of exosomes
The supernatant from THP-1 cells was initially subjected to centrifugation at 2000g for 30 min at 4°C. Subsequently, it underwent filtration through a 0.22 μm syringe filter (no. SLGVR13SL, Millipore) and was further subjected to ultracentrifugation at 120,000 g overnight at 4°C utilizing an Optima XPN-100 high-speed freezing centrifuge (no. CP100MX, Hitachi), facilitating the precipitation of exosomes. The resultant exosomal pellet was resuspended in ice-cold phosphate buffered saline (PBS, pH = 7.4) and ultracentrifuged once more at 120,000 g for 90 min at 4°C. The final exosomal pellet was resuspended in either ice-cold PBS or SDT lysate buffer (no. P0015 F, Beyotime, 4% SDS, 100 mM Tris-HCl, pH = 7.6), and subsequently stored at −80°C for future analyses.
Electron microscopy
The supernatant containing purified exosomes was resuspended in PBS. A volume of 20 μL of the exosomal suspension was meticulously applied onto a copper grid, followed by careful blotting and staining with 2% phosphotungstic acid (PTA). After the staining process, the sample was subjected to high-resolution imaging utilizing a transmission electron microscope (no. HT-7700, Hitachi).
Nanoparticle tracking analysis
The isolated exosome sample was diluted to a volume of 1 mL in Tris-Phosphate-Modified Saline (TPM) to prepare it for subsequent analyses. The size of distribution of the exosomes was meticulously determined utilizing Nanosight Tracking Analysis, which employs nanoparticle tracking analysis (NTA, NanoFCM, N30E), in strict adherence to the established protocols. 39
Western blot
Western blot analysis was conducted in accordance with the methodology previously described. 40 In brief, protein samples were extracted from THP-1 cells and THP-1-derived exosomes using RIPA lysis buffer (no. HY-K1001, Med. Chem Express). The protein concentrations were quantified using the BCA Protein Assay Kit (no. 23225, ThermoFisher). Subsequently, 5 μg of proteins isolated from exosomes was combined with 2 × SDS sample buffer and resolved on 12% SDS-acrylamide gels. This was followed by electrotransfer onto PVDF membranes. The membranes were then blocked with 5% non-fat milk and incubated overnight at 4°C with primary antibodies: rabbit anti-Calnexin (1:1000 dilution, no. ab133615, Abcam), rabbit anti-TSG101 (1:1000 dilution, no.102286-T38, Sino Biological), mouse anti-CD81 (1:1000 dilution, no. ab79559, Abcam), and rabbit anti-CD63 (1:1000 dilution, no. ab216130, Abcam). As secondary antibodies, horseradish peroxidase-conjugated goat anti-mouse IgG (1:5000 dilution; cat. no. BA1051; Boster) and horseradish peroxidase-conjugated mouse anti-rabbit IgG (1:5000 dilution; no. BM2006; Boster) were used for 1.5 h at room temperature. Detection of immunoreactive bands was achieved using an ECL hypersensitive chemiluminescence kit (no. P0018 M; Beyotime Institute of Biotechnology) and visualized with an Odyssey Scanning System (version 3.0, LI-COR Biosciences). The protein gray value was calculated by Image J (Version 1.8.0, NIH Image J system).
Proteins were extracted by RIPA lysis buffer from miR-141 mimic and inhibitor-treated HUVECs. The following primary antibodies were employed: rabbit anti-pro-caspase-1 (1:1000 dilution, no. ab179515, Abcam), rabbit anti-Caspase-1 (1:1000 dilution, no. 22915-1-AP, Proteintech), rabbit anti- IL-1β (1:1000 dilution, no. A1112, ABclonal), rabbit anti-IL-18 (1:1000 dilution, no. A1115, ABclonal), rabbit anti-ASC (1:1000 dilution, no. ab283684, Abcam), rabbit anti-GSDMD (1:1000 dilution, no. ab215203, Abcam), rabbit anti-NLRP3 (1:1000 dilution, no. ab263899, Abcam), and mouse anti-GAPDH (1:5000 dilution, no. 60004-1-lg, Proteintech).
miR-141 mimic and inhibitor transfections
miRNA mimic and inhibitor transfection were performed as described previously. 41 The miR-141 mimic (5′-UAACACUGUCUGGUA AAGAUGG-3′), miR-141 inhibitor (5′-CCAUCUUUACCAGACAGUGUUA-3′), and negative control (NC, 5′-GGGAGUGAAGACACG GAGCCAGA-3′) were obtained from Shanghai GenePharma Company. Transfection of the miR-141 mimic and inhibitor, as well as the NC, into HUVECs and the miR-141 inhibitor and NC into THP-1 cells, was executed utilizing Oligofectamine™, adhering to the guidelines specified by the manufacturer.
