Abstract
MicroRNAs (miRNAs) exhibit a crucial role in the regulation of angiogenesis and tumor progression, of which miR-199a-5p (miR-199a) has been reported to function as a tumor suppressor in multiple malignancies. However, the precise mechanisms underlying miR-199a in hemangiomas (HAs) remain elusive. In this study, we found that miR-199a had low expression level, while proliferating cell nuclear antigen (PCNA) had high expression level in proliferating-phase HAs compared with the involuting-phase HAs and normal tissues. Spearman correlation analysis revealed the negative correlation of miR-199a with PCNA expression in proliferating-phase HAs. In vitro experiments showed that restoration of miR-199a suppressed cell proliferation capability and induced cell apoptosis in HA-derived endothelial cells (HDEC) and CRL-2586 EOMA cells, followed with decreased PCNA expression and increased cleaved caspase-3 expression, but miR-199a inhibitor reversed these effects. Furthermore, HIF1A was identified as a target of miR-199a and had negative correlation with miR-199a expression in proliferating-phase HAs. Overexpression of HIF1A attenuated the anti-proliferation effect of miR-199a mimic in HAs cells. Taken together, our findings demonstrate that miR-199a may inhibit proliferation and induce apoptosis in HAs cells via targeting HIF1A and provide a potential therapeutic target for HAs.
Introduction
Hemangiomas (HAs) as noncancerous tumors are mainly formed by the proliferating endothelial cells of blood vessels, and their life cycles consisting of rapid growth and spontaneous regression distinguish them as proliferating-phase HAs and involuting-phase HAs. 1 HAs are also the most common benign tumors in infants and children, and their incidence approximately approaches 2%–3.2%. 2 Current advances in the identification of microRNAs (miRNAs) and interplay with their target genes have enhanced our awareness of the pathogenesis of HAs. 3
MiRNA, a subgroup of small noncoding RNAs with 21–25 nucleotides in length, are partially complementary to messenger RNA (mRNA) molecules, reduce the expression of their target genes by translational repression and mRNA cleavage, 4 and participate in cell proliferation and apoptosis of HAs.5,6 Among the miRNAs, miR-199 family members including miR-199a-1/-2 and miR-199b have been reported as potential prognostic markers in multiple cancers,7 –9 of which miR-199a is downregulated in a variety of malignant tumors including papillary thyroid carcinoma, 10 osteosarcoma (OS), 11 esophageal cancer, 12 prostate cancer, 13 breast cancer, 14 and hepatocellular carcinoma (HCC). 15 MiR-199a impairs cell proliferation, invasion, and tumorigenic capabilities10 –15 via regulation of AKT and mitogen-activated protein kinase (MAPK) pathways11,12 and inhibits HCC glucose consumption and lactate production by targeting hexokinase 2. 15 These studies indicate a tumor-suppressive role of miR-199a in tumors.
Nevertheless, some studies show that miR-199a expression is upregulated in colorectal cancer (CRC), 8 gastric cancer, 16 and OS 17 and predicts poor survival of CRC patients 8 and promotes tumor growth and progression16,17 inferring a tumor-promoting role in tumors. In addition, proliferating cell nuclear antigen (PCNA) is expressed by most of the vascular endothelial cells and can be used to distinguish the HA progression stage in proliferative or involuting phase. 18 Caspase-3 as the core member of the apoptosis cascade signaling is implicated in the development and involution of HAs. 19 Herein, to delineate the function and mechanisms of miR-199a in HAs, we examined the expression levels of miR-199a and PCNA in HAs and evaluated the effects of miR-199a on cell proliferation and apoptosis of HAs cells and found that miR-199a suppressed proliferation and induced apoptosis in HAs cells via targeting HIF1A and might represent a potential therapeutic target for HAs.
Materials and methods
Materials
HA-derived endothelial cells (HDEC) and CRL-2586 EOMA cell lines used in our study were from Laboratory of General Surgery, Xinhua Hospital. The lentivirus vector Lv-HIF1A overexpression vector, negative control vector, and virion-packaging elements were from Genechem (Shanghai, China). The primers of miR-199a and HIF1A were provided by Genepharma (Shanghai, China); HIF1A, PCNA, and cleaved caspase-3 antibodies were from Abcam (Cambridge, MA, USA).
