Abstract
Background
Lung adenocarcinoma (LUAD) remains one of the leading causes of cancer-related deaths worldwide, with limited therapeutic efficacy despite advances in treatment. Our study investigated the role of LINC00578 and its regulatory interaction with miR-153-5p in LUAD progression.
Methods
The expression levels of LINC00578 and miR-153-5p were assessed in LUAD tissues and adjacent normal tissues using quantitative real-time polymerase chain reaction. Associations between LINC00578 expression and clinicopathological features and patient prognosis were analyzed. Luciferase reporter assays, proliferation, migration, invasion, and apoptosis analyses, were conducted in LUAD cell lines (PC-9 and H1299) following modulation of LINC00578 and miR-153-5p expression.
Results
LINC00578 was significantly upregulated in LUAD tissues compared to adjacent normal tissues, whereas miR-153-5p was markedly downregulated. High LINC00578 expression was associated with lymph node metastasis, tumor-node-metastasis stage, and poor overall survival. Functional studies revealed that silencing LINC00578 inhibited LUAD cell proliferation, migration, and invasion while promoting apoptosis. Mechanistically, LINC00578 exerted its oncogenic effects by negatively regulating miR-153-5p expression. Inhibition of miR-153-5p reversed the tumor-suppressive effects induced by LINC00578 knockdown.
Conclusion
LINC00578 functions as an oncogenic long non-coding RNA in LUAD by promoting proliferation and metastasis through suppression of miR-153-5p. LINC00578 may serve as a novel prognostic biomarker for LUAD.
Introduction
Globally, lung cancer remains the most prevalent malignancy and the primary contributor to cancer mortality. In terms of incidence, it holds the second position among cancers, preceded by prostate cancer in males and breast cancer in females. 1 Lung cancer is responsible for nearly 1.8 million annual fatalities, representing roughly 18% of global cancer mortality and ranking as the deadliest form of the disease. 2 Pathologically, it is classified into two main subtypes: small cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC), the latter comprising about 85% of diagnoses.3,4 Lung adenocarcinoma (LUAD) is one of the most common subtypes of NSCLC, mostly originating from the bronchial mucosal epithelium, with a few originating from the mucous glands of the large bronchioles, and mainly from the mucous glands of the small bronchioles, with more than three-quarters of the patients’ foci occurring in the periphery, and rarely with cavitary formation. 5 Lung adenocarcinomas are relatively common in nonsmokers, women, and younger patients, with the average age of patients typically between 40 and 50 years. 6 This characteristic makes lung adenocarcinoma somewhat different from other types of lung cancer and requires individualized treatment strategies. 7
Long non-coding RNAs (lncRNAs), which are transcribed by RNA polymerase II, play a regulatory role in numerous cellular and biological processes. 8 Functionally, lncRNAs are involved in cell cycle regulation, modulation of cell death, influence on cell differentiation, and the regulation of immune responses, further underscoring the complexity of lncRNA-mediated biological activities. 9 The regulatory mechanisms of lncRNAs are considered potential factors in tumor malignant transformation primarily through the modulation of oncogenes, thereby driving cancer phenotypes. A growing body of research has confirmed lncRNAs can serve as biomarkers for tumor prediction and prognostic assessment, making them promising candidates for novel diagnostic tools and therapeutic targets in cancer.10–12
Recent studies have revealed that LINC00578, an immune-related lncRNA, is dysregulated in NSCLC. 13 Moreover, aberrant expression and functional roles of LINC00578 have been identified in various human malignancies, including pancreatic cancer, breast cancer, and hepatocellular carcinoma (HCC).14–16 In pancreatic cancer, LINC00578 is highly expressed and promotes malignant tumor progression; its elevated levels are associated with poor patient prognosis, suggesting its potential as a diagnostic and prognostic biomarker as well as a therapeutic target. 14 Bioinformatics analyses have identified a binding site between LINC00578 and miR-153-5p. In addition, it was found that miR-153-5p was involved in the progression of colorectal, breast, and gastric cancers.17–19 In NSCLC, miR-153-5p expression is suppressed by the lncRNA HIF1A-AS2, and its downregulation is associated with poor prognosis in patients. 20 Mechanistically, miR-153-5p promotes proliferation, migration, and invasion of lung cancer cells by targeting and inhibiting ROCK1 expression, thereby activating the epithelial–mesenchymal transition (EMT) signaling pathway. 21
This investigation examines the expression characteristics and functional roles of LINC00578 and miR-153-5p in LUAD. Using comprehensive experimental approaches and mechanistic analyses, we seek to characterize their expression profiles in LUAD specimens and evaluate potential correlations with clinical–pathological parameters and patient outcomes. The ultimate goal is to assess their potential as novel prognostic biomarkers and to provide new theoretical foundations and potential therapeutic targets for targeted treatment of LUAD.