Flow cytometry
The assessment of cell death in HUVECs was meticulously conducted utilizing an Annexin V-FITC and PI staining kit (no. A211-01/02, Vazyme), following the precise protocols delineated by the manufacturer. 42 After the staining procedure, flow cytometric analysis was executed employing a FACS CaliburTM flow cytometer. The quantification of the proportion of dead cells (PI+) was accomplished using the advanced analytical capabilities of FlowJo software (Tree star, Ashland, Oregon).
Cell viability Assay
HUVECs were meticulously seeded onto the 96-well plates at a density of 1 × 103 cells per well and subsequently incubated for designated durations spanning 24, 48, 72, 96, and 120 h. Following these incubation periods, each well was treated with 10 µL of Cell Counting Kit-8 (CCK-8) regent (no. C0039, Beyotime), and the cells were incubated at 37°C for an additional 2 h. 43 Absorbance measurements at 450 nm were then performed using an Eppendorf BioPhotometer® D30 (BioPhotometer D30, Germany). The calculation of cell viability was based on the absorbance data collected over the 5-day period.
Cell migration assay
The Transwell migration assay was utilized to evaluate alterations in the migratory capacity of HUVECs post-treatment with miR-141 mimic, inhibitor, or macrophage-derived exosomal miR-141 inhibitor treatment, as described previously. 44 HUVECs were seeded in a 12-well plate (7 × 103 cells/well). Subsequently, 150 μL of a serum-free cell suspension and 600 μL of DMEM were, respectively, introduced into the upper and lower chambers of the Transwell apparatus. After incubation at 37°C for 48 h, cells situated on the basolateral chamber were meticulously washed twice with PBS and then stained with 1% crystal violet for a duration of 40 min. Stained HUVECs were visualized and photographed under the microscope (magnification, ×200; Olympus cX2; Olympus) after washing using PBS twice.
Quantitative real-time polymerase chain reaction (qPCR)
Primer sequences for the real-time reverse transcriptase polymerase chain reaction.

Dual-luciferase reporter assay
To ascertain the interaction between miR-141 and NLRP3, wild-type (WT) and mutant (MUT) sequences of the NLRP3 3′-untranslated regions (UTR) were cloned into the pmirGLO luciferase reporter vector (Promega). Subsequently, 293T cells were co-transfected with either WT or MUT reporter plasmids, alongside miR-41 mimic or a negative control (NC) mimic, employing Lipofectamine® 3000 (Invitrogen) for transfection, and incubated for 48 h. Then the cells were harvested and luciferase activity was quantified in accordance with the guidelines of the TransDetect Double Luciferase Reporter Assay Kit (no. FR201-01, TransGen Biotech). The luminescence signal was measured using Eppendorf BioSpectrometer fluorescence (Eppendorf), with the luminescent signal indicative of reporter gene activation quantified based on the ratio of firefly to Renilla luciferase activity.
Statistical analysis
Statistical evaluations were performed utilizing GraphPad Prism software (version 8.0; California. USA). Experimental procedures were replicated on three separate occasions, and the resultant data are expressed as mean ± standard deviation. Comparative analysis between two distinct groups was executed employing the unpaired student’s t-test. For multiple group comparisons, one-way Analysis of Variance (ANOVA) followed by Tukey’s post hoc test was implemented. A threshold of p < .05 was established to denote statistical significance.
Results
Characterization of isolated exosomes
The exosomes obtained from the macrophage supernatant were examined using electron microscopy, revealing discernible small membrane-bound vesicles (Figure 1(a)). The majority of these vesicles were found to be within the size range of 50–120 nm in diameter (Figure 1(b)), aligning with established exosomal dimensions reported in extant literature.
46
To confirm the identity of these exosomes, Western blot analysis was employed to evaluate the presence of quintessential exosomal biomarkers, encompassing cluster of differentiation (CD) 81 antigen (CD81), CD63 antigen (CD63), tumor susceptibility gene 101 (TSG101), and endoplasmic reticulum protein marker Calnexin.