Tissue samples
According to the Mulliken classification in combination with the protein level of PCNA, 20 the patients with HAs were divided into groups: normal skin tissues (n = 15), involuting-phase HAs (n = 15), and proliferating-phase HAs (n = 15), which were collected at the Department of General Surgery. Tissues and clinical information were obtained as part of an approved study at Shanghai Jiao Tong University School of Medicine. Among the patients with HAs, the HA tissues originated from the infants and children, of which there were 19 males and 11 females, and their age ranged from 2 months to 11 years (average age 21 months). HA tissues were diagnosed by two independent pathologists. We promised that the parents of the infants or children gave written informed consent for the HA patients.
Cell culture and transfection
HDEC and CRL-2586 EOMA cell lines were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Sigma Aldrich, St Louis, MO, USA) supplemented with 10% heat-inactivated fetal bovine serum (FBS; Sigma Aldrich, USA), 100 U/mL of penicillin, and 100 μg/mL of streptomycin (Invitrogen, Carlsbad, CA, USA) and were placed in a humidified atmosphere containing 5% CO2 at 37°C. Lv-HIF1A overexpression vector and miR-199a mimic or inhibitor (Genepharma) were transfected into HAs cells, and further functional experiments were performed.
Quantitative real-time polymerase chain reaction
Total RNA for HIF1A was extracted with TRIzol reagent kit (Ambion, USA) and the total RNA for miR-199a was extracted using a mirVana microRNA Isolation Kit (Ambion) according to the manufacturer’s protocol. After added poly(A) tail on miRNA, the complementary DNAs (cDNAs) were acquired based on oligo(dT)18 primers and Moloney Murine Leukemia Virus (M-MLV) reverse transcriptase utilizing SYBR Green Master Mix kit (Applied Biosystems). qRT-PCR assay was conducted with quantitative polymerase chain reaction (qPCR) instrument (Fermentas, Burlington, ON, Canada) to detect relative expression levels of miR-199a and HIF1A mRNA. HIF1A gene was amplified using specific oligonucleotide primer and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) or U6 was used as an endogenous control. The PCR primer sequences of HIF1A gene were as follows: 5′-GAACGTCGAAAAGAAAAGTCTCG-3′ and 5′-CCTTATCAAGATGCGAACTCACA-3′; GAPDH, 5′-CAACGAATT TG GCTACAGC A-3′ and 5′-AGGGGTCTACATGGCAACTG-3′. miR-199a, 5′-CAATCGCTTTCAAATAG-3′ and 5′-CAGGAGATGCTGTC ATC-3′. U6, 5′-CTCGCTTCGGCAGCACA-3′ and 5′-AACGCTTCACGAATT TGCGT-3′. For the detection of miRNA, miRNA-specific looped RT-primers and TaqMan probes were used as described by the manufacturer’s protocol (Applied Biosystems). PCRs were performed in triplicate using a 7300 Real-Time PCR system (Applied Biosystems), and the data were analyzed using the comparative Ct method (2-∆∆Ct).
Western blot assay
HDEC and CRL-2586 EOMA cell lines were harvested and extracted using lysis buffer, and then equal amount of cell extracts were separated on 15% sodium dodecyl sulphate–polyacrylamide gel electrophoresis (SDS-PAGE) gels. Separated protein bands were transferred into polyvinylidene fluoride (PVDF) membranes from Millipore Corporation (Billerica, MA, USA), which were blocked in 5% skim milk powder. The primary antibodies against HIF1A, PCNA, and cleaved caspase-3 were diluted according to the instructions of antibodies and incubated overnight at 4°C. Then, horseradish peroxidase–linked secondary antibodies (Abcam, Cambridge, MA, USA) were added at a dilution ratio of 1:1000 and incubated at room temperature for 2 h. The membranes were washed with phosphate-buffered saline (PBS) and the bands were visualized using ECL-PLUS/Kit (GE Healthcare, Piscataway, NJ, USA) according to the kit’s instruction.