Materials and methods
Study population and follow-up
A total of 97 patients diagnosed with LUAD and treated surgically at our hospital were included in this study. All surgical specimens met the following criteria: all patients had a single primary tumor with no other antitumor therapy, and the pathology results had been reviewed by two senior pathologists and finally determined to be primary lung adenocarcinoma. The exclusion criteria for the collection of specimens were as follows: patients who had received any neoadjuvant therapy prior to surgery; patients with a history of other tumors in the past; and patients diagnosed with other immune system disorders during the treatment period. Ethical approval for this research was obtained from the Institutional Review Board of Jinling Hospital, The Affiliated Hospital of Medical School, Nanjing University. Surgical specimens, including tumor tissues and matched normal adjacent tissues, were collected intraoperatively and histologically confirmed by two certified pathologists. All specimens were rapidly frozen at −80°C for long-term preservation prior to molecular analyses. For outcome assessment, participants received systematic postoperative surveillance via scheduled clinical evaluations or telephone follow-ups over a period ranging from 3 to 60 months. Survival metrics were prospectively recorded and statistically evaluated.
Clinicopathological staging for all patients was rigorously determined according to the American Joint Committee on Cancer (AJCC) Cancer Staging Manual, 8th edition. 22 The tumor-node-metastasis (TNM) stage is a composite indicator encompassing: the extent of the primary tumor (T stage), lymph node involvement (N stage), and distant metastasis (M stage). Tumor differentiation grade (well, moderate, poor) was assessed independently by the pathologists as a measure of histological aggressiveness, which is related to but distinct from the T stage. All staging and grading information was extracted from the final, reviewed pathology reports.
Gene expression assessment by quantitative real-time polymerase chain reaction
Total RNA was extracted from tissues and cell lines using Trizol reagent (Invitrogen, USA) per the manufacturer's protocol. The concentration and purity of RNA were measured using a NanoDrop 2000, and only samples with an OD260/280 ratio between 1.8 and 2.0 were considered to be of high quality. For LINC00578 mRNA detection, cDNA was synthesized from 1 µg of total RNA using the PrimeScript™ RT Reagent Kit with gDNA Eraser (TaKaRa, Japan) according to the manufacturer's instructions, which includes genomic DNA removal and reverse transcription with oligo(dT) and random hexamer primers. For the specific detection of mature hsa-miR-153-5p (MIMAT0026480), cDNA was synthesized using the Mir-X™ miRNA First-Strand Synthesis Kit (TaKaRa, Japan), which employs a stem-loop reverse transcription primer specific to miR-153-5p, following the kit protocol. quantitative real-time polymerase chain reaction (qRT-PCR) for both LINC00578 and miR-153-5p was performed on a CFX96 Real-Time PCR System (Bio-Rad) using SYBR® Premix Ex Taq™ II (TaKaRa, Kusatsu, Shiga, Japan). The thermocycling conditions were: 95°C for 30 s, followed by 40 cycles of 95°C for 5 s and 60°C for 30 s. GAPDH and U6 snRNA were used as endogenous controls for LINC00578 and miR-153-5p, respectively. Gene expression levels were calculated using the 2^(-ΔΔCt) method. The sequences of all primers used are listed in Supplemental Table S1. 23
Cell culture and transfection
LUAD cell lines (A549, H1299, PC-9, and H2009) and the normal human bronchial epithelial (HBE) cell line were cultured in DMEM (Gibco, USA) supplemented with 10% fetal bovine serum (FBS) and 0.1% penicillin-streptomycin. Cells were transfected with miR-153-5p inhibitors to modulate miR-153-5p expression. The LINC00578-targeting siRNA (5′-GCTGTCCACATTTTCACTATCT-3′) and a negative control siRNA (5′-UUCUCCGAACGUGUCACGUTT-3′) were designed and synthesized by Sangon Biotech (Shanghai, China). Transfections were performed at room temperature using Lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions. After 48 h, the expression levels of LINC00578 and miR-153-5p were measured to evaluate transfection efficiency.
Cell counting kit-8 assay
For cell viability assessment, cultures were plated in 96-well plates (100 μL/well) and incubated for 24 h to ensure proper attachment prior to experimental treatments. Following intervention, 10 μL of cell counting kit-8 (CCK-8) reagent (Dojindo, Kumamoto, Japan) was introduced into each well, followed by a 1–4 h dark incubation period. Absorbance measurements at 450 nm were then obtained using a microplate spectrophotometer to quantify viable cells.
Transwell assay
Cell migration was assessed using Transwell chambers without Matrigel coating. After 48 h of transfection, cells were digested and resuspended in serum-free medium. Cells were incubated at 37°C in a 5% CO₂ atmosphere for 24 h. Migrated cells on the lower surface were fixed with 4% paraformaldehyde for 30 min and stained with 0.1% crystal violet for 20 min.
For the invasion assay, the upper chambers were pre-coated with 50 μL of Matrigel and incubated at 37°C for 4 h to allow solidification. The subsequent steps were identical to those used in the migration assay, except that the incubation time was extended to 36–48 h to account for the slower invasion process. Stained cells were counted in five randomly selected fields per well under a light microscope to quantify migration and invasion capacities.
Cell apoptosis assay
Apoptosis rates were determined by flow cytometric analysis of cells double-labeled with Annexin V-FITC and propidium iodide (PI). Following transfection, harvested cells underwent fixation in 3.7% paraformaldehyde (15 min, RT) and permeabilization with 0.1% Triton X-100 (5 min), then were suspended in binding buffer. Quantitative assessment of apoptotic populations was performed using a BD FACScan.