47
As anticipated, exosomal hallmarks, specifically CD63, CD81, and TSG101, were markedly prevalent in the exosomal fraction, confirming their presence in the isolated vesicles (Figure 1(c)). In contrast, Calnexin, which is specific to the endoplasmic reticulum, was only detected in the macrophage samples and not in the exosomes (Figure 1(c)). Characterization of exosomes after isolation. (a) Electron micrographs of exosomes isolated from THP-1 cells. (b) NTA (NanoSight) was applied to determine the size distribution of isolated exosomes in diameter. (c) Western blot was performed for the indicated proteins.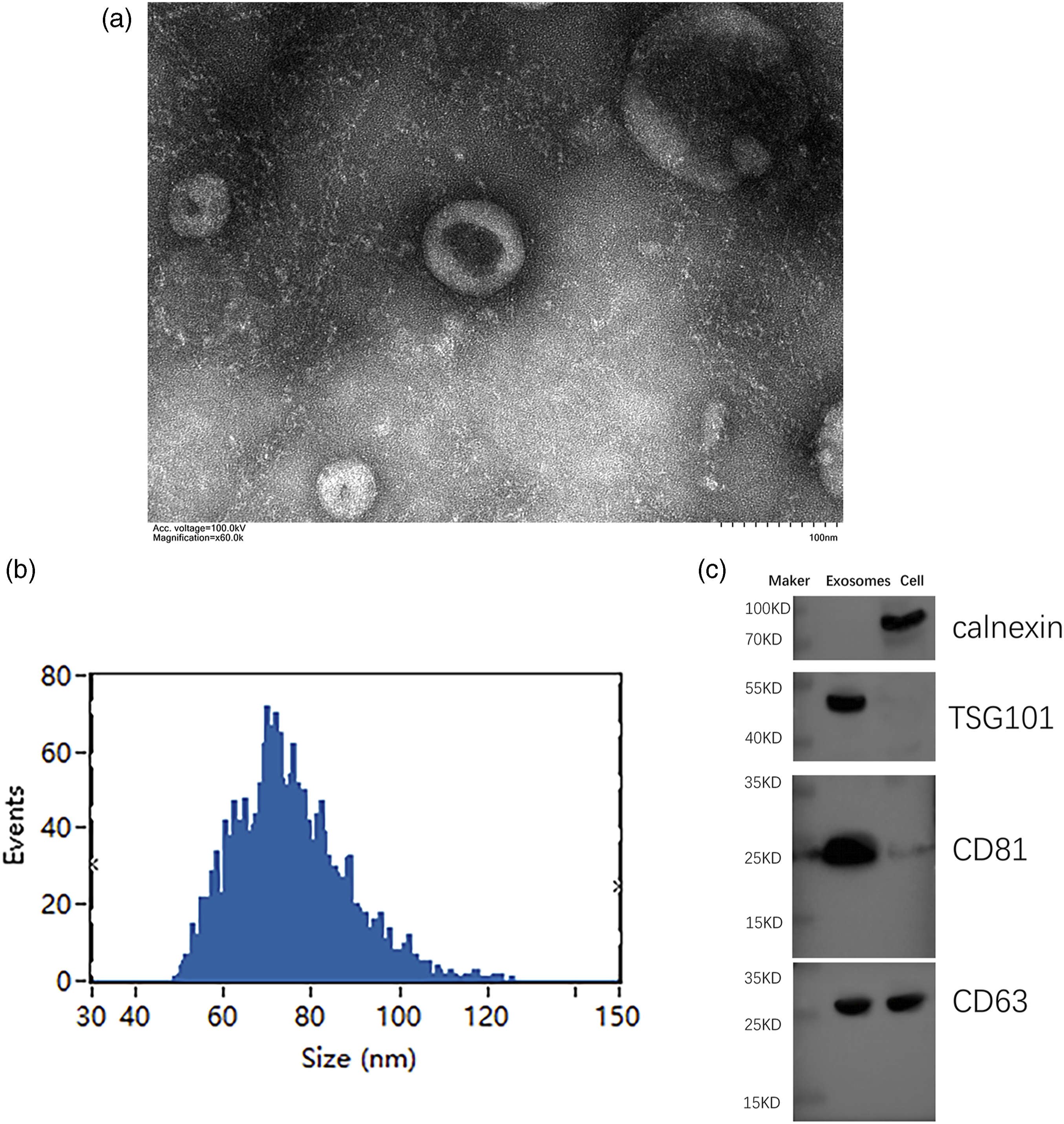
Macrophages-derived exosomal miR-141 inhibitor triggered HUVECs death
To delineate the putative role of miR-141 in sepsis, we performed transfection experiments in HUVECs employing miR-141 mimic, miR-141 inhibitor, and miR-NC. Subsequently, the application of miR-141 mimic significantly decreased HUVECs death rate, whereas miR-141 inhibitor remarkably promoted HUVECs death compared with control (Figure 2(a) and (b)). In concordance, miR-141 mimic transfection resulted in an augmented cell proliferation rate, in stark contrast to the miR-141 inhibitor, which exerted an inhibitory impact on cellular proliferation (Figure 2(c)). Furthermore, the migration ability of HUVECs was enhanced following miR-141 mimic treatment, whereas the miR-141 inhibitor significantly impeded cellular migration (Figure 2(d)). Moreover, we investigated the effects of macrophage-derived exosomal miR-141 inhibitor on HUVECs. Treatment with exosomes carrying miR-141 inhibitor derived from macrophages significantly contributed to a significant enhanced cell death rate, a remarkable decreased cell proliferation, and migration rate in HUVECs compared with the control group (Figure 3(a)–(c)). These observations indicate that miR-141 exerts a promotive effect on HUVECs death and affects their proliferation and migration abilities. miR-141 inhibits HUVECs death and promotes HUVECs viability and migration. miR-141 mimic, inhibitor, and miRNA negative control (NC) were transfected into HUVECs. (AB) Flow cytometry analysis of death in HUVECs stained with Annexin V-FITC. (C) The viability of HUVECs was detected by CCK-8. (D) Invasion ability of