Cell viability assay
Cell viability was analyzed with the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay. Briefly, cells transfected with miR-199a mimic or inhibitor were incubated in 96-well plates with DMEM medium supplemented with 10% FBS. Cells were treated with 10 μL MTT dye (Sigma-Aldrich, St. Louis, MO, USA) at each day and then incubated with 100 μL of dimethyl sulfoxide (DMSO) for 15 min. The color reaction was measured at 570 nm with enzyme immunoassay analyzer (Bio-Rad, Hercules, CA, USA).
Cell apoptosis analysis
HDEC and CRL-2586 EOMA was trypsinized, washed with PBS, and resuspended in binding buffer according to the instruction of the apoptosis kit with fluorescein isothiocyante (FITC)–Annexin V and propidium iodide (PI; BD Biosciences, San Diego, CA, USA), which were added to the fixed cells for 15 min in darkness at room temperature. Then, Annexin V binding buffer was added to the mixture before the fluorescence was measured on FACsort flow cytometer (BD Biosciences).
Luciferase reporter assay
HA cells were seeded into 24-well plates. After 24 h incubation, pmirGLO report vector carrying wild-type (WT) 3′-UTR or mutated 3′-UTR of HIF1A was co-transfected with miR-199a mimic or inhibitor into HDEC and CRL-2586 EOMA cells. Luciferase activities were examined with a Dual-luciferase Reporter System (Promega, Beijing, PR, China) 48 h after transfection.
Statistical analysis
SPSS 15.0 was used for the statistical analysis. Independent T-test was used to analyze the differences between groups. Statistical significance was P < 0.05.
Results
The expression levels of miR-199a and PCNA in different-phase HAs
The expression levels of miR-199a and PCNA were examined in distinct-phase HAs by qRT-PCR analysis, which showed that the expression level of miR-199a was decreased (Figure 1(a)), while that of PCNA was increased in proliferating-phase HAs compared with the normal skin tissues and the involuting-phase HAs (Figure 1(b)). In addition, spearman correlation analysis indicated that miR-199a had a negative correlation with PCNA expression in proliferating-phase HAs (Figure 1(c)), but they had no correlation in involuting-phase HAs (Figure 1(d)).

The expression of miR-199a in distinct-phase HAs. (a and b) qRT-PCR analysis of the expression levels of miR-199a and PCNA in normal skin tissues, involuting-phase HAs and proliferating-phase HAs. (c and d) Spearman correlation analysis of the association of miR-199a with PCNA expression in involuting-phase HAs and proliferating-phase HAs.
miR-199a mimic inhibited cell proliferation and induced apoptosis
To delineate the functions of miR-199a in HAs cells, cell viability assay and flow cytometry analysis were conducted. After HDEC and CRL-2586 EOMA cells were transfected with miR-199a mimic for 48 h, the transfection efficiency of miR-199a mimic was determined by qRT-PCR analysis (Figure 2(a)). We then found that miR-199a mimic suppressed cell proliferation activity (Figure 2(b)) and induced cell apoptosis (Figure 2(c) and (d)) in HDEC and CRL-2586 EOMA cells. The expression levels of PCNA and cleaved caspase-3 were detected by western blotting, indicating a decreased expression of PCNA and increased expression of cleaved caspased-3 caused by miR-199a mimic (Figure 2(e)).

The effects of miR-199a mimic on cell proliferation and apoptosis. (a) qRT-PCR analysis of transfection efficiency of miR-199a mimic in HDEC and CRL-2586 EOMA cells. (b) MTT assay evaluation of the cell proliferation activity after transfection with miR-199a mimic. (c and d) Flow cytometry analysis of the cell apoptosis index after transfection with miR-199a mimic. (e) Western blotting analysis of the expression levels of PCNA and cleaved caspase-3 in miR-199a mimic–transfected HAs cells (**P < 0.01).
miR-199a inhibitor promoted proliferation and prevented apoptosis
We further investigated the effects of miR-199a inhibitor on HAs cell proliferation and apoptosis. After HDEC and CRL-2586 EOMA cells were pretreated with miR-199a inhibitor for 48 h, the transfection efficiency of miR-199a inhibitor was assessed by qRT-PCR analysis (Figure 3(a)). Then, we found that miR-199a inhibitor increased cell proliferation activity (Figure 3(b)) but reduced cell apoptosis (Figure 3(c) and (d)) in these two cell lines. Western blotting analysis showed an increased expression of PCNA but decreased expression of cleaved caspased-3 induced by miR-199a inhibitor (Figure 3(e)).