Dual-Luciferase reporter assay
The interaction between LINC00578 and miR-153-5p was predicted by bioinformatics tools (lncRNASNP2: https://guolab.wchscu.cn/lncRNASNP#!/lncrna_info?lncrna = NONHSAT093344.2). To validate this, a DNA fragment containing the wild-type (WT) predicted binding site was amplified and inserted into the pmirGLO dual-luciferase vector (Promega, Madison, WI, USA), downstream of the firefly luciferase gene. A corresponding mutant (MT) reporter construct was generated by introducing specific nucleotide substitutions within the binding site using site-directed mutagenesis. PC-9 and H1299 cells Pirlo were co-transfected with the reporter plasmid (WT or MT) and either a synthetic miR-153-5p mimic (miR-mimic, to augment cellular miR-153-5p levels), a miR-153-5p inhibitor (antagomiR, a chemically modified antisense oligonucleotide, miR-inhibitor), or a negative control (miR-NC), (all purchased from GenePharma, Suzhou, China). After 48 h, firefly and Renilla luciferase activities were measured sequentially using the Dual-Luciferase Reporter Assay System (Promega). Firefly luminescence was normalized to Renilla luminescence for each sample. The MUT reporter serves as a critical negative control to rule off-target effects and confirm the binding site specificity of miR-153-5p.
Statistical analysis
Statistical analyses were performed with SPSS 26.0 (IBM) and GraphPad Prism 7.0. Data are expressed as mean ± standard deviation. The specific sample size (n) for each experiment is provided in the corresponding figure legends. Differences between two groups were analyzed using the two-tailed Student's t-test. Comparisons among multiple groups were performed with one-way analysis of variance (ANOVA) followed by Tukey's post hoc test. Associations between