Abstract
Keywords
Introduction
Cervical squamous cell carcinoma and endocervical adenocarcinoma (CESC) are the second most prevalent gynecological cancer. 1 According to the available data, approximately 500,000 women are diagnosed with cervical squamous cell carcinoma and endocervical adenocarcinoma (CESC), and the disorder causes more than 300,000 fatalities worldwide annually.2–4 The condition of the immune microenvironment is intently interrelated to the CESC development. Many studies have deeply explored the relationship between cervical cancer and immune cells.5–7 Immunotherapies using anti-CTLA4 8 and anti-PD1 9 drugs have been successful in CESC treatment. However, to date, none of them can significantly improve overall survival.10–12 There is still an imperative need for effective diagnosis and prognostic indicators of CESC tumor occurrence and recurrence to assist treatment.
Sphingosine-1-phosphate (S1P) is a pluripotency and extensive biologically active molecule be classified to the sphingolipid family, which is an intricate lipid group appeared on all eukaryotic cells. In the past, it was thought to only perform structural functions, while recently sphingolipids are regarded as pivotal conditioners of countless cell functions in pathophysiological processes.13,14 S1PR1, S1PR2, and S1PR3, S1P receptors 1–3, there are expressed general, while S1PR4 and S1PR5 are mainly restricted expression to central nervous system, lymphatics and hematopoietic tissues. Binding to different G proteins activates several downstream pathways that help regulate many cellular mechanisms. 15 Therefore, there has been considerable interest in the S1P/S1PRs axis as a potential therapeutic target for regulating various cellular processes. In contrast, so far, the other S1P acceptors function in the cervix has obtained rare concern. S1PR2 is the key acceptor for the occurrence and development of different genre of cancer. Even though its role varies from tissue to tissue, most data support anti-tumor function. 16 In fact, S1PR2 adjusts negatively the invasion and migration of human glioblastoma, 17 melanoma, 18 oral squamous cell carcinoma, and gastric cell lines 19 and cell proliferation in human kidney tumor cells. 20 In addition, the gene decomposition of S1PR2 promotes the in vivo growth of Lewis lung and melanoma cancer, 21 standing by S1PR2 is a key acceptor of cell proliferation. Recently research showed that S1PR2 impeding proliferate CD4 + T cell to adjusts the epithelial barrier on intestinal epithelial cells. 22 Therefore, its features in epithelial cells still to be elaborated. Nevertheless, in CESC, the S1PR2 prognostic value and its expression has not been fully elucidated. In addition, in CESC, the relationship between TRIM27 and tumor immune infiltration remains indistinct.
In the study, we executed survival and expression analysis of S1PR2 in various human cancers. We found that S1PR2 is down-regulated in CESC, and the down-regulation of S1PR2 is related to the adverse clinical characteristics and risk factors of CESC patients. We discovered that the decreased S1PR2 expression is interrelated to the low survival rate of CESC patients. We further explored diagnostic and prognostic value of S1PR2 and the connection between S1PR2 biomarkers and expression of immune cells, immune cell infiltration or immune checkpoints in CESC.
Methods
The cancer genome atlas dataset analysis
We downloaded the expression transcriptional data of S1PR2 and the information of corresponding clinical from The Cancer Genome Atlas (TCGA) database (https://genome-cancer.ucsc.edu/). After normalizing these data, the variant expression of S1PR2 was analyzed by the R package limma (3.6.3). 23
GEPIA database analysis
GEPIA (http://gepia.cancer-pku.cn/) is a network instrument for normal or cancer gene expression profiles, Genotype Tissue Expression data and TCGA are used to analysis the interactive. 24 CESC patients survival analysis, including analyses of disease-free survival (DFS) and overall survival (OS) were performed by GEPIA.