HUVECs was assessed by Transwell assay. Data is represented as means ± SD (n = 3, **p < .01, ***p < .001). Macrophages-derived exosomal miR-141 inhibitor triggered HUVECs death. miR-141 inhibitor and miRNA negative control (NC) were transfected into macrophages THP-1. Exosomes isolated from the supernatant of miR-141 inhibitor-treated THP-1 cells were applied to treat HUVECs. (A) Flow cytometry analysis of death in macrophages-derived exosomal miR-141 inhibitor treated HUVECs stained with Annexin V-FITC. (B) The viability of macrophages-derived exosomal miR-141 inhibitor treated HUVECs was detected by CCK-8. (C) Invasion ability of macrophages-derived exosomal miR-141 inhibitor treated HUVECs was assessed by Transwell assay. Data is represented as means ± SD (n = 3, ***p < .001).

HUVECs displayed characteristic features of pyroptosis after miR-141 inhibitor treatment
Further, overexpressing miR-141 mimic in HUVECs resulted in a significant downregulation of several key pyroptosis-related proteins, including caspase-1, IL-18, IL-1β, NLRP3, ASC, and cleaved-GSDMD (Figure 4(a)). Remarkably, the suppressive impact of the miR-141 mimic on these pyroptosis-related proteins was almost completely nullified when HUVECs were treated with miR-141 inhibitor, evidenced by the ensuing upsurge in the expression of these proteins (Figure 4(a)). What is more, macrophages-derived exosomal miR-141 inhibitor can also significantly promote the protein expression of caspase-1, IL-18, IL-1β, NLRP3, ASC, and cleaved-GSDMD in HUVECs (Figure 4(b)), indicating that decreased miR-141 induced pyroptosis in HUVECs. HUVECs displayed characteristic features of pyroptosis after miR-141 inhibitor (a) miR-141 mimic, inhibitor, and miRNA negative control (NC) were transfected into HUVECs. Western blot analysis evaluating the expression of pro-caspase-1, caspase-1, IL-18, IL-1β, NLRP3, ASC, and cleaved-GSDMD in HUVECs. (b) Exosomes isolated from the supernatant of miR-141 inhibitor-treated THP-1 cells were applied to treat HUVECs. pro-caspase-1, caspase-1, IL-18, IL-1β, NLRP3, ASC, and cleaved-GSDMD protein level in HUVECs was detected by western blot. Grayscale analysis was performed by ImageJ software. Data is represented as means ± SD (n = 3, *p < .05, ***p < .001, ****p < .0001).