LINC00578 expression and clinicopathological variables were evaluated with the Chi-square test. Survival curves were generated by the Kaplan–Meier method and compared using the log-rank test. Prognostic factors were identified by multivariate Cox proportional hazards regression analyses. The correlation between LINC00578 and miR-153-5p expression levels was determined using Pearson's correlation analysis. A
Results
Association between LINC00578 expression, clinicopathological features, and prognosis in LUAD patients
Data retrieved from the UALCAN database (The Cancer Genome Atlas (TCGA)-LUAD cohort) showed that LINC00578 was significantly upregulated in primary LUAD tumors compared with normal lung tissues (
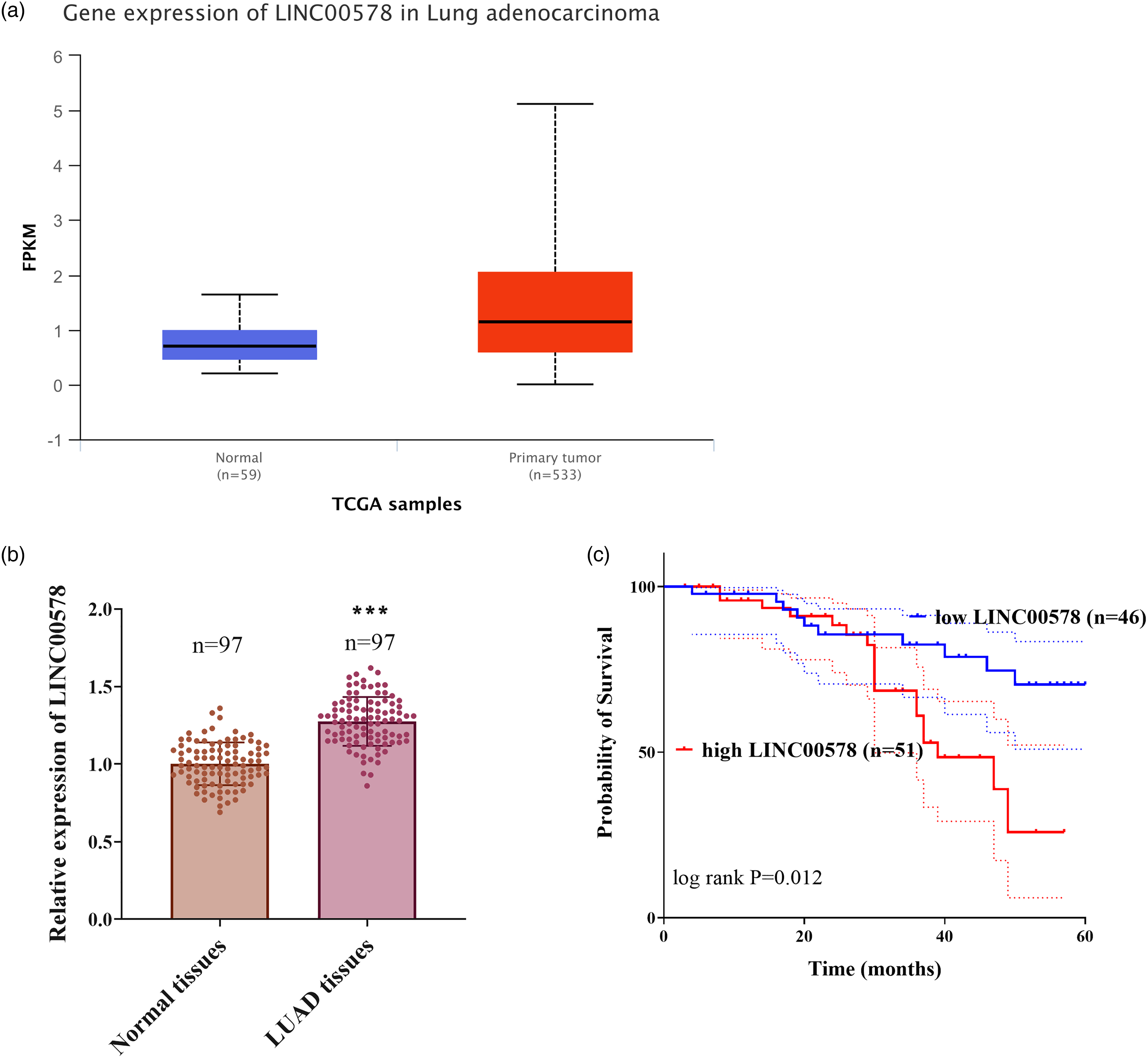
LINC00578 was upregulated in LUAD and associated with poor prognosis. (a) LINC00578 expression levels in normal lung tissues (n = 59) and primary LUAD tumors (n = 533) from the TCGA-LUAD cohort (data sourced from UALCAN database: https://ualcan.path.uab.edu/cgi-bin/TCGA-lncRNA-Result.pl?genenam = LINC00578&ctype = LUAD. Statistical significance was analyzed by unpaired Student's t-test,
Regulatory relationship between LINC00578 and miR-153-5p
Conversely, miR-153-5p expression showed substantial downregulation in malignant specimens (
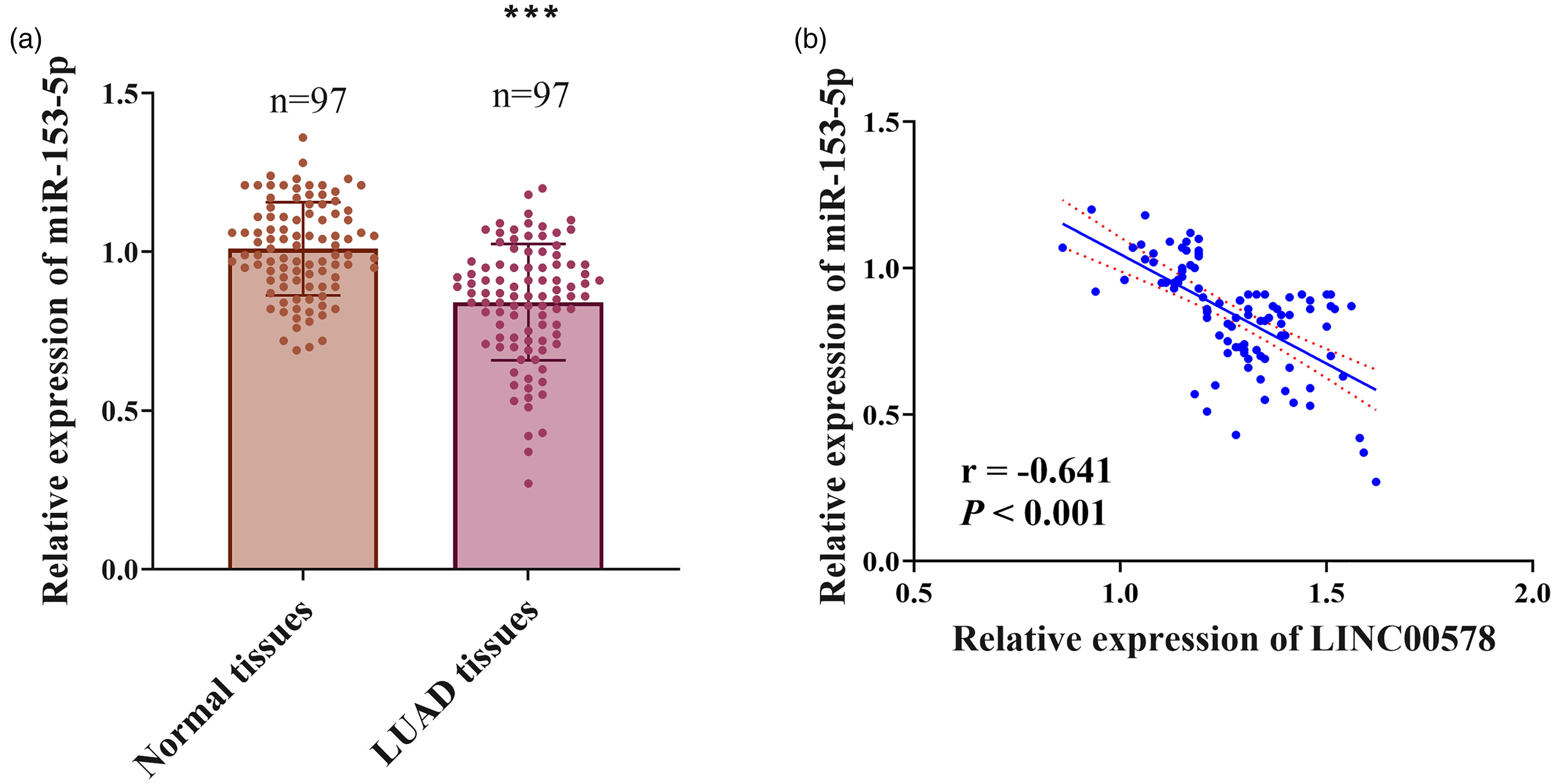
miR-153-5p was downregulated in LUAD and inversely correlated with LINC00578 expression. (a) Relative expression levels of miR-153-5p in 97 paired LUAD tissues and adjacent normal tissues. Data are presented as individual data points with mean ± SD. ***
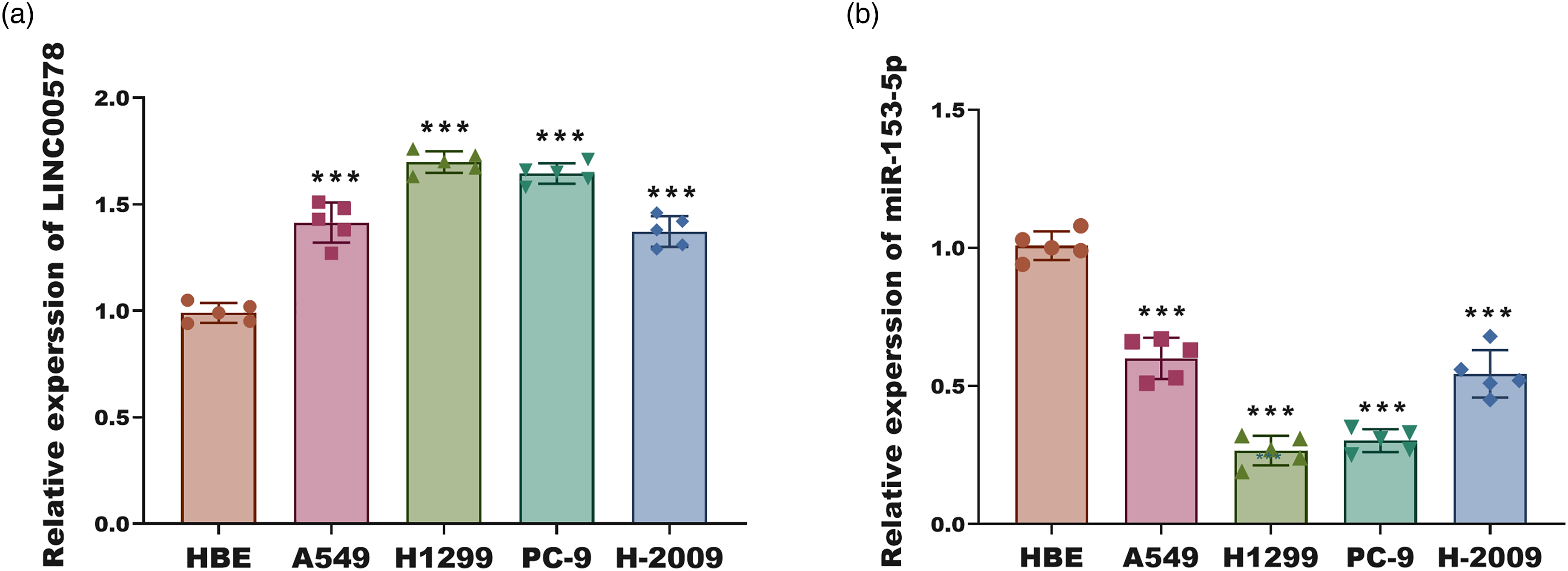