Protein-protein interaction networks and functional enrichment analysis
STRING (https://www.string-db.org/) for searching the genes of for interacting to construct Protein-Protein Interaction (PPI) networks. 25 We performed with an interaction score >0.4 of a STRING search for co-expressing genes of S1PR2 and constructed a PPI network. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis of co-expressing genes and Gene Ontology (GO) enrichment were performed using the “clusterProfiler” package and visualized by the “ggplot2” package. 26
The tumor immune estimation resource database
The Tumor Immune Estimation Resource (TIMER) (https://cistrome.shinyapps.io/timer/) is a resource online for the immune infiltration systemic analyzed in miscellaneous cancer types. 27 The connection between S1PR2 expression level and the level of immune cell infiltration in CESC were determined by TIMER.
Statistical analysis
Statistical analyses were calculated by the above-mentioned online databases. A log rank p value of <0.05 or p value <0.05 was considered statistically significant.
Results
Expression of S1PR2 in pan-cancer
To make a thorough inquiry the roles of S1PR2 in carcinogenesis, first, we inquired the expression of S1PR2 in 32 human cancers. As presented in Figure 1(a), compared to normal samples, S1PR2 was markedly up-regulated in 7 of all 32 cancer types, including STAD, CHOL, ESCA, HNSC, LIHC, GBM and THCA, and was significantly down-regulated in 9 cancer types, involving KIRP, BLCA, BRCA, PRAD, KICH, KIRC, LUAD, LUSC, and CESC. However, no obviously difference expression of S1PR2 in COAD, PAAD, PCPG, READ, or UCEC was observed. Next, we used TCGA database to further verify the S1PR2 expression in these 16 types of cancer. As shown in Figure 1(b) to (h), S1PR2 expression in BLCA, BRCA, CESC, LUAD, LUSC, PRAD or ESCA was reduced compared to the homologous normal contrasts. In CHOL, GBM, HNSC, LIHC, STAD, or THCA, S1PR2 was obviously increased (Figure 1(i) to (n)). Taken together, S1PR2 was up-regulated in CHOL, GBM, HNSC, LIHC, STAD, or THCA, and down-regulated in BLCA, BRCA, CESC, LUAD, LUSC, and PRAD, demonstrating that S1PR2 may as a key regulatory role in the carcinogenesis of 12 cancer types. Expression of S1PR2 in Pan-cancer. (a) The expression of S1PR2 in 32 human cancers based on TCGA data of normal and cancer. 