miR-141 inhibiter facilitated pyroptosis by sponging NLRP3
miRNAs have been demonstrated to regulate pyroptosis by modulating NLRP3 expression [33]. Based on this knowledge, we speculated that miR-141 might have a similar function in macrophages. To substantiate our conjecture regarding the potential target genes of miR-141, we employed an online prediction tool (RNAhybrid v2.2, https://bibiserv.cebitec.uni-bielefeld.de/rnahybrid) for analytical exploration. Surprisingly, the results favor our speculation, suggesting that NLRP3 is a target of miR-141 (Figure 5(a)). To further validate this interaction, we engineered luciferase reporter constructs harboring either the wild-type or mutated NLRP3 3′-UTR and conducted dual-luciferase reporter assays. The introduction of miR-141 mimic significantly diminished the luciferase activity associated with the wild-type NLRP3 3′-UTR, in contrast to the neutral effect observed with the NC mimic. Notably, no discernible impact on the luciferase activity was observed in the mutated NLRP3 3′UTR constructs, irrespective of the presence of NC mimic or miR-141 mimic (Figure 5(b)). Additionally, the miR-141 mimic markedly decreased NLRP3 mRNA level, while the miR-141 inhibitor significantly increased NLRP3 mRNA level (Figure 5(c)). Furthermore, the macrophages-derived exosomal miR-141 inhibitor effectively promoted NLRP3 expression, evident at both mRNA and protein levels (Figure 5DE). Collectively, these results indicate that miR-141 inhibitor facilitates pyroptosis in HUVECs by targeting NLRP3. miR-141 regulated pyroptosis in HUVECs by sponging NLRP3. (a) Bioinformatics analysis was used to predict the binding site of NLRP3 to miR-141. (b) Analyzed results of dual-luciferase reporter assay evaluating the interaction between miR-141 and NLRP3. (c) Analyzed qPCR results detecting the expression of miR-141 and NLRP3 in HUVECs with miR-141 mimic, inhibitor, and NC treatment. (d) Analyzed qPCR results detecting the expression of miR-141 and NLRP3 in HUVECs with macrophages-derived exosomal miR-141 inhibitor and NC treatment. (e) Western blot analysis evaluating the expression of NLRP3 in HUVECs with macrophages-derived exosomal miR-141 inhibitor and NC treatment. Grayscale analysis was performed by ImageJ software. Data is represented as means ± SD (n = 3, *p < .05, ***p < .001).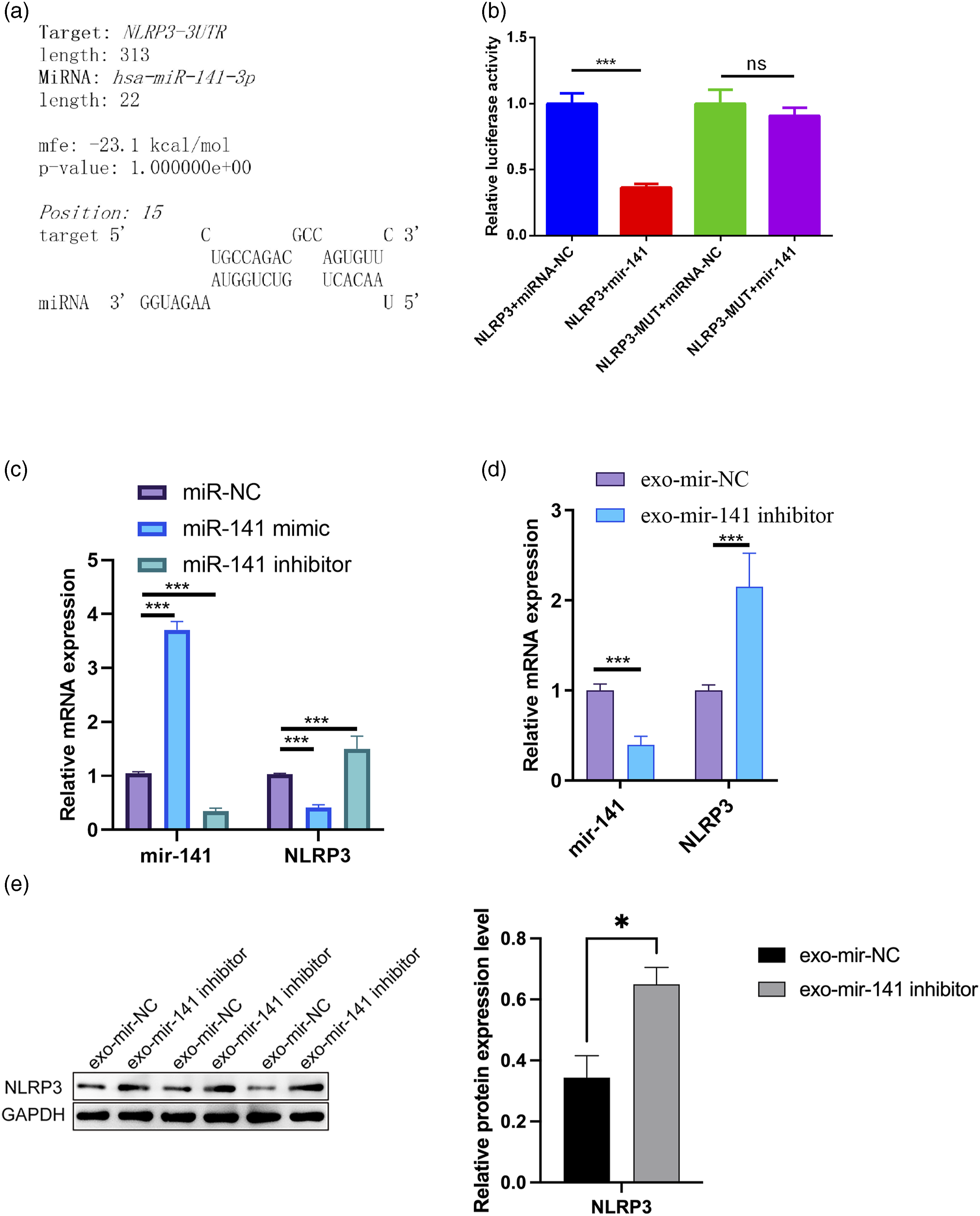
Discussion