Expression patterns of LINC00578 and miR-153-5p in LUAD cell lines. (a) Relative expression of LINC00578 in normal human bronchial epithelial (HBE) cells and four LUAD cell lines (A549, H1299, PC-9, H2009). (b) Relative expression of miR-153-5p in the same panel of cells as in (a). Data in (a) and (b) are from five independent experiments (n = 5) and presented as mean ± SD. ***
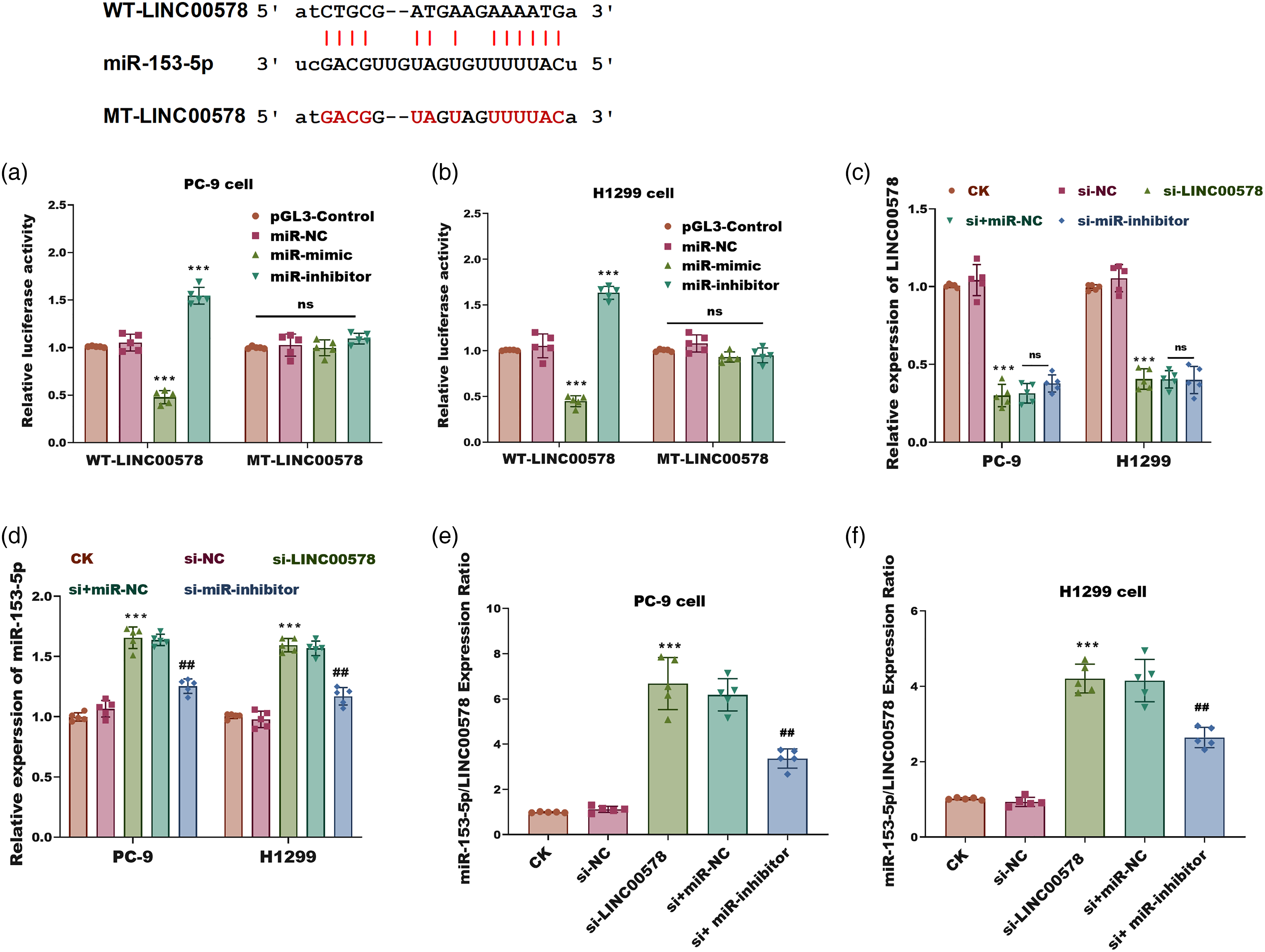
MiR-153-5p directly binds to and regulates LINC00578 in LUAD cells. (a) and (b) Dual-luciferase reporter assay in PC-9 (a) and H1299 (b) cells. Cells were co-transfected with either the wild-type (WT) or mutant (MT) LINC00578 reporter plasmid, along with the miR-153-5p mimic, miR-153-5p inhibitor (antagomir), or their negative controls (miR-NC). The pGL3-Control group represents cells transfected with the empty pGL3-basic vector to establish baseline activity. Firefly luciferase activity was normalized to Renilla luciferase activity. (c) Relative expression of LINC00578 in PC-9 and H1299 cells transfected with si-NC, si-LINC00578, or si-LINC00578 + miR-153-5p inhibitor. (d) Relative expression of miR-153-5p in the same groups as (c). (e) and (f) The miR-153-5p/LINC00578 expression ratio in PC-9 (e) and H1299 (f) cells. CK: Blank Control. Data are presented as mean ± SD (n = 5). ***
Effects of LINC00578 and miR-153-5p on LUAD cell
In PC-9 and H1299 cells, LINC00578 knockdown significantly suppressed cell proliferation compared to controls (
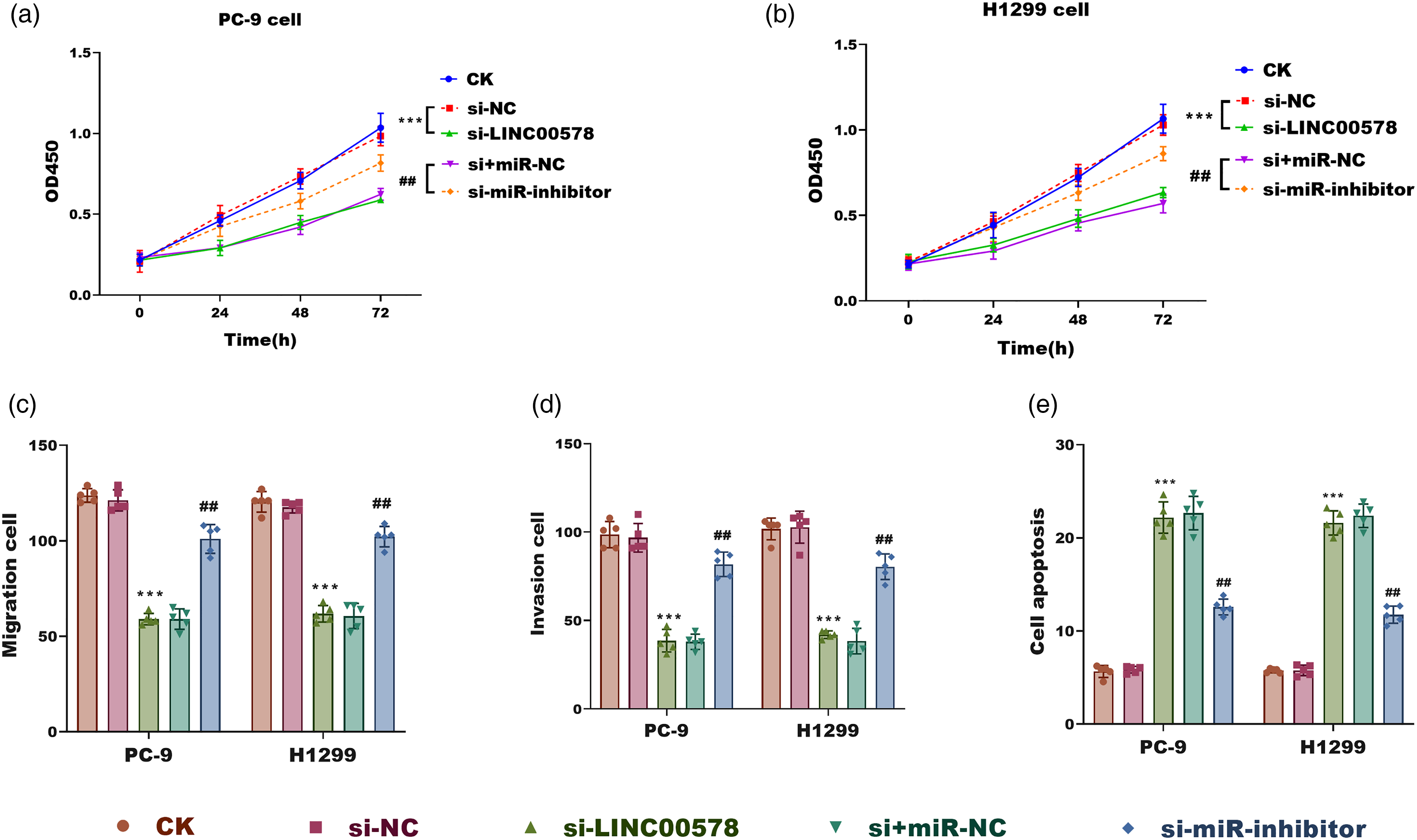
LINC00578 promotes malignant phenotypes in LUAD cells by sponging miR-153-5p. (a) and (b) Cell proliferation assessed by CCK-8 assay in PC-9 (a) and H1299 (b) cells transfected as indicated. Absorbance was measured at 450 nm. (c, d) Cell migration (c) and invasion (d) abilities assessed by Transwell assays in PC-9 and H1299 cells under the indicated treatments. (e) Flow cytometry analysis of apoptosis in PC-9 and H1299 cells under the indicated treatments. Data are mean ± SD (n = 5). ***