Prognostic values of S1PR2 in human cancer
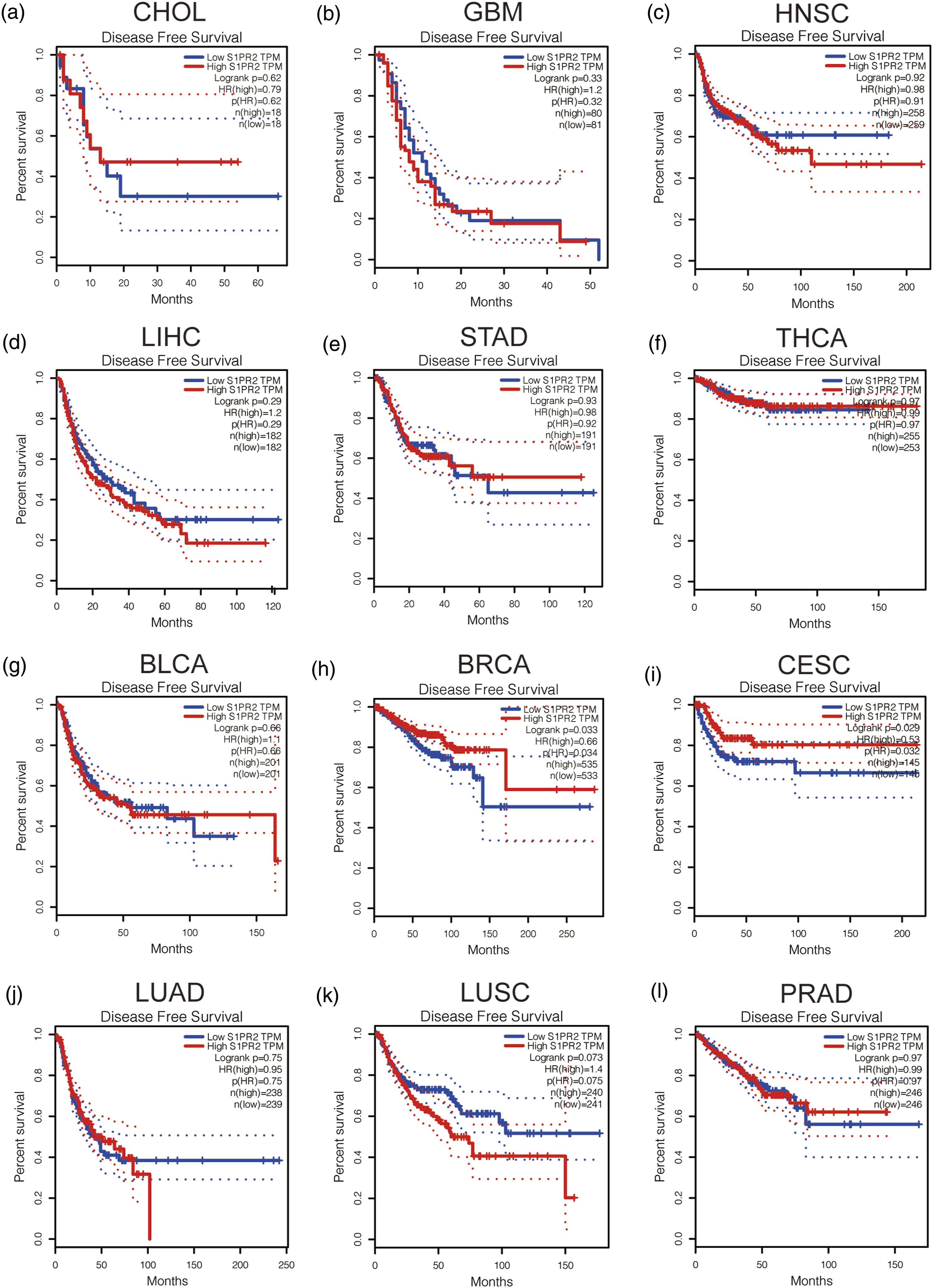
We analyzed the survival of S1PR2 in HNSC, CHOL, STAD, LIHC, GBM, or THCA, and down-regulated in BLCA, BRCA, CESC, LUAD, LUSC, and PRAD was conducted. Including two prognostic indicators: one is disease-free survival (DFS) and another is overall survival (OS). As presented in For OS, low S1PR2 expression in CESC has a poor prognosis, but BLCA patients with low S1PR2 expression have a better prognosis (Figure 2). As presented in Figure 3, for RFS, reduced S1PR2 expression in all cancer types indicates poor prognosis for BRCA and CESC. No statistical significance of S1PR2 in predicting the patients prognosis with other cancers was inquired. Integrating OS and RFS, S1PR2 can be used as disadvantageous prognostic biomarker for CESC patients. S1PR2 overall survival (OS) analysis in diverse human cancers established by GEPIA database. (a–i) The OS plot of S1PR2 in CHOL (a), GBM (b), HNSC (c), LIHC (d), STAD (e), THCA (f), BLCA (g), BRCA (h), CESC (i), LUAD (j), LUCS (k), and PRAD (l). S1PR2 disease-free survival (RFS) analysis in diverse human cancers established by GEPIA database. (a–i) The RFS plot of S1PR2 in CHOL (a), GBM (b), HNSC (c), LIHC (d), STAD (e), THCA (f), BLCA (g), BRCA (h), CESC (i), LUAD (J), LUCS (k), and PRAD (l).
The prospective biomarker of S1PR2 in CESC patients
We implemented ROC curve analysis to probe the values for S1PR2 in distinguishing CESC from normal samples. As presented in Figure 4, the ROC curve analysis indicated that S1PR2 with an AUC value about 0.870 (95% CI: 0.713–1.000). With a 2.689 cutoff, S1PR2 had a specificity, sensitivity of 84.6 and 90.5%. And the negative predictive value was 27.5%, the 99.3% positive predictive value. These results indicate that S1PR2 may be a prospective biomarker for distinguishing CESC tissues from normal tissues. ROC curves for S1PR2. ROC curve for CESC patients based on S1PR2 expression.