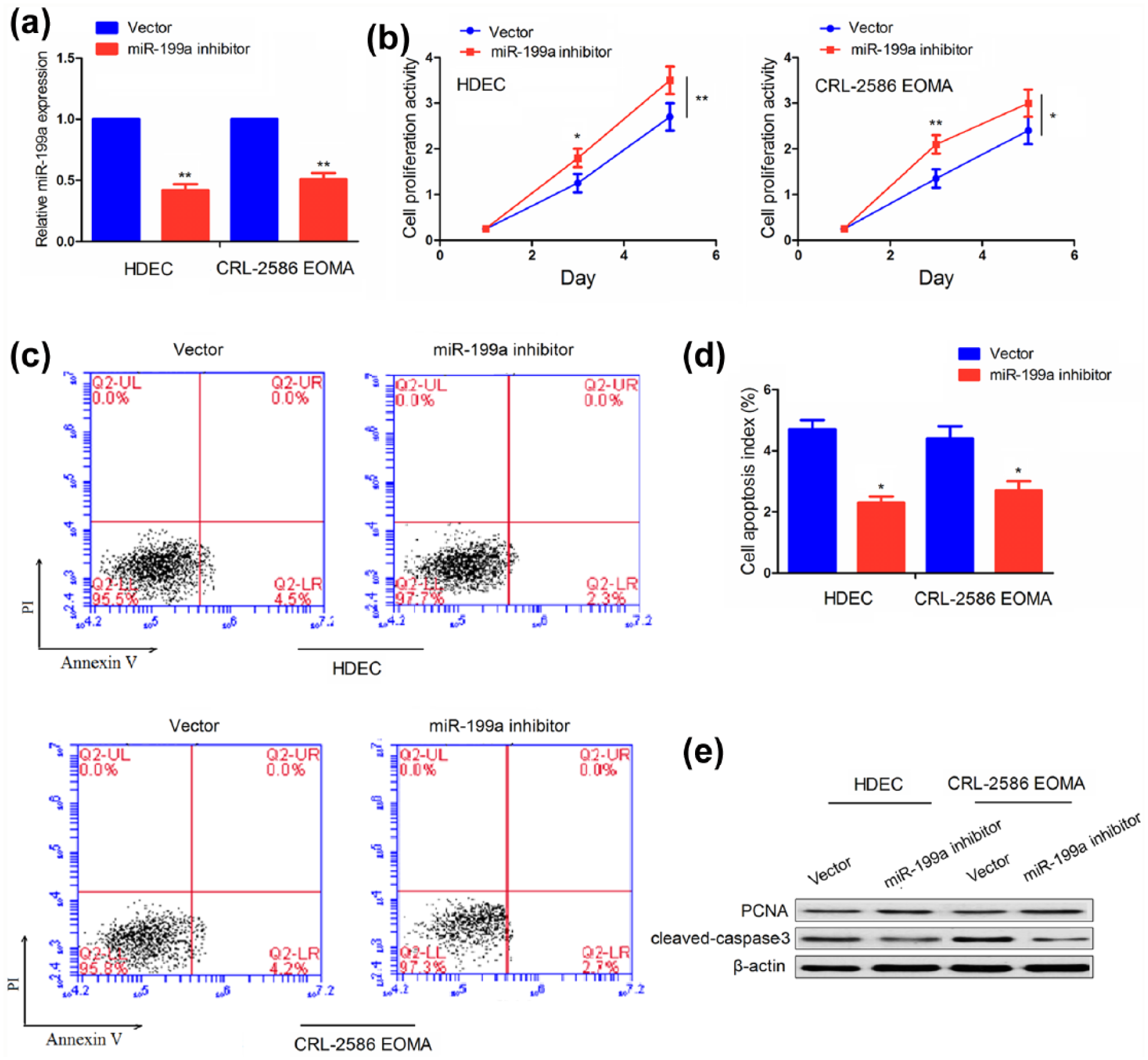
The effects of miR-199a inhibitor on cell proliferation and apoptosis. (a) qRT-PCR analysis of transfection efficiency of miR-199a inhibitor in HDEC and CRL-2586 EOMA cells. (b) MTT assay evaluation of the cell proliferation activity after transfection with miR-199a inhibitor. (c and d) Flow cytometry analysis of the cell apoptosis index after transfection with miR-199a inhibitor. (e) Western blotting analysis of the expression levels of PCNA and cleaved caspase-3 in miR-199a inhibitor pretreated HA cells (*P < 0.05; **P < 0.01).