Discussion
In recent years, LUAD's incidence and mortality rates have been rising globally due to the combined effects of environmental factors, genetic predispositions, and adverse lifestyle habits. 24 LUAD is characterized by high malignancy, rapid growth, and strong invasive potential into surrounding tissues and organs, complicating treatment strategies. In our study, we found that elevated LINC00578 expression might predict poor prognosis and malignant progression in LUAD patients, with miR-153-5p mediating the oncogenic effects of LINC00578 on LUAD cells.
Our study comprised 97 histologically confirmed LUAD cases, with comparative analysis revealing significantly elevated LINC00578 expression in tumor tissues relative to paired normal adjacent tissue samples. This observation aligns with existing literature documenting LINC00578 overexpression in LUAD pathogenesis, reinforcing its potential role as a consistent molecular marker in this malignancy. 13 However, studies investigating the role of LINC00578 in cancer development are still relatively limited. As a differentially expressed, subtype-specific lncRNA in breast cancer, LINC00578 is implicated in tumor progression through pathways such as NOTCH/NF-κB signaling. 25 These findings suggest LINC00578 may as a promoter of tumorigenesis. Moreover, LINC00578 expression levels are significantly associated with T stage and patient mortality status in HCC, 16 highlighting its potential as a prognostic biomarker. However, the apparent differences in the role of LINC00578 across different cancer types emphasize the complex, context-dependent nature of its function. Building on previous studies emphasizing the oncogenic role of LINC00578 