Sepsis, characterized by systemic inflammation, causes a dysfunction of ECs due to elevated levels of inflammatory cytokines. 48 However, excessive and prolonged pro-inflammatory activation of ECs may precipitate compromised microcirculatory blood flow, tissue hypoperfusion, and potentially catastrophic organ failure. 6 Consequently, a rigorous investigation into the molecular pathways that precipitate sepsis-induced endothelial dysfunction is imperative for preventing organ failure in sepsis. Macrophages have been implicated in the regulation of vascular inflammation. 49 However, the mechanisms by which macrophages regulate inflammatory responses in the vasculature remain poorly understood. In this manuscript, we present a novel connection between macrophages and endothelial cells, wherein macrophage-derived exosomal miR-141 targets NLRP3 in ECs. This interaction precipitates pyroptosis, a profoundly inflammatory variant of programmed cell death, thereby advancing endothelial dysfunction, a pivotal factor in the pathophysiology of sepsis.
Macrophages are pivotal orchestrators in the pathophysiology of a myriad of chronic inflammatory disorders and autoimmune diseases.13–16 It is increasingly evident that uncontrolled macrophage activation is a fundamental mechanism underlying sepsis.20,21 Exhibiting remarkable phenotypic plasticity, macrophages can oscillate between pro-inflammatory M1 and anti-inflammatory M2 states in response to varying environmental cues. 50 At the onset of sepsis, macrophages are excessively activated by inflammatory stimuli and differentiate into a pro-inflammatory M1 phenotype. These activated macrophages release massive amounts of pro-inflammatory cytokines, including tumor necrosis factor-alpha (TNF-α), IL-1β, IL-12, IL-18, and IL-23,24,25 which can induce and exaggerate the inflammatory response and immune dysfunction.21,51 There is accumulating evidence underscoring the role of macrophages in disseminating exosomal miRNAs laden with epigenetic information, thereby modulating gene expression in target cells and influencing disease progression.30–32 For example, exosomes derived from M2 macrophages containing miR-690 have been shown to decrease inflammation and improve insulin signaling in obese mice. 9 Likewise, the release of miR-21-3p in exosomes from macrophages activated by nicotine exposure is associated with the hastening of atherosclerotic development. 30 In our study, the transfection of miR-141 mimic dramatically reduced cell death and promoted the viability and migration of HUVECs, in stark contrast to the miR-141 inhibitor. These observations underscore the salient role of miR-141 in the developmental dynamics of HUVECs. Additionally, the administration of macrophage-derived exosomes containing the miR-141 inhibitor to HUVECs mirrored these effects, thereby positing miR-141 as a critical intermediary in the macrophage-mediated modulation of endothelial cells.