Association of S1PR2 levels with clinical characteristics in CESC patients
Clinical characteristics of the CESC patients (TCGA).

*p value <0.05; ***p value <0.001.
PPI network and functional annotation about S1PR2
We implemented GO, KEGG analyses and STRING database to elevate functional annotations and PPI networks. As presented in Figure 5(a), a network of S1PR2 and its 10 co-expression genes. Figure 5(b) shown transforms in the biological circuit of S1PR2 were related with sphingolipid signaling pathway, and parathyriod hormone synthesis, secretion, and action. The correlation analysis between S1PR2 expression and CESE co-expressed genes from TCGA is shown in Figure 5(c)–(l). PPI enrichment analyzed of networks and functional. (a) S1PR2 network and the co-expression genes of S1PR2. (b) Enrichment function analyzed of 10 involved genes. (c–l) The correlation analyzed about the S1PR2 expression with co-expressed genes of CESC.
S1PR2 positively correlates with immune cell infiltration in CESC
We next examined the relationship between S1PR2 expression and seven tumor types of infiltrating immune cells from the TIMER database. The SCNA module shown the association between the CESC tumor immune cell infiltration and copy number of different somatic cells of RRAGB through the Wilcoxon rank sum test (Figure 6(a)). Connection analyzed could supply important clues for researching the machine-processed and foundation of S1PR2. Therefore, the relationship about the level of immune cell infiltration and the expression level of S1PR2 was assessed. As shown in Figure 6(b), the expression of S1PR2 was markedly positively related with immune cells, counting dendritic cell, neutrophil, CD4+ T cell, macrophage, and B cell in CESC. The correlation between immune cell infiltration and the level of S1PR2 in CESC. (a) Different kinds immune cell infiltration levels under different S1PR2 copy numbers of CESC. (b) The relationship of the expression level of S1PR2 with dendritic cell, neutrophil, macrophage, CD4+ T cell, CD8+ T cell, or B cell infiltration level of CESC.
Correlation between S1PR2 and the expression of immune cells in CESC
Correlation analysis between S1PR2 and biomarkers of immune cells in CESC determined by GEPIA database.

**p value <0.01; ***p value <0.001.
Correlation between S1PR2 and immune checkpoints in CESC
PD1 and CTLA-4 are key immune checkpoints there are liable for tumor immune escape. The correlation of S1PR2 with PD1 and CTLA-4 was evaluated while premeditating the latent oncogenic role of S1PR2 in CESC. There was markedly positively related between S1PR2 with CTLA-4 and PD1 in CESC (Figure 7(a) to (d)). These foundings indicate that tumor immune escape could be concerned in S1PR2 intermediary carcinogenesis of CESC. Relationship in CESC of the expression of S1PR2 with the expression of PD-1 and CTLA-4. Using TIMER to analyzed the Spearman relationship of S1PR27 with PD-1 (a), CTLA-4 (b) expression in CESC adjusted by purity. The expression relationship between S1PR2 and PD1 (c), CTLA-4 (d) in CESC determined by GEPIA database.
Discussion
CESC is a general tumor in women. With the upgrowth of standard treatment options for CESC patients with concurrent chemotherapy and brachytherapy, it has improved of the 5-year survival rate, while the prognosis is poor. 28 Nevertheless, there is a pressing need for promising prognostic biomarkers to assess pressing patients risk with metastatic CESC and corresponding effective therapeutic targets that can improve clinical outcomes.