in various malignancies, we further investigated its association with the clinicopathological features of LUAD. Our analysis identified LINC00578 expression, along with TNM stage and lymph node metastasis, as independent prognostic indicators. Additionally, previous studies have demonstrated that LINC00578 may modulate pro-tumorigenic polarization of M2 macrophages through regulation of Notch, transforming growth factor-beta (TGF-β), and interferon-α/γ response pathways, further underlining its involvement in lung cancer prognosis prediction models. 26
Prior research has reported that LINC00578 participates in several cellular processes relevant to disease progression. Li et al. reported that LINC00578 enhances proliferation and tumorigenicity of pancreatic cancer cells by stabilizing SLC7A11 protein expression through the UBE2 K/SLC7A11 axis, thereby inhibiting ferroptosis and promoting cancer progression. 27 Conversely, our findings suggest that silencing LINC00578 affects LUAD cell apoptosis, invasion, and proliferation in an opposing manner. Specifically, knockdown of LINC00578 suppressed LUAD cell invasiveness while markedly promoting apoptosis, providing further evidence that LINC00578 may function through inhibiting apoptotic pathways. Mechanistically, miR-153-5p was identified as a downstream competing endogenous RNA (ceRNA) of LINC00578. Importantly, our results demonstrate that miR-153-5p could counteract the oncogenic effects of LINC00578 in LUAD cells. For instance, in colorectal cancer, miR-153-5p acts as a tumor suppressor by inhibiting MYO6, with its function being compromised by circ_0000395-mediated sponging. 28 Therefore, we propose that the tumor-promoting role of LINC00578 in LUAD is at least partially mediated by miR-153-5p suppression.