HIF1A was identified as a direct target of miR-199a
To dissect the molecular mechanism by which miR-199a regulates HAs proliferation, we screened the target genes of miR-199a by using the miRWalk 2.0 (http://zmf.umm.uni-heidelberg.de/apps/zmf/mirwalk2/genepub.html) for bioinformatic analysis including targetScan (http://www.targetscan.org/vert_71/), miRanda, picTar, and PITA, which identified about 13 target genes of miR-199a (Figure 4(a)). Among them, HIF1A ranked the first place and was considered to have the greatest potential to bind with miR-199a (Figure 4(b)). Besides, qRT-PCR (Figure 4(c)) and western blotting analyses (Figure 4(d)) showed that miR-199a mimic decreased the expression of HIF1A, whereas miR-199a inhibitor increased its expression in HDEC and CRL-2586 EOMA cells. Then, the WT 3′-UTR or mutant 3′-UTR target sequences of HIF1A (Figure 4(e)) were cloned into the luciferase reporter vector and co-transfected with miR-199a mimic or inhibitor into HAs cells. Our results illustrated that miR-199a mimic reduced the luciferase activity of WT 3′-UTR of HIF1A, while miR-199a inhibitor increased this effect, but both of them had no effect on the luciferase activity of mutation 3′-UTR of HIF1A (Figure 4(f)).

HIF1A was identified as a target of miR-199a. (a and b) Bioinformatic analysis of the target genes of miR-199a. (c and d) qRT-PCR and western blotting analysis of the expression level of HIF1A in HDEC and CRL-2586 EOMA cells transfected with miR-199a mimic or inhibitor. (e) Diagrams showed the binding sites of miR-199a with WT and mutant 3′-UTR of HIF1A. (f) Luciferase activity of WT or mutant HIF1A 3′-UTR in HDEC and CRL-2586 EOMA cells transfected with miR-199a mimic or inhibitor (**P < 0.01).
HIF1A overexpression reversed the anti-proliferation effect of miR-199a
We first detected the expression level of HIF1A in different-phase HAs by qRT-PCR and found that HIF1A expression was elevated in proliferating-phase HAs, and spearman correlation analysis revealed the negative correlation of HIF1A with miR-199a expression in proliferating-phase HAs (Figure 5(a)). Furthermore, qRT-PCR (Figure 5(b)) and western blotting analysis (Figure 5(c)) showed the transfection efficiency of HIF1A overexpression vector or HIF1A siRNA (si-HIF1A) in HDEC and CRL-2586 EOMA cells. After transfection for 5 days, cell viability assay displayed increased cell proliferation activity by HIF1A overexpression but decreased proliferation activity caused by HIF1A knockdown (Figure 5(d)). Moreover, HIF1A overexpression attenuated the anti-proliferating effect of miR-199a mimic (Figure 5(e)), while knockdown of HIF1A lowered the proliferation promoting effect by miR-199a inhibitor in HAs cells (Figure 5(f)).

HIF1A overexpression reversed the anti-proliferation effect of miR-199a. (a) qRT-PCR analysis of the expression level of HIF1A and its correlation with miR-199a in proliferating-phase HAs. (b and c) qRT-PCR and western blot analysis of the expression level of HIF1A in HDEC and CRL-2586 EOMA cells transfected with HIF1A overexpression plasmid or HIF1A siRNA. (d) MTT analysis of the cell proliferation activity after transfection with HIF1A overexpression plasmid or HIF1A siRNA for 5 days. (e and f) MTT analysis of the cell proliferation activity after co-transfection with miR-199a mimic + HIF1A or miR-199a inhibitor + si-HIF1A for 5 days (*P < 0.05; **P < 0.01).