Pyroptosis is initiated by an inflammasome, such as NLRP3 inflammasomes when exposure to stimuli.52,53 Once NLRP3 oligomerization occurs, NLRP3 interacts with ASC, an adaptor protein that facilitates the interaction between NLRP3 and caspase-1.54,55 This engagement leads to the activation of caspase-1 by NLRP3, which catalyzes the cleavage of GSDMD to release its N-terminal fragment (GSDMD-N), a pivotal step in pyroptosis. 56 GSDMD-N then assembles into the plasma membrane, binding to the lipids of the inner membrane to form pores. 57 These pores induce cellular swelling and membrane rupture, culminating in the extracellular release of pro-inflammatory cytokines and the execution of pyroptotic cell death. 58 In our study, miR-141 mimic downregulated the expression of NLRP3, ASC, caspase-1, as well as inflammasome-dependent effectors including pro-caspase 1, caspase 1, IL-18, IL-1β, and cleaved-GSDMD in HUVECs. Conversely, miR-141 inhibitor exhibited opposite effects, suggesting that miR-141 protects HUVECs from cell inflammation-related pyroptosis. Consistently, macrophages-derived exosomal miR-141 inhibitor also augmented the expression of pyroptosis-related proteins in HUVECs. These findings collectively underscore the crucial role of miR-141 in attenuating pyroptotic cell death in sepsis.
It has been well-documented that miRNAs mediate gene silencing at the post-transcriptional level.59–62 They are recognized as key modulators across various cellular and pathological mechanisms. 63 Increasing evidence has linked miRNAs to pyroptosis. For example, miR-214 has been shown to enhance pyroptosis in H9c2 cells by targeting caspase-1. 64 Similarly, miR-21 promotes pyroptosis through the upregulation of NLRP3, ASC, and caspase-1 expression. 65 In this study, we have identified that miR-141 inhibits NLRP3 transcription in HUVECs. Moreover macrophages-derived exosomal miR-141 inhibitor significantly increased NLRP3 level in HUVECs. Therefore, these findings indicate that inflammatory macrophages can induce pyroptosis in ECs, promoting the progression of sepsis through the secretion of exosomes containing downregulated miR-141.
While our study provides valuable insights, there are some limitations to consider. Sepsis is a systemic condition that involves various cells and tissues, necessitating the use of a broader range of cellular models for comprehensive investigation. 1 In our research, we focused on macrophages and ECs, which narrows the applicability of our findings. Additionally, in vivo sepsis models provide advantages in the in-depth investigation of the pathogenesis of sepsis, drug development, and evaluation of new therapeutic strategies. 66 The absence of animal models limits the generalizability of our results. The utilization of in vivo models will enhance the credibility of the findings in this study, and future studies should address this limitation. Our understanding of the interaction between miR-141 and NLRP3 is still preliminary and requires further exploration. We observed that the inhibition of miR-141 in macrophage-derived exosomes notably enhanced pyroptosis in HUVECs. However, it is important to acknowledge that the present study lacks experiments involving the introduction of exosomal miR-141 mimic. The inclusion of such experiments would provide additional and compelling evidence to substantiate the role of miR-141 in HUVECs pyroptosis, warranting further exploration and validation in future studies. Lastly, future research should focus on the practical implications, particularly the therapeutic effects of miR-141 on sepsis induced by EC pyroptosis, to enhance the relevance of our findings in sepsis treatment.
Conclusion
The administration of exosomal miR-141 inhibitors, derived from macrophages, significantly augmented cell death, while concurrently diminishing cell viability and migratory capabilities in endothelial cells. This treatment also promoted the upregulation of proteins associated with pyroptosis. Furthermore, miR-141 was observed to inhibit NLRP3 transcription. The secretion of diminished levels of miR-141-containing exosomes by macrophages may represent a promising therapeutic target in sepsis management, primarily by activating NLRP3-mediated pyroptosis in endothelial cells.
Footnotes
Author contributions
Feng Zhan and Yanhong Ouyang conceived and designed this study. Ping He and Wenteng Chen carried out the analysis and participated in the study design. Feng Zhan and Jun Zhang wrote the manuscript. Feng Zhan and Yanhong Ouyang have seen and can confirm the authenticity of the raw data. All authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Hainan Natural Science Foundation of China (821MS122).
Data Availability Statement
Data sharing not applicable to this article as no datasets were generated or analyzed during the current study.