More and more evidence shows that S1P is associated with angiogenesis, cell proliferation, chemotaxis, migration and differentiation, and is also associated with the cancer biology. These S1PRs seems to be specific of tissues, has been shown to be consist of the cell proliferation regular pattern, surviving in various cancer types. S1PR2 roles in cancer remains agonistical. According to reports, S1PR2 can act as an anti-cancer and cancer-promoting receiver. For example, in B-cell lymphoma, glioblastoma, and melanoma, S1PR2 plays an anti-cancer receiver. On the other hand, it is reported that in prostate cancer, S1PR2 plays a carcinogenic receiver. A review introduced that on the roles of S1PR2 in cancers referred to the affect of this receiver on developing of tumor and headway is the specific of cell-type, because of its taking part with the specific G proteins to regulate physiological functions. 29 Therefore, despite the existence of context-specific and controversial evidence, the knowledge of S1PR2 in CESC is still insufficient and further research is needed. In this study, we first performed a pan-cancer analyzed of S1PR2 expression used Cancer Genome Atlas (TCGA) data, and used it to verify the S1PR2 expression. Survival analysis of S1PR2 in these cancer types of interest indicated that patients of CESC with low S1PR2 expression have a poorer prognosis. And we further found down-regulated S1PR2 expression is positively correlated with patients with high clinical stages, more histological types of squamous cell carcinoma, and poor primary treatment outcomes. ROC curve analysis indicates S1PR2 might be a promising diagnostic biomarker in distinguishing CESC from normal tissues. S1PR2 might be a promising biomarker for impoverished prognosis of CESC.
S1PR2 participates in differentiation, cell proliferation, angiogenesis, migration, and chemotaxis through the sphingolipid signaling pathway. TP inhibits the SPHK-S1P signaling pathway and effectively reduces the levels of S1P and the expression of SPHK1/S1PR1/S1PR2, and markedly suppressing the S1P-mediated phosphorylation activation of ERK protein in macrophages. 30 Our co-expression analysis indicated that the expression of S1PR2 was markedly related to the expression of RHOA, GNA11, GNA12, GNAI1, GNAI2, and GNAQ. We speculate that the down-regulation of S1PR2 will effect the entire pathway, and it possibility could be examined in future findings.
Numerous studies have convinced that the tumor immune cell infiltration could effect the usefulness of immunotherapy, radiotherapy, or chemotherapy and the prognostication of cancer patients.31–33 This work shows that S1PR2 is markedly positively related to various immune cells, involving dendritic cells, macrophages, CD4+ T cells, neutrophils, and B cells in CESC. In addition, S1PR2 is also significantly positively correlated with these biomarkers of infiltrating immune cells. These results indicate tumor immune infiltration and may not wholly explain the carcinogenic effects of S1PR2 mediated in HCC.
The usefulness of immunotherapy requires sufficient immune cells to infiltrate the tumor microenvironment, and relies on the full expression of immune checkpoints. 34 Therefore, this study more evaluated the related about S1PR2 with immune checkpoints. The findings indicate that S1PR2 is closely related to CTLA-4 or PD1 in CESC, showing that aiming S1PR2 may enhance the efficacious of immunotreatment in CESC.
There are definitely limitations to our study. A database used by use to confirm the correlation between the expression of S1PR2 and CESE to make the sample size enlargely and make sure the accurateness of the experimental outcomes. The data we owned from multiplex databases to narrow the deviation that a single database might cause. Future studies need to carry out related animal and cell experiments, forward to study the potential roles of S1PR2 in CESC.
Conclusion
We demonstrated that S1PR2 is under-expressed in many types of human cancers, including CESC, and showed that S1PR2 represents a possible poor prognostic biomarker which could be used to identify CESC patients with poor clinical outcomes. Our research results also indicate that S1PR2 may exert its anti-cancer effect by increasing the expression of immune checkpoint and tumor immune cell infiltration. The findings could be verified through additional experiments and large-scale clinical trials.
Footnotes
Acknowledgements
Authors’ contributions
ZY and WHC drafted the manuscript and performed the data analysis. YL and GZR designed the experiments. LQ, LJ, and YB collected the data, which was supervised by YL. All authors have reviewed and approve the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by the Science and Technology Development Foundation of Pudong New Area (PKJ2020-Y26).
Data availability
Data and material availability can be obtained from corresponding author on request.