Discussion
Accumulating data demonstrate that miR-199a confers a dual role in tumor progression. On one hand, miR-199a has been proved to be downregulated in most of the tumor tissues.10 –15 On the other hand, it is also thought to be upregulated in a small amount of tumor tissues.16,17 Some studies indicate PCNA as the biomarker in response to the proliferating- or involuting-phase HAs, 18 and our previous studies showed the correlation of PCNA expression with the proliferating HAs. 20 But, there is little knowledge about the correlation of miR-199a with PCNA expression in HAs. In the present study, we explored the expression of miR-199a in human HAs samples, and found miR-199a was downregulated but PCNA was upregulated in proliferating-phase HAs compared with the normal tissues and involuting-phase HAs. Likewise, we found that miR-199a had the negative correlation with PCNA expression in proliferating-phase HAs rather than in the involuting-phase HAs.
Functionally, many studies have shown that miRNAs exert a critical role in the development of HAs. For example, circular C19MC miRNAs can be utilized as the biomarkers of infantile HAs. 21 miR-21a/Nox-4 axis mediates the Dicer to promote the formation of HAs. 22 Overexpression of miR-424 or knockdown of miR-382 by propranolol inhibits the progression of HAs via MEK1 or Phosphatase and tensin homolog (PTEN)/AKT/mechanistic target of rapamycin (mTOR) pathway.23,24 Moreover, the connection between miR-199a and tumor have shown that restoration of miR-199a either restrains tumor proliferation and invasion10 –15 or promotes tumor growth and aggressiveness.16,17 In addition, miR-199a facilitates breast cancer cell sensitivity to cisplatin by targeting TFAM 25 and inhibition of miR-199a results in cisplatin resistance of ovarian cancer by increasing DDR1 expression. 26 To illuminate the functions of miR-199a in HAs cells, we utilized the miR-199a mimic and inhibitor to pretreat HAs cells and found that miR-199a mimic suppressed cell proliferation and induced cell apoptosis with the decreased PCNA expression and increased cleaved caspase-3 expression, while miR-199 inhibitor reversed these effects, indicating the tumor-suppressive role of miR-199a in HAs cells, which was analogous to the previously reported studies.10 –15
Furthermore, increased expression of HIF1A is connected to cellular apoptosis, angiogenesis, unfavorable survival in rectal carcinoma, 27 and esophageal carcinoma 28 and reduces cell apoptosis and accelerates growth and invasion in tongue squamous cell carcinoma 29 and neuroblastoma. 30 Our and other previous studies showed that inhibition of HIF1A signaling represses HA cell proliferation and induces cell apoptosis.31,32 Moreover, it is demonstrated that miR-199a has the negative correlation with HIF-1 expression and suppresses the tumor progression in multiple malignant tumors.33,34 But, the association between miR-199a and HIF1A in HAs is still undefined. We here found that HIF1A was just a direct target of miR-199a and overexpression of HIF1A attenuated the anti-proliferation effect of miR-199a in HAs cells. Our findings were consistent with the previously reported results, which showed that miR-199a inhibited cell proliferation by targeting HIF1A in melanoma, 33 pancreatic ductal adenocarcinoma, 34 lung cancer, 35 and endometrial stromal cells. 36
In conclusion, our findings demonstrate that miR-199a suppresses proliferation and induces apoptosis in HA cells via targeting HIF1A and provide a potential therapeutic target for HAs.
Footnotes
Acknowledgements
Yang Wang and Yu-Xin Dai contributed equally to this article. Especially, we thank ZW Quan, YB Liu, and JM Ou for providing help and guide for our work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the Medical-Engineering Cross Fund Project of Shanghai Jiao Tong University (No. YG2015MS66), Shanghai Municipal Science and Technology Commission (No. 15140901600), and Natural Science Foundation of China (81572673).