In addition to identifying potential novel biomarkers for LUAD, our study highlights the significant advantages of using lncRNAs for monitoring tumor progression. First, we confirmed the prognostic significance of LINC00578 and elucidated its potential molecular mechanisms, underscoring the critical role of lncRNAs in cancer biology. Furthermore, by evaluating the expression of LINC00578 and miR-153-5p not only in tissue samples and cell lines using qRT-PCR but also in plasma specimens from LUAD patients, we expanded the potential application of these molecules as circulating biomarkers for disease diagnosis and prognosis.
Several study limitations warrant consideration. First, the restricted cohort size may limit the statistical power and broader applicability of our clinical findings. Although we validated LINC00578's upregulation using the TCGA dataset, the prognostic value of the LINC00578/miR-153-5p axis, particularly the clinical relevance of miR-153-5p itself, was not independently validated in a larger, multi-center population. Second, while our loss-of-function experiments robustly demonstrated the oncogenic role of LINC00578, the study lacked gain-of-function validation through LINC00578 overexpression, which would have provided complementary evidence for its oncogenic function. Third, our findings are primarily based on in vitro models. Validation through in vivo animal models is essential to confirm the functional roles of LINC00578 and miR-153-5p within a complex tumor microenvironment. Finally, the origin and transport mechanisms of these molecules, such as their presence in extracellular vesicles or exosomes, remain unexplored and could have significant biological implications.29,30 Future research should address these limitations. Efforts will focus on validating the prognostic signature in larger, multi-center cohorts to assess its clinical utility. Gain-of-function studies with LINC00578 overexpression will be performed to further solidify its oncogenic role. Moreover, extensive in vivo experiments are planned to corroborate the tumor-promoting functions of the LINC00578/miR-153-5p axis and to explore its potential as a therapeutic target. Investigating the extracellular vesicle packaging and cell–cell communication mediated by this axis will also be a key direction to understand its broader role in LUAD progression.
In conclusion, elevated expression of LINC00578 in LUAD serves as an independent prognostic marker and is closely associated with disease progression. Mechanistically, LINC00578 promotes LUAD cell proliferation and metastasis, at least in part, through inhibition of miR-153-5p expression.
Supplemental Material
sj-docx-1-jbm-10.1177_03936155261417698 - Supplemental material for LINC00578 as a prognostic biomarker in lung adenocarcinoma via regulation of miR-153-5p
Supplemental material, sj-docx-1-jbm-10.1177_03936155261417698 for LINC00578 as a prognostic biomarker in lung adenocarcinoma via regulation of miR-153-5p by Wenju Pan, Jun Hong, Mingwei Zhang, Sumeng Wang, Xin Lu, Tingting Ji, Jing Zhang and Yaoman Guo in The International Journal of Biological Markers
Supplemental Material
sj-docx-2-jbm-10.1177_03936155261417698 - Supplemental material for LINC00578 as a prognostic biomarker in lung adenocarcinoma via regulation of miR-153-5p
Supplemental material, sj-docx-2-jbm-10.1177_03936155261417698 for LINC00578 as a prognostic biomarker in lung adenocarcinoma via regulation of miR-153-5p by Wenju Pan, Jun Hong, Mingwei Zhang, Sumeng Wang, Xin Lu, Tingting Ji, Jing Zhang and Yaoman Guo in The International Journal of Biological Markers
Supplemental Material
sj-docx-3-jbm-10.1177_03936155261417698 - Supplemental material for LINC00578 as a prognostic biomarker in lung adenocarcinoma via regulation of miR-153-5p
Supplemental material, sj-docx-3-jbm-10.1177_03936155261417698 for LINC00578 as a prognostic biomarker in lung adenocarcinoma via regulation of miR-153-5p by Wenju Pan, Jun Hong, Mingwei Zhang, Sumeng Wang, Xin Lu, Tingting Ji, Jing Zhang and Yaoman Guo in The International Journal of Biological Markers
Footnotes
Ethics approval and consent to participate
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committee of Jinling Hospital, The Affiliated Hospital of Medical School, Nanjing University. In addition, for investigations involving human subjects, informed consent has been obtained from the participants involved.
Consent to publish
All patients provided written informed consent.
Author contributions
Wenju Pan, Xin Lu, Tingting Ji, Jing Zhang contributed to the study conception and design. Material preparation, data collection and analysis were performed by Wenju Pan, Jun Hong, Sumeng Wang, Yaoman Guo and Mingwei Zhang. The first draft of the manuscript was written by Wenju Pan and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
All data generated or analyzed during this study are included in this article and its supplementary material files. Further enquiries can be directed to the corresponding author.
Supplemental material
Supplemental material for this article is available online